

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.8 - phase: 0
(1078 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
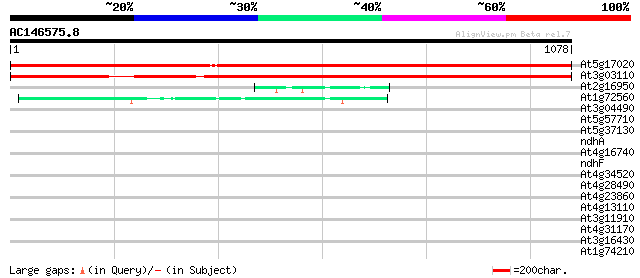
Score E
Sequences producing significant alignments: (bits) Value
At5g17020 Exportin1 (XPO1) protein 1828 0.0
At3g03110 exportin1 (XPO1) like protein 1605 0.0
At2g16950 transportin like protein 45 2e-04
At1g72560 PAUSED 45 2e-04
At3g04490 hypothetical protein 40 0.005
At5g57710 101 kDa heat shock protein; HSP101-like protein 33 0.67
At5g37130 putative protein 33 0.87
ndhA -chloroplast genome- NADH dehydrogenase ND1 32 1.9
At4g16740 limonene cyclase like protein 31 4.3
ndhF -chloroplast genome- NADH dehydrogenase ND5 30 7.4
At4g34520 fatty acid elongase 1 30 7.4
At4g28490 receptor-like protein kinase 5 precursor (RLK5) 30 7.4
At4g23860 unknown protein 30 7.4
At4g13110 putative protein 30 7.4
At3g11910 ubiquitin carboxyl-terminal hydrolase like protein 30 7.4
At4g31170 protein kinase - like protein 30 9.7
At3g16430 putative lectin 30 9.7
At1g74210 glycerophosphodiester phosphodiesterase like protein 30 9.7
>At5g17020 Exportin1 (XPO1) protein
Length = 1075
Score = 1828 bits (4734), Expect = 0.0
Identities = 918/1078 (85%), Positives = 993/1078 (91%), Gaps = 3/1078 (0%)
Query: 1 MAAEKLRDLSQPIDVPLLDATVAAFYGTGSKEQRTAADQILRELQNNPDMWLQVMHILQN 60
MAAEKLRDLSQPIDV +LDATVAAF+ TGSKE+R AADQILR+LQ NPDMWLQV+HILQN
Sbjct: 1 MAAEKLRDLSQPIDVGVLDATVAAFFVTGSKEERAAADQILRDLQANPDMWLQVVHILQN 60
Query: 61 TQNLNTKFFALQVLEGVIKYRWNALPVEQRDGMKNFISDVIVQLSRNEASFRTERLYVNK 120
T +L+TKFFALQVLEGVIKYRWNALPVEQRDGMKN+IS+VIVQLS NEASFR+ERLYVNK
Sbjct: 61 TNSLDTKFFALQVLEGVIKYRWNALPVEQRDGMKNYISEVIVQLSSNEASFRSERLYVNK 120
Query: 121 LNIILVQILKHEWPARWRNFIPDLVSAAKTSETICENCMAILKLLSEEVFDFSRGEMTQL 180
LN+ILVQI+KH+WPA+W +FIPDLV+AAKTSETICENCMAILKLLSEEVFDFSRGEMTQ
Sbjct: 121 LNVILVQIVKHDWPAKWTSFIPDLVAAAKTSETICENCMAILKLLSEEVFDFSRGEMTQQ 180
Query: 181 KIKELKQSLNSEFQLIHELCLFVLSVSQRTELIRATLSTLHAFLSWIPLGYIFESPLLET 240
KIKELKQSLNSEF+LIHELCL+VLS SQR +LIRATLS LHA+LSWIPLGYIFES LLET
Sbjct: 181 KIKELKQSLNSEFKLIHELCLYVLSASQRQDLIRATLSALHAYLSWIPLGYIFESTLLET 240
Query: 241 LLKFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPPTTNIPEAYA 300
LLKFF + AYRNLT+QCLTEVA+L FG+FY+ QYVKMY IF+ QL+ ILPP+T IPEAY+
Sbjct: 241 LLKFFPVPAYRNLTIQCLTEVAALNFGDFYNVQYVKMYTIFIGQLRIILPPSTKIPEAYS 300
Query: 301 HGSSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYVDDTEVFKVC 360
GS +EQAFIQNLALFFT FFK HIR+LEST E +S LL GLEYLINISYVDDTEVFKVC
Sbjct: 301 SGSGEEQAFIQNLALFFTSFFKFHIRVLESTPEVVSLLLAGLEYLINISYVDDTEVFKVC 360
Query: 361 LDYWNTLVSELFQPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQRRQLYAGPVS 420
LDYWN+LV ELF H + +N A +A+ +MG Q L PGMVDGLG Q++QRRQLY+ P+S
Sbjct: 361 LDYWNSLVLELFDAHHNSDNPAVSAS-LMGLQPFL--PGMVDGLGSQVMQRRQLYSHPMS 417
Query: 421 KLRMLMICRMAKPEEVLIVEDENGNIVRETMKDSDVLVQYKIMRETLIYLSHLDHDDTEK 480
KLR LMI RMAKPEEVLIVEDENGNIVRETMKD+DVLVQYKIMRETLIYLSHLDHDDTEK
Sbjct: 418 KLRGLMINRMAKPEEVLIVEDENGNIVRETMKDNDVLVQYKIMRETLIYLSHLDHDDTEK 477
Query: 481 QMLGKLSKQLSGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEITKGK 540
QML KL+KQLSG++W WNNLNTLCWAIGSISGSM E+QENRFLVMVIRDLLNLCEITKGK
Sbjct: 478 QMLRKLNKQLSGEEWAWNNLNTLCWAIGSISGSMAEDQENRFLVMVIRDLLNLCEITKGK 537
Query: 541 DNKAVIASNIMYVVGQYPRFLRAHWKFLKTVVNKLFEFMHETHPGVQDMACDTFLKIVQK 600
DNKAVIASNIMYVVGQYPRFLRAHWKFLKTVVNKLFEFMHETHPGVQDMACDTFLKIVQK
Sbjct: 538 DNKAVIASNIMYVVGQYPRFLRAHWKFLKTVVNKLFEFMHETHPGVQDMACDTFLKIVQK 597
Query: 601 CRRKFVITQVGENEPFVSELLSGLSTTIADLEPHQIHTFYESVGSMIQAESDSQKRDEYL 660
C+RKFVI QVGENEPFVSELL+GL+TT+ DLEPHQIH+FYESVG+MIQAESD QKRDEYL
Sbjct: 598 CKRKFVIVQVGENEPFVSELLTGLATTVQDLEPHQIHSFYESVGNMIQAESDPQKRDEYL 657
Query: 661 QRLMVLPNQKWMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLPQITLIF 720
QRLM LPNQKW EIIGQAR + +FLKDQ VIRTVLN+LQTNTS A+SLGTYFL QI+LIF
Sbjct: 658 QRLMALPNQKWAEIIGQARHSVEFLKDQVVIRTVLNILQTNTSAATSLGTYFLSQISLIF 717
Query: 721 LDMLNVYRMYSELISKSIAEGTPFTSRTSYVKLLRSVKRETLKLIETFLDKAEDQPQIGK 780
LDMLNVYRMYSEL+S +I EG P+ S+TS+VKLLRSVKRETLKLIETFLDKAEDQP IGK
Sbjct: 718 LDMLNVYRMYSELVSTNITEGGPYASKTSFVKLLRSVKRETLKLIETFLDKAEDQPHIGK 777
Query: 781 QFVPPMMDPVLGDYARNVPDARESEVLSLFATIINKYKASMIEDIPHIFEAVFQCTLEMI 840
QFVPPMM+ VLGDYARNVPDARESEVLSLFATIINKYKA+M++D+PHIFEAVFQCTLEMI
Sbjct: 778 QFVPPMMESVLGDYARNVPDARESEVLSLFATIINKYKATMLDDVPHIFEAVFQCTLEMI 837
Query: 841 TKNFEDYPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSIIWAFRHTERNIAETGL 900
TKNFEDYPEHRLKFFSLLRAIAT CFPALI LSS QLK VMDSIIWAFRHTERNIAETGL
Sbjct: 838 TKNFEDYPEHRLKFFSLLRAIATFCFPALIKLSSPQLKLVMDSIIWAFRHTERNIAETGL 897
Query: 901 NLLLEMLNKFQASEFCNQFYRTYFLTIEQEIFAVLTDTFHKPGFKLHVLVLQHLLCLAES 960
NLLLEML FQ SEFCNQFYR+YF+ IEQEIFAVLTDTFHKPGFKLHVLVLQ L CL ES
Sbjct: 898 NLLLEMLKNFQQSEFCNQFYRSYFMQIEQEIFAVLTDTFHKPGFKLHVLVLQQLFCLPES 957
Query: 961 GALTEPLWDAATNSYPYPSNGAFVREYTIKLLSTSFPNMTAAEVTQFVNGLFESTNDLST 1020
GALTEPLWDA T YPYP N AFVREYTIKLLS+SFPNMTAAEVTQFVNGL+ES ND S
Sbjct: 958 GALTEPLWDATTVPYPYPDNVAFVREYTIKLLSSSFPNMTAAEVTQFVNGLYESRNDPSG 1017
Query: 1021 FKTHIRDFLIQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPVELQDEMVDS 1078
FK +IRDFL+QSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAP E+QDEMVDS
Sbjct: 1018 FKNNIRDFLVQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPNEIQDEMVDS 1075
>At3g03110 exportin1 (XPO1) like protein
Length = 1022
Score = 1605 bits (4157), Expect = 0.0
Identities = 831/1080 (76%), Positives = 909/1080 (83%), Gaps = 60/1080 (5%)
Query: 1 MAAEKLRDLSQPIDVPLLDATVAAFYGTGSKEQRTAADQILRELQNNPDMWLQVMHILQN 60
MAAEKLRDLSQPIDV LLDATV AFY TGSKE+R +AD ILR+L+ NPD WLQV+HILQN
Sbjct: 1 MAAEKLRDLSQPIDVVLLDATVEAFYSTGSKEERASADNILRDLKANPDTWLQVVHILQN 60
Query: 61 TQNLNTKFFALQVLEGVIKYRWNALPVEQRDGMKNFISDVIVQLSRNEASFRTERLYVNK 120
T + +TKFFALQVLEGVIKYRWNALPVEQRDGMKN+ISDVIVQLSR+EASFRTERLYVNK
Sbjct: 61 TSSTHTKFFALQVLEGVIKYRWNALPVEQRDGMKNYISDVIVQLSRDEASFRTERLYVNK 120
Query: 121 LNIILVQILKHEWPARWRNFIPDLVSAAKTSETICENCMAILKLLSEEVFDFSRGEMTQL 180
LNIILVQI+K EWPA+W++FIPDLV AAKTSETICENCMAILKLLSEEVFDFS+GEMTQ
Sbjct: 121 LNIILVQIVKQEWPAKWKSFIPDLVIAAKTSETICENCMAILKLLSEEVFDFSKGEMTQQ 180
Query: 181 KIKELKQSLNSEFQLIHELCLFVLSVSQRTELIRATLSTLHAFLSWIPLGYIFESPLLET 240
KIKELKQSLN + LE
Sbjct: 181 KIKELKQSLNRQ---------------------------------------------LEI 195
Query: 241 LLKFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPPTTNIPEAYA 300
LLKFF + AYRNLTLQCL+EVASL FG+FYD QYVKMY+IFM QLQ+ILP NIPEAY+
Sbjct: 196 LLKFFPVPAYRNLTLQCLSEVASLNFGDFYDMQYVKMYSIFMNQLQAILPLNLNIPEAYS 255
Query: 301 HGSSDEQA--FIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYVDDTEVFK 358
GSS+EQA FIQNLALFFT FFK+HI+ILES ENIS LL GL YLI+ISYVDDTEVFK
Sbjct: 256 TGSSEEQASAFIQNLALFFTSFFKLHIKILESAPENISLLLAGLGYLISISYVDDTEVFK 315
Query: 359 VCLDYWNTLVSELFQPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQRRQLYAGP 418
P S++ + ++ S+ P VDG+ ++ +R++LY+ P
Sbjct: 316 N-------------GPMLSVDALETDSFCLLTSEQMAFLPSTVDGVKSEVTERQKLYSDP 362
Query: 419 VSKLRMLMICRMAKPEEVLIVEDENGNIVRETMKDSDVLVQYKIMRETLIYLSHLDHDDT 478
+SKLR LMI R AKPEEVLIVEDENGNIVRETMKD+DVLVQYKIMRETLIYLSHLDH+DT
Sbjct: 363 MSKLRGLMISRTAKPEEVLIVEDENGNIVRETMKDNDVLVQYKIMRETLIYLSHLDHEDT 422
Query: 479 EKQMLGKLSKQLSGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEITK 538
EKQML KLSKQLSG++W WNNLNTLCWAIGSISGSM EQENRFLVMVIRDLL+LCE+ K
Sbjct: 423 EKQMLSKLSKQLSGEEWAWNNLNTLCWAIGSISGSMVVEQENRFLVMVIRDLLSLCEVVK 482
Query: 539 GKDNKAVIASNIMYVVGQYPRFLRAHWKFLKTVVNKLFEFMHETHPGVQDMACDTFLKIV 598
GKDNKAVIASNIMYVVGQY RFLRAHWKFLKTVV+KLFEFMHETHPGVQDMACDTFLKIV
Sbjct: 483 GKDNKAVIASNIMYVVGQYSRFLRAHWKFLKTVVHKLFEFMHETHPGVQDMACDTFLKIV 542
Query: 599 QKCRRKFVITQVGENEPFVSELLSGLSTTIADLEPHQIHTFYESVGSMIQAESDSQKRDE 658
QKC+RKFVI QVGE+EPFVSELLSGL+T + DL+PHQIHTFYESVGSMIQAESD QKR E
Sbjct: 543 QKCKRKFVIVQVGESEPFVSELLSGLATIVGDLQPHQIHTFYESVGSMIQAESDPQKRGE 602
Query: 659 YLQRLMVLPNQKWMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLPQITL 718
YLQRLM LPNQKW EIIGQARQ+AD LK+ DVIRTVLN+LQTNT VA+SLGT+FL QI+L
Sbjct: 603 YLQRLMALPNQKWAEIIGQARQSADILKEPDVIRTVLNILQTNTRVATSLGTFFLSQISL 662
Query: 719 IFLDMLNVYRMYSELISKSIAEGTPFTSRTSYVKLLRSVKRETLKLIETFLDKAEDQPQI 778
IFLDMLNVYRMYSEL+S SIA G P+ SRTS VKLLRSVKRE LKLIETFLDKAE+QP I
Sbjct: 663 IFLDMLNVYRMYSELVSSSIANGGPYASRTSLVKLLRSVKREILKLIETFLDKAENQPHI 722
Query: 779 GKQFVPPMMDPVLGDYARNVPDARESEVLSLFATIINKYKASMIEDIPHIFEAVFQCTLE 838
GKQFVPPMMD VLGDYARNVPDARESEVLSLFATIINKYK M +++P IFEAVFQCTLE
Sbjct: 723 GKQFVPPMMDQVLGDYARNVPDARESEVLSLFATIINKYKVVMRDEVPLIFEAVFQCTLE 782
Query: 839 MITKNFEDYPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSIIWAFRHTERNIAET 898
MITKNFEDYPEHRLKFFSLLRAIAT CF ALI LSS+QLK VMDS+IWAFRHTERNIAET
Sbjct: 783 MITKNFEDYPEHRLKFFSLLRAIATFCFRALIQLSSEQLKLVMDSVIWAFRHTERNIAET 842
Query: 899 GLNLLLEMLNKFQASEFCNQFYRTYFLTIEQEIFAVLTDTFHKPGFKLHVLVLQHLLCLA 958
GLNLLLEML FQ S+FCN+FY+TYFL IEQE+FAVLTDTFHKPGFKLHVLVLQHL L
Sbjct: 843 GLNLLLEMLKNFQKSDFCNKFYQTYFLQIEQEVFAVLTDTFHKPGFKLHVLVLQHLFSLV 902
Query: 959 ESGALTEPLWDAATNSYPYPSNGAFVREYTIKLLSTSFPNMTAAEVTQFVNGLFESTNDL 1018
ESG+L EPLWDAAT +PY +N AFV EYT KLLS+SFPNMT EVTQFVNGL+ES ND+
Sbjct: 903 ESGSLAEPLWDAATVPHPYSNNVAFVLEYTTKLLSSSFPNMTTTEVTQFVNGLYESRNDV 962
Query: 1019 STFKTHIRDFLIQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPVELQDEMVDS 1078
FK +IRDFLIQSKEFSAQDNKDLYAEEAAAQ ERERQRMLSIPGLIAP E+QD+M DS
Sbjct: 963 GRFKDNIRDFLIQSKEFSAQDNKDLYAEEAAAQMERERQRMLSIPGLIAPSEIQDDMADS 1022
>At2g16950 transportin like protein
Length = 827
Score = 45.4 bits (106), Expect = 2e-04
Identities = 51/273 (18%), Positives = 111/273 (39%), Gaps = 43/273 (15%)
Query: 470 LSHLDHDDTEKQMLGKLSKQLSGK-DWTWNNLNTLCWAIGSIS-----GSMGEEQENRFL 523
LS++ D+ ++ + K LS D W A+G+I+ G E +
Sbjct: 379 LSNVFGDEILPALMPLIQKNLSASGDEAWKQREAAVLALGAIAEGCMNGLYPHLSEASLI 438
Query: 524 VMVIRDLLNLCEITKGKDNKAVIASNIMYVVGQYPRFL-------RAHWKFLKTVVNKLF 576
V + LL+ D +I S + + ++ ++L + + +F K ++ L
Sbjct: 439 VAFLLPLLD--------DKFPLIRSISCWTLSRFGKYLIQESGNPKGYEQFEKVLMGLLR 490
Query: 577 EFMHETHPGVQDMACDTFLKIVQKCRRKFVITQVGENEPFVSELLSGLSTTIADLEPHQI 636
+ +T+ VQ+ AC F + + + V P + +L L + +
Sbjct: 491 RLL-DTNKRVQEAACSAFATVEEDAAEELV--------PHLGVILQHLMCAFGKYQRRNL 541
Query: 637 HTFYESVGSMIQAESDSQKRDEYLQRLMVLPNQKWMEIIGQARQNADFLKDQDVIRTVLN 696
Y+++G++ + + + YL+ LM KW ++ N+D + +
Sbjct: 542 RIVYDAIGTLADSVREELNKPAYLEILMPPLVAKWQQL-----SNSD--------KDLFP 588
Query: 697 VLQTNTSVASSLGTYFLPQITLIFLDMLNVYRM 729
+L+ TS++ +LG F P +F +++ ++
Sbjct: 589 LLECFTSISQALGVGFAPFAQPVFQRCMDIIQL 621
>At1g72560 PAUSED
Length = 988
Score = 45.1 bits (105), Expect = 2e-04
Identities = 128/746 (17%), Positives = 294/746 (39%), Gaps = 93/746 (12%)
Query: 18 LDATVAAFYGTGSKEQ--RTAADQILRELQNNPDMWLQVMHILQNTQNLNTKFFALQVLE 75
L+ + + TG+ + ++ A ++++ P + + L ++ + +F+ LQ L+
Sbjct: 4 LEQAIVISFETGAVDSALKSQAVTYCQQIKETPSICSICIEKLWFSKLVQVQFWCLQTLQ 63
Query: 76 GVIKYRWNALPVEQRDGM-KNFISDVIVQLSRNEASFRTER---LYVNKLNIILVQILKH 131
V++ ++ ++ ++++ + K+ S +++ NE + R NKL +L ++ +
Sbjct: 64 DVLRVKYGSMSLDEQSYVRKSVFSMACLEVIDNENAGRVVEGPPFVKNKLAQVLATLIYY 123
Query: 132 EWPARWRNFIPDLVSAAKTSETICENCMAILKLLSEEV--FDFSRGEMTQLKIKELKQSL 189
E+P W + D + + + +L L +E+ D+ R +K ++
Sbjct: 124 EYPLIWSSVFLDFMLHLCKGAVVIDMFCRVLNALDDELISLDYPRTPEEISVAARVKDAM 183
Query: 190 NSE-FQLIHELCLFVLSVSQRT--ELIRATLSTLHAFLSWIPLGYI----FESPLLETLL 242
+ I ++S+ + + +L L + F+SWI +G + F L E +L
Sbjct: 184 RQQCVPQIARAWYDIVSMYKNSDPDLSATVLDCMRRFVSWIDIGLVANDAFVPLLFELIL 243
Query: 243 KFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPPTTNIPEAYAHG 302
+ R C+ + S + + P + +P
Sbjct: 244 SDGLSEQVRGAAAGCVLAMVSKR-----------------------MDPQSKLP------ 274
Query: 303 SSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLG--LEYLINISYVDDTEVFKVC 360
+Q L + +F V + +SALL G +E L ++ + V
Sbjct: 275 ------LLQTLQI-SRVFGLVSGDVDSDLVSKVSALLTGYAVEVLECHKRLNSEDTKAVS 327
Query: 361 LDYWNTLVSELFQPHRSLE-NSAAAATNMMGSQVSLMPPGMVDGLGPQLLQRRQLYAGPV 419
+D N ++ +F + E +S + + VS + GL P L +++ L+ +
Sbjct: 328 MDLLNEVLPSVFYVMQKCEVDSTFSIVQFLLGYVSTL-----KGL-PALKEKQLLHITQI 381
Query: 420 SKLRMLMICRMAKPEEVLIVEDENGNIVRETMKDSDVLVQY-KIMRETLIYLSHLDHDDT 478
++ + IC L D+ G +++ D + ++ K + L + + + T
Sbjct: 382 LEVIRIQICYDPMYRNNLNSLDKTG------LEEEDRMSEFRKDLFVLLRTVGRVAPEVT 435
Query: 479 EKQMLGKLSKQL-SGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEIT 537
+ + L+ + S + + + S SM EE + + L
Sbjct: 436 QHFIRNSLANAVESSSESNVEEVEAALSLLYSFGESMTEEAMKTGSGCLSELIPMLLTTQ 495
Query: 538 KGKDNKAVIASNIMYVVGQYPRFLRAHWKFLKTVVNKLFE--FMHETHPGVQDMACDTFL 595
+ ++A + + +Y +F++ + +++ V+ + +H + V A F+
Sbjct: 496 FPGHSHRLVALVYLENITRYMKFIQENSQYIPNVLGAFLDDRGLHHQNFYVSRRAGYLFM 555
Query: 596 KIVQKCRRKFVITQVGENEPFVSELLSGLSTTIADLEPHQIHT-----------FYESVG 644
++V+ + K V PF+ ++L L T++ L + +E++G
Sbjct: 556 RVVKLLKSKLV--------PFIDKILQNLQDTLSQLTTMNFASRELTGTEDGSHIFEAIG 607
Query: 645 SMIQAES-DSQKRDEYLQRLMVLPNQKWMEIIGQAR--QNADF-LKDQDVIRTVLNVLQT 700
+I E ++K+ +YL L+ Q+ + QA+ + DF +K ++ ++ +
Sbjct: 608 IIIGLEDVPAEKQSDYLSLLLTPLCQQIEAGLVQAKVASSEDFPVKIANIQFAIVAINAL 667
Query: 701 NTSVASSLGTYFLPQITLIFLDMLNV 726
+ L T P I L+F L+V
Sbjct: 668 SKGFNERLVTASRPGIGLMFKQTLDV 693
>At3g04490 hypothetical protein
Length = 1248
Score = 40.4 bits (93), Expect = 0.005
Identities = 31/139 (22%), Positives = 59/139 (42%), Gaps = 16/139 (11%)
Query: 36 AADQILRELQNNPDMWLQVMHILQNTQNLNTKFFALQVLEGVIKYRWNALPVEQRDGMKN 95
AA+ + L +P + +IL+N+Q N +F A + W+ L + + G+ +
Sbjct: 256 AAEATILSLHQSPQPYKACRYILENSQVANARFQAAAAIRESAIREWSFLATDDKGGLIS 315
Query: 96 FISDVIVQLSRNEASFRTERLYVNKLNIILVQILKHEW----PARWR-------NFIPDL 144
F ++Q + + +E ++K++ + Q++K W PA+ F P
Sbjct: 316 FCLGYVMQHANS-----SEGYVLSKVSSVAAQLMKRGWLEFTPAQKEVFFYQVSEFSPST 370
Query: 145 VSAAKTSETICENCMAILK 163
SA ENC L+
Sbjct: 371 SSAMGLPREFHENCRKSLE 389
>At5g57710 101 kDa heat shock protein; HSP101-like protein
Length = 990
Score = 33.5 bits (75), Expect = 0.67
Identities = 46/182 (25%), Positives = 73/182 (39%), Gaps = 17/182 (9%)
Query: 329 ESTQENISALLLGLEYLINISYVDDTEVFKVCLDYWNTLVSELFQPHRSLENSAAAATNM 388
E Q+ + A+ + LE LI IS +DD V +V + + + +SL NS
Sbjct: 117 EQQQQPLLAVKVELEQLI-ISILDDPSVSRVMREASFSSPAVKATIEQSLNNSVTPTPIP 175
Query: 389 MGSQVSLM-------PPGMVDGLGPQLLQRRQLYAGPVSKLR-----MLMICRMAKPEEV 436
S V L P L P+L Q VSK M ++ R K V
Sbjct: 176 SVSSVGLNFRPGGGGPMTRNSYLNPRLQQNASSVQSGVSKNDDVERVMDILGRAKKKNPV 235
Query: 437 LIVEDENGNIVRETMKDSDV--LVQYKIMRETLIYLSHLDHDDT--EKQMLGKLSKQLSG 492
L+ + E G ++RE +K +V + + ++ L + D K++ G L +L
Sbjct: 236 LVGDSEPGRVIREILKKIEVGEVGNLAVKNSKVVSLEEISSDKALRIKELDGLLQTRLKN 295
Query: 493 KD 494
D
Sbjct: 296 SD 297
>At5g37130 putative protein
Length = 887
Score = 33.1 bits (74), Expect = 0.87
Identities = 48/199 (24%), Positives = 82/199 (41%), Gaps = 26/199 (13%)
Query: 30 SKEQRTAADQILRELQNNPDMWLQVMHILQNTQNLNTKFFALQVLEGVIKYRWNALPVEQ 89
SKE A ++L+ +++ +W H+ + N + F A+Q + + + + ++ +
Sbjct: 629 SKESFIAFKEVLKLNRDSWQIWENFSHVAMDVGNTDQAFEAIQQIMRLTQNKSISVVLLD 688
Query: 90 RDGMKNFISDVIVQLSRNE----ASFRTERLYVNKLNIILVQILKHE-----WP--ARWR 138
R ++ + S NE TERLY+ I+ QI+K E W ARW
Sbjct: 689 RLMTDLENRNISYESSSNELIKTKPTTTERLYIELFGKIIQQIVKTESTFENWGLYARWS 748
Query: 139 NFIPDLVSAAKTSETICENCMAILKLLSEEVFDFSRGEMTQLKIKELKQSLNSEFQLIHE 198
DL TIC A+LK +V + EM + K + K+ + +L
Sbjct: 749 RINGDL--------TICSE--ALLK----QVRSYLGVEMWKDK-ERFKKFARASLELCRV 793
Query: 199 LCLFVLSVSQRTELIRATL 217
SV + EL A +
Sbjct: 794 YIEISASVESKRELFSAEM 812
>ndhA -chloroplast genome- NADH dehydrogenase ND1
Length = 360
Score = 32.0 bits (71), Expect = 1.9
Identities = 51/196 (26%), Positives = 86/196 (43%), Gaps = 35/196 (17%)
Query: 199 LCLFVLSVSQRTELIRATLSTL--------HAFLSWI----PLGYIFE--SPLLET-LLK 243
L L VLS+S L+ +LST+ + F W P+G+I S L E L
Sbjct: 175 LTLCVLSIS----LLSNSLSTVDIVEAQSKYGFWGWNLWRQPIGFIIFLISSLAECERLP 230
Query: 244 FFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMY--NIFMVQLQ----SILPPTTNIPE 297
F +A L TE + ++FG FY A Y+ + ++F+ L +I P +I E
Sbjct: 231 FDLPEAEEELIAGYQTEYSGIKFGLFYVASYLNLLISSLFVTVLYLGGWNISIPYISILE 290
Query: 298 AYAHGSSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYVDDTEVF 357
+ +Q F + +F TL K ++ + S + L ++ L+N+ + F
Sbjct: 291 LFQR----DQIFGTTIGIFITL-AKTYLFLFVSIATRWTLPRLRMDQLLNLGW-----KF 340
Query: 358 KVCLDYWNTLVSELFQ 373
+ + N L++ FQ
Sbjct: 341 LLPISLGNLLLTTSFQ 356
>At4g16740 limonene cyclase like protein
Length = 1024
Score = 30.8 bits (68), Expect = 4.3
Identities = 12/38 (31%), Positives = 22/38 (57%)
Query: 805 EVLSLFATIINKYKASMIEDIPHIFEAVFQCTLEMITK 842
E L LF TI+ K+ + +E++P+ + F C + I +
Sbjct: 277 EELQLFTTIVEKWDVNRLEELPNYMKLCFLCLVNEINQ 314
>ndhF -chloroplast genome- NADH dehydrogenase ND5
Length = 746
Score = 30.0 bits (66), Expect = 7.4
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 9/65 (13%)
Query: 885 IWAFRHTERNIAETGLNLLLEMLNKFQASEFCNQFYR-------TYFLTIEQEIFAVLTD 937
+W ++ GL LL M N +AS FCN+ Y+ F+T+E F + T
Sbjct: 478 LWGKEEEKKLNKNFGLVPLLTMNNTKRASFFCNKTYKISNNVRNQIFITVEN--FGLNTR 535
Query: 938 TFHKP 942
TF+ P
Sbjct: 536 TFYYP 540
>At4g34520 fatty acid elongase 1
Length = 506
Score = 30.0 bits (66), Expect = 7.4
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 8/71 (11%)
Query: 177 MTQLKIKELKQSLNSEFQLIHELCLFVLSVSQRTELIRATLSTLHAFLSWIPLGYIFESP 236
MT + +K L + + + F LCLF L+ + R T++ LH FLS Y+ +
Sbjct: 1 MTSVNVKLLYRYVLTNF---FNLCLFPLTAFLAGKASRLTINDLHNFLS-----YLQHNL 52
Query: 237 LLETLLKFFTM 247
+ TLL FT+
Sbjct: 53 ITVTLLFAFTV 63
>At4g28490 receptor-like protein kinase 5 precursor (RLK5)
Length = 999
Score = 30.0 bits (66), Expect = 7.4
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 5/54 (9%)
Query: 465 ETLIYLSHLDHDDTEK-QMLGKLSKQLSGKDWTWNNLNTLCWAIGSISGSMGEE 517
E+L+ L L D K Q+ G++ ++L G W NLN L A +SG + +E
Sbjct: 493 ESLVKLKQLSRLDLSKNQLSGEIPRELRG----WKNLNELNLANNHLSGEIPKE 542
>At4g23860 unknown protein
Length = 452
Score = 30.0 bits (66), Expect = 7.4
Identities = 14/46 (30%), Positives = 24/46 (51%)
Query: 569 KTVVNKLFEFMHETHPGVQDMACDTFLKIVQKCRRKFVITQVGENE 614
K+VV K E + ++ PG + + +VQKC K ++ G+ E
Sbjct: 246 KSVVGKCSETISDSEPGQPENGTEAEKSVVQKCSEKIDESEAGQPE 291
>At4g13110 putative protein
Length = 365
Score = 30.0 bits (66), Expect = 7.4
Identities = 29/144 (20%), Positives = 62/144 (42%), Gaps = 6/144 (4%)
Query: 634 HQIHTFYESVGSMIQAESDSQKRDEYLQRLMVLPNQKWMEIIGQARQN-----ADFLKDQ 688
H+ S M ++ + S L+ + K +IG R++ ++ K+
Sbjct: 48 HKDFLIGSSDSPMHESTTSSSSSSWSFGNLIKTLSTKSESVIGSYRRDLVEFGSELKKET 107
Query: 689 DVIRTVLNVLQTNTSVASSLGTYFLPQITLIFLDM-LNVYRMYSELISKSIAEGTPFTSR 747
+IR V + L + + +S+ + L + + D+ V++ +++IS+ P R
Sbjct: 108 SIIRRVASRLPDSLEIGASVASESLESVGQVIDDIGATVWKSTAKIISRGKESLEPNRDR 167
Query: 748 TSYVKLLRSVKRETLKLIETFLDK 771
T+ V L+ +R + L+ DK
Sbjct: 168 TNQVLSLKPYRRFEMMLLALQSDK 191
>At3g11910 ubiquitin carboxyl-terminal hydrolase like protein
Length = 1089
Score = 30.0 bits (66), Expect = 7.4
Identities = 13/32 (40%), Positives = 21/32 (65%)
Query: 659 YLQRLMVLPNQKWMEIIGQARQNADFLKDQDV 690
Y+ RLMV + K M+I+GQ + A F D+++
Sbjct: 681 YVGRLMVKSSSKPMDIVGQLNKMAGFAPDEEI 712
>At4g31170 protein kinase - like protein
Length = 412
Score = 29.6 bits (65), Expect = 9.7
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 1/55 (1%)
Query: 783 VPPMMDPVLGDYARNVPDARESEVLSLFATIINKYKASMIEDIPHIFEAVFQCTL 837
VP PVLG+ DA + EV FA I+N +A+ E + ++ +A F+C +
Sbjct: 352 VPADCLPVLGEIMTRCWDA-DPEVRPCFAEIVNLLEAAETEIMTNVRKARFRCCM 405
>At3g16430 putative lectin
Length = 296
Score = 29.6 bits (65), Expect = 9.7
Identities = 19/60 (31%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Query: 655 KRDEYLQRLMVLPNQKWMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLP 714
K D +Q L + N+K IG R FL+ QD + ++ + +SLG YF P
Sbjct: 85 KPDGLIQGLRFISNKKTSRFIGYDRGTRSFLQVQD--KKIIGFHGSAGDNLNSLGAYFAP 142
>At1g74210 glycerophosphodiester phosphodiesterase like protein
Length = 392
Score = 29.6 bits (65), Expect = 9.7
Identities = 34/153 (22%), Positives = 65/153 (42%), Gaps = 27/153 (17%)
Query: 170 FDFSRGEMTQLKIKELKQSLNSEFQLIHELCLF--VLSVSQRT--------ELIRATLST 219
FDF+ E+ QL+IK+ + ++ ++ + F L++++ E+ L
Sbjct: 127 FDFTLKELKQLRIKQRYAFRDQQYNGMYPIITFEEFLTIARDAPRVVGIYPEIKNPVLMN 186
Query: 220 LHAFLSWIPLGYIFESPLLETLLKFFTMQAY-------RNLTLQCLTEVASLQFGNFYDA 272
H + W P G FE ++ETL K+ +Y + L +Q + + N D+
Sbjct: 187 QH--VKW-PGGKKFEDKVVETLKKYGYGGSYLSKKWLKKPLFIQSFAPTSLVYISNLTDS 243
Query: 273 QYVKMYNIFMVQLQSILPPTTNIPEAYAHGSSD 305
V + + + PT + + YA +SD
Sbjct: 244 PKVLL-------IDDVTMPTQDTNQTYAEITSD 269
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,156,318
Number of Sequences: 26719
Number of extensions: 904098
Number of successful extensions: 2432
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2416
Number of HSP's gapped (non-prelim): 23
length of query: 1078
length of database: 11,318,596
effective HSP length: 110
effective length of query: 968
effective length of database: 8,379,506
effective search space: 8111361808
effective search space used: 8111361808
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146575.8


