

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.2 + phase: 0
(305 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
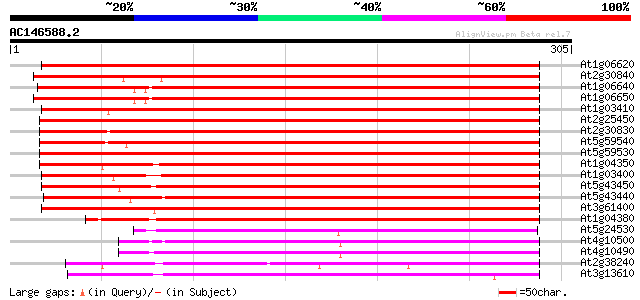
Score E
Sequences producing significant alignments: (bits) Value
At1g06620 oxidoreductase-like protein 286 7e-78
At2g30840 putative dioxygenase 285 2e-77
At1g06640 putative Iron/Ascorbate oxidoreductase family protein 284 5e-77
At1g06650 oxidoreductase, putative 283 6e-77
At1g03410 putative 1-aminocyclopropane-1-carboxylate oxidase 280 9e-76
At2g25450 putative dioxygenase 278 3e-75
At2g30830 putative dioxygenase 278 3e-75
At5g59540 1-aminocyclopropane-1-carboxylate oxidase - like protein 273 1e-73
At5g59530 1-aminocyclopropane-1-carboxylate oxidase - like protein 270 7e-73
At1g04350 putative 1-aminocyclopropane-1-carboxylate oxidase 265 2e-71
At1g03400 putative 1-aminocyclopropane-1-carboxylate oxidase 265 3e-71
At5g43450 1-aminocyclopropane-1-carboxylate oxidase 257 5e-69
At5g43440 1-aminocyclopropane-1-carboxylate oxidase 253 9e-68
At3g61400 1-aminocyclopropane-1-carboxylate oxidase-like protein 253 9e-68
At1g04380 putative 1-aminocyclopropane-1-carboxylate oxidase 226 1e-59
At5g24530 flavanone 3-hydroxylase-like protein 157 7e-39
At4g10500 putative Fe(II)/ascorbate oxidase 157 9e-39
At4g10490 putative flavanone 3-beta-hydroxylase 153 1e-37
At2g38240 putative anthocyanidin synthase 144 6e-35
At3g13610 unknown protein 139 1e-33
>At1g06620 oxidoreductase-like protein
Length = 365
Score = 286 bits (733), Expect = 7e-78
Identities = 134/272 (49%), Positives = 193/272 (70%), Gaps = 1/272 (0%)
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIK-NPIKNSMLNVPIIDLKDIH 76
DR+ LK FD++K GV+GL++ G+T+IP +F + + P +S ++P IDLK
Sbjct: 12 DRSTLLKAFDETKTGVKGLIDAGITEIPSIFRAPPATLTSPKPPSSSDFSIPTIDLKGGG 71
Query: 77 IDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRD 136
D R ++ +I A ++WGFFQVINHGIP+DVL++ IDGIR FHEQD EV+K FY+RD
Sbjct: 72 TDSITRRSLVEKIGDAAEKWGFFQVINHGIPMDVLEKMIDGIREFHEQDTEVKKGFYSRD 131
Query: 137 MKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGST 196
K+VY S L+ +ANWRD++GC+ AP+PP+ E+LP ++++EYSK+V LG
Sbjct: 132 PASKMVYSSNFDLFSSPAANWRDTLGCYTAPDPPRPEDLPATCGEMMIEYSKEVMKLGKL 191
Query: 197 ISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQ 256
+ EL SE LGL+ ++LK+ + + + GHYYPPCP+P+LT+G + H+D +F+TI+LQ+
Sbjct: 192 LFELLSEALGLNTNHLKDMDCTNSLLLLGHYYPPCPQPDLTLGLTKHSDNSFLTILLQDH 251
Query: 257 LAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ GLQVL D W +V PV GALVVN+GDLLQ+
Sbjct: 252 IGGLQVLHDQYWVDVPPVPGALVVNVGDLLQL 283
>At2g30840 putative dioxygenase
Length = 362
Score = 285 bits (729), Expect = 2e-77
Identities = 139/279 (49%), Positives = 198/279 (70%), Gaps = 4/279 (1%)
Query: 14 DSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPI--KNSMLNVPIID 71
++TYDR E+K FD+ K+GV+GL++ GVT+IPR+F+ +N+ + + ++ + +P ID
Sbjct: 2 EATYDRASEVKAFDELKIGVKGLLDAGVTQIPRIFHHPHLNLTDSNLLLSSTTMVIPTID 61
Query: 72 LKDIHIDPSR--RVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVR 129
LK D R VI IR A + +GFFQVINHGI DV+++ DGIR FHEQD +VR
Sbjct: 62 LKGGVFDEYTVTRESVIAMIRDAVERFGFFQVINHGISNDVMEKMKDGIRGFHEQDSDVR 121
Query: 130 KQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKK 189
K+FY RD+ K + Y S LY SANWRD++ CFMAP+ P+ E+LP++ +I++EY+K+
Sbjct: 122 KKFYTRDVTKTVKYNSNFDLYSSPSANWRDTLSCFMAPDVPETEDLPDICGEIMLEYAKR 181
Query: 190 VATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFM 249
V LG I EL SE LGL+P++LKE + +G+ + HYYPPCPEP LT G S H+D +F+
Sbjct: 182 VMKLGELIFELLSEALGLNPNHLKEMDCTKGLLMLSHYYPPCPEPGLTFGTSPHSDRSFL 241
Query: 250 TIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
TI+LQ+ + GLQV ++ W +V PV GAL+VN+GDLLQ+
Sbjct: 242 TILLQDHIGGLQVRQNGYWVDVPPVPGALLVNLGDLLQL 280
>At1g06640 putative Iron/Ascorbate oxidoreductase family protein
Length = 369
Score = 284 bits (726), Expect = 5e-77
Identities = 133/279 (47%), Positives = 197/279 (69%), Gaps = 7/279 (2%)
Query: 16 TYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIK-NPIKNSMLN---VPIID 71
++DR ELK FD++K GV+GL++ G++KIPR+F+ V + P+ + +L+ +P ID
Sbjct: 9 SFDRASELKAFDETKTGVKGLVDSGISKIPRIFHHSSVELANPKPLPSDLLHLKTIPTID 68
Query: 72 L--KDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVR 129
L +D D + I I+ A +WGFFQVINHG+ +++L++ DG+R FHEQ PEVR
Sbjct: 69 LGGRDFQ-DAIKHKNAIEGIKEAAAKWGFFQVINHGVSLELLEKMKDGVRDFHEQPPEVR 127
Query: 130 KQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKK 189
K Y+RD +K +YLS LY +ANWRD+ C+MAP+PP+ ++LPE+ RD++MEYSK+
Sbjct: 128 KDLYSRDFGRKFIYLSNFDLYTAAAANWRDTFYCYMAPDPPEPQDLPEICRDVMMEYSKQ 187
Query: 190 VATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFM 249
V LG + EL SE LGL+P++LK+ +G+ + HY+PPCPEP+LT G S H+D +F+
Sbjct: 188 VMILGEFLFELLSEALGLNPNHLKDMECLKGLRMLCHYFPPCPEPDLTFGTSKHSDGSFL 247
Query: 250 TIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
T++L + + GLQV R+ WF+V V GAL++NIGDLLQ+
Sbjct: 248 TVLLPDNIEGLQVCREGYWFDVPHVPGALIINIGDLLQL 286
>At1g06650 oxidoreductase, putative
Length = 369
Score = 283 bits (725), Expect = 6e-77
Identities = 133/281 (47%), Positives = 198/281 (70%), Gaps = 7/281 (2%)
Query: 14 DSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNI-IKNPIKNSMLN---VPI 69
D +DR ELK FD++K GV+GL++ GV+++PR+F+ V + P+ + +L+ +P
Sbjct: 7 DPLFDRASELKAFDETKTGVKGLVDSGVSQVPRIFHHPTVKLSTPKPLPSDLLHLKTIPT 66
Query: 70 IDL--KDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPE 127
IDL +D D +R I +I+ A +WGFFQVINHG+ +++L++ G+R FHEQ E
Sbjct: 67 IDLGGRDFQ-DAIKRNNAIEEIKEAAAKWGFFQVINHGVSLELLEKMKKGVRDFHEQSQE 125
Query: 128 VRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYS 187
VRK+FY+RD ++ +YLS L+ +ANWRD+ C MAP+ PK ++LPE+ RDI+MEYS
Sbjct: 126 VRKEFYSRDFSRRFLYLSNFDLFSSPAANWRDTFSCTMAPDTPKPQDLPEICRDIMMEYS 185
Query: 188 KKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPA 247
K+V LG + EL SE LGL P++L + + +G+ + HYYPPCPEP+LT+G S H+D +
Sbjct: 186 KQVMNLGKFLFELLSEALGLEPNHLNDMDCSKGLLMLSHYYPPCPEPDLTLGTSQHSDNS 245
Query: 248 FMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
F+T++L +Q+ GLQV R+ WF+V V GAL++NIGDLLQ+
Sbjct: 246 FLTVLLPDQIEGLQVRREGHWFDVPHVSGALIINIGDLLQL 286
>At1g03410 putative 1-aminocyclopropane-1-carboxylate oxidase
Length = 398
Score = 280 bits (715), Expect = 9e-76
Identities = 131/275 (47%), Positives = 196/275 (70%), Gaps = 4/275 (1%)
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGE---VNIIKNPIKNSMLNVPIIDLKD 74
DR+ + K FD++K GV+GL+ G+ +IP MF++ ++ + + L +P +DLK
Sbjct: 42 DRSSQAKAFDETKTGVKGLVASGIKEIPAMFHTPPDTLTSLKQTAPPSQQLTIPTVDLKG 101
Query: 75 IHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYN 134
+D R V+ +I A + WGFFQV+NHGI ++V++ +GIRRFHEQDPEV+K+FY+
Sbjct: 102 GSMDLISRRSVVEKIGDAAERWGFFQVVNHGISVEVMERMKEGIRRFHEQDPEVKKRFYS 161
Query: 135 RDMKKKIVYLSTTSLYR-DKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATL 193
RD + ++Y S L+ +K+ANWRD++ C+MAP+PPK ++LP V +I+MEYSK++ TL
Sbjct: 162 RDHTRDVLYYSNIDLHTCNKAANWRDTLACYMAPDPPKLQDLPAVCGEIMMEYSKQLMTL 221
Query: 194 GSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVL 253
G + EL SE LGL+P++LK+ + + G YYPPCP+P+LT+G S HTD +F+TI+L
Sbjct: 222 GEFLFELLSEALGLNPNHLKDMGCAKSHIMFGQYYPPCPQPDLTLGISKHTDFSFITILL 281
Query: 254 QEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
Q+ + GLQV+ D W +V+PV GALV+NIGDLLQ+
Sbjct: 282 QDNIGGLQVIHDQCWVDVSPVPGALVINIGDLLQL 316
>At2g25450 putative dioxygenase
Length = 359
Score = 278 bits (711), Expect = 3e-75
Identities = 128/272 (47%), Positives = 189/272 (69%)
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIH 76
YDR ELK FD+ K+GV+GL++ GVTK+PR+F++ VN+ ++++ +P IDL +
Sbjct: 5 YDRASELKAFDEMKIGVKGLVDAGVTKVPRIFHNPHVNVANPKPTSTVVMIPTIDLGGVF 64
Query: 77 IDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRD 136
R V+ +++ A +++GFFQ INHG+P+DV+++ I+GIRRFH+QDPEVRK FY RD
Sbjct: 65 ESTVVRESVVAKVKDAMEKFGFFQAINHGVPLDVMEKMINGIRRFHDQDPEVRKMFYTRD 124
Query: 137 MKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGST 196
KK+ Y S LY +A+WRD++ C MAP+ PK ++LPEV +I++EYSK+V L
Sbjct: 125 KTKKLKYHSNADLYESPAASWRDTLSCVMAPDVPKAQDLPEVCGEIMLEYSKEVMKLAEL 184
Query: 197 ISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQ 256
+ E+ SE LGL P++LKE + +G+++ H +PPCPEP T G + HTD +F+TI+L +
Sbjct: 185 MFEILSEALGLSPNHLKEMDCAKGLWMLCHCFPPCPEPNRTFGGAQHTDRSFLTILLNDN 244
Query: 257 LAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
GLQVL D W +V P AL+ N+GD LQ+
Sbjct: 245 NGGLQVLYDGYWIDVPPNPEALIFNVGDFLQL 276
>At2g30830 putative dioxygenase
Length = 358
Score = 278 bits (710), Expect = 3e-75
Identities = 134/273 (49%), Positives = 192/273 (70%), Gaps = 2/273 (0%)
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKD-I 75
YDR E+K FD K+GV+GL++ G+TK+PR+F+ +V + NP +S L +P ID+ +
Sbjct: 5 YDRAGEVKAFDQMKIGVKGLVDAGITKVPRIFHHQDV-AVTNPKPSSTLEIPTIDVGGGV 63
Query: 76 HIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNR 135
R VI ++R A +++GFFQVINHGIP++V++ DGIR FHEQD EV+K FY+R
Sbjct: 64 FESTVTRKSVIAKVRAAVEKFGFFQVINHGIPLEVMESMKDGIRGFHEQDSEVKKTFYSR 123
Query: 136 DMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGS 195
D+ KK+ Y + LY ++ANWRD++ MAP+ P+ +LP + R+I++EYSK++ LG
Sbjct: 124 DITKKVKYNTNFDLYSSQAANWRDTLTMVMAPDVPQAGDLPVICREIMLEYSKRMMKLGE 183
Query: 196 TISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQE 255
I EL SE LGL P++LKE N + + + HYYPPCPEP+ T G S+HTD +F+TI+LQ+
Sbjct: 184 LIFELLSEALGLKPNHLKELNCAKSLSLLSHYYPPCPEPDRTFGISSHTDISFITILLQD 243
Query: 256 QLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ GLQVL D W +V P AL+VN+GDLLQ+
Sbjct: 244 HIGGLQVLHDGYWIDVPPNPEALIVNLGDLLQL 276
>At5g59540 1-aminocyclopropane-1-carboxylate oxidase - like protein
Length = 366
Score = 273 bits (697), Expect = 1e-73
Identities = 129/277 (46%), Positives = 190/277 (68%), Gaps = 6/277 (2%)
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKN---SMLNVPIIDLK 73
+D E K FD++K GV+GL++ +T++PR+F+ + +I+ N + S L +PIID
Sbjct: 9 FDPYIERKAFDETKQGVKGLVDAKITEVPRIFHHRQ-DILTNKKPSASVSDLEIPIIDFA 67
Query: 74 DIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFY 133
+H D + R ++ +++ A + WGFFQVINH IP++VL+E DG+RRFHE+DPEV+K F+
Sbjct: 68 SVHADTASREAIVEKVKYAVENWGFFQVINHSIPLNVLEEIKDGVRRFHEEDPEVKKSFF 127
Query: 134 NRDM-KKKIVYLSTTSLYRDK-SANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVA 191
+RD KK VY S LY S NWRDS C++AP+PP EE+PE RD + EYSK V
Sbjct: 128 SRDAGNKKFVYNSNFDLYSSSPSVNWRDSFSCYIAPDPPAPEEIPETCRDAMFEYSKHVL 187
Query: 192 TLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTI 251
+ G + EL SE LGL L+ + + + + HYYPPCP+P+LT+G + H+D +F+T+
Sbjct: 188 SFGGLLFELLSEALGLKSQTLESMDCVKTLLMICHYYPPCPQPDLTLGITKHSDNSFLTL 247
Query: 252 VLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+LQ+ + GLQ+L + W +V+P+HGALVVNIGD LQ+
Sbjct: 248 LLQDNIGGLQILHQDSWVDVSPIHGALVVNIGDFLQL 284
>At5g59530 1-aminocyclopropane-1-carboxylate oxidase - like protein
Length = 364
Score = 270 bits (690), Expect = 7e-73
Identities = 127/275 (46%), Positives = 186/275 (67%), Gaps = 3/275 (1%)
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIH 76
+DR E K FD++K GV+GL++ +T+IPR+F+ + + S L +P ID ++
Sbjct: 8 FDRYIERKAFDNTKEGVKGLIDAKITEIPRIFHVPQDTLPDKKRSVSDLEIPTIDFASVN 67
Query: 77 IDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQ-DPEVRKQFYNR 135
+D R ++ +++ A + WGFFQVINHG+P++VL+E DG+RRFHE+ DPEV+K +Y+
Sbjct: 68 VDTPSREAIVEKVKYAVENWGFFQVINHGVPLNVLEEIKDGVRRFHEEEDPEVKKSYYSL 127
Query: 136 DM-KKKIVYLSTTSLYRDK-SANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATL 193
D K K Y S LY S WRDS+ C+MAP+PP EELPE RD ++EYSK V +L
Sbjct: 128 DFTKNKFAYSSNFDLYSSSPSLTWRDSISCYMAPDPPTPEELPETCRDAMIEYSKHVLSL 187
Query: 194 GSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVL 253
G + EL SE LGL LK + + + + HYYPPCP+P+LT+G S H+D +F+T++L
Sbjct: 188 GDLLFELLSEALGLKSEILKSMDCLKSLLMICHYYPPCPQPDLTLGISKHSDNSFLTVLL 247
Query: 254 QEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
Q+ + GLQ+L + W +V+P+ GALVVN+GD LQ+
Sbjct: 248 QDNIGGLQILHQDSWVDVSPLPGALVVNVGDFLQL 282
>At1g04350 putative 1-aminocyclopropane-1-carboxylate oxidase
Length = 360
Score = 265 bits (678), Expect = 2e-71
Identities = 124/275 (45%), Positives = 183/275 (66%), Gaps = 6/275 (2%)
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMFY--SGEVNIIKNPIKNSMLNVPIIDLKD 74
+D E K FD++K GV+GL++ +T+IPR+F G ++ K + + +PIID +
Sbjct: 6 FDSYSERKAFDETKTGVKGLIDAHITEIPRIFCLPQGSLSDKKPFVSTTDFAIPIIDFEG 65
Query: 75 IHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYN 134
+H+ R +++ +I+ A WGFFQVINHG+P++VL E DG+RRFHE+ PEV+K ++
Sbjct: 66 LHVS---REDIVGKIKDAASNWGFFQVINHGVPLNVLQEIQDGVRRFHEEAPEVKKTYFT 122
Query: 135 RDMKKKIVYLSTTSLYRDKSA-NWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATL 193
RD K+ VY S LY S NWRDS C+MAP+PP E+LP R + EYSK + L
Sbjct: 123 RDATKRFVYNSNFDLYSSSSCVNWRDSFACYMAPDPPNPEDLPVACRVAMFEYSKHMMRL 182
Query: 194 GSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVL 253
G + EL SE LGL LK + +G+ + HYYPPCP+P+LT+G + H+D +F+TI+L
Sbjct: 183 GDLLFELLSEALGLRSDKLKSMDCMKGLLLLCHYYPPCPQPDLTIGTNNHSDNSFLTILL 242
Query: 254 QEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
Q+Q+ GLQ+ + W +V+P+ GALV+N+GD LQ+
Sbjct: 243 QDQIGGLQIFHQDCWVDVSPIPGALVINMGDFLQL 277
>At1g03400 putative 1-aminocyclopropane-1-carboxylate oxidase
Length = 351
Score = 265 bits (676), Expect = 3e-71
Identities = 118/273 (43%), Positives = 188/273 (68%), Gaps = 10/273 (3%)
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNI--IKNPIKNSMLNVPIIDLKDI 75
DR+ + K FD++K+GV+GL++ G+T+IP +F + + +K+P L +P +DLK
Sbjct: 5 DRSSQAKAFDEAKIGVKGLVDSGITEIPALFRATPATLASLKSPPPPKHLTIPTVDLKG- 63
Query: 76 HIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNR 135
V+ +I A ++WG F ++NHGIP++VL+ I GIR FHEQ+PE +K+FY+R
Sbjct: 64 -------ASVVEKIGEAAEKWGLFHLVNHGIPVEVLERMIQGIRGFHEQEPEAKKRFYSR 116
Query: 136 DMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGS 195
D + ++Y S L ++A+WRD++GC+ AP PP+ E+LP V +I++EYSK++ +LG
Sbjct: 117 DHTRDVLYFSNHDLQNSEAASWRDTLGCYTAPEPPRLEDLPAVCGEIMLEYSKEIMSLGE 176
Query: 196 TISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQE 255
+ EL SE LGL+ +LK+ + + ++ G +YPPCP+P+LT+G + HTD +F+T++LQ+
Sbjct: 177 RLFELLSEALGLNSHHLKDMDCAKSQYMVGQHYPPCPQPDLTIGINKHTDISFLTVLLQD 236
Query: 256 QLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ GLQV + W +V PV GALV+NIGD LQ+
Sbjct: 237 NVGGLQVFHEQYWIDVTPVPGALVINIGDFLQL 269
>At5g43450 1-aminocyclopropane-1-carboxylate oxidase
Length = 362
Score = 257 bits (657), Expect = 5e-69
Identities = 127/274 (46%), Positives = 180/274 (65%), Gaps = 5/274 (1%)
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKN---PIKNSMLNVPIIDLKD 74
DR ++L F +K GV+GL++ +T++P MF+ + N I L VPIIDL D
Sbjct: 9 DRLNDLTTFISTKTGVKGLVDAEITEVPSMFHVPSSILSNNRPSDISGLNLTVPIIDLGD 68
Query: 75 IHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYN 134
+ S R VI++I+ A + WGFFQVINH +P+ VL+E + +RRFHEQDP V+ Q+
Sbjct: 69 RNT--SSRNVVISKIKDAAENWGFFQVINHDVPLTVLEEIKESVRRFHEQDPVVKNQYLP 126
Query: 135 RDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLG 194
D K+ VY + LY NWRDS C++AP+PP EE+P R ++EY+K V LG
Sbjct: 127 TDNNKRFVYNNDFDLYHSSPLNWRDSFTCYIAPDPPNPEEIPLACRSAVIEYTKHVMELG 186
Query: 195 STISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQ 254
+ + +L SE LGL LK + +G+F+ HYYPPCP+P+LT+G S HTD +F+T++LQ
Sbjct: 187 AVLFQLLSEALGLDSETLKRIDCLKGLFMLCHYYPPCPQPDLTLGISKHTDNSFLTLLLQ 246
Query: 255 EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+Q+ GLQVL ++ W +V PV GALVVNIGD +Q+
Sbjct: 247 DQIGGLQVLHEDYWVDVPPVPGALVVNIGDFMQL 280
>At5g43440 1-aminocyclopropane-1-carboxylate oxidase
Length = 365
Score = 253 bits (646), Expect = 9e-68
Identities = 122/275 (44%), Positives = 181/275 (65%), Gaps = 6/275 (2%)
Query: 19 RNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSM----LNVPIIDLKD 74
R +ELK F +K GV+GL++ +T++PR+F+ + + N + + L VPIIDL D
Sbjct: 10 RLNELKAFVSTKAGVKGLVDTKITEVPRIFHIPSSSTLSNNKPSDIFGLNLTVPIIDLGD 69
Query: 75 IHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYN 134
+ +R V ++++I+ A + WGFFQVINHGIP+ VL + G+RRFHE+DPEV+KQ++
Sbjct: 70 GNTSAARNV-LVSKIKEAAENWGFFQVINHGIPLTVLKDIKQGVRRFHEEDPEVKKQYFA 128
Query: 135 RDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPP-KHEELPEVFRDIIMEYSKKVATL 193
D + Y + ++ NW+DS C+ P P K EE+P RD+++EYSK V L
Sbjct: 129 TDFNTRFAYNTNFDIHYSSPMNWKDSFTCYTCPQDPLKPEEIPLACRDVVIEYSKHVMEL 188
Query: 194 GSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVL 253
G + +L SE LGL LK + +G+ + HYYPPCP+P+LT+G S HTD +F+TI+L
Sbjct: 189 GGLLFQLLSEALGLDSEILKNMDCLKGLLMLCHYYPPCPQPDLTLGISKHTDNSFITILL 248
Query: 254 QEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
Q+Q+ GLQVL + W +V PV GALV++IGD +Q+
Sbjct: 249 QDQIGGLQVLHQDSWVDVTPVPGALVISIGDFMQL 283
>At3g61400 1-aminocyclopropane-1-carboxylate oxidase-like protein
Length = 370
Score = 253 bits (646), Expect = 9e-68
Identities = 128/279 (45%), Positives = 186/279 (65%), Gaps = 8/279 (2%)
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNII-KNPIKNSMLNVPIIDLKDIH 76
DR +L F+++ GV+GL++ G+ ++P MF + + P +P IDL
Sbjct: 10 DRLSQLNSFEETMTGVKGLVDSGIKEVPAMFREPPAILASRKPPLALQFTIPTIDLNGGV 69
Query: 77 I-----DPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQ 131
+ D R ++ +I A ++WGFFQV+NHGIP+DVL++ +GIR FHEQD E++K+
Sbjct: 70 VYYKNQDSVTRRSMVEKIGDAAEKWGFFQVVNHGIPLDVLEKVKEGIRAFHEQDAELKKR 129
Query: 132 FYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVA 191
FY+RD +K+VY S L+ A+WRD++ +MAP+PP E+LPEV +I+MEY+K++
Sbjct: 130 FYSRDHTRKMVYYSNLDLFTAMKASWRDTMCAYMAPDPPTSEDLPEVCGEIMMEYAKEIM 189
Query: 192 TLGSTISELFSETLGLHPS-YLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMT 250
LG I EL SE LGL+ S +LK+ + + + + G YYPPCP+P+ T+G S HTD +F+T
Sbjct: 190 NLGELIFELLSEALGLNNSNHLKDMDCSKSLVLFGQYYPPCPQPDHTLGLSKHTDFSFLT 249
Query: 251 IVLQEQLAGLQVLRDNQ-WFNVAPVHGALVVNIGDLLQV 288
IVLQ L GLQVL D Q W ++ PV GALVVN+GDLLQ+
Sbjct: 250 IVLQGNLGGLQVLHDKQYWIDIPPVPGALVVNLGDLLQL 288
>At1g04380 putative 1-aminocyclopropane-1-carboxylate oxidase
Length = 345
Score = 226 bits (576), Expect = 1e-59
Identities = 107/247 (43%), Positives = 158/247 (63%), Gaps = 4/247 (1%)
Query: 42 TKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQV 101
TK+P +F + + + S VPIID +H R V+ +I+ A + WG FQV
Sbjct: 21 TKVPPIF-GLPPDALDDKKPTSDFAVPIIDFAGVH---KSREAVVEKIKAAAENWGIFQV 76
Query: 102 INHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSV 161
INHG+P+ VL+E +G+ RFHE+DPEV+K +++ D+ K +Y + LY + NWRDS
Sbjct: 77 INHGVPLSVLEEIQNGVVRFHEEDPEVKKSYFSLDLTKTFIYHNNFELYSSSAGNWRDSF 136
Query: 162 GCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGI 221
C+M P+P E+LP RD ++ YSK V +LG + EL SE LGL+ LK +G+
Sbjct: 137 VCYMDPDPSNPEDLPVACRDAMIGYSKHVMSLGGLLFELLSEALGLNSDTLKSMGCMKGL 196
Query: 222 FIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVN 281
+ HYYPPCP+P+ T+G S H+D F+TI+LQ+ + GLQ+L + W +V+P+ GAL++N
Sbjct: 197 HMICHYYPPCPQPDQTLGTSKHSDNTFITILLQDNIGGLQILHQDCWVDVSPLPGALIIN 256
Query: 282 IGDLLQV 288
IGD LQ+
Sbjct: 257 IGDFLQL 263
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 157 bits (397), Expect = 7e-39
Identities = 89/224 (39%), Positives = 127/224 (55%), Gaps = 9/224 (4%)
Query: 68 PIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPE 127
P+IDL + R +I QI AC +GFFQVINHG+ ++DE + R F E
Sbjct: 39 PLIDLSS-----TDRSFLIQQIHQACARFGFFQVINHGVNKQIIDEMVSVAREFFSMSME 93
Query: 128 VRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPE---VFRDIIM 184
+ + Y+ D K ++ ++ +++ NWRD + P E P F++I+
Sbjct: 94 EKMKLYSDDPTKTTRLSTSFNVKKEEVNNWRDYLRLHCYPIHKYVNEWPSNPPSFKEIVS 153
Query: 185 EYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHT 244
+YS++V +G I EL SE+LGL Y+K+ +G + +YYPPCPEPELT G HT
Sbjct: 154 KYSREVREVGFKIEELISESLGLEKDYMKKVLGEQGQHMAVNYYPPCPEPELTYGLPAHT 213
Query: 245 DPAFMTIVLQE-QLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQ 287
DP +TI+LQ+ + GLQ+L D QWF V P A V+NIGD LQ
Sbjct: 214 DPNALTILLQDTTVCGLQILIDGQWFAVNPHPDAFVINIGDQLQ 257
>At4g10500 putative Fe(II)/ascorbate oxidase
Length = 349
Score = 157 bits (396), Expect = 9e-39
Identities = 84/232 (36%), Positives = 132/232 (56%), Gaps = 5/232 (2%)
Query: 60 IKNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIR 119
+++S ++P+IDL+D+H P+R V ++ Q+ +AC +GFFQ+ NHG+P +++ R
Sbjct: 37 VESSGDSIPLIDLRDLH-GPNRAV-IVQQLASACSTYGFFQIKNHGVPDTTVNKMQTVAR 94
Query: 120 RFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEV- 178
F Q R + Y+ D K ++ ++ DK NWRD + P EE P
Sbjct: 95 EFFHQPESERVKHYSADPTKTTRLSTSFNVGADKVLNWRDFLRLHCFPIEDFIEEWPSSP 154
Query: 179 --FRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPEL 236
FR++ EY+ V L + E SE+LGL ++ + +YYPPCPEPEL
Sbjct: 155 ISFREVTAEYATSVRALVLRLLEAISESLGLESDHISNILGKHAQHMAFNYYPPCPEPEL 214
Query: 237 TMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
T G H DP +T++LQ+Q++GLQV +D++W V+P+ +VNIGD +QV
Sbjct: 215 TYGLPGHKDPTVITVLLQDQVSGLQVFKDDKWVAVSPIPNTFIVNIGDQMQV 266
>At4g10490 putative flavanone 3-beta-hydroxylase
Length = 348
Score = 153 bits (387), Expect = 1e-37
Identities = 81/232 (34%), Positives = 127/232 (53%), Gaps = 5/232 (2%)
Query: 60 IKNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIR 119
++ S ++P+IDL D+H R ++INQ AC GFFQ+ NHG+P + + + ++ R
Sbjct: 35 VQTSGDSIPLIDLHDLH--GPNRADIINQFAHACSSCGFFQIKNHGVPEETIKKMMNAAR 92
Query: 120 RFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEV- 178
F Q R + Y+ D KK ++ ++ ++K +NWRD + P E P
Sbjct: 93 EFFRQSESERVKHYSADTKKTTRLSTSFNVSKEKVSNWRDFLRLHCYPIEDFINEWPSTP 152
Query: 179 --FRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPEL 236
FR++ EY+ V L T+ E SE+LGL + G + +YYP CP+PEL
Sbjct: 153 ISFREVTAEYATSVRALVLTLLEAISESLGLAKDRVSNTIGKHGQHMAINYYPRCPQPEL 212
Query: 237 TMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
T G H D +T++LQ++++GLQV +D +W V PV +VN+GD +QV
Sbjct: 213 TYGLPGHKDANLITVLLQDEVSGLQVFKDGKWIAVNPVPNTFIVNLGDQMQV 264
>At2g38240 putative anthocyanidin synthase
Length = 353
Score = 144 bits (363), Expect = 6e-35
Identities = 84/268 (31%), Positives = 145/268 (53%), Gaps = 15/268 (5%)
Query: 31 LGVRGLMERGVTKIPRMFY--SGEVNIIKNPIKNSMLNVPIIDLKDIHIDPSRRVEVINQ 88
+ V+ L + GV +P + + + + ++ + +P++D+ D+ P E +
Sbjct: 10 VSVQSLSQTGVPTVPNRYVKPAHQRPVFNTTQSDAGIEIPVLDMNDVWGKP----EGLRL 65
Query: 89 IRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTS 148
+R+AC+EWGFFQ++NHG+ +++ R F E E ++++ N + Y S
Sbjct: 66 VRSACEEWGFFQMVNHGVTHSLMERVRGAWREFFELPLEEKRKYANSPDTYE-GYGSRLG 124
Query: 149 LYRDKSANWRDSVGCFMAP----NPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSET 204
+ +D +W D P NP K P R++I +Y ++V L ++E SE+
Sbjct: 125 VVKDAKLDWSDYFFLNYLPSSIRNPSKWPSQPPKIRELIEKYGEEVRKLCERLTETLSES 184
Query: 205 LGLHPSYLKER---NYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVL-QEQLAGL 260
LGL P+ L + G ++ ++YP CP+P+LT+G S+H+DP +TI+L E++AGL
Sbjct: 185 LGLKPNKLMQALGGGDKVGASLRTNFYPKCPQPQLTLGLSSHSDPGGITILLPDEKVAGL 244
Query: 261 QVLRDNQWFNVAPVHGALVVNIGDLLQV 288
QV R + W + V AL+VNIGD LQ+
Sbjct: 245 QVRRGDGWVTIKSVPNALIVNIGDQLQI 272
>At3g13610 unknown protein
Length = 361
Score = 139 bits (351), Expect = 1e-33
Identities = 79/262 (30%), Positives = 133/262 (50%), Gaps = 10/262 (3%)
Query: 32 GVRGLMERGVTKIPRMFYSG-EVNIIKNPIKNSMLNVPIIDLKDIHIDPSRRVEVINQIR 90
GV+GL E G+ +P + E +I + + +P+ID+ + D V +
Sbjct: 26 GVKGLSETGIKALPEQYIQPLEERLINKFVNETDEAIPVIDMSNPDED-----RVAEAVC 80
Query: 91 TACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLSTT-SL 149
A ++WGFFQVINHG+P++VLD+ +F E +++F + V T+ S
Sbjct: 81 DAAEKWGFFQVINHGVPLEVLDDVKAATHKFFNLPVEEKRKFTKENSLSTTVRFGTSFSP 140
Query: 150 YRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHP 209
+++ W+D + F + P++ R+ +EY K + + E + L +
Sbjct: 141 LAEQALEWKDYLSLFFVSEAEAEQFWPDICRNETLEYINKSKKMVRRLLEYLGKNLNVKE 200
Query: 210 -SYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQV--LRDN 266
KE + I + +YYP CP P+LT+G H+D + +TI+LQ+Q+ GL V L
Sbjct: 201 LDETKESLFMGSIRVNLNYYPICPNPDLTVGVGRHSDVSSLTILLQDQIGGLHVRSLASG 260
Query: 267 QWFNVAPVHGALVVNIGDLLQV 288
W +V PV G+ V+NIGD +Q+
Sbjct: 261 NWVHVPPVAGSFVINIGDAMQI 282
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,300,357
Number of Sequences: 26719
Number of extensions: 320842
Number of successful extensions: 1044
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 764
Number of HSP's gapped (non-prelim): 116
length of query: 305
length of database: 11,318,596
effective HSP length: 99
effective length of query: 206
effective length of database: 8,673,415
effective search space: 1786723490
effective search space used: 1786723490
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146588.2


