

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146651.7 - phase: 0 /pseudo
(394 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
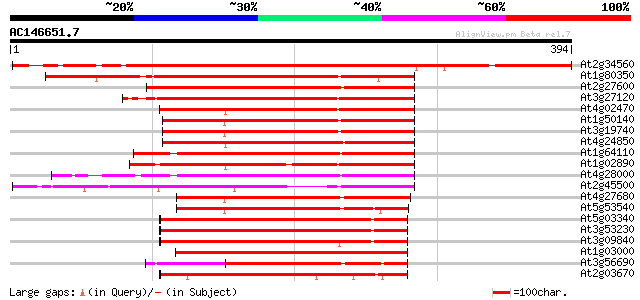
Score E
Sequences producing significant alignments: (bits) Value
At2g34560 katanin like protein 431 e-121
At1g80350 CAD ATPase (AAA1) 237 7e-63
At2g27600 putative AAA-type ATPase 223 1e-58
At3g27120 spastin protein, putative 192 4e-49
At4g02470 172 3e-43
At1g50140 hypothetical protein, 5' partial 170 1e-42
At3g19740 ATPase, putative 168 4e-42
At4g24850 unknown protein (At4g24850) 167 7e-42
At1g64110 unknown protein 164 8e-41
At1g02890 hypothetical protein 159 3e-39
At4g28000 putative protein 157 1e-38
At2g45500 hypothetical protein 153 1e-37
At4g27680 unknown protein 152 4e-37
At5g53540 26S proteasome regulatory particle chain RPT6-like pro... 150 1e-36
At5g03340 transitional endoplasmic reticulum ATPase 147 8e-36
At3g53230 CDC48 - like protein 146 2e-35
At3g09840 putative transitional endoplasmic reticulum ATPase 145 3e-35
At1g03000 putative peroxisome assembly factor-2 135 3e-32
At3g56690 calmodulin-binding protein 134 7e-32
At2g03670 putative AAA-type ATPase 129 4e-30
>At2g34560 katanin like protein
Length = 384
Score = 431 bits (1108), Expect = e-121
Identities = 248/403 (61%), Positives = 283/403 (69%), Gaps = 32/403 (7%)
Query: 3 ADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDRAI 62
A DEP TRWSF GRKKL E VSN + S+ ++G V ++ +
Sbjct: 2 ATDEPSQTRWSF---------LFGRKKLPEED---VSNKDQPEDGSSNGNNGDVNNNSS- 48
Query: 63 YDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWE 122
+Q N NG N + EKPKKS+ PPFESAE RTLAESLSRDIIRG+PN+KWE
Sbjct: 49 --PVTNQDGNTALANG---NVIREKPKKSMFPPFESAETRTLAESLSRDIIRGNPNIKWE 103
Query: 123 SIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTT 182
SIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKAVATECNTT
Sbjct: 104 SIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKAVATECNTT 163
Query: 183 FFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR-GEGRSEHEASR 241
FFNISASS+VSKWRGDSEKL++VLF+LARHHAP+TIFLDEIDAIISQR GEGRSEHEASR
Sbjct: 164 FFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEGRSEHEASR 223
Query: 242 RLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR--NQKQEEQCLRNSFLC 299
RLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKR + + R F
Sbjct: 224 RLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPEARRGMFEM 283
Query: 300 SLMRN-------RCPMIYWWTGQKDTQARIFAYFVKKQQCSH*DA**HNLNKNQMWCPK- 351
+ ++ G + RI Q A L + P+
Sbjct: 284 LIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLA---ILEDREDVVPED 340
Query: 352 KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
+LPK+GP++PED++ AL NTRPSAHL AH YD FN DYGSQIL
Sbjct: 341 ELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQIL 383
>At1g80350 CAD ATPase (AAA1)
Length = 523
Score = 237 bits (605), Expect = 7e-63
Identities = 124/270 (45%), Positives = 177/270 (64%), Gaps = 16/270 (5%)
Query: 26 GRKKLVENGENAVSNGNSSGIASNGNSHGKVTSD----RAIYDQFQSQGQNPTHTNGFGP 81
G +K ++G A +G AS G G + R+ +
Sbjct: 143 GTRKSPQDGAWARGPTTRTGPASRGGRGGATSKSTAGARSSTAGKKGAASKSNKAESMNG 202
Query: 82 NGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMP 141
+ D K K+ L +E + LA L RD++ +P V+W+ + GL AKRLL+EAVV+P
Sbjct: 203 DAEDGKSKRGL---YEGPD-EDLAAMLERDVLDSTPGVRWDDVAGLSEAKRLLEEAVVLP 258
Query: 142 IKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEK 201
+ P+YF G+ PWKG+L+FGPPGTGKT+LAKAVATEC TTFFN+S++++ SKWRG+SE+
Sbjct: 259 LWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESER 318
Query: 202 LVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLART---- 257
+V+ LF+LAR +AP+TIF+DEID++ + RG G EHE+SRR+K+ELL+Q+DG++ T
Sbjct: 319 MVRCLFDLARAYAPSTIFIDEIDSLCNSRG-GSGEHESSRRVKSELLVQVDGVSNTATNE 377
Query: 258 ---DELVFVLAATNLPWELDAAMLRRLEKR 284
++V VLAATN PW++D A+ RRLEKR
Sbjct: 378 DGSRKIVMVLAATNFPWDIDEALRRRLEKR 407
>At2g27600 putative AAA-type ATPase
Length = 435
Score = 223 bits (569), Expect = 1e-58
Identities = 107/188 (56%), Positives = 142/188 (74%), Gaps = 1/188 (0%)
Query: 97 ESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 156
E E L L+ I+R PN+KW + GLE+AK+ L+EAV++P+K+P++FTG PW+
Sbjct: 107 EDPEQSKLRAGLNSAIVREKPNIKWSDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWR 166
Query: 157 GILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPA 216
LL+GPPGTGK+ LAKAVATE ++TFF++S+S +VSKW G+SEKLV LFE+AR AP+
Sbjct: 167 AFLLYGPPGTGKSYLAKAVATEADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPS 226
Query: 217 TIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAA 276
IF+DEID++ RGEG +E EASRR+KTELL+QM G+ DE V VLAATN P+ LD A
Sbjct: 227 IIFVDEIDSLCGTRGEG-NESEASRRIKTELLVQMQGVGHNDEKVLVLAATNTPYALDQA 285
Query: 277 MLRRLEKR 284
+ RR +KR
Sbjct: 286 IRRRFDKR 293
>At3g27120 spastin protein, putative
Length = 327
Score = 192 bits (487), Expect = 4e-49
Identities = 105/205 (51%), Positives = 137/205 (66%), Gaps = 9/205 (4%)
Query: 80 GPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVV 139
GP+G E P+K + E R L E +S +I+ PNV+W+ I GLE+AK+ + E V+
Sbjct: 16 GPDG--ELPEK-----LRNLEPR-LIEHVSNEIMDRDPNVRWDDIAGLEHAKKCVTEMVI 67
Query: 140 MPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDS 199
P+ P F G SP KG+LLFGPPGTGKTM+ KA+A E TFF ISASS+ SKW G+
Sbjct: 68 WPLLRPDIFKGCRSPGKGLLLFGPPGTGKTMIGKAIAGEAKATFFYISASSLTSKWIGEG 127
Query: 200 EKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDE 259
EKLV+ LF +A PA IF+DEID+++SQR + EHE+SRRLKT+ LI+M+G E
Sbjct: 128 EKLVRALFGVASCRQPAVIFVDEIDSLLSQR-KSDGEHESSRRLKTQFLIEMEGFDSGSE 186
Query: 260 LVFVLAATNLPWELDAAMLRRLEKR 284
+ ++ ATN P ELD A RRL KR
Sbjct: 187 QILLIGATNRPQELDEAARRRLTKR 211
>At4g02470
Length = 371
Score = 172 bits (436), Expect = 3e-43
Identities = 96/183 (52%), Positives = 123/183 (66%), Gaps = 5/183 (2%)
Query: 106 ESLSRDIIRGSP-NVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFG 162
+ L D+I S V ++ I LEN K LKE V++P++ P+ F L P KGILLFG
Sbjct: 52 KKLLSDVIPPSDIGVSFDDIGALENVKETLKELVMLPLQRPELFDKGQLTKPTKGILLFG 111
Query: 163 PPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDE 222
PPGTGKTMLAKAVATE F NIS SSI SKW G+ EK VK +F LA AP+ IF+DE
Sbjct: 112 PPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDE 171
Query: 223 IDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTD-ELVFVLAATNLPWELDAAMLRRL 281
+D+++ +R E EHEA R++K E ++ DGL D E V VLAATN P++LD A++RRL
Sbjct: 172 VDSMLGRR-ENPGEHEAMRKMKNEFMVNWDGLRTKDRERVLVLAATNRPFDLDEAVIRRL 230
Query: 282 EKR 284
+R
Sbjct: 231 PRR 233
>At1g50140 hypothetical protein, 5' partial
Length = 601
Score = 170 bits (431), Expect = 1e-42
Identities = 89/180 (49%), Positives = 122/180 (67%), Gaps = 4/180 (2%)
Query: 108 LSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPG 165
+S + G VK+E I LE+ K+ L E V++P++ P+ F LL P KGILLFGPPG
Sbjct: 298 VSAVVAPGEIGVKFEDIGALEDVKKALNELVILPMRRPELFARGNLLRPCKGILLFGPPG 357
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKA+ATE F +I+ S++ SKW GD+EKL K LF A AP IF+DEID+
Sbjct: 358 TGKTLLAKALATEAGANFISITGSTLTSKWFGDAEKLTKALFSFATKLAPVIIFVDEIDS 417
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGLARTD-ELVFVLAATNLPWELDAAMLRRLEKR 284
++ RG G SEHEA+RR++ E + DGL D + + +L ATN P++LD A++RRL +R
Sbjct: 418 LLGARG-GSSEHEATRRMRNEFMAAWDGLRSKDSQRILILGATNRPFDLDDAVIRRLPRR 476
>At3g19740 ATPase, putative
Length = 439
Score = 168 bits (426), Expect = 4e-42
Identities = 87/180 (48%), Positives = 122/180 (67%), Gaps = 4/180 (2%)
Query: 108 LSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPG 165
+S + G VK++ I LE+ K+ L E V++P++ P+ FT LL P KGILLFGPPG
Sbjct: 136 VSAVVAPGEIGVKFDDIGALEHVKKTLNELVILPMRRPELFTRGNLLRPCKGILLFGPPG 195
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKA+ATE F +I+ S++ SKW GD+EKL K LF A AP IF+DE+D+
Sbjct: 196 TGKTLLAKALATEAGANFISITGSTLTSKWFGDAEKLTKALFSFASKLAPVIIFVDEVDS 255
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGLARTD-ELVFVLAATNLPWELDAAMLRRLEKR 284
++ RG G EHEA+RR++ E + DGL D + + +L ATN P++LD A++RRL +R
Sbjct: 256 LLGARG-GAFEHEATRRMRNEFMAAWDGLRSKDSQRILILGATNRPFDLDDAVIRRLPRR 314
>At4g24850 unknown protein (At4g24850)
Length = 442
Score = 167 bits (424), Expect = 7e-42
Identities = 92/180 (51%), Positives = 123/180 (68%), Gaps = 4/180 (2%)
Query: 108 LSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPG 165
LS I+ +V ++ I LE K +LKE V++P++ P+ F L P KGILLFGPPG
Sbjct: 126 LSDVILPSDIDVTFDDIGALEKVKDILKELVMLPLQRPELFCKGELTKPCKGILLFGPPG 185
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKTMLAKAVA E + F NIS SSI SKW G+ EK VK +F LA +P+ IF+DE+D+
Sbjct: 186 TGKTMLAKAVAKEADANFINISMSSITSKWFGEGEKYVKAVFSLASKMSPSVIFVDEVDS 245
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGLARTD-ELVFVLAATNLPWELDAAMLRRLEKR 284
++ +R R EHEASR++K E ++ DGL + E V VLAATN P++LD A++RRL +R
Sbjct: 246 MLGRREHPR-EHEASRKIKNEFMMHWDGLTTQERERVLVLAATNRPFDLDEAVIRRLPRR 304
>At1g64110 unknown protein
Length = 824
Score = 164 bits (415), Expect = 8e-41
Identities = 92/199 (46%), Positives = 126/199 (63%), Gaps = 7/199 (3%)
Query: 88 PKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKY 147
PK + P E R E + + I NV ++ I L+ K L+E V++P++ P
Sbjct: 486 PKAPEVAPDNEFEKRIRPEVIPAEEI----NVTFKDIGALDEIKESLQELVMLPLRRPDL 541
Query: 148 FTG-LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVL 206
FTG LL P +GILLFGPPGTGKTMLAKA+A E +F N+S S+I SKW G+ EK V+ L
Sbjct: 542 FTGGLLKPCRGILLFGPPGTGKTMLAKAIAKEAGASFINVSMSTITSKWFGEDEKNVRAL 601
Query: 207 FELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLA 265
F LA +P IF+DE+D+++ QR EHEA R++K E + DGL + E + VLA
Sbjct: 602 FTLASKVSPTIIFVDEVDSMLGQRTR-VGEHEAMRKIKNEFMSHWDGLMTKPGERILVLA 660
Query: 266 ATNLPWELDAAMLRRLEKR 284
ATN P++LD A++RR E+R
Sbjct: 661 ATNRPFDLDEAIIRRFERR 679
>At1g02890 hypothetical protein
Length = 1240
Score = 159 bits (402), Expect = 3e-39
Identities = 100/204 (49%), Positives = 126/204 (61%), Gaps = 12/204 (5%)
Query: 85 DEKPKKSLLPPFESAEMRTLAESLSRDIIRGSP-NVKWESIKGLENAKRLLKEAVVMPIK 143
++ KKSL E + L D+I S V + I LEN K LKE V++P++
Sbjct: 907 NKSTKKSLKDVVTENEFE---KKLLSDVIPPSDIGVSFSDIGALENVKDTLKELVMLPLQ 963
Query: 144 YPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEK 201
P+ F L P KGILLFGPPGTGKTMLAKAVATE F NIS SSI SK EK
Sbjct: 964 RPELFGKGQLTKPTKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSK----GEK 1019
Query: 202 LVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTD-EL 260
VK +F LA AP+ IF+DE+D+++ +R E EHEA R++K E +I DGL D E
Sbjct: 1020 YVKAVFSLASKIAPSVIFVDEVDSMLGRR-ENPGEHEAMRKMKNEFMINWDGLRTKDKER 1078
Query: 261 VFVLAATNLPWELDAAMLRRLEKR 284
V VLAATN P++LD A++RRL +R
Sbjct: 1079 VLVLAATNRPFDLDEAVIRRLPRR 1102
>At4g28000 putative protein
Length = 726
Score = 157 bits (397), Expect = 1e-38
Identities = 101/257 (39%), Positives = 141/257 (54%), Gaps = 21/257 (8%)
Query: 30 LVENGENAVSNGNSSGIASNGNSHGKVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPK 89
L+ N E NG I+SN SHG +GQ + +D K
Sbjct: 340 LMNNKEPEYKNGRLV-ISSNSLSHGL---------NILQEGQGCFEDSLKLDTNIDSKE- 388
Query: 90 KSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT 149
+ P E R E + + I V + I L+ K L+E V++P++ P F
Sbjct: 389 ---VAPDNEFEKRIRPEVIPANEI----GVTFADIGSLDETKESLQELVMLPLRRPDLFK 441
Query: 150 G-LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFE 208
G LL P +GILLFGPPGTGKTM+AKA+A E +F N+S S+I SKW G+ EK V+ LF
Sbjct: 442 GGLLKPCRGILLFGPPGTGKTMMAKAIANEAGASFINVSMSTITSKWFGEDEKNVRALFT 501
Query: 209 LARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLAAT 267
LA +P IF+DE+D+++ QR EHEA R++K E + DGL + + + VLAAT
Sbjct: 502 LAAKVSPTIIFVDEVDSMLGQRTR-VGEHEAMRKIKNEFMTHWDGLMSNAGDRILVLAAT 560
Query: 268 NLPWELDAAMLRRLEKR 284
N P++LD A++RR E+R
Sbjct: 561 NRPFDLDEAIIRRFERR 577
>At2g45500 hypothetical protein
Length = 481
Score = 153 bits (387), Expect = 1e-37
Identities = 117/298 (39%), Positives = 155/298 (51%), Gaps = 50/298 (16%)
Query: 3 ADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNG-NSSGIASNGN-------SHG 54
A P P+ S E +K VR R+K + N +N VS + G+ + N S
Sbjct: 92 ATSTPSPSYISSSEKEK---VRSYREK-ISNWQNQVSERLQALGVGMSENKRTVAYPSSA 147
Query: 55 KVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRT--LAESLSRDI 112
V+S + Y + SQ + P G + S P ES + L E ++ I
Sbjct: 148 SVSSTASRYRKTLSQ-KTPVARGGVATPRNPKDAAASPKPVKESGNVYDDKLVEMINTTI 206
Query: 113 IRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK-----GILLFGPPGTG 167
+ SP+VKW+ + GL AK+ L E V++P K FTGL P + G+LLFGPPG G
Sbjct: 207 VDRSPSVKWDDVAGLNGAKQALLEMVILPAKRRDLFTGLRRPARVTSLLGLLLFGPPGNG 266
Query: 168 KTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAII 227
KTMLAKAVA+E TFFN+SASS+ SKW ID+I+
Sbjct: 267 KTMLAKAVASESQATFFNVSASSLTSKW---------------------------IDSIM 299
Query: 228 SQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLRRLEKR 284
S R SE+EASRRLK+E LIQ DG+ + D+LV ++ ATN P ELD A+LRRL KR
Sbjct: 300 STR--STSENEASRRLKSEFLIQFDGVTSNPDDLVIIIGATNKPQELDDAVLRRLVKR 355
>At4g27680 unknown protein
Length = 398
Score = 152 bits (383), Expect = 4e-37
Identities = 82/167 (49%), Positives = 111/167 (66%), Gaps = 5/167 (2%)
Query: 118 NVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPGTGKTMLAKAV 175
+V++ SI GLE K+ L E V++P+K P+ F LL P KG+LL+GPPGTGKTMLAKA+
Sbjct: 80 DVEFGSIGGLETIKQALYELVILPLKRPELFAYGKLLGPQKGVLLYGPPGTGKTMLAKAI 139
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRS 235
A E F N+ S+++SKW GD++KLV +F LA PA IF+DE+++ + QR +
Sbjct: 140 AKESGAVFINVRVSNLMSKWFGDAQKLVSAVFSLAYKLQPAIIFIDEVESFLGQRRS--T 197
Query: 236 EHEASRRLKTELLIQMDGLARTDEL-VFVLAATNLPWELDAAMLRRL 281
+HEA +KTE + DG + V VLAATN P ELD A+LRRL
Sbjct: 198 DHEAMANMKTEFMALWDGFSTDPHARVMVLAATNRPSELDEAILRRL 244
>At5g53540 26S proteasome regulatory particle chain RPT6-like
protein
Length = 403
Score = 150 bits (379), Expect = 1e-36
Identities = 83/167 (49%), Positives = 113/167 (66%), Gaps = 7/167 (4%)
Query: 118 NVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPGTGKTMLAKAV 175
+V++ SI GLE+ K+ L E V++P+K P+ F LL P KG+LL+GPPGTGKTMLAKA+
Sbjct: 83 DVEFGSIGGLESIKQALYELVILPLKRPELFAYGKLLGPQKGVLLYGPPGTGKTMLAKAI 142
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRS 235
A E F N+ S+++SKW GD++KLV +F LA PA IF+DE+D+ + QR +
Sbjct: 143 ARESEAVFINVKVSNLMSKWFGDAQKLVSAVFSLAYKLQPAIIFIDEVDSFLGQRRS--T 200
Query: 236 EHEASRRLKTELLIQMDGLARTDE--LVFVLAATNLPWELDAAMLRR 280
++EA +KTE + DG TD+ V VLAATN P ELD A+LRR
Sbjct: 201 DNEAMSNMKTEFMALWDGFT-TDQNARVMVLAATNRPSELDEAILRR 246
>At5g03340 transitional endoplasmic reticulum ATPase
Length = 810
Score = 147 bits (372), Expect = 8e-36
Identities = 75/175 (42%), Positives = 112/175 (63%), Gaps = 3/175 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QRG + A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR
Sbjct: 585 IATQRGNSAGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLR 638
Score = 117 bits (294), Expect = 8e-27
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 311 SIAPKR--EKTNGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 362
>At3g53230 CDC48 - like protein
Length = 815
Score = 146 bits (369), Expect = 2e-35
Identities = 76/175 (43%), Positives = 111/175 (63%), Gaps = 3/175 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 466 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 525
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F +I +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 526 CGKTLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 585
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QRG + A+ R+ +LL +MDG+ + VF++ ATN P +D A+LR
Sbjct: 586 IATQRGNSVGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDPALLR 639
Score = 118 bits (295), Expect = 6e-27
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 192 EPIKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 251
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 252 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 311
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 312 SIAPKR--EKTHGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 363
>At3g09840 putative transitional endoplasmic reticulum ATPase
Length = 809
Score = 145 bits (367), Expect = 3e-35
Identities = 75/176 (42%), Positives = 111/176 (62%), Gaps = 4/176 (2%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV W I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 226 IISQR--GEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QR G G A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR
Sbjct: 585 IATQRGGGSGGDGGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLR 639
Score = 118 bits (295), Expect = 6e-27
Identities = 67/175 (38%), Positives = 108/175 (61%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ +V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDDVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 311 SIAPKR--EKTNGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 362
>At1g03000 putative peroxisome assembly factor-2
Length = 941
Score = 135 bits (341), Expect = 3e-32
Identities = 64/163 (39%), Positives = 104/163 (63%)
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVA 176
PNVKW+ + GLE+ K + + V +P+ + F+ L G+LL+GPPGTGKT+LAKAVA
Sbjct: 653 PNVKWDDVGGLEDVKTSILDTVQLPLLHKDLFSSGLRKRSGVLLYGPPGTGKTLLAKAVA 712
Query: 177 TECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSE 236
TEC+ F ++ +++ + G+SEK V+ +FE AR P IF DE+D++ RG
Sbjct: 713 TECSLNFLSVKGPELINMYIGESEKNVRDIFEKARSARPCVIFFDELDSLAPARGASGDS 772
Query: 237 HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
R+ +++L ++DGL+ + + +F++ A+N P +D A+LR
Sbjct: 773 GGVMDRVVSQMLAEIDGLSDSSQDLFIIGASNRPDLIDPALLR 815
Score = 33.9 bits (76), Expect = 0.16
Identities = 18/67 (26%), Positives = 29/67 (42%)
Query: 158 ILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPAT 217
+LL G PG GK + K VA S S+++ + + F +AR ++P
Sbjct: 380 VLLHGIPGCGKRTVVKYVARRLGLHVVEFSCHSLLASSERKTSTALAQTFNMARRYSPTI 439
Query: 218 IFLDEID 224
+ L D
Sbjct: 440 LLLRHFD 446
>At3g56690 calmodulin-binding protein
Length = 1022
Score = 134 bits (338), Expect = 7e-32
Identities = 77/185 (41%), Positives = 112/185 (59%), Gaps = 3/185 (1%)
Query: 96 FESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSP 154
FE+A+ + + S R++I P V WE + G K L EAV P K+ F + P
Sbjct: 699 FENAKTK-IRPSAMREVILEVPKVNWEDVGGQNEVKNQLMEAVEWPQKHQDAFKRIGTRP 757
Query: 155 WKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHA 214
GIL+FGPPG KT++A+AVA+E F + + SKW G+SEK V+ LF AR +A
Sbjct: 758 PSGILMFGPPGCSKTLMARAVASEAKLNFLAVKGPELFSKWVGESEKAVRSLFAKARANA 817
Query: 215 PATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELD 274
P+ IF DEID++ S RG+ S R+ ++LL+++DGL + V V+AATN P ++D
Sbjct: 818 PSIIFFDEIDSLASIRGKENDGVSVSDRVMSQLLVELDGLHQRVG-VTVIAATNRPDKID 876
Query: 275 AAMLR 279
+A+LR
Sbjct: 877 SALLR 881
Score = 105 bits (261), Expect = 6e-23
Identities = 56/128 (43%), Positives = 82/128 (63%), Gaps = 3/128 (2%)
Query: 152 LSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELAR 211
L P KG+L+ GPPGTGKT LA+ A FF+++ I+S++ G+SEK + +F A
Sbjct: 415 LRPTKGVLIHGPPGTGKTSLARTFARHSGVNFFSVNGPEIISQYLGESEKALDEVFRSAS 474
Query: 212 HHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPW 271
+ PA +F+D++DAI R EG E S+R+ LL MDG++RTD +V V+AATN P
Sbjct: 475 NATPAVVFIDDLDAIAPARKEG--GEELSQRMVATLLNLMDGISRTDGVV-VIAATNRPD 531
Query: 272 ELDAAMLR 279
++ A+ R
Sbjct: 532 SIEPALRR 539
>At2g03670 putative AAA-type ATPase
Length = 603
Score = 129 bits (323), Expect = 4e-30
Identities = 71/176 (40%), Positives = 110/176 (62%), Gaps = 4/176 (2%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S++R I P V W+ + GL++ K+ L++AV PIK+ F + +SP +GILL GPPG
Sbjct: 271 SINRGITVEIPKVTWDDVGGLKDLKKKLQQAVEWPIKHSAAFVKMGISPMRGILLHGPPG 330
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
KT LAKA A +FF++S + + S + G+ E L++ F+ AR +P+ IF DE D
Sbjct: 331 CSKTTLAKAAANAAQASFFSLSCAELFSMYVGEGEALLRNTFQRARLASPSIIFFDEADV 390
Query: 226 IISQRGEGRSEHEAS--RRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+ +RG+ S + ++ RL + LL +MDGL + + VLAATN P+ +DAA++R
Sbjct: 391 VACKRGDESSSNSSTVGERLLSTLLTEMDGLEEA-KGILVLAATNRPYAIDAALMR 445
Score = 93.6 bits (231), Expect = 2e-19
Identities = 67/184 (36%), Positives = 96/184 (51%), Gaps = 15/184 (8%)
Query: 106 ESLSRDIIRGSPNVKWES---IKGLENAKRLLKEAVVMPIKYPKYFTGLLSPW-KGILLF 161
ES D I G N KW + I G E A + L+E ++ P +YP L W +G+LL+
Sbjct: 5 ESSVCDNIAG--NEKWRAEAEIGGNERALQALRELIIFPFRYPLEARTLGLKWPRGLLLY 62
Query: 162 GPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHA----PAT 217
GPPGTGKT L +AV EC+ +S S+ G+SEK+++ F A HA P+
Sbjct: 63 GPPGTGKTSLVRAVVQECDAHLIVLSPHSVHRAHAGESEKVLREAFAEASSHAVSDKPSV 122
Query: 218 IFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDEL--VFVLAATNLPWELDA 275
IF+DEID + +R R E R+ ++L MD + V V+A+TN +D
Sbjct: 123 IFIDEIDVLCPRRDARR---EQDVRIASQLFTLMDSNKPSSSAPRVVVVASTNRVDAIDP 179
Query: 276 AMLR 279
A+ R
Sbjct: 180 ALRR 183
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,043,916
Number of Sequences: 26719
Number of extensions: 380661
Number of successful extensions: 1493
Number of sequences better than 10.0: 151
Number of HSP's better than 10.0 without gapping: 129
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1217
Number of HSP's gapped (non-prelim): 178
length of query: 394
length of database: 11,318,596
effective HSP length: 101
effective length of query: 293
effective length of database: 8,619,977
effective search space: 2525653261
effective search space used: 2525653261
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146651.7


