

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146574.11 + phase: 0 /pseudo
(738 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
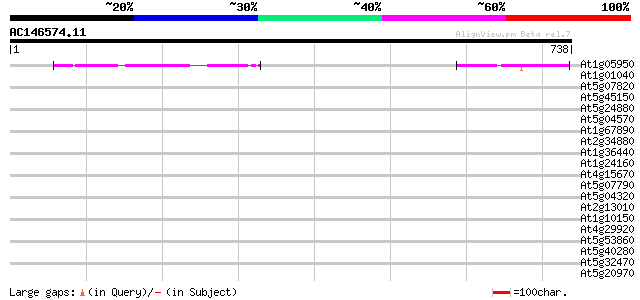
Score E
Sequences producing significant alignments: (bits) Value
At1g05950 hypothetical protein 187 1e-47
At1g01040 CAF protein 39 0.008
At5g07820 putative protein 38 0.018
At5g45150 putative protein 32 1.7
At5g24880 glutamic acid-rich protein 32 1.7
At5g04570 putative protein 32 1.7
At1g67890 putative protein kinase (At1g67890) 31 2.2
At2g34880 unknown protein 31 2.9
At1g36440 hypothetical protein 31 2.9
At1g24160 unknown protein 31 2.9
At4g15670 glutaredoxin 30 3.7
At5g07790 putative protein 30 4.9
At5g04320 putative protein 30 4.9
At2g13010 pseudogene 30 4.9
At1g10150 unknown protein 30 4.9
At4g29920 putative protein 30 6.4
At5g53860 unknown protein 29 8.3
At5g40280 farnesyltransferase beta subunit (ERA1) 29 8.3
At5g32470 putative protein 29 8.3
At5g20970 small heat shock protein -like 29 8.3
>At1g05950 hypothetical protein
Length = 590
Score = 187 bits (476), Expect = 1e-47
Identities = 113/273 (41%), Positives = 162/273 (58%), Gaps = 35/273 (12%)
Query: 58 SDVCPTADAVRAFVEHLVDPMLPAKASIRDAPSISQQQKVAKQVHSVVLLYNYYHRKRHP 117
+D CPT DA+RA +E LVDP+LP+K + D PS S ++ VAKQVH+VVLLYNYYHRK +P
Sbjct: 14 TDSCPTEDAIRALLESLVDPLLPSKPT-DDLPSTSIRESVAKQVHAVVLLYNYYHRKDNP 72
Query: 118 ELEFVAFKDFCKLIVGLRPALLIYMKFTQKPDQTDLVDVEQQLSLTEKAIASSCDICTCL 177
LE ++F+ F L ++PALL ++K + V Q L EK I +C + L
Sbjct: 73 HLECLSFESFRSLATVMKPALLQHLK--------EDGGVSGQTVLLEKVIVDACSLSMSL 124
Query: 178 DASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVGSSDQISEDMNE 237
DAS ++ + P+ +VA+LLVDS+ ++C L+ S T GVWS++EK + E+ E
Sbjct: 125 DASSDLFILNKCPIRRVAVLLVDSEKKSCYLQHSSITQGVWSLLEKPIEKEKAARENQKE 184
Query: 238 LKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSNDIMLLESYTVYSQRKEK 297
+ F +V ++ VKEATGVN DI++LE + V S +EK
Sbjct: 185 ----------------------EGVFQKVAFAVVKEATGVNHKDIVILERHLVCSLSEEK 222
Query: 298 TASRFYIMKC-SQSTADGSIQVPIKDLIESFRG 329
TA RFYIMKC SQ G + P+++++ S+RG
Sbjct: 223 TAVRFYIMKCTSQDKFSG--ENPVEEVL-SWRG 252
Score = 109 bits (272), Expect = 6e-24
Identities = 69/180 (38%), Positives = 97/180 (53%), Gaps = 36/180 (20%)
Query: 589 NANSSNKNLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIEDEMAVCNMKIHRWL 648
+A++SN NLE++QT + SK T LS+TAL L+ KR+ L QQR IEDE+A C+ + +
Sbjct: 407 SAHNSNPNLEELQTSLLSKATSLSETALKVLLCKRDKLTRQQRNIEDEIAKCD----KCI 462
Query: 649 AGEEDDFELKLESVIEGCNGTWLR--------------------------------KLDG 676
+ D+EL+LE+V+E CN T+ R +LD
Sbjct: 463 QNIKGDWELQLETVLECCNETYPRRNLQESLDKSACQSNKRLKLSETLPSTKSLCQRLDD 522
Query: 677 ICHENNWILPTYSVSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQMLTKFR 736
IC NNW+LP Y V+ SDG + A VR+ G + G EAR+SAAA +LTK +
Sbjct: 523 ICLMNNWVLPNYRVAPSDGGYEAEVRITGNHVACTIHGEEKSDAEEARESAAACLLTKLQ 582
>At1g01040 CAF protein
Length = 1909
Score = 39.3 bits (90), Expect = 0.008
Identities = 45/152 (29%), Positives = 67/152 (43%), Gaps = 15/152 (9%)
Query: 592 SSNKNLEKIQTFIASKGTILSQTALNALIRK---RNALALQQRAIEDEMAVCNMK-IHRW 647
S + N ++ FI ++Q + +K RNALA + E E+A K I+
Sbjct: 1754 SRSGNTATVEVFIDGVQVGVAQNPQKKMAQKLAARNALAALK---EKEIAESKEKHINNG 1810
Query: 648 LAGEEDDFELKLESVIEGCNGTWLRKLDGICHENNWILPTYSVSLSDGEFHAT-----VR 702
AGE D E + + G + L+ IC NW +P+Y G HA VR
Sbjct: 1811 NAGE-DQGENENGNKKNGHQPFTRQTLNDICLRKNWPMPSYRCVKEGGPAHAKRFTFGVR 1869
Query: 703 VKGVDFEYS--CEGNTCPFPREARDSAAAQML 732
V D ++ C G P ++A+DSAA +L
Sbjct: 1870 VNTSDRGWTDECIGEPMPSVKKAKDSAAVLLL 1901
>At5g07820 putative protein
Length = 561
Score = 38.1 bits (87), Expect = 0.018
Identities = 43/166 (25%), Positives = 73/166 (43%), Gaps = 17/166 (10%)
Query: 455 VSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKEC 514
V +K I + P D L KEK RCD V E D++V K S+ E
Sbjct: 180 VVNKASRISQNKSPKEDLSKNLKNKEKGKIVEPVRCDDVLEKTDLEVKK----VSRISEN 235
Query: 515 QKHTANTLHISEDQKIENP-----SVQHHSNECTRPSK-AEKAVSKRKHITEGGIKDQSA 568
+ +TL E KI+ P ++ S + + S+ +E SK + + K+++
Sbjct: 236 KSSKEDTLKNKEKAKIDEPVRCDDVLEKTSLDAQKVSRISENKNSKEERLKNLKNKEKTN 295
Query: 569 FDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQT 614
D+ +D+VEK + SS +EK + +++K +S+T
Sbjct: 296 IDE----PVRPDDAVEKTLYVVESS---VEKKKKKMSTKSVKISET 334
>At5g45150 putative protein
Length = 957
Score = 31.6 bits (70), Expect = 1.7
Identities = 18/72 (25%), Positives = 33/72 (45%), Gaps = 7/72 (9%)
Query: 673 KLDGICHENNWILPTYSVSLSDGEFH-------ATVRVKGVDFEYSCEGNTCPFPREARD 725
KL IC +N W P +SV G+ + + + ++ + +G+ P +EA +
Sbjct: 841 KLFEICAKNKWPNPIFSVEEERGQQNEQKIVCSVKIEIPNIEGTFHIKGDAKPTKKEAEN 900
Query: 726 SAAAQMLTKFRS 737
S+A M+ S
Sbjct: 901 SSADHMIRALES 912
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 31.6 bits (70), Expect = 1.7
Identities = 33/157 (21%), Positives = 63/157 (40%), Gaps = 21/157 (13%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
Q +K E++ K ++ E+ K +D ++K + +KE + VKE
Sbjct: 295 QANKSEEEEDVKKKIDENETPEK---VDTESKEVESVEETTQEKE----------EEVKE 341
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKR 555
+ V++ K KE K + E++K + E + + E A K+
Sbjct: 342 EGKERVEEE----EKEKEKVKEDDQKEKVEEEEKEKVKG----DEEKEKVKEEESAEGKK 393
Query: 556 KHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANS 592
K + +G + SA++ + A EN K + A +
Sbjct: 394 KEVVKGKKESPSAYNDVIASKMQENPRKNKVLALAGA 430
>At5g04570 putative protein
Length = 555
Score = 31.6 bits (70), Expect = 1.7
Identities = 30/116 (25%), Positives = 57/116 (48%), Gaps = 18/116 (15%)
Query: 507 LPSKNKECQKHTANTLHISEDQKIENPSVQ-------HHSNEC-TRPSKAEKAVSKRKHI 558
L + Q+H +T H +D+ + +Q + S EC TR S S +++I
Sbjct: 293 LSGSSSAVQEHQDDTQHNQQDEMNKASHLQKTFLDLLNSSEECLTRQS------STKQNI 346
Query: 559 TEGGI-KDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQ 613
T+G + +D++A D + N S++ ++ +NSSNK ++ + + TIL +
Sbjct: 347 TDGCLPRDRTAEDVV--DPLSNNSSLQNILVESNSSNKEQTAVE-YKETNATILRE 399
>At1g67890 putative protein kinase (At1g67890)
Length = 765
Score = 31.2 bits (69), Expect = 2.2
Identities = 36/156 (23%), Positives = 56/156 (35%), Gaps = 18/156 (11%)
Query: 431 FSNSLQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRC 490
FS+ +D++ + P+ + +S G+++ D+K S N + + K+
Sbjct: 317 FSDQREDAETNDASTPRGNLIQSPF---GVFLCNDDKSSSKASGESNDENDRNSVVPKKL 373
Query: 491 DSVKEDWDMDVDKSLVLPSKNKECQKHTANTLHI------SEDQKIENPSVQHHSNECTR 544
S E+W V K L P K E + H +E QK E HHSN
Sbjct: 374 TSKTEEW--MVKKGLSWPWKGNEREGLERRNAHSVWPWVHNEQQKEE----AHHSNSYNS 427
Query: 545 PSKAEKAVSKRKHITE---GGIKDQSAFDKICAGTT 577
A K G + SA G+T
Sbjct: 428 VKSESLASESNKPANNENMGSVNVNSASSASSCGST 463
>At2g34880 unknown protein
Length = 806
Score = 30.8 bits (68), Expect = 2.9
Identities = 34/143 (23%), Positives = 61/143 (41%), Gaps = 19/143 (13%)
Query: 542 CTRPSKAEKAVSKRKHITEGGIKDQ-SAFDKICAGTTFENDSVEKCI-----LNANSSN- 594
C + KA+ R + E I+ + F + F+++ +CI L+ +++
Sbjct: 469 CGKDGIITKAIEARLRMEEKRIEALGNGFSLVKMDKDFDSNCERECISCFSDLHLSATGC 528
Query: 595 KNLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIEDEMAVCNMKIHRWLAGEEDD 654
KN ++ + +K I S + I R + DE++ + R L GE DD
Sbjct: 529 KNCSSLEEYGCTKHDICSCEGKDRFIFLRYTI--------DELS----SLVRALEGESDD 576
Query: 655 FELKLESVIEGCNGTWLRKLDGI 677
+ L V+EGC+ T + GI
Sbjct: 577 LKAWLSKVMEGCSETQKGESSGI 599
>At1g36440 hypothetical protein
Length = 409
Score = 30.8 bits (68), Expect = 2.9
Identities = 33/142 (23%), Positives = 55/142 (38%), Gaps = 14/142 (9%)
Query: 468 PGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTANTLHISED 527
P +D K+ L + CG+ D + + W + + L K K S+
Sbjct: 111 PTADEKLLLERTMDVECGVNDVDDVIVDGWKKRLLRLLWSRLSEKPVWKK-------SKV 163
Query: 528 QKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCI 587
+ + P++ + + VS K + E G KD S K C+ T+ END EK +
Sbjct: 164 KVAQTPTIASNDG-------LTEVVSGLKTLMENGFKDISKKMKDCSKTSKENDVREKEM 216
Query: 588 LNANSSNKNLEKIQTFIASKGT 609
+ N+ E KG+
Sbjct: 217 GTKDGENEAEEGSDANGGEKGS 238
>At1g24160 unknown protein
Length = 540
Score = 30.8 bits (68), Expect = 2.9
Identities = 41/162 (25%), Positives = 69/162 (42%), Gaps = 15/162 (9%)
Query: 578 FENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTAL----NALIRKRNALALQQRAI 633
FENDS+ +A S NK LE++ A+ G++ + A I +R A + Q +
Sbjct: 44 FENDSLSWEKFSAFSPNKYLEEVGK-CATPGSVAQKKAYFEAHYKKIAERKAEIIDQEKL 102
Query: 634 EDEMAVCNMKIHRWLAGEEDDFELKLESVIE-GCNGTWLRKLDGICHENNWILPTYSVSL 692
DE A + + E ++ + +ES ++ GCNG D H + VS+
Sbjct: 103 MDENASFRSIVSDQESVECENGGVVVESEVDNGCNGQ--LSCDEDKHVTDITAKVNQVSI 160
Query: 693 SDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQMLTK 734
+ T+ VK C+ + E +DS + +L K
Sbjct: 161 DESN-EETIVVK------ECQSSVDTVKDEVKDSVDSPVLEK 195
Score = 29.3 bits (64), Expect = 8.3
Identities = 32/175 (18%), Positives = 72/175 (40%), Gaps = 20/175 (11%)
Query: 439 KLAEKQ---FPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
K+AE++ + ++ + + S + + D ++ + V + + NGC CD K
Sbjct: 88 KIAERKAEIIDQEKLMDENASFRSIVSDQESVECENGGVVVESEVDNGCNGQLSCDEDKH 147
Query: 496 DWD-------MDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKA 548
D + +D+S KECQ + +T+ +++P ++ K
Sbjct: 148 VTDITAKVNQVSIDESNEETIVVKECQS-SVDTVKDEVKDSVDSPVLEKAEEIALEEEKI 206
Query: 549 EKAVSKRKH----ITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEK 599
E V ++ + E ++ +++ + +ND+V+ AN + K + K
Sbjct: 207 EMVVHVQERSEEVLQEDEKEETEVREEVRDDISLQNDTVD-----ANETTKKVVK 256
>At4g15670 glutaredoxin
Length = 102
Score = 30.4 bits (67), Expect = 3.7
Identities = 19/56 (33%), Positives = 31/56 (54%), Gaps = 4/56 (7%)
Query: 499 MDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSK 554
M +KSLV+ SKN C HT TL + NP++ + +E R + E+A+++
Sbjct: 7 MTSEKSLVIFSKNSCCMSHTIKTLFLDLG---VNPTI-YELDEINRGKEIEQALAQ 58
>At5g07790 putative protein
Length = 616
Score = 30.0 bits (66), Expect = 4.9
Identities = 36/170 (21%), Positives = 60/170 (35%), Gaps = 6/170 (3%)
Query: 455 VSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKEC 514
VS+ +D+ K G +T + KN SV ED +D L K+
Sbjct: 433 VSAAEAIVDISRKSGRETAACITSLSKNLLWFADISSSVAEDNKID-----TLQLTEKKL 487
Query: 515 QKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICA 574
++H N S D S+ +R + ++ +HI + + ++ D
Sbjct: 488 EEHETNLRLNSSDNIAAVSSIVLKKQRRSRARQGKRKCQDDQHIVDTFSEWEANDDLQVI 547
Query: 575 GTTFE-NDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTALNALIRKR 623
G E +D C N S + K S G + AL ++R
Sbjct: 548 GKLMEASDLKWNCWFPKNKSKSSPPKPVDAFTSSGEEKTGVDWGALRKRR 597
>At5g04320 putative protein
Length = 446
Score = 30.0 bits (66), Expect = 4.9
Identities = 17/71 (23%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Query: 525 SEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFENDSVE 584
S + +E +H + E TR +++ S R+ T+G + A +I + N V+
Sbjct: 338 SRSESVEPSESRHETKEITR----KRSFSTRRQSTKGKSQTDEAIKEIATDPSLVNTIVQ 393
Query: 585 KCILNANSSNK 595
+C S +K
Sbjct: 394 ECDQETESKDK 404
>At2g13010 pseudogene
Length = 1040
Score = 30.0 bits (66), Expect = 4.9
Identities = 16/54 (29%), Positives = 29/54 (53%)
Query: 97 VAKQVHSVVLLYNYYHRKRHPELEFVAFKDFCKLIVGLRPALLIYMKFTQKPDQ 150
V K +++ + NYY RH ++ +A + +++V +LI M+ Q PDQ
Sbjct: 229 VKKHLYNRGFMPNYYVWLRHGDVVEIAHELVIQIMVSSNLVILIMMRIMQVPDQ 282
>At1g10150 unknown protein
Length = 414
Score = 30.0 bits (66), Expect = 4.9
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 4/48 (8%)
Query: 197 LLVDSK----MENCLLRFCSTTDGVWSVIEKDVGSSDQISEDMNELKH 240
LL D K + +C+ RF ST GV+ D+ + DQI E + KH
Sbjct: 200 LLGDDKCRELLADCIERFTSTAIGVYLDKTMDINTYDQIFEGLTNPKH 247
>At4g29920 putative protein
Length = 1017
Score = 29.6 bits (65), Expect = 6.4
Identities = 26/95 (27%), Positives = 46/95 (48%), Gaps = 7/95 (7%)
Query: 593 SNKNLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIEDEMAVCNMKIHRWLAGEE 652
S++N+ KI SK + + NAL +K + L +R + N+ + R+ AG+
Sbjct: 705 SHENMLKIN-LRTSKASEACEELKNALKKKEEVVILIERVDLADAQFMNILVDRFEAGDL 763
Query: 653 DDFELKLESVIEGCNGTWLRKLDGICHEN-NWILP 686
D F+ K +I L + D C EN ++++P
Sbjct: 764 DGFQGKKSQII-----FLLTREDDECVENEHFVIP 793
>At5g53860 unknown protein
Length = 422
Score = 29.3 bits (64), Expect = 8.3
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 1/62 (1%)
Query: 262 EFL-QVGYSAVKEATGVNSNDIMLLESYTVYSQRKEKTASRFYIMKCSQSTADGSIQVPI 320
EF+ ++ Y +V + + DI L V QR KT + + ++C+ GS+ I
Sbjct: 64 EFISKISYLSVFAVATLGTYDIALDLGKKVICQRDCKTCNGWQALRCTMCKGTGSVHYQI 123
Query: 321 KD 322
KD
Sbjct: 124 KD 125
>At5g40280 farnesyltransferase beta subunit (ERA1)
Length = 482
Score = 29.3 bits (64), Expect = 8.3
Identities = 32/149 (21%), Positives = 59/149 (39%), Gaps = 19/149 (12%)
Query: 182 NVPNIEGWPVSKVAILLVDSKMENCLLRFCST---------TDGVWSVIEKDVGSSDQIS 232
N+ ++ W V + + + N L+ C T ++S + DV S IS
Sbjct: 277 NLDSLMNWAVHRQGVEMGFQGRTNKLVDGCYTFWQAAPCVLLQRLYSTNDHDVHGSSHIS 336
Query: 233 EDMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSNDIMLLESYTVYS 292
E NE H + + + + + + D+DE G+ +T +N ++ +S +
Sbjct: 337 EGTNEEHHAHDEDDLEDSDDDDDSDEDNDEDSVNGHRIHHTSTYINRRMQLVFDSLGL-- 394
Query: 293 QRKEKTASRFYIMKCSQSTADGSIQVPIK 321
QR Y++ CS+ G P K
Sbjct: 395 QR--------YVLLCSKIPDGGFRDKPRK 415
>At5g32470 putative protein
Length = 529
Score = 29.3 bits (64), Expect = 8.3
Identities = 13/39 (33%), Positives = 23/39 (58%)
Query: 471 DTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPS 509
D K+A++ K+ K DS +DWD+D++K + + S
Sbjct: 83 DDKLAISDLRKSVMEELKMHDSFVQDWDLDINKEVSVNS 121
>At5g20970 small heat shock protein -like
Length = 249
Score = 29.3 bits (64), Expect = 8.3
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 20/81 (24%)
Query: 216 GVWSVIEKDVGS-SDQISEDMNELKHTYQK---------RRVIQN-------PTKNGLNV 258
G +++EK G +DQ+ D +LKH Q+ R V+Q+ K L+
Sbjct: 125 GPKAMVEKPSGGKTDQLKHDAQQLKHDAQQLKHDAQQKAREVVQSGKNKLTGEPKGPLSS 184
Query: 259 DDDEFLQVG---YSAVKEATG 276
DDE +VG + KEATG
Sbjct: 185 KDDEKDKVGAKWFEKYKEATG 205
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,244,302
Number of Sequences: 26719
Number of extensions: 691620
Number of successful extensions: 2386
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 2365
Number of HSP's gapped (non-prelim): 33
length of query: 738
length of database: 11,318,596
effective HSP length: 107
effective length of query: 631
effective length of database: 8,459,663
effective search space: 5338047353
effective search space used: 5338047353
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146574.11


