

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.4 - phase: 0
(165 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
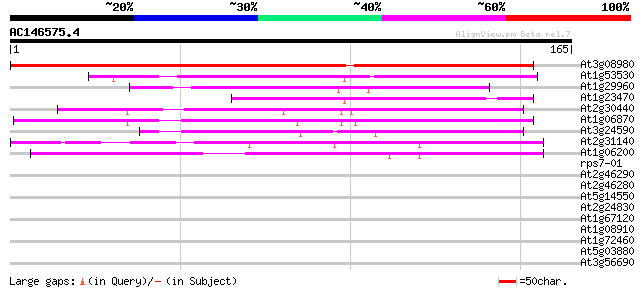
Sequences producing significant alignments: (bits) Value
At3g08980 putative mitochondrial inner membrane protease subunit 2 207 2e-54
At1g53530 unknown protein 87 4e-18
At1g29960 hypothetical protein 74 3e-14
At1g23470 putative polygalacturonase precursor 70 7e-13
At2g30440 signal peptidase I like protein 58 2e-09
At1g06870 chloroplast thylakoidal processing peptidase like protein 58 2e-09
At3g24590 chloroplast thylakoidal processing peptidase, putative 54 5e-08
At2g31140 unknown protein 49 1e-06
At1g06200 unknown protein (At1g06200) 49 2e-06
rps7-01 -chloroplast genome- 28 2.2
At2g46290 eukaryotic translation initiation factor 3 delta subunit 28 2.9
At2g46280 eukaryotic translation initiation factor 3 delta subunit 28 2.9
At5g14550 putative protein 27 5.0
At2g24830 unknown protein 27 5.0
At1g67120 hypothetical protein 27 5.0
At1g08910 hypothetical protein 27 5.0
At1g72460 putative receptor-like protein kinase 27 6.5
At5g03880 unknown protein 26 8.5
At3g56690 calmodulin-binding protein 26 8.5
>At3g08980 putative mitochondrial inner membrane protease subunit 2
Length = 154
Score = 207 bits (527), Expect = 2e-54
Identities = 91/154 (59%), Positives = 122/154 (79%), Gaps = 2/154 (1%)
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL 60
MG +N++W V KK T +I T+SDR +VVPVRG SMSPTFNP+ NS+ DDYV V+K
Sbjct: 1 MGIQNILWQVAKKSFTGSIIGLTISDRCCSVVPVRGDSMSPTFNPQRNSYLDDYVLVDKF 60
Query: 61 CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNA 120
CL +KF+ GD+V+FSSP++F + +IKRI+ +PGEW + ++DV++VPEGHCWVEGDN
Sbjct: 61 CLKDYKFARGDVVVFSSPTHFGDRYIKRIVGMPGEWISS--SRDVIRVPEGHCWVEGDNK 118
Query: 121 ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAV 154
SS DS+S+GP+PLGL++GRVT V+WPPQRI +
Sbjct: 119 TSSLDSRSFGPIPLGLIQGRVTRVMWPPQRISKI 152
>At1g53530 unknown protein
Length = 168
Score = 87.0 bits (214), Expect = 4e-18
Identities = 53/140 (37%), Positives = 74/140 (52%), Gaps = 14/140 (10%)
Query: 24 VSDRYA-TVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFK 82
V+DRY + V G SM PT N T D + E L K GD+V+ SP + K
Sbjct: 35 VTDRYIISTTHVHGPSMLPTLN-----LTGDVILAEHLSHRFGKIGLGDVVLVRSPRDPK 89
Query: 83 ETHIKRIIALPGEWF-------VNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLG 135
KRI+ L G+ V + VL VP+GH W++GDN +STDS+ +GPVP
Sbjct: 90 RMVTKRILGLEGDRLTFSADPLVGDASVSVL-VPKGHVWIQGDNLYASTDSRHFGPVPYS 148
Query: 136 LVRGRVTHVVWPPQRIGAVK 155
L+ G+ VWPP+ G+++
Sbjct: 149 LIEGKALLRVWPPEYFGSLR 168
>At1g29960 hypothetical protein
Length = 310
Score = 74.3 bits (181), Expect = 3e-14
Identities = 42/112 (37%), Positives = 65/112 (57%), Gaps = 11/112 (9%)
Query: 36 GASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGE 95
G SM+PT +P N + E++ K S GDIV+ SP N +T IKR+I + G+
Sbjct: 45 GPSMTPTLHPSGN-----VLLAERISKRYQKPSRGDIVVIRSPENPNKTPIKRVIGIEGD 99
Query: 96 ---WFVNRHNQD---VLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRV 141
+ ++ D + VP+GH +V+GD +S DS+++G VP GL++GRV
Sbjct: 100 CISFVIDSRKSDESQTIVVPKGHVFVQGDYTHNSRDSRNFGTVPYGLIQGRV 151
>At1g23470 putative polygalacturonase precursor
Length = 309
Score = 69.7 bits (169), Expect = 7e-13
Identities = 37/95 (38%), Positives = 54/95 (55%), Gaps = 9/95 (9%)
Query: 66 KFSHGDIVIFSSPSNFKETHIKRIIALPGEWF------VNRHNQDVLKVPEGHCWVEGDN 119
K S GDIV+ SP N +T IKR++ + G+ V + VP+GH +V+GD
Sbjct: 91 KPSRGDIVVIRSPENPNKTPIKRVVGVEGDCISFVIDPVKSDESQTIVVPKGHVFVQGDY 150
Query: 120 AASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAV 154
+S DS+++GPVP GL++GR V+W G V
Sbjct: 151 THNSRDSRNFGPVPYGLIQGR---VLWRDNTTGIV 182
>At2g30440 signal peptidase I like protein
Length = 340
Score = 58.2 bits (139), Expect = 2e-09
Identities = 48/170 (28%), Positives = 72/170 (42%), Gaps = 39/170 (22%)
Query: 15 ATIGLITFTVSDRYATVVP--VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDI 72
A +T ++ R A P + SM PT + D V EK+ K DI
Sbjct: 159 AAFTAVTVSILFRSALAEPKSIPSTSMYPTLDK------GDRVMAEKVSYFFRKPEVSDI 212
Query: 73 VIFSSPS----------NFKETHIKRIIALPGEW--------FVN-------------RH 101
VIF +P + + IKRI+A G+W FVN +
Sbjct: 213 VIFKAPPILLEYPEYGYSSNDVFIKRIVASEGDWVEVRDGKLFVNDIVQEEDFVLEPMSY 272
Query: 102 NQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
+ + VP+G+ +V GDN S DS ++GP+P+ + GR WPP ++
Sbjct: 273 EMEPMFVPKGYVFVLGDNRNKSFDSHNWGPLPIENIVGRSVFRYWPPSKV 322
>At1g06870 chloroplast thylakoidal processing peptidase like protein
Length = 367
Score = 58.2 bits (139), Expect = 2e-09
Identities = 51/185 (27%), Positives = 76/185 (40%), Gaps = 38/185 (20%)
Query: 2 GTRNLVWNVTKKLATIGLITFTVSDRYATVVP----VRGASMSPTFNPKTNSFTDDYVFV 57
G N + N+ + A TVS + + + + SM PT + D V
Sbjct: 174 GWVNKLLNICSEDAKAAFTAVTVSLLFRSALAEPKSIPSTSMLPTLD------VGDRVIA 227
Query: 58 EKLCLDKFKFSHGDIVIFSSPSNFKE-------THIKRIIALPGEW--------FVNR-- 100
EK+ K DIVIF +P E IKRI+A G+W VN
Sbjct: 228 EKVSYFFRKPEVSDIVIFKAPPILVEHGYSCADVFIKRIVASEGDWVEVCDGKLLVNDTV 287
Query: 101 -----------HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQ 149
+ + + VPEG+ +V GDN S DS ++GP+P+ + GR WPP
Sbjct: 288 QAEDFVLEPIDYEMEPMFVPEGYVFVLGDNRNKSFDSHNWGPLPIKNIIGRSVFRYWPPS 347
Query: 150 RIGAV 154
++ +
Sbjct: 348 KVSDI 352
>At3g24590 chloroplast thylakoidal processing peptidase, putative
Length = 149
Score = 53.5 bits (127), Expect = 5e-08
Identities = 45/142 (31%), Positives = 60/142 (41%), Gaps = 36/142 (25%)
Query: 39 MSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKET-------HIKRIIA 91
M PTF+ D + EK+ K DIVIF SP +E IKRI+A
Sbjct: 1 MYPTFD------VGDRLVAEKVSYYFRKPCANDIVIFKSPPVLQEVGYTDADVFIKRIVA 54
Query: 92 LPGEWFVNRHNQDVL----------------------KVPEGHCWVEGDNAASSTDSKSY 129
G+ V HN ++ +VPE +V GDN +S DS +
Sbjct: 55 KEGD-LVEVHNGKLMVNGVARNEKFILEPPGYEMTPIRVPENSVFVMGDNRNNSYDSHVW 113
Query: 130 GPVPLGLVRGRVTHVVWPPQRI 151
GP+PL + GR WPP R+
Sbjct: 114 GPLPLKNIIGRSVFRYWPPNRV 135
>At2g31140 unknown protein
Length = 205
Score = 48.9 bits (115), Expect = 1e-06
Identities = 40/163 (24%), Positives = 73/163 (44%), Gaps = 20/163 (12%)
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL 60
+ R L+ + K L G IT+ SD+ G MSPT N+ + + K+
Sbjct: 31 LSDRELIQIICKNLF-YGKITYLHSDK--------GPEMSPTMTANENT-----LLIRKI 76
Query: 61 CLDKFKFSH-GDIVIFSSPSNFKETHIKRIIALPG-EWFVNRHNQDVLKVPEGHCWVEGD 118
+ +F GD V+ P++ + ++R+ A+ G E ++ + + CWV +
Sbjct: 77 PIANTRFVFIGDAVVLKDPNDSDKYLVRRLAAVEGFEMVSGDEKEEPFVLEKNQCWVTAE 136
Query: 119 N----AASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNT 157
N A + DS+++GPV + GR + + G V+N+
Sbjct: 137 NQELKAKEAYDSRTFGPVSTADIVGRAIYCLRTAVDHGPVRNS 179
>At1g06200 unknown protein (At1g06200)
Length = 206
Score = 48.5 bits (114), Expect = 2e-06
Identities = 38/156 (24%), Positives = 66/156 (41%), Gaps = 17/156 (10%)
Query: 7 VWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFK 66
++ V K G I++ SD+ + P G + S K Y+FV
Sbjct: 36 LFGVVMKNLFYGRISYLHSDKGKEMAPTMGTNESTLLVRKLPVVDTRYIFV--------- 86
Query: 67 FSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPE-GHCWVEGDN----AA 121
GD V+ P+ + ++R+ AL G V+ +D V E CWV +N +
Sbjct: 87 ---GDAVVLKDPNETNKYIVRRLAALEGSEMVSSDEKDEPFVLEKDQCWVVAENQEMKSK 143
Query: 122 SSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNT 157
+ DS+++GP+ + + GR + + G V N+
Sbjct: 144 EAYDSRTFGPISMADIVGRAIYCLRTAVDHGPVSNS 179
>rps7-01 -chloroplast genome-
Length = 1786
Score = 28.1 bits (61), Expect = 2.2
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 9/72 (12%)
Query: 80 NFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRG 139
N K+ + RI AL EW V +++L+ C+ E A K Y P G+ RG
Sbjct: 443 NLKKEFLNRIEALDKEWSV----ENILEKTTRFCYNE---AKKEYLPKIYDPFLHGISRG 495
Query: 140 RVTHVVWPPQRI 151
R+ + PP +I
Sbjct: 496 RIKKL--PPFQI 505
>At2g46290 eukaryotic translation initiation factor 3 delta subunit
Length = 355
Score = 27.7 bits (60), Expect = 2.9
Identities = 12/24 (50%), Positives = 14/24 (58%)
Query: 21 TFTVSDRYATVVPVRGASMSPTFN 44
T T+ Y TVVPV +MSP N
Sbjct: 251 TLTLIKTYTTVVPVNAVAMSPLLN 274
>At2g46280 eukaryotic translation initiation factor 3 delta subunit
Length = 328
Score = 27.7 bits (60), Expect = 2.9
Identities = 12/24 (50%), Positives = 14/24 (58%)
Query: 21 TFTVSDRYATVVPVRGASMSPTFN 44
T T+ Y TVVPV S+SP N
Sbjct: 224 TLTLLKTYTTVVPVNAVSLSPLLN 247
>At5g14550 putative protein
Length = 364
Score = 26.9 bits (58), Expect = 5.0
Identities = 11/37 (29%), Positives = 22/37 (58%)
Query: 48 NSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKET 84
+SF +VF+ C+ + FS+ I S+P++F ++
Sbjct: 147 DSFNHRFVFLSDSCIPLYSFSYTYNYIMSTPTSFVDS 183
>At2g24830 unknown protein
Length = 497
Score = 26.9 bits (58), Expect = 5.0
Identities = 18/72 (25%), Positives = 32/72 (44%), Gaps = 2/72 (2%)
Query: 45 PKTNSFTDDYVFVEKLCL--DKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHN 102
P + S F+++ C + SHG V SS N+++T K+++ W V+
Sbjct: 134 PTSESMMICKFFMQQRCRFGSSCRSSHGLDVPISSLKNYEQTEWKQLMVGSKIWAVSGSK 193
Query: 103 QDVLKVPEGHCW 114
D+ + E W
Sbjct: 194 YDIWRKAELESW 205
>At1g67120 hypothetical protein
Length = 5138
Score = 26.9 bits (58), Expect = 5.0
Identities = 17/57 (29%), Positives = 26/57 (44%)
Query: 73 VIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSY 129
+ FS S+ KE+++ I EWF H L V + WV N A+ + +Y
Sbjct: 1414 IAFSGLSSLKESNVVDPIINFWEWFNRLHTGRTLTVRDLLSWVAFVNMATESLGPAY 1470
>At1g08910 hypothetical protein
Length = 600
Score = 26.9 bits (58), Expect = 5.0
Identities = 18/59 (30%), Positives = 25/59 (41%), Gaps = 4/59 (6%)
Query: 56 FVEKLCL---DKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHN-QDVLKVPE 110
F KLC + K + F + F +T +I+ G W V N +DV VPE
Sbjct: 72 FEAKLCFLLAARIKDDDEGLPYFKEDATFSQTESYVVISADGTWMVETENDEDVELVPE 130
>At1g72460 putative receptor-like protein kinase
Length = 644
Score = 26.6 bits (57), Expect = 6.5
Identities = 18/78 (23%), Positives = 34/78 (43%), Gaps = 4/78 (5%)
Query: 31 VVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRII 90
V P++ T + NSF+ D +L K + G+ + PS++ ET ++
Sbjct: 83 VAPLKDLPSLRTISIMNNSFSGDIPEFNRLTALKSLYISGNRFSGNIPSDYFET----MV 138
Query: 91 ALPGEWFVNRHNQDVLKV 108
+L W N H ++ +
Sbjct: 139 SLKKAWLSNNHFSGLIPI 156
>At5g03880 unknown protein
Length = 339
Score = 26.2 bits (56), Expect = 8.5
Identities = 17/68 (25%), Positives = 29/68 (42%), Gaps = 14/68 (20%)
Query: 30 TVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRI 89
T+V V+ +S P+ + ++ T D G +V+F++P FK KR
Sbjct: 30 TLVMVKASSSEPSESVSVSTKTSD--------------DTGAVVVFTAPPGFKPPEPKRF 75
Query: 90 IALPGEWF 97
G+ F
Sbjct: 76 AVKSGKLF 83
>At3g56690 calmodulin-binding protein
Length = 1022
Score = 26.2 bits (56), Expect = 8.5
Identities = 18/49 (36%), Positives = 23/49 (46%), Gaps = 5/49 (10%)
Query: 24 VSDRYATVVPVRGASMSPTFNPKTNSFTDD-----YVFVEKLCLDKFKF 67
VS R V PV G S+S FN S DD + ++LCL+ F
Sbjct: 168 VSGRTVFVYPVSGPSLSDQFNGNGRSRYDDVNHLSLLACKELCLELTPF 216
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,095,544
Number of Sequences: 26719
Number of extensions: 177087
Number of successful extensions: 327
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 308
Number of HSP's gapped (non-prelim): 19
length of query: 165
length of database: 11,318,596
effective HSP length: 92
effective length of query: 73
effective length of database: 8,860,448
effective search space: 646812704
effective search space used: 646812704
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146575.4


