

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146649.7 - phase: 0 /pseudo
(859 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
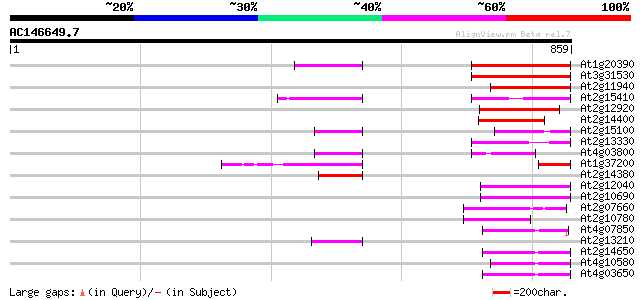
Sequences producing significant alignments: (bits) Value
At1g20390 hypothetical protein 140 3e-33
At3g31530 hypothetical protein 122 9e-28
At2g11940 putative retroelement gag/pol polyprotein 107 2e-23
At2g15410 putative retroelement pol polyprotein 100 5e-21
At2g12920 pseudogene 98 2e-20
At2g14400 putative retroelement pol polyprotein 92 1e-18
At2g15100 putative retroelement pol polyprotein 79 8e-15
At2g13330 F14O4.9 76 7e-14
At4g03800 74 3e-13
At1g37200 hypothetical protein 73 8e-13
At2g14380 putative retroelement pol polyprotein 61 2e-09
At2g12040 T10J7.2 60 7e-09
At2g10690 putative retroelement pol polyprotein 56 8e-08
At2g07660 putative retroelement pol polyprotein 56 1e-07
At2g10780 pseudogene 55 1e-07
At4g07850 putative polyprotein 55 2e-07
At2g13210 pseudogene 52 2e-06
At2g14650 putative retroelement pol polyprotein 50 4e-06
At4g10580 putative reverse-transcriptase -like protein 50 5e-06
At4g03650 putative reverse transcriptase 50 7e-06
>At1g20390 hypothetical protein
Length = 1791
Score = 140 bits (353), Expect = 3e-33
Identities = 70/152 (46%), Positives = 102/152 (67%)
Query: 707 RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEK 766
+Q+ +I P + +Q L G + +L RF+++S LPF+NLLK+ +F+W + E+
Sbjct: 991 KQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEE 1050
Query: 767 ALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGP 826
A + LK+ LS PP+L +P+ GETLYLY++VS AVS L E + Q+P ++TSK+L+
Sbjct: 1051 AFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEA 1110
Query: 827 ETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
ETRY IEK ALA+VT+AR YF +HTIAV
Sbjct: 1111 ETRYPVIEKAALAVVTSARKLRPYFQSHTIAV 1142
Score = 80.9 bits (198), Expect = 3e-15
Identities = 43/103 (41%), Positives = 59/103 (56%)
Query: 437 DPTKCFKLGKGIPEHARAHLIACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVA 496
DPT+C +G I R LIA L N+ FA SI DM IDP++ H+L V P+ V
Sbjct: 720 DPTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKPVK 779
Query: 497 QRRKKQFPEKAEGTEKFVKDLAEEKFIAEAKYTTWLSNILLVK 539
Q+R+K PE+A + V+ L + I E KY WL+N ++VK
Sbjct: 780 QKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVK 822
>At3g31530 hypothetical protein
Length = 831
Score = 122 bits (306), Expect = 9e-28
Identities = 60/152 (39%), Positives = 97/152 (63%)
Query: 707 RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEK 766
+Q+ ++ G K+ +Q L G + +L RF+++S + L F+++L++ KFEWT CE+
Sbjct: 198 KQIEALLGMASPQNKREVQCLTGRVAALNRFISRSTEKCLAFYDVLRRNKKFEWTTRCEE 257
Query: 767 ALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGP 826
A Q LK L+ PP+L++P GE YLY++VS AVSG L E + QK ++ S+
Sbjct: 258 AFQELKKYLATPPILAKPVIGEPQYLYVAVSDTAVSGELVREDRGEQKLIFYVSQTFTSA 317
Query: 827 ETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
E+RY ++EK+ALA+V +A+ YF +H+I V
Sbjct: 318 ESRYPQMEKLALAVVMSAQKLRPYFQSHSIIV 349
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 107 bits (268), Expect = 2e-23
Identities = 56/123 (45%), Positives = 79/123 (63%), Gaps = 1/123 (0%)
Query: 737 FVAKSAQHALPFFNLLKKETK-FEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLS 795
F+++S LPF LL+K K F W CE+A + LK+ LS P VL++P+ GE L+LY+S
Sbjct: 564 FISRSTDKCLPFCTLLRKVNKGFVWDERCEEAFRQLKSYLSEPLVLAKPEFGERLFLYIS 623
Query: 796 VSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFMAHT 855
VS A+SG L + ++P ++ SK+ ETRY +EK+ALA+VTAAR YF +H
Sbjct: 624 VSENAISGVLVRVERSDERPIFYVSKSFSDVETRYPMMEKLALAVVTAARKLRPYFQSHP 683
Query: 856 IAV 858
I V
Sbjct: 684 IVV 686
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/74 (35%), Positives = 38/74 (51%), Gaps = 1/74 (1%)
Query: 450 EHARAHLIACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEG 509
++ R LI L N FA +I +M I V H+L V P+ V Q+R+K P++AE
Sbjct: 358 QNFRRELIQFLEKNKSTFAWTIREMTGISTEVISHELNVDPTFKPVKQKRQKHGPDRAEA 417
Query: 510 TE-KFVKDLAEEKF 522
K V+ L + F
Sbjct: 418 VNVKVVRLLKADCF 431
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 100 bits (248), Expect = 5e-21
Identities = 56/152 (36%), Positives = 83/152 (53%), Gaps = 20/152 (13%)
Query: 707 RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEK 766
+Q+ +I P + +Q L G + +L RF+++S LPF+ LL+ +FEW +CE
Sbjct: 996 KQISAIIDLPSPRNTREVQRLIGRIAALNRFISRSTDKCLPFYQLLRANKRFEWDEKCE- 1054
Query: 767 ALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGP 826
EGETLYLY++VS+ AVSG L E + Q P ++ SK L G
Sbjct: 1055 -------------------EGETLYLYIAVSTSAVSGVLVREDRGEQHPIFYVSKTLDGA 1095
Query: 827 ETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
E RY +EK+A A+V +AR YF ++T+ V
Sbjct: 1096 ELRYPTLEKLAFAVVISARKLRPYFKSYTVEV 1127
Score = 74.7 bits (182), Expect = 2e-13
Identities = 41/133 (30%), Positives = 69/133 (51%), Gaps = 4/133 (3%)
Query: 410 LHKVIVERAEEGKQGPTQRGNTSPFG---DDPTKCFKLGKGIPEHARAHLIACLWLNTDL 466
++ VI+ ++ + P Q+G DP++C + +P + L+ L N
Sbjct: 696 IYNVILVPTQDERPNP-QKGTVVQVNIDESDPSRCVGIRIDLPSELQNELVNFLRQNAAT 754
Query: 467 FARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDLAEEKFIAEA 526
FA S+ DMP ID +V CH+L V P+ + Q+R+K P++ + + VK L + I E
Sbjct: 755 FAWSVEDMPGIDSAVTCHELNVDPTYKPLKQKRRKLGPDRTKDVNEEVKKLLDAGSIVEV 814
Query: 527 KYTTWLSNILLVK 539
+Y WL N ++VK
Sbjct: 815 RYPDWLRNPVVVK 827
>At2g12920 pseudogene
Length = 863
Score = 98.2 bits (243), Expect = 2e-20
Identities = 48/122 (39%), Positives = 77/122 (62%)
Query: 720 KKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPP 779
K++ +N + +L RF ++S F+++L+ KFEWT +CE+A Q LK L++PP
Sbjct: 372 KQRLSTAINRRVAALNRFFSRSTNKCWAFYDVLRGNKKFEWTTQCEEAFQELKKYLASPP 431
Query: 780 VLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALA 839
+L++ E LYLY+SVS VS L E + QKP ++ S+ G E+RY ++EK+ALA
Sbjct: 432 ILAKLVIREPLYLYVSVSDTTVSKVLVREDRGEQKPIFYVSQTFTGAESRYPQMEKLALA 491
Query: 840 LV 841
++
Sbjct: 492 VI 493
Score = 33.1 bits (74), Expect = 0.68
Identities = 22/63 (34%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Query: 478 DPSVACHQLTVSPSASVVAQRRKKQFP-EKAEGTEKFVKDLAEEKFIAEAKYTTWLSNIL 536
DPS Q S S+++ ++ KK+ E+AE + V+ L I EAKY L+N +
Sbjct: 189 DPSKQTPQADKSQSSTLSQKKEKKKIGWERAEAVKAEVEKLLRNGSITEAKYPDRLANPV 248
Query: 537 LVK 539
+VK
Sbjct: 249 VVK 251
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 92.0 bits (227), Expect = 1e-18
Identities = 47/102 (46%), Positives = 67/102 (65%), Gaps = 1/102 (0%)
Query: 718 SPKK-KCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLS 776
SPK K +Q L G + +L RF+++S +LPF+ +LK +F W +CE+A LK L+
Sbjct: 688 SPKNFKEVQRLTGRIAALNRFISRSTDKSLPFYQILKGNKEFLWDEKCEEAFGQLKAYLT 747
Query: 777 APPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYF 818
+PPVL +P+ E LYLY+SVS+ AVSG L E + QKP Y+
Sbjct: 748 SPPVLFKPELDEKLYLYVSVSNHAVSGVLIREDRGEQKPIYY 789
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 79.3 bits (194), Expect = 8e-15
Identities = 45/117 (38%), Positives = 67/117 (56%), Gaps = 8/117 (6%)
Query: 743 QHALPFFNLLKKETK-FEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAV 801
+ PF + K +K F W CE+A + LK LS P VL++ + GE L+LY++VS AV
Sbjct: 613 RQVFPFLHFTAKISKGFIWNESCEEAFKQLKRYLSEPSVLAKREFGEQLFLYIAVSESAV 672
Query: 802 SGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
+G + ++P ++ +TRY +EK+ALA+VTAAR YF +H I V
Sbjct: 673 TGVQVRVERSDKRPIFYV-------KTRYPMMEKLALAVVTAARKLRPYFQSHPIVV 722
Score = 47.0 bits (110), Expect = 5e-05
Identities = 26/73 (35%), Positives = 37/73 (50%)
Query: 467 FARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDLAEEKFIAEA 526
F + M I V H+L V P+ V Q+R+K P++A+ V L E I E
Sbjct: 405 FPTPVGKMTGISTEVISHELNVDPTFKPVKQKRRKLGPDRAQAVNIEVVRLLEVGRIREV 464
Query: 527 KYTTWLSNILLVK 539
KY WL+N ++VK
Sbjct: 465 KYPEWLANPVVVK 477
>At2g13330 F14O4.9
Length = 889
Score = 76.3 bits (186), Expect = 7e-14
Identities = 43/152 (28%), Positives = 74/152 (48%), Gaps = 30/152 (19%)
Query: 707 RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEK 766
+Q+ + P + +Q L G + +L F+++SA +PF+ L+K +F+W +CE+
Sbjct: 457 KQIAAFIEMPSPKMAREVQRLTGRIPALNGFISRSANKCVPFYQPLRKGKEFDWNKDCEQ 516
Query: 767 ALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGP 826
A + LK L+ PP+L++P++GE LYLY +
Sbjct: 517 AFKQLKAYLTEPPILAKPEKGEPLYLYTN------------------------------R 546
Query: 827 ETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
+ R +EK+AL +V + R YF +H I V
Sbjct: 547 DKRCPAMEKLALTVVMSVRKLRLYFQSHPIVV 578
>At4g03800
Length = 637
Score = 73.9 bits (180), Expect = 3e-13
Identities = 40/98 (40%), Positives = 56/98 (56%), Gaps = 6/98 (6%)
Query: 708 QMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKA 767
Q+ + TP K +Q L G R ++S +LPF+ +LK F W +CE+A
Sbjct: 541 QINAFLKTPSLRNFKEVQRLTG------RIASRSTDKSLPFYQILKGNNGFLWDEKCEEA 594
Query: 768 LQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTL 805
+ LK L+ PPVLS+P+ E LYLY+ VS+ AVSG L
Sbjct: 595 FRQLKAYLTTPPVLSKPEADEKLYLYVFVSNHAVSGVL 632
Score = 52.0 bits (123), Expect = 1e-06
Identities = 25/72 (34%), Positives = 41/72 (56%)
Query: 468 ARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDLAEEKFIAEAK 527
+R+ MPDIDPS+ CH+L V P + Q+R+K E+A+ + L + I E +
Sbjct: 315 SRTSAYMPDIDPSIICHELNVDPRFKPLKQKRRKLGVERAKAVNGKIDKLLKIGSIREVQ 374
Query: 528 YTTWLSNILLVK 539
Y W++ ++VK
Sbjct: 375 YPDWVAITVVVK 386
Score = 32.0 bits (71), Expect = 1.5
Identities = 18/54 (33%), Positives = 29/54 (53%), Gaps = 1/54 (1%)
Query: 314 IPFLVIEVTFLYNCIIGMTGLVQLGAVC*TAHLKLKYHADKNIITTLYGDFEAA 367
+ F+V + YN I+G + Q+ AV T H LK+ N + T++GD E +
Sbjct: 263 VDFVVFDKPATYNIILGTPWIFQMKAVPSTYHQWLKF-PTSNGVETIWGDLEGS 315
>At1g37200 hypothetical protein
Length = 1564
Score = 72.8 bits (177), Expect = 8e-13
Identities = 60/218 (27%), Positives = 99/218 (44%), Gaps = 20/218 (9%)
Query: 325 YNCIIGMTGLVQLGAVC*TAHLKLKYHADKNIITTLYGDFEAA*RCFLQGE*TPELGFPL 384
+N I+G L + AV T H +K+ ++K I +YG ++ RC++ EL
Sbjct: 532 FNAILGRPWLHAMKAVPSTYHQCIKFPSEKGI-AVIYGSQRSSRRCYMGSY---ELIKKA 587
Query: 385 KVVCYG*GENCYEYSGVQPDRSRSTLHKVIVERAEEGKQGPTQRGNTSPFGD--DPTKCF 442
++V + E V+ ++ + GP + T + D DP +C
Sbjct: 588 ELVVLMIEDKLAEMKTVRS--------------SDPSQCGPQKSSITQVYIDESDPKQCV 633
Query: 443 KLGKGIPEHARAHLIACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQ 502
+G+ + R I L N D FA S ++ I V H+L V P+ + Q+R+K
Sbjct: 634 GVGQDLDPAIREDFITFLKENKDSFAWSSANLQGISLEVTSHELNVDPTYRPIKQKRRKL 693
Query: 503 FPEKAEGTEKFVKDLAEEKFIAEAKYTTWLSNILLVKN 540
PE+A+ + V L + I E KY WL+N ++VKN
Sbjct: 694 GPERAKAVQDEVDRLLKIGSIREVKYPDWLANPVVVKN 731
Score = 51.2 bits (121), Expect = 2e-06
Identities = 24/48 (50%), Positives = 33/48 (68%)
Query: 811 EGQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFMAHTIAV 858
EGQKP Y+ SK L+ ET Y+ +EK+ALA+V +AR +Y +H I V
Sbjct: 929 EGQKPIYYVSKTLIDAETHYRAMEKLALAVVMSARKLRQYLQSHPIVV 976
Score = 29.6 bits (65), Expect = 7.5
Identities = 12/36 (33%), Positives = 19/36 (52%)
Query: 76 KAHSQEKRRVDKTKYCRFYKCHGHVTDHCIHLKDAI 111
+A E +D KYC+++ GH T+ C +K I
Sbjct: 309 EAAEDESPELDLGKYCKYHNKRGHSTEECRAVKKLI 344
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 61.2 bits (147), Expect = 2e-09
Identities = 29/67 (43%), Positives = 41/67 (60%)
Query: 473 DMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDLAEEKFIAEAKYTTWL 532
DM IDP VACH+L V P+ +V Q+R+K PE+++ V L + I E KY WL
Sbjct: 476 DMVGIDPEVACHELNVDPTFKLVKQKRRKLGPERSKAVNDEVDKLLDAGSIVEVKYPEWL 535
Query: 533 SNILLVK 539
+N ++VK
Sbjct: 536 ANPVVVK 542
Score = 37.4 bits (85), Expect = 0.036
Identities = 18/51 (35%), Positives = 28/51 (54%)
Query: 707 RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETK 757
+Q+ +I KK +Q L G + L RF+A+S +LPF+ LL+ K
Sbjct: 711 KQIRAILDLQSPRNKKEVQRLTGRIAGLNRFIARSTDKSLPFYQLLRSANK 761
>At2g12040 T10J7.2
Length = 976
Score = 59.7 bits (143), Expect = 7e-09
Identities = 39/137 (28%), Positives = 60/137 (43%)
Query: 722 KCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVL 781
K +++ G RF+ ++ A P +LL KE KFE+T EC A Q +K +L + P++
Sbjct: 173 KAVRSFLGHAGFYRRFIKDFSKIARPLTSLLCKEVKFEFTQECHDAFQQIKQALISAPIV 232
Query: 782 SRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALV 841
PD + S AV L + Y+ S+ L + Y EK LA+V
Sbjct: 233 QPPDWDLPFEVMCDASDFAVGAVLGQRKDKKLHAIYYASRTLDDAKRNYATTEKEFLAVV 292
Query: 842 TAARTH*KYFMAHTIAV 858
A Y + + V
Sbjct: 293 FAFEKFRSYLVGLKVIV 309
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 56.2 bits (134), Expect = 8e-08
Identities = 37/137 (27%), Positives = 61/137 (44%)
Query: 722 KCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVL 781
K I++ G RF+ ++ A P LL KET+F + +C KA +K +L + P+
Sbjct: 282 KDIRSFLGHAEFYRRFIKDFSKIARPLTRLLCKETEFNFDEDCLKAFHLIKETLVSAPIF 341
Query: 782 SRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALV 841
P+ + AV L+ + + Y+ S+ + +TRY IEK LA+V
Sbjct: 342 QAPNWDHPFEIMCDAFDYAVGAVLDQKIDDKLHVIYYASRTMDEAQTRYATIEKELLAVV 401
Query: 842 TAARTH*KYFMAHTIAV 858
A Y + + V
Sbjct: 402 FAFEKIRSYLVGFKVKV 418
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 55.8 bits (133), Expect = 1e-07
Identities = 50/161 (31%), Positives = 70/161 (43%), Gaps = 7/161 (4%)
Query: 695 FLSNKIWD*GQS---RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNL 751
FL + I D G S ++ SI P I++ G+ RFV A A P L
Sbjct: 352 FLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRL 411
Query: 752 LKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQE 811
K+T F W++ECEK+ LK L PVL P+EGE +Y S + G + Q+
Sbjct: 412 TGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDAS---IVGLGCVLMQK 468
Query: 812 GQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFM 852
G Y S+ L E Y + A+V A + Y +
Sbjct: 469 GSVIAY-ASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLI 508
>At2g10780 pseudogene
Length = 1611
Score = 55.5 bits (132), Expect = 1e-07
Identities = 38/106 (35%), Positives = 51/106 (47%), Gaps = 3/106 (2%)
Query: 695 FLSNKIWD*GQS---RQMPSIYGTPYSPKKKCIQTLNGMLTSLFRFVAKSAQHALPFFNL 751
FL + I D G S ++ SI P I++ G+ RFV A A P L
Sbjct: 857 FLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRL 916
Query: 752 LKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSVS 797
K+T F W++ECEK+ LK L+ PVL P+EGE +Y S
Sbjct: 917 TGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDAS 962
>At4g07850 putative polyprotein
Length = 1138
Score = 55.1 bits (131), Expect = 2e-07
Identities = 40/137 (29%), Positives = 60/137 (43%), Gaps = 9/137 (6%)
Query: 724 IQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSR 783
+++ +G+ RFV + A P ++KK F+W E A Q LK L+ PVLS
Sbjct: 559 VRSFHGLAGFYRRFVRDFSTLAAPLTEVIKKNVGFKWEQAPEDAFQALKEKLTHAPVLSL 618
Query: 784 PDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTA 843
PD +T + + L + +KP + S+ L G Y +K ALV A
Sbjct: 619 PDFLKTFEIECDAPGVGIGAVL----MQDKKPIAYFSEKLGGATLNYPTYDKELYALVRA 674
Query: 844 ARTH*KY-----FMAHT 855
+T Y F+ HT
Sbjct: 675 LQTWQHYLWPKEFVIHT 691
>At2g13210 pseudogene
Length = 152
Score = 51.6 bits (122), Expect = 2e-06
Identities = 26/78 (33%), Positives = 42/78 (53%)
Query: 462 LNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDLAEEK 521
++ F S D+P + + H+L V P+ V Q+R+K E+AE + V+ L
Sbjct: 1 MDVSTFVWSPEDLPGVSVEIVAHELNVDPTFKPVKQKRRKLGRERAEAVKAEVEKLLRIG 60
Query: 522 FIAEAKYTTWLSNILLVK 539
I +AKY WL+N ++VK
Sbjct: 61 SITKAKYPEWLANPVVVK 78
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 50.4 bits (119), Expect = 4e-06
Identities = 38/135 (28%), Positives = 57/135 (42%), Gaps = 4/135 (2%)
Query: 724 IQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSR 783
I++ G+ RFV A A P L K+ F W+ ECE+ LK L++ PVL+
Sbjct: 696 IRSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLAL 755
Query: 784 PDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTA 843
P+ GE +Y S + L Q G K + S+ L E Y + A++ A
Sbjct: 756 PEHGEPYMVYTDASGVGLGCVL---MQRG-KVIAYASRQLRKHEGNYPTHDLEMAAVIFA 811
Query: 844 ARTH*KYFMAHTIAV 858
+ Y + V
Sbjct: 812 LKIWRSYLYGGNVQV 826
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 50.1 bits (118), Expect = 5e-06
Identities = 35/123 (28%), Positives = 53/123 (42%), Gaps = 4/123 (3%)
Query: 736 RFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLS 795
RFV A A P L K+ F W+ ECE+ LK L++ PVL+ P+ G+ +Y
Sbjct: 733 RFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTD 792
Query: 796 VSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFMAHT 855
S + L Q G K + S+ L+ E Y + A++ A + Y
Sbjct: 793 ASRVGLGCVL---MQHG-KVIAYASRQLMKHEGNYPTHDLEMAAVIFALKIWRSYLYGGK 848
Query: 856 IAV 858
+ V
Sbjct: 849 VQV 851
>At4g03650 putative reverse transcriptase
Length = 839
Score = 49.7 bits (117), Expect = 7e-06
Identities = 37/135 (27%), Positives = 57/135 (41%), Gaps = 4/135 (2%)
Query: 724 IQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSR 783
I++ G+ RF+ A A P L K+ F W+ ECE+ LK L++ PVL+
Sbjct: 646 IRSFLGLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLAL 705
Query: 784 PDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTA 843
P+ GE +Y S + L Q G K + S+ L E Y + A++ A
Sbjct: 706 PEHGEPYMVYTDASGVGLGCAL---MQRG-KVIAYASRQLRKHEGNYPTHDLEMAAVIFA 761
Query: 844 ARTH*KYFMAHTIAV 858
+ Y + V
Sbjct: 762 LKIWRSYLYGGKVQV 776
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.335 0.146 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,379,029
Number of Sequences: 26719
Number of extensions: 757530
Number of successful extensions: 2719
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 2640
Number of HSP's gapped (non-prelim): 81
length of query: 859
length of database: 11,318,596
effective HSP length: 108
effective length of query: 751
effective length of database: 8,432,944
effective search space: 6333140944
effective search space used: 6333140944
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146649.7


