

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
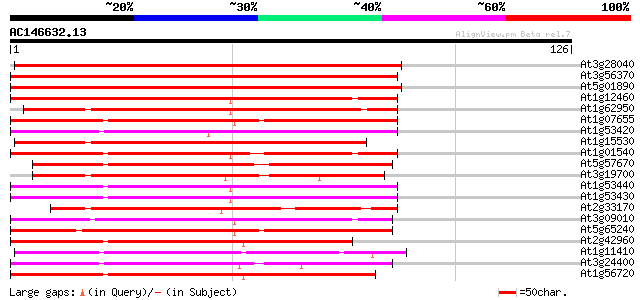
Sequences producing significant alignments: (bits) Value
At3g28040 unknown protein 145 5e-36
At3g56370 putative protein 115 8e-27
At5g01890 unknown protein 113 3e-26
At1g12460 unknown protein 94 2e-20
At1g62950 protein kinase like protein 89 6e-19
At1g07655 unknown protein 68 1e-12
At1g53420 hypothetical protein, 5' partial 67 3e-12
At1g15530 hypothetical protein 66 4e-12
At1g01540 unknown protein 63 3e-11
At5g57670 putative protein 62 6e-11
At3g19700 receptor-like protein kinase, putative 61 1e-10
At1g53440 receptor-like serine/threonine kinase, putative 61 2e-10
At1g53430 receptor-like serine/threonine kinase, putative 60 2e-10
At2g33170 putative receptor-like protein kinase 60 3e-10
At3g09010 putative receptor ser/thr protein kinase 60 4e-10
At5g65240 receptor-like protein kinase 59 5e-10
At2g42960 putative protein kinase 59 5e-10
At1g11410 putative brassinosteroid insensitive protein; similar ... 59 6e-10
At3g24400 protein kinase, putative 59 8e-10
At1g56720 protein kinase, putative 59 8e-10
>At3g28040 unknown protein
Length = 997
Score = 145 bits (366), Expect = 5e-36
Identities = 66/87 (75%), Positives = 78/87 (88%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
+NRFQ+ALGYVAPEL CQ LRVNEKCDVYGFGV+ILE+VTG+RPVEYGED+ +IL+DHVR
Sbjct: 870 NNRFQNALGYVAPELECQNLRVNEKCDVYGFGVLILELVTGRRPVEYGEDSFVILSDHVR 929
Query: 62 VLLEHGNALECVDPSLMSEYPEDEVLP 88
V+LE GN LEC+DP + +Y EDEVLP
Sbjct: 930 VMLEQGNVLECIDPVMEEQYSEDEVLP 956
>At3g56370 putative protein
Length = 964
Score = 115 bits (287), Expect = 8e-27
Identities = 50/87 (57%), Positives = 70/87 (79%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LE+VTGK+PVEY ED+V++L D V
Sbjct: 835 LSSKIQSALGYMAPEFACRTVKITEKCDVYGFGVLVLEVVTGKKPVEYMEDDVVVLCDMV 894
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVL 87
R LE G A EC+DP L ++P +E +
Sbjct: 895 REALEDGRADECIDPRLQGKFPVEEAV 921
>At5g01890 unknown protein
Length = 967
Score = 113 bits (282), Expect = 3e-26
Identities = 48/88 (54%), Positives = 67/88 (75%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S + QSALGY APE AC+ +++ ++CDVYGFG+++LE+VTGKRPVEY ED+V++L + V
Sbjct: 843 LSGKVQSALGYTAPEFACRTVKITDRCDVYGFGILVLEVVTGKRPVEYAEDDVVVLCETV 902
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R LE G ECVDP L +P +E +P
Sbjct: 903 REGLEEGRVEECVDPRLRGNFPAEEAIP 930
>At1g12460 unknown protein
Length = 882
Score = 94.0 bits (232), Expect = 2e-20
Identities = 47/88 (53%), Positives = 68/88 (76%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
++ +F +A+GY+APELA Q LR +EKCDVY +GV++LE+VTG++PVE E+ VLIL D+
Sbjct: 760 LTKKFHNAVGYIAPELAQQSLRASEKCDVYSYGVVLLELVTGRKPVESPSENQVLILRDY 819
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VR LLE G+A +C D L E+ E+E++
Sbjct: 820 VRDLLETGSASDCFDRRL-REFEENELI 846
>At1g62950 protein kinase like protein
Length = 890
Score = 89.0 bits (219), Expect = 6e-19
Identities = 46/85 (54%), Positives = 66/85 (77%), Gaps = 3/85 (3%)
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDHVRV 62
+F +A+GY+APELA Q LRV++KCDVY +GV++LE+VTG++PVE E+ V+IL DHVR
Sbjct: 772 KFHNAVGYIAPELA-QSLRVSDKCDVYSYGVVLLELVTGRKPVESPSENEVVILRDHVRN 830
Query: 63 LLEHGNALECVDPSLMSEYPEDEVL 87
LLE G+A +C D L + E+E++
Sbjct: 831 LLETGSASDCFDRRLRG-FEENELI 854
>At1g07655 unknown protein
Length = 577
Score = 67.8 bits (164), Expect = 1e-12
Identities = 41/89 (46%), Positives = 55/89 (61%), Gaps = 4/89 (4%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG--EDNVLILND 58
+S R +GY+APE A + + EK DVY FGV+ LEIV+GK + ED V +L D
Sbjct: 400 ISTRIAGTIGYMAPEYAMRGY-LTEKADVYSFGVVALEIVSGKSNTNFRPTEDFVYLL-D 457
Query: 59 HVRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E G+ LE VDP+L S+Y E+E +
Sbjct: 458 WAYVLQERGSLLELVDPTLASDYSEEEAM 486
>At1g53420 hypothetical protein, 5' partial
Length = 856
Score = 66.6 bits (161), Expect = 3e-12
Identities = 38/88 (43%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGK-RPVEYGEDNVLILNDH 59
+S R GY+APE A + + +K DVY FG++ LEIV G+ +E ++N L D
Sbjct: 685 ISTRIAGTFGYMAPEYAMRG-HLTDKADVYSFGIVALEIVHGRSNKIERSKNNTFYLIDW 743
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
V VL E N LE VDP L SEY +E +
Sbjct: 744 VEVLREKNNLLELVDPRLGSEYNREEAM 771
>At1g15530 hypothetical protein
Length = 656
Score = 66.2 bits (160), Expect = 4e-12
Identities = 34/79 (43%), Positives = 48/79 (60%), Gaps = 1/79 (1%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
+ R LGY+APELA E DVY FGV++LE+V+G+RP+EY E+ ++L D VR
Sbjct: 518 TTRVVGTLGYLAPELA-SASAPTEASDVYSFGVVVLEVVSGRRPIEYAEEEDMVLVDWVR 576
Query: 62 VLLEHGNALECVDPSLMSE 80
L G ++ D + SE
Sbjct: 577 DLYGGGRVVDAADERVRSE 595
>At1g01540 unknown protein
Length = 472
Score = 63.2 bits (152), Expect = 3e-11
Identities = 35/91 (38%), Positives = 56/91 (61%), Gaps = 9/91 (9%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLIL 56
++ R GYVAPE AC + +NEK D+Y FG++I+EI+TG+ PV+Y GE N++
Sbjct: 312 VTTRVMGTFGYVAPEYACTGM-LNEKSDIYSFGILIMEIITGRNPVDYSRPQGETNLV-- 368
Query: 57 NDHVRVLLEHGNALECVDPSLMSEYPEDEVL 87
D ++ ++ + + E VDP + E P + L
Sbjct: 369 -DWLKSMVGNRRSEEVVDPKI-PEPPSSKAL 397
>At5g57670 putative protein
Length = 416
Score = 62.4 bits (150), Expect = 6e-11
Identities = 30/81 (37%), Positives = 50/81 (61%), Gaps = 4/81 (4%)
Query: 6 QSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLE 65
+ GY+APE Q ++EK D+Y FG+++LEI+TG+RPV + ++L+ + +E
Sbjct: 267 EGTFGYLAPESLMQGT-IDEKTDIYAFGILLLEIITGRRPVNPTQKHILL---WAKPAME 322
Query: 66 HGNALECVDPSLMSEYPEDEV 86
GN E VDP L +Y + ++
Sbjct: 323 TGNTSELVDPKLQDKYDDQQM 343
>At3g19700 receptor-like protein kinase, putative
Length = 991
Score = 61.2 bits (147), Expect = 1e-10
Identities = 35/84 (41%), Positives = 55/84 (64%), Gaps = 8/84 (9%)
Query: 6 QSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE--YGEDNVLILNDHVRVL 63
+ LGY+APE A +VNEK DVY FGV+++E+VTGK+P+E +GE+N +++ V +
Sbjct: 854 KGTLGYIAPEYA-YTTKVNEKSDVYSFGVVLMELVTGKKPLETDFGENNDIVM--WVWSV 910
Query: 64 LEHGN---ALECVDPSLMSEYPED 84
+ N ++ +D S+ EY ED
Sbjct: 911 SKETNREMMMKLIDTSIEDEYKED 934
>At1g53440 receptor-like serine/threonine kinase, putative
Length = 1035
Score = 60.8 bits (146), Expect = 2e-10
Identities = 35/88 (39%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FGV+ LEIV+GK Y ++ + L D
Sbjct: 825 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGVVCLEIVSGKSNTNYRPKEEFIYLLDW 883
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E G+ LE VDP L + + + E +
Sbjct: 884 AYVLQEQGSLLELVDPDLGTSFSKKEAM 911
>At1g53430 receptor-like serine/threonine kinase, putative
Length = 1030
Score = 60.5 bits (145), Expect = 2e-10
Identities = 35/88 (39%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FGV+ LEIV+GK Y ++ + L D
Sbjct: 819 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGVVCLEIVSGKSNTNYRPKEEFVYLLDW 877
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E G+ LE VDP L + + + E +
Sbjct: 878 AYVLQEQGSLLELVDPDLGTSFSKKEAM 905
>At2g33170 putative receptor-like protein kinase
Length = 1124
Score = 60.1 bits (144), Expect = 3e-10
Identities = 35/81 (43%), Positives = 49/81 (60%), Gaps = 9/81 (11%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV---EYGEDNVLILNDHVRVLLEH 66
GY+APE A ++V EKCD+Y FGV++LE++TGK PV E G D +H+R +H
Sbjct: 993 GYIAPEYA-YTMKVTEKCDIYSFGVVLLELLTGKAPVQPLEQGGDLATWTRNHIR---DH 1048
Query: 67 GNALECVDPSLMSEYPEDEVL 87
E +DP L ED+V+
Sbjct: 1049 SLTSEILDPYLTK--VEDDVI 1067
>At3g09010 putative receptor ser/thr protein kinase
Length = 393
Score = 59.7 bits (143), Expect = 4e-10
Identities = 33/87 (37%), Positives = 52/87 (58%), Gaps = 3/87 (3%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG-EDNVLILNDH 59
+S R +GY+APE A + ++ +K DVY FG+++LE+++G D ++L +
Sbjct: 204 VSTRVAGTVGYLAPEYAL-LGQLTKKADVYSFGILVLEVISGNSSTRAAFGDEYMVLVEW 262
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEV 86
V L E LECVDP L +++P DEV
Sbjct: 263 VWKLREERRLLECVDPEL-TKFPADEV 288
>At5g65240 receptor-like protein kinase
Length = 607
Score = 59.3 bits (142), Expect = 5e-10
Identities = 32/90 (35%), Positives = 57/90 (62%), Gaps = 6/90 (6%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG----EDNVLIL 56
++ + + +G++APE + +EK DV+G+G+M+LE+VTG+R +++ ED+VL+L
Sbjct: 443 VTTQVRGTMGHIAPE-CISTGKSSEKTDVFGYGIMLLELVTGQRAIDFSRLEEEDDVLLL 501
Query: 57 NDHVRVLLEHGNALECVDPSLMSEYPEDEV 86
DHV+ L + VD L +Y ++EV
Sbjct: 502 -DHVKKLEREKRLEDIVDKKLDEDYIKEEV 530
>At2g42960 putative protein kinase
Length = 494
Score = 59.3 bits (142), Expect = 5e-10
Identities = 33/78 (42%), Positives = 49/78 (62%), Gaps = 2/78 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED-NVLILNDH 59
++ R GYVAPE A L +NEK D+Y FGV++LE +TG+ PV+YG N + L +
Sbjct: 341 ITTRVMGTFGYVAPEYANTGL-LNEKSDIYSFGVLLLEAITGRDPVDYGRPANEVNLVEW 399
Query: 60 VRVLLEHGNALECVDPSL 77
+++++ A E VDP L
Sbjct: 400 LKMMVGTRRAEEVVDPRL 417
>At1g11410 putative brassinosteroid insensitive protein; similar to
EST gb|W43102
Length = 840
Score = 58.9 bits (141), Expect = 6e-10
Identities = 39/90 (43%), Positives = 54/90 (59%), Gaps = 5/90 (5%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
+NR GY++PE A + + K DVY FGV+ILEI+TGKR + E++ L L H+
Sbjct: 677 TNRVVGTYGYMSPEYAMDG-QFSIKSDVYSFGVLILEIITGKRNSAFYEES-LNLVKHIW 734
Query: 62 VLLEHGNALECVDPSLMSE--YPEDEVLPC 89
E+G A+E +D LM E Y E EV+ C
Sbjct: 735 DRWENGEAIEIID-KLMGEETYDEGEVMKC 763
>At3g24400 protein kinase, putative
Length = 694
Score = 58.5 bits (140), Expect = 8e-10
Identities = 37/93 (39%), Positives = 54/93 (57%), Gaps = 10/93 (10%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE---DNVLILN 57
+S R GY+APE A ++ EK DV+ FGV++LE++TG+RP++ DN L+
Sbjct: 487 VSTRVMGTFGYLAPEYASSG-KLTEKSDVFSFGVVLLELITGRRPIDVNNVHADNSLV-- 543
Query: 58 DHVRVLL----EHGNALECVDPSLMSEYPEDEV 86
D R LL E GN VD L +EY ++E+
Sbjct: 544 DWARPLLNQVSELGNFEVVVDKKLNNEYDKEEM 576
>At1g56720 protein kinase, putative
Length = 492
Score = 58.5 bits (140), Expect = 8e-10
Identities = 33/83 (39%), Positives = 52/83 (61%), Gaps = 2/83 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED-NVLILNDH 59
++ R GYVAPE A L +NEK DVY FGV++LE +TG+ PV+YG + + L D
Sbjct: 337 VTTRVMGTFGYVAPEYANSGL-LNEKSDVYSFGVVLLEAITGRDPVDYGRPAHEVNLVDW 395
Query: 60 VRVLLEHGNALECVDPSLMSEYP 82
+++++ + E VDP++ + P
Sbjct: 396 LKMMVGTRRSEEVVDPNIEVKPP 418
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,137,585
Number of Sequences: 26719
Number of extensions: 77000
Number of successful extensions: 1054
Number of sequences better than 10.0: 723
Number of HSP's better than 10.0 without gapping: 667
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 274
Number of HSP's gapped (non-prelim): 729
length of query: 126
length of database: 11,318,596
effective HSP length: 87
effective length of query: 39
effective length of database: 8,994,043
effective search space: 350767677
effective search space used: 350767677
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146632.13


