

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
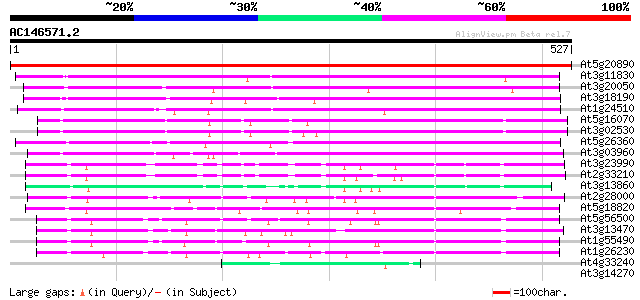
Query= AC146571.2 - phase: 0
(527 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g20890 T-complex protein 1, beta subunit 912 0.0
At3g11830 putative T-complex protein 1, ETA subunit 318 6e-87
At3g20050 t-complex polypeptide 1 homologue 311 4e-85
At3g18190 chaperonin subunit, putative 293 2e-79
At1g24510 T-complex chaperonin protein , epsilon subunit 276 2e-74
At5g16070 TCP-1 chaperonin-like protein 221 8e-58
At3g02530 putative chaperonin 221 1e-57
At5g26360 chaperonin gamma chain - like protein 219 2e-57
At3g03960 putative T-complex protein 1, theta subunit (TCP-1-Theta) 197 1e-50
At3g23990 mitochondrial chaperonin hsp60 109 4e-24
At2g33210 mitochondrial chaperonin (HSP60) 108 7e-24
At3g13860 chaperonin like protein 98 1e-20
At2g28000 putative rubisco subunit binding-protein alpha subunit 91 1e-18
At5g18820 chaperonin 60 alpha chain - like protein 86 4e-17
At5g56500 RuBisCO subunit binding-protein beta subunit precursor... 84 2e-16
At3g13470 chaperonin 60 beta - like protein 84 2e-16
At1g55490 Rubisco subunit binding-protein beta subunit 84 3e-16
At1g26230 chaperonin precursor, putative 75 7e-14
At4g33240 putative protein 46 4e-05
At3g14270 unknown protein 41 0.001
>At5g20890 T-complex protein 1, beta subunit
Length = 527
Score = 912 bits (2357), Expect = 0.0
Identities = 467/527 (88%), Positives = 500/527 (94%)
Query: 1 MTIGNIFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTN 60
M I IFKD+ASEEKGERARM+SFVGAMAI+DLVK+TLGPKGMDKILQSTGRG VTVTN
Sbjct: 1 MPIDKIFKDDASEEKGERARMASFVGAMAISDLVKSTLGPKGMDKILQSTGRGHAVTVTN 60
Query: 61 DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMT 120
DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVA+KIHPMT
Sbjct: 61 DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVASKIHPMT 120
Query: 121 IIAGFRMAAECARSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVD 180
IIAG+RMA+ECAR+AL+++V+DNK +AEKFRSDL+ IA TTL SKILSQDKEHFA++AVD
Sbjct: 121 IIAGYRMASECARNALLKRVIDNKDNAEKFRSDLLKIAMTTLCSKILSQDKEHFAEMAVD 180
Query: 181 AVMRLKGSTNLESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMD 240
AV RLKGSTNLE++QIIKKPGGSL DSFLDEGFILDKKIG+GQPKRIENA ILVANTAMD
Sbjct: 181 AVFRLKGSTNLEAIQIIKKPGGSLKDSFLDEGFILDKKIGIGQPKRIENANILVANTAMD 240
Query: 241 TDKVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFA 300
TDKVKIYGARVRVDSM KVAEIEGAEKEKMK+KV KIIGHGINCFVNRQLIYNFPEELFA
Sbjct: 241 TDKVKIYGARVRVDSMTKVAEIEGAEKEKMKDKVKKIIGHGINCFVNRQLIYNFPEELFA 300
Query: 301 DAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGV 360
DAGILAIEHADF+GIERL LVTGGEIASTFDNPESVKLG C+LIEEIMIGEDKLI FSG
Sbjct: 301 DAGILAIEHADFEGIERLGLVTGGEIASTFDNPESVKLGHCKLIEEIMIGEDKLIHFSGC 360
Query: 361 AMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALA 420
MGQAC+IVLRGASHHVLDEAERSLHDALCVLSQTVND+RVLLGGGWPEMVMAKE+D LA
Sbjct: 361 EMGQACSIVLRGASHHVLDEAERSLHDALCVLSQTVNDTRVLLGGGWPEMVMAKEVDELA 420
Query: 421 RKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSV 480
RKT GKKS A+EAFSRAL+AIPTTIADNAGLDSAEL++QLRAEH EGC AGIDVI+G+V
Sbjct: 421 RKTAGKKSHAIEAFSRALVAIPTTIADNAGLDSAELVAQLRAEHHTEGCNAGIDVITGAV 480
Query: 481 GDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRREDRM 527
GDM ERGI EAFKVKQAVLLSATEA+EMILRVD+IITCAPRRREDRM
Sbjct: 481 GDMEERGIYEAFKVKQAVLLSATEASEMILRVDEIITCAPRRREDRM 527
>At3g11830 putative T-complex protein 1, ETA subunit
Length = 557
Score = 318 bits (814), Expect = 6e-87
Identities = 187/519 (36%), Positives = 297/519 (57%), Gaps = 11/519 (2%)
Query: 6 IFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATI 65
+ K+ +G+ +S+ A+ D+V+TTLGP+GMDK++ +G VT++NDGATI
Sbjct: 11 LLKEGTDTSQGKAQLVSNINACTAVGDVVRTTLGPRGMDKLIHDD-KG-SVTISNDGATI 68
Query: 66 LKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGF 125
+K L I +PAAK+LVDI+K QD EVGDGTT+VV+LA E L+EA+ + +H +I +
Sbjct: 69 MKLLDIVHPAAKILVDIAKSQDSEVGDGTTTVVLLAAEFLKEAKPFIEDGVHAQNLIRSY 128
Query: 126 RMAAECARSALVEKVVDNKGDA-EKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMR 184
R A+ A + + E V +G + E+ + L A TTLSSK++ +KE FA + VDAVM
Sbjct: 129 RTASTLAIAKVKELAVSIEGKSVEEKKGLLAKCAATTLSSKLIGGEKEFFATMVVDAVMA 188
Query: 185 LKGSTNLESVQIIKKPGGSLIDSFLDEGFILDKKIGLG----QPKRIENAKILVANTAMD 240
+ L + I K PGG++ DSFL +G K QPK+ N KIL+ N ++
Sbjct: 189 IGNDDRLNLIGIKKVPGGNMRDSFLVDGVAFKKTFSYAGFEQQPKKFLNPKILLLNIELE 248
Query: 241 TDKVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFA 300
K A +R+ ++ I AE + +K++K + G ++R I + + FA
Sbjct: 249 LKSEK-ENAEIRLSDPSQYQSIVDAEWNIIYDKLDKCVESGAKVVLSRLAIGDLATQYFA 307
Query: 301 DAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGV 360
D I + + R+A GG + ++ +N LG CE+ EE +G ++ FSG
Sbjct: 308 DRDIFCAGRVAEEDLNRVAAAAGGTVQTSVNNIIDEVLGTCEIFEEKQVGGERFNIFSGC 367
Query: 361 AMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALA 420
G+ TIVLRG + ++EAERSLHDA+ ++ + V +S V+ GGG +M ++K + +
Sbjct: 368 PSGRTATIVLRGGADQFIEEAERSLHDAIMIVRRAVKNSTVVPGGGAIDMEISKYLRQHS 427
Query: 421 RKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEH---QNEGCTAGIDVIS 477
R GK L + ++++AL IP + DNAG D+ +++++LR +H EG + G+D+ +
Sbjct: 428 RTIAGKSQLFINSYAKALEVIPRQLCDNAGFDATDVLNKLRQKHAMQSGEGASYGVDINT 487
Query: 478 GSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
G + D + E VK + +ATEAA +IL VD+ +
Sbjct: 488 GGIADSFANFVWEPAVVKINAINAATEAACLILSVDETV 526
>At3g20050 t-complex polypeptide 1 homologue
Length = 545
Score = 311 bits (798), Expect = 4e-85
Identities = 186/527 (35%), Positives = 310/527 (58%), Gaps = 31/527 (5%)
Query: 14 EKGERARMSSFVGAMAIADLVKTTLGPKGMDKIL-QSTGRGREVTVTNDGATILKSLHID 72
+ G+ R + + A++++VKT+LGP G+DK+L G +VT+TNDGATIL+ L ++
Sbjct: 15 QSGQDVRTQNVMACQAVSNIVKTSLGPVGLDKMLVDDIG---DVTITNDGATILRMLEVE 71
Query: 73 NPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECA 132
+PAAKVLV+++++QD EVGDGTTSVV++A ELL+ A LV KIHP +II+G+R+A +
Sbjct: 72 HPAAKVLVELAELQDREVGDGTTSVVIVAAELLKRANDLVRNKIHPTSIISGYRLAMRES 131
Query: 133 RSALVEKVVDNKGDAEKF-RSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGSTN- 190
+ EK+V EK + L+N A+T++SSK++S D + FA L V+AV+ +K +
Sbjct: 132 CKYIEEKLVTK---VEKLGKVPLINCAKTSMSSKLISGDSDFFANLVVEAVLSVKMTNQR 188
Query: 191 ------LESVQIIKKPGGSLIDSFLDEGFILDK-KIGLGQPKRIENAKILVANTAMDTDK 243
++ + I+K G S DS+L G+ L+ + G P R+ AKI + + K
Sbjct: 189 GEIKYPIKGINILKAHGQSARDSYLLNGYALNTGRAAQGMPLRVSPAKIACLDFNLQKTK 248
Query: 244 VKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAG 303
+++ G +V V+ ++ +I E + KE++ K++ G N + + I + + F +AG
Sbjct: 249 MQL-GVQVVVNDPRELEKIRQREADMTKERIEKLLKAGANVILTTKGIDDMALKYFVEAG 307
Query: 304 ILAIEHADFDGIERLALVTGGEIASTFDNPES------VKLGQCELIEEIMIGEDKLIKF 357
+A+ + + +A TG + +TF + E LG + + E I +D +I
Sbjct: 308 AIAVRRVRKEDMRHVAKATGATLVTTFADMEGEETFDPAHLGSADEVVEERIADDDVILI 367
Query: 358 SGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEID 417
G A +++LRGA+ ++LDE ER+LHDALC++ +T+ + V+ GGG E ++ ++
Sbjct: 368 KGTKTSSAVSLILRGANDYMLDEMERALHDALCIVKRTLESNTVVAGGGAVESALSVYLE 427
Query: 418 ALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTA------ 471
LA ++ LA+ F+ ALL IP +A NA D+ EL+++LRA H A
Sbjct: 428 HLATTLGSREQLAIAEFADALLIIPKVLAVNAAKDATELVAKLRAYHHTAQTKADKKHYS 487
Query: 472 --GIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
G+D+++G++ + +E G+ E K ++ ATEAA ILR+DD+I
Sbjct: 488 SMGLDLVNGTIRNNLEAGVIEPAMSKVKIIQFATEAAITILRIDDMI 534
>At3g18190 chaperonin subunit, putative
Length = 536
Score = 293 bits (749), Expect = 2e-79
Identities = 183/517 (35%), Positives = 292/517 (56%), Gaps = 19/517 (3%)
Query: 14 EKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDN 73
++ E R ++ A A++D V+T+LGPKGMDK++ ST G EV +TNDGATIL + +
Sbjct: 24 KRREDIRFANINSARAVSDAVRTSLGPKGMDKMI-STANG-EVIITNDGATILNKMEVLQ 81
Query: 74 PAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECAR 133
PAAK+LV++SK QD GDGTT+VVV+AG LL+E + L+ IHP I A A
Sbjct: 82 PAAKMLVELSKSQDSAAGDGTTTVVVIAGALLKECQSLLTNGIHPTVISDSLHKACGKAI 141
Query: 134 SALVEKVVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGS----- 188
L V + R L+ A T+L+SK++SQ A LAVDAV+ +
Sbjct: 142 DILTAMAVPVELTD---RDSLVKSASTSLNSKVVSQYSTLLAPLAVDAVLSVIDPEKPEI 198
Query: 189 TNLESVQIIKKPGGSLIDSFLDEGFILDKKIG--LGQPKRIENAKILVANTAMDTDKVKI 246
+L ++I+KK GG++ D+ +G + DKK+ G P R+ENAKI V + K I
Sbjct: 199 VDLRDIKIVKKLGGTVDDTHTVKGLVFDKKVSRAAGGPTRVENAKIAVIQFQISPPKTDI 258
Query: 247 YGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCF-----VNRQLIYNFPEELFAD 301
+ V V ++ I E+ + + KI G N + R + + A
Sbjct: 259 EQSIV-VSDYTQMDRILKEERNYILGMIKKIKATGCNVLLIQKSILRDAVTDLSLHYLAK 317
Query: 302 AGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVA 361
A I+ I+ + D IE + + ++ + KLG +L+EE +G+ K++K +G+
Sbjct: 318 AKIMVIKDVERDEIEFVTKTLNCLPIANIEHFRAEKLGHADLVEEASLGDGKILKITGIK 377
Query: 362 -MGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALA 420
MG+ ++++RG++ VLDEAERSLHDALCV+ V+ ++ GGG PE+ +++++ A A
Sbjct: 378 DMGRTTSVLVRGSNQLVLDEAERSLHDALCVVRCLVSKRFLIAGGGAPEIELSRQLGAWA 437
Query: 421 RKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSV 480
+ G + +++F+ AL IP T+A+NAGL+ ++++LR +H AGI+V G +
Sbjct: 438 KVLHGMEGYCVKSFAEALEVIPYTLAENAGLNPIAIVTELRNKHAQGEINAGINVRKGQI 497
Query: 481 GDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIIT 517
+++E + + V + + ATE MIL++DDI+T
Sbjct: 498 TNILEENVVQPLLVSTSAITLATECVRMILKIDDIVT 534
>At1g24510 T-complex chaperonin protein , epsilon subunit
Length = 535
Score = 276 bits (707), Expect = 2e-74
Identities = 173/524 (33%), Positives = 291/524 (55%), Gaps = 24/524 (4%)
Query: 8 KDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILK 67
+D+ + +G A+ ++ A+A +++++LGPKGMDK+LQ G ++T+TNDGATIL+
Sbjct: 18 QDQKTRLRGIDAQKANIAAGKAVARILRSSLGPKGMDKMLQ--GPDGDITITNDGATILE 75
Query: 68 SLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRM 127
+ +DN AK++V++S+ QD E+GDGTT VVV+AG LL +AE+ + IHP+ I G+ M
Sbjct: 76 QMDVDNQIAKLMVELSRSQDYEIGDGTTGVVVMAGALLEQAERQLDRGIHPIRIAEGYEM 135
Query: 128 AAECARSALVEKVVDNKGDAEKFRSD------LMNIARTTLSSKILSQDKEHFAKLAVDA 181
A+ A L E++ A+KF D L+ TTLSSKI+++ K A++AV A
Sbjct: 136 ASRVAVEHL-ERI------AQKFEFDVNNYEPLVQTCMTTLSSKIVNRCKRSLAEIAVKA 188
Query: 182 VMRL----KGSTNLESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQ-PKRIENAKILVAN 236
V+ + + NL+ +++ K GG L D+ L G ++DK + Q PK+IE+A I +
Sbjct: 189 VLAVADLERRDVNLDLIKVEGKVGGKLEDTELIYGILIDKDMSHPQMPKQIEDAHIAILT 248
Query: 237 TAMDTDKVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPE 296
+ K K +V +D++ K + E++ E V K G + + +
Sbjct: 249 CPFEPPKPKT-KHKVDIDTVEKFETLRKQEQQYFDEMVQKCKDVGATLVICQWGFDDEAN 307
Query: 297 ELFADAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIG--EDKL 354
L + A+ +E +A+ TGG I F KLG+ ++ E G ++++
Sbjct: 308 HLLMHRNLPAVRWVGGVELELIAIATGGRIVPRFQELTPEKLGKAGVVREKSFGTTKERM 367
Query: 355 IKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAK 414
+ A +A T+ +RG + +++E +RS+HDALCV + + ++ GGG E+ +
Sbjct: 368 LYIEHCANSKAVTVFIRGGNKMMIEETKRSIHDALCVARNLIRNKSIVYGGGAAEIACSL 427
Query: 415 EIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCT-AGI 473
+DA A K PG + A+ AF+ AL ++P +A+N+GL E +S ++++ E GI
Sbjct: 428 AVDAAADKYPGVEQYAIRAFAEALDSVPMALAENSGLQPIETLSAVKSQQIKENIPFYGI 487
Query: 474 DVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIIT 517
D DM E+ + E KQ +L AT+ +MIL++DD+I+
Sbjct: 488 DCNDVGTNDMREQNVFETLIGKQQQILLATQVVKMILKIDDVIS 531
>At5g16070 TCP-1 chaperonin-like protein
Length = 535
Score = 221 bits (563), Expect = 8e-58
Identities = 158/514 (30%), Positives = 266/514 (51%), Gaps = 23/514 (4%)
Query: 27 AMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAAKVLVDISKVQ 86
A + D++K+ LGPKG K+L G ++ +T DG T+LK + I NP A ++ + Q
Sbjct: 26 AKGLQDVLKSNLGPKGTIKML--VGGSGDIKLTKDGNTLLKEMQIQNPTAIMIARTAVAQ 83
Query: 87 DDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVE-KVVDNKG 145
DD GDGTTS V+ GEL++++E+ + +HP ++ GF +A L K G
Sbjct: 84 DDISGDGTTSTVIFIGELMKQSERCIDEGMHPRVLVDGFEIAKRATLQFLDNFKTPVVMG 143
Query: 146 DAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLK---GSTNLESVQIIKKPGG 202
D E + L +ARTTL +K+ + + V++V+ ++ + +L V+I+
Sbjct: 144 D-EVDKEILKMVARTTLRTKLYEGLADQLTDIVVNSVLCIRKPEEAIDLFMVEIMHMRHK 202
Query: 203 SLIDSFLDEGFILDKKIGLGQP---KRIENAKILVANTAMDTDKVKIYGARVRVDSMAKV 259
+D+ L EG +LD G P +R EN IL N +++ +K +I ++ +
Sbjct: 203 FDVDTRLVEGLVLDH--GSRHPDMKRRAENCHILTCNVSLEYEKSEINAGFFYSNAEQRE 260
Query: 260 AEIEGAEKEKMKEKVNKII-------GHGINCFVNRQLIYNFPE-ELFADAGILAIEHAD 311
A + AE+ + E+V KII G N V Q + P +L A GI+ + A
Sbjct: 261 AMVT-AERRSVDERVKKIIELKKKVCGDNDNFVVINQKGIDPPSLDLLAREGIIGLRRAK 319
Query: 312 FDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQACTIVLR 371
+ERL L GGE ++ D+ LG L+ E ++GE+K V +CTI+++
Sbjct: 320 RRNMERLVLACGGEAVNSVDDLTPESLGWAGLVYEHVLGEEKYTFVEQVKNPNSCTILIK 379
Query: 372 GASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKT-PGKKSLA 430
G + H + + + ++ D L + T+ D V+LG G E+ + + +KT G+ L
Sbjct: 380 GPNDHTIAQIKDAVRDGLRSVKNTIEDECVVLGAGAFEVAARQHLLNEVKKTVQGRAQLG 439
Query: 431 MEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICE 490
+EAF+ ALL +P T+A+NAGLD+ ++I L +EH ++G G+++ G D GI +
Sbjct: 440 VEAFANALLVVPKTLAENAGLDTQDVIISLTSEH-DKGNVVGLNLQDGEPIDPQLAGIFD 498
Query: 491 AFKVKQAVLLSATEAAEMILRVDDIITCAPRRRE 524
+ VK+ ++ S A +L VD++I R+
Sbjct: 499 NYSVKRQLINSGPVIASQLLLVDEVIRAGRNMRK 532
>At3g02530 putative chaperonin
Length = 535
Score = 221 bits (562), Expect = 1e-57
Identities = 159/515 (30%), Positives = 267/515 (50%), Gaps = 24/515 (4%)
Query: 27 AMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAAKVLVDISKVQ 86
A + D++K+ LGPKG K+L G ++ +T DG T+LK + I NP A ++ + Q
Sbjct: 26 AKGLQDVLKSNLGPKGTIKML--VGGSGDIKLTKDGNTLLKEMQIQNPTAIMIARTAVAQ 83
Query: 87 DDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSAL-VEKVVDNKG 145
DD GDGTTS V+ GEL++++E+ + +HP ++ GF +A L K G
Sbjct: 84 DDISGDGTTSTVIFIGELMKQSERCIDEGMHPRVLVDGFEIAKRATLQFLDTFKTPVVMG 143
Query: 146 DAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLK---GSTNLESVQIIKKPGG 202
D E + L +ARTTL +K+ + + V++V+ ++ +L V+I+
Sbjct: 144 D-EPDKEILKMVARTTLRTKLYEGLADQLTDIVVNSVLCIRKPQEPIDLFMVEIMHMRHK 202
Query: 203 SLIDSFLDEGFILDKKIGLGQP---KRIENAKILVANTAMDTDKVKIYGARVRVDSMAKV 259
+D+ L EG +LD G P +R EN IL N +++ +K +I ++ +
Sbjct: 203 FDVDTRLVEGLVLDH--GSRHPDMKRRAENCHILTCNVSLEYEKSEINAGFFYSNAEQRE 260
Query: 260 AEIEGAEKEKMKEKV-------NKIIGHGINCFV--NRQLIYNFPEELFADAGILAIEHA 310
A + AE+ + E+V NK+ N FV N++ I +L A GI+A+ A
Sbjct: 261 AMVT-AERRSVDERVQKIIELKNKVCAGNDNSFVILNQKGIDPPSLDLLAREGIIALRRA 319
Query: 311 DFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQACTIVL 370
+ERL L GGE ++ D+ LG L+ E ++GE+K V +CTI++
Sbjct: 320 KRRNMERLVLACGGEAVNSVDDLTPDCLGWAGLVYEHVLGEEKYTFVEQVKNPHSCTILI 379
Query: 371 RGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKT-PGKKSL 429
+G + H + + + ++ D L + T+ D V+LG G E+ + + +KT G+ L
Sbjct: 380 KGPNDHTIAQIKDAVRDGLRSVKNTLEDECVVLGAGAFEVAARQHLINEVKKTVQGRAQL 439
Query: 430 AMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGIC 489
+EAF+ ALL +P T+A+NAGLD+ ++I L +EH ++G G+D+ G D GI
Sbjct: 440 GVEAFANALLVVPKTLAENAGLDTQDVIISLTSEH-DKGNIVGLDLQDGEPVDPQLAGIF 498
Query: 490 EAFKVKQAVLLSATEAAEMILRVDDIITCAPRRRE 524
+ + VK+ ++ S A +L VD++I R+
Sbjct: 499 DNYSVKRQLINSGPVIASQLLLVDEVIRAGRNMRK 533
>At5g26360 chaperonin gamma chain - like protein
Length = 555
Score = 219 bits (559), Expect = 2e-57
Identities = 155/526 (29%), Positives = 271/526 (51%), Gaps = 22/526 (4%)
Query: 6 IFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATI 65
+ D E G + + + A+AD+++TTLGP+ M K+L G G + VTNDG I
Sbjct: 7 VLSDSLKRESGSKVHHGNIQASKAVADIIRTTLGPRSMLKMLLDAGGG--IVVTNDGNAI 64
Query: 66 LKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGF 125
L+ L + +PAAK ++++S+ QD+EVGDGTTSV+VLAGE+L AE + HP I +
Sbjct: 65 LRELDVAHPAAKSMIELSRTQDEEVGDGTTSVIVLAGEMLHVAEAFLEKNYHPTVICRAY 124
Query: 126 RMAAECARSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAV--- 182
A E + A+++K+ + + RS ++ + ++ + +K SQ + A LA+DA
Sbjct: 125 IKALEDS-IAVLDKIAMSIDIND--RSQVLGLVKSCIGTKFTSQFGDLIADLAIDATTTV 181
Query: 183 -----MRLKGSTNLESVQIIKKPGGSLIDSFLDEGFILDKK-IGLGQPKR-IENAKILVA 235
L+ + +++ K PGG DS + +G + +K + G+ KR I N +I++
Sbjct: 182 GVDLGQGLREVDIKKYIKVEKVPGGQFEDSEVLKGVMFNKDVVAPGKMKRKIVNPRIILL 241
Query: 236 NTAMDTDK--VKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYN 293
+ ++ K + VR + + ++ E+E ++ +I+ + + + + +
Sbjct: 242 DCPLEYKKGENQTNAELVREEDWEVLLKL---EEEYIENICVQILKFKPDLVITEKGLSD 298
Query: 294 FPEELFADAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQ-CELIEEIMIGED 352
F+ AG+ AI R+A G I + D + +G L E IG+D
Sbjct: 299 LACHYFSKAGVSAIRRLRKTDNNRIAKACGAVIVNRPDELQESDIGTGAGLFEVKKIGDD 358
Query: 353 KLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVM 412
+ACT++LRG S ++E ER+L DA+ V + + +++ GGG E+ +
Sbjct: 359 FFSFIVDCKEPKACTVLLRGPSKDFINEVERNLQDAMSVARNIIKNPKLVPGGGATELTV 418
Query: 413 AKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQN-EGCTA 471
+ + + G + EA + A AIP T+A N G++ ++ L+ +H N E
Sbjct: 419 SATLKQKSATIEGIEKWPYEAAAIAFEAIPRTLAQNCGVNVIRTMTALQGKHANGENAWT 478
Query: 472 GIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIIT 517
GID +G++ DM E I +++ VK +A EAA M+LR+DDI++
Sbjct: 479 GIDGNTGAIADMKESKIWDSYNVKAQTFKTAIEAACMLLRIDDIVS 524
>At3g03960 putative T-complex protein 1, theta subunit (TCP-1-Theta)
Length = 549
Score = 197 bits (501), Expect = 1e-50
Identities = 146/511 (28%), Positives = 250/511 (48%), Gaps = 13/511 (2%)
Query: 17 ERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAA 76
+ A + + ++ + +T+LGP GM+K++ ++ VTND ATI+ L I +PAA
Sbjct: 26 DEAVIKNIEACKELSTITRTSLGPNGMNKMV--INHLDKLFVTNDAATIVNELEIQHPAA 83
Query: 77 KVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSAL 136
K+LV +K Q +E+GDG + AGELL+ AE+L+ +HP II+G+ A A L
Sbjct: 84 KLLVLAAKAQQEEIGDGANLTISFAGELLQNAEELIRMGLHPSEIISGYTKAVSKAVEIL 143
Query: 137 VEKVVDNKGDAEKFRS--DLMNIARTTLSSKILSQDKEHFAKLAVDAVMRL--KGSTN-- 190
E++V+ + R+ ++++ R ++SK Q+ E L DA +++ K TN
Sbjct: 144 -EQLVETGSETMDVRNKDEVISRMRAAVASKQFGQE-EIICSLVTDACIQVCPKNPTNFN 201
Query: 191 LESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMDTDKVKIYGAR 250
+++V++ K GG L +S + G +L K +G KR+E AK+ V +DT + G
Sbjct: 202 VDNVRVSKLLGGGLHNSCIVRGMVL-KSDAVGSIKRMEKAKVAVFAGGVDTTATETKGT- 259
Query: 251 VRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHA 310
V + S ++ E+ K++E + + G V+ I I+ ++ +
Sbjct: 260 VLIHSAEQLENYAKTEEAKVEELIKAVAESGAKVIVSGGSIGEMALHFCERYKIMVLKIS 319
Query: 311 DFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQAC-TIV 369
+ R G P LG + I IG + G + T+V
Sbjct: 320 SKFELRRFCRTAGAVAHLKLSRPSPEDLGYVDSISVEEIGGVTVTIARNEEGGNSISTVV 379
Query: 370 LRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSL 429
LRG++ +LD+ ER++ D + DSR++ G E+ +A+ + A G
Sbjct: 380 LRGSTDSILDDLERAVDDGVNTYKAMCRDSRIVPGAAATEIELAQRLKEYANAEIGLDKY 439
Query: 430 AMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGIC 489
A+ ++ + +P T+ADNAGL++ E+I+ L H + GID+ G+ D+ E +
Sbjct: 440 AITKYAESFEFVPKTLADNAGLNAMEIIAALYTGHGSGNTKLGIDLEEGACKDVSETKVW 499
Query: 490 EAFKVKQAVLLSATEAAEMILRVDDIITCAP 520
+ F K L A++AA +LRVD II P
Sbjct: 500 DLFATKLFALKYASDAACTVLRVDQIIMAKP 530
>At3g23990 mitochondrial chaperonin hsp60
Length = 577
Score = 109 bits (272), Expect = 4e-24
Identities = 136/556 (24%), Positives = 227/556 (40%), Gaps = 87/556 (15%)
Query: 16 GERARMSSFVGAMAIADLVKTTLGPKGMDKIL-QSTGRGREVTVTNDGATILKSLH---- 70
G AR G +AD VK T+GPKG + ++ QS G + VT DG T+ KS+
Sbjct: 39 GVEARALMLKGVEDLADAVKVTMGPKGRNVVIEQSWGAPK---VTKDGVTVAKSIEFKDK 95
Query: 71 IDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAE 130
I N A ++ ++ +D GDGTT VL + E K VAA ++ M + G MA
Sbjct: 96 IKNVGASLVKQVANATNDVAGDGTTCATVLTRAIFAEGCKSVAAGMNAMDLRRGISMA-- 153
Query: 131 CARSALVEKVVDNKGDAEKFRSDLMNIART-TLSSKILSQDKEHFAKLAVDAVMRLKGST 189
V+ VV N + S IA+ T+S+ + E AK M G
Sbjct: 154 ------VDAVVTNLKSKARMISTSEEIAQVGTISANGEREIGELIAK-----AMEKVGKE 202
Query: 190 NLESVQIIKKPGGSLIDSF-LDEGFILDKKIGLGQPKRIENAKILVANTAMDTDKVKIYG 248
+ ++Q G +L + + EG LD+ G P I N K +D + I+
Sbjct: 203 GVITIQ----DGKTLFNELEVVEGMKLDR--GYTSPYFITNQK--TQKCELDDPLILIHE 254
Query: 249 ARV-RVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAI 307
++ ++S+ KV E+ +++ I+ + LI N +L A + AI
Sbjct: 255 KKISSINSIVKVLEL-----ALKRQRPLLIVSEDVESDALATLILN---KLRAGIKVCAI 306
Query: 308 EHADFD-----GIERLALVTGGEIAS--TFDNPESVKLGQCELIEEIMIGEDKLIKFSGV 360
+ F ++ LA +TGGE+ + N E V L +++ + +D + G
Sbjct: 307 KAPGFGENRKANLQDLAALTGGEVITDELGMNLEKVDLSMLGTCKKVTVSKDDTVILDGA 366
Query: 361 A-----------------------------------MGQACTIVLRGASHHVLDEAERSL 385
G + + GAS + E + +
Sbjct: 367 GDKKGIEERCEQIRSAIELSTSDYDKEKLQERLAKLSGGVAVLKIGGASEAEVGEKKDRV 426
Query: 386 HDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTI 445
DAL V + +L GGG + A+E++ L +K + ++ AL TI
Sbjct: 427 TDALNATKAAVEEG-ILPGGGVALLYAARELEKLPTANFDQK-IGVQIIQNALKTPVYTI 484
Query: 446 ADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEA 505
A NAG++ A ++ +L + + G D G DMV+ GI + KV + L+ A
Sbjct: 485 ASNAGVEGAVIVGKLL---EQDNPDLGYDAAKGEYVDMVKAGIIDPLKVIRTALVDAASV 541
Query: 506 AEMILRVDDIITCAPR 521
+ ++ + ++ P+
Sbjct: 542 SSLLTTTEAVVVDLPK 557
>At2g33210 mitochondrial chaperonin (HSP60)
Length = 585
Score = 108 bits (270), Expect = 7e-24
Identities = 134/560 (23%), Positives = 233/560 (40%), Gaps = 93/560 (16%)
Query: 16 GERARMSSFVGAMAIADLVKTTLGPKGMDKIL-QSTGRGREVTVTNDGATILKSLH---- 70
G AR G +AD VK T+GPKG + I+ QS G + VT DG T+ KS+
Sbjct: 40 GVEARALMLRGVEDLADAVKVTMGPKGRNVIIEQSWGAPK---VTKDGVTVAKSIEFKDR 96
Query: 71 IDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAE 130
I N A ++ ++ +D GDGTT VL + E K VAA ++ M + G ++A
Sbjct: 97 IKNVGASLVKQVANATNDVAGDGTTCATVLTRAIFTEGCKSVAAGMNAMDLRRGIKLA-- 154
Query: 131 CARSALVEKVVDNKGDAEKFRSDLMNIART-TLSSKILSQDKEHFAKLAVDAVMRLKGST 189
V+ VV N + S IA+ T+S+ + E AK M G
Sbjct: 155 ------VDTVVTNLQSRARMISTSEEIAQVGTISANGDREIGELIAK-----AMETVGKE 203
Query: 190 NLESVQIIKKPGGSLIDSF-LDEGFILDKKIGLGQPKRIENAKILVANTAMDTDKVKIYG 248
+ ++Q G +L + + EG +D+ G P I N K ++ + I+
Sbjct: 204 GVITIQ----DGKTLFNELEVVEGMKIDR--GYISPYFITNPK--TQKCELEDPLILIHE 255
Query: 249 ARV-RVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAI 307
++ +++M KV E+ K++ I+ + LI N +L A+ + A+
Sbjct: 256 KKISNINAMVKVLEL-----ALKKQRPLLIVAEDVESDALATLILN---KLRANIKVCAV 307
Query: 308 EHADFD-----GIERLALVTGG-----EIASTFDNPESVKLGQCELIEEIMIGEDKLIKF 357
+ F + LA +TG E+ DN + G C +++ + +D +
Sbjct: 308 KAPGFGENRKANLHDLAALTGAQVITEELGMNLDNIDLSMFGNC---KKVTVSKDDTVVL 364
Query: 358 SGV----AMGQAC-------------------------------TIVLRGASHHVLDEAE 382
G A+G+ C + + GAS + E +
Sbjct: 365 DGAGDKQAIGERCEQIRSMVEASTSDYDKEKLQERLAKLSGGVAVLKIGGASETEVSEKK 424
Query: 383 RSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIP 442
+ DAL V + ++ GGG + +KE++ L+ +K + ++ AL
Sbjct: 425 DRVTDALNATKAAVEEG-IVPGGGVALLYASKELEKLSTANFDQK-IGVQIIQNALKTPV 482
Query: 443 TTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSA 502
TIA NAG++ A ++ +L + + G D G DM++ GI + KV + L+ A
Sbjct: 483 YTIASNAGVEGAVVVGKLL---EQDNPDLGYDAAKGEYVDMIKAGIIDPLKVIRTALVDA 539
Query: 503 TEAAEMILRVDDIITCAPRR 522
+ ++ + ++T P +
Sbjct: 540 ASVSSLLTTTEAVVTEIPTK 559
>At3g13860 chaperonin like protein
Length = 572
Score = 97.8 bits (242), Expect = 1e-20
Identities = 127/540 (23%), Positives = 218/540 (39%), Gaps = 79/540 (14%)
Query: 16 GERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHID--- 72
G AR + G +A+ VK T+GPKG + I++S+ G ++T DG T+ KS+
Sbjct: 39 GIGARAAMLQGVSEVAEAVKVTMGPKGRNVIIESSYGGPKIT--KDGVTVAKSISFQAKA 96
Query: 73 -NPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAEC 131
N A+++ ++ + GDGTT VL +L E K VAA ++ M + G MA
Sbjct: 97 KNIGAELVKQVASATNKVAGDGTTCATVLTQAILIEGCKSVAAGVNVMDLRVGINMAIAA 156
Query: 132 ARSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGSTNL 191
S L + V E + ++ +++++ E K V V G+T
Sbjct: 157 VVSDLKSRAVMISTPEEITQVATISANGEREIGELIARAMEKVGKEGVITV--ADGNTLD 214
Query: 192 ESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMDTDKVKIYGARV 251
+++++ G L ++ FI D+K Q +EN IL+ +
Sbjct: 215 NELEVVE--GMKLARGYISPYFITDEKT---QKCELENPIILIHEKKISD---------- 259
Query: 252 RVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHAD 311
++S+ KV +E A K + I+ + LI N + + AI+
Sbjct: 260 -INSLLKV--LEAAVK---SSRPLLIVAEDVESDALAMLILN---KHHGGLKVCAIKAPG 310
Query: 312 FD-----GIERLALVTGGEIAS---------------------TFDNPESVKL---GQCE 342
F ++ LA++TG E+ S T +++ L G +
Sbjct: 311 FGDNRKASLDDLAVLTGAEVISEERGLSLEKIRPELLGTAKKVTVTRDDTIILHGGGDKK 370
Query: 343 LIEE-------------IMIGEDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDAL 389
LIEE ++K + G + GAS + E + + DAL
Sbjct: 371 LIEERCEELRSANEKSTSTFDQEKTQERLSKLSGGVAVFKVGGASESEVGERKDRVTDAL 430
Query: 390 CVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNA 449
V + ++ GGG + K +D L + ++ ++ AL A TIA NA
Sbjct: 431 NATRAAVEEG-IIPGGGVALLYATKALDNLQTENEDQRR-GVQIVQNALKAPAFTIAANA 488
Query: 450 GLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMI 509
G D + ++ +L + + C G D G DMV+ GI + KV + L A + ++
Sbjct: 489 GYDGSLVVGKLL---EQDDCNFGFDAAKGKYVDMVKAGIIDPVKVIRTALTDAASVSLLL 545
>At2g28000 putative rubisco subunit binding-protein alpha subunit
Length = 586
Score = 91.3 bits (225), Expect = 1e-18
Identities = 111/536 (20%), Positives = 228/536 (41%), Gaps = 49/536 (9%)
Query: 17 ERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHI----D 72
+ +R + G +AD V TLGP+G + +L G + V NDG TI +++ + +
Sbjct: 55 QHSRAALQAGIDKLADCVGLTLGPRGRNVVLDEFGSPK---VVNDGVTIARAIELPNAME 111
Query: 73 NPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECA 132
N A ++ +++ +D GDGTT+ +LA E+++ V + +P+++ G +
Sbjct: 112 NAGAALIREVASKTNDSAGDGTTTASILAREIIKHGLLSVTSGANPVSLKRGIDKTVQ-- 169
Query: 133 RSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKIL--SQDKEHFAKLAVDAVMRLKGSTN 190
L+E+ + K K R D+ +A + + L S + K+ D V+ ++ S++
Sbjct: 170 --GLIEE-LQKKARPVKGRDDIRAVASISAGNDDLIGSMIADAIDKVGPDGVLSIESSSS 226
Query: 191 LESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAM----DTDKVKI 246
E+ +++ G + ++ F+ + + L + ENA++L+ + + D +
Sbjct: 227 FETTVEVEE-GMEIDRGYISPQFVTNPEKLLAE---FENARVLITDQKITAIKDIIPILE 282
Query: 247 YGARVRVDSMAKVAEIEGAE-----KEKMKEKVNKI------IGHGINCFVNRQLIYNFP 295
++R + ++ G K++ +N + G + I
Sbjct: 283 KTTQLRAPLLIIAEDVTGEALATLVVNKLRGVLNVVAVKAPGFGERRKAMLQDIAILTGA 342
Query: 296 EELFADAGILAIEHADFD--GIERLALVTG------GEIASTFDNPESVKLGQCELIE-E 346
E L D +L +E+A D GI R ++ + AS + + + EL E +
Sbjct: 343 EYLAMDMSLL-VENATIDQLGIARKVTISKDSTTLIADAASKDELQARIAQLKKELFETD 401
Query: 347 IMIGEDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGG 406
+ +KL + G I + A+ L++ + + DA + + ++ GGG
Sbjct: 402 SVYDSEKLAERIAKLSGGVAVIKVGAATETELEDRKLRIEDAKNATFAAIEEG-IVPGGG 460
Query: 407 WPEMVMAKEIDALARK-TPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQ 465
+ ++ I A+ + L + +ALL+ IA NAG++ ++ ++
Sbjct: 461 AALVHLSTVIPAIKETFEDADERLGADIVQKALLSPAALIAQNAGVEGEVVVEKIMFSDW 520
Query: 466 NEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPR 521
G A D ++ E G+ + KV + L +A A M+L I+ P+
Sbjct: 521 ENGYNAMTDTYE----NLFEAGVIDPAKVTRCALQNAASVAGMVLTTQAIVVDKPK 572
>At5g18820 chaperonin 60 alpha chain - like protein
Length = 575
Score = 86.3 bits (212), Expect = 4e-17
Identities = 117/530 (22%), Positives = 224/530 (42%), Gaps = 44/530 (8%)
Query: 16 GERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLH----I 71
G+ +R G +AD V TLGP+G + +L + V NDG TI KS+ I
Sbjct: 41 GKDSREKLQAGIDKLADAVSITLGPRGRNVVLAEKDT---IKVINDGVTIAKSIELPDTI 97
Query: 72 DNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGF-RMAAE 130
+N A ++ +++ ++ GDGTT+ ++LA E+++ +A + +++ G + E
Sbjct: 98 ENAGATLIQEVAIKMNESAGDGTTTAIILAREMIKAGSLAIAFGANAVSVKNGMNKTVKE 157
Query: 131 CARSALVEKV-VDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGST 189
R ++ + V K D + S ++ ++++ E K+ D V+ ++ S+
Sbjct: 158 LVRVLQMKSIPVQGKNDIKAVAS--ISAGNDEFVGNLIAETVE---KIGPDGVISIESSS 212
Query: 190 NLESVQIIKKPGGSLIDSFLDEGFI---------LDKKIGLGQPKRIENAKILVANTAMD 240
E+ +I + G ++ FI DK L ++I +AK LV
Sbjct: 213 TSET-SVIVEEGMKFDKGYMSPHFITNQEKSTVEFDKAKILVTDQKITSAKELVP-LLEK 270
Query: 241 TDKVKIYGARVRVDSMAKVAEIEGAEKEK--MKEKVNKIIG--HGINCFVNRQLIYNFPE 296
T ++ + + D A+V EI K++ + V K G G + + +
Sbjct: 271 TSQLSVPLLIIAEDISAEVLEILVVNKKQGLINVAVVKCPGMLDGKKALLQDIALMTGAD 330
Query: 297 ELFADAGI-LAIEHADFDGIERLALVTGGE---IASTFDNPE-SVKLGQC--ELIE-EIM 348
L D G+ L +D G+ R ++T +A PE ++ Q +L E +
Sbjct: 331 YLSGDLGMSLMGATSDQLGVSRRVVITANSTTIVADASTKPEIQARIAQMKKDLAETDNS 390
Query: 349 IGEDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWP 408
K+ + G I + G + L++ + + DA + + ++ GGG
Sbjct: 391 YLSKKIAERIAKLTGGVAVIKVGGHTETELEDRKLRIEDAKNATFAAMREG-IVPGGGAT 449
Query: 409 EMVMAKEIDALARK--TPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQN 466
+ + EI + + + + + + AL A IA NAG+D + ++ + R
Sbjct: 450 YIHLLDEIPRIKKNLMEDSYEQIGADIVAMALTAPAMAIATNAGVDGSVVVQKTRELEWR 509
Query: 467 EGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
G A +SG D++ GI + +V + L +A A +IL ++
Sbjct: 510 SGYNA----MSGKYEDLLNAGIADPCRVSRFALQNAVSVAGIILTTQAVL 555
>At5g56500 RuBisCO subunit binding-protein beta subunit precursor;
chaperonin, 60 kDa
Length = 596
Score = 84.0 bits (206), Expect = 2e-16
Identities = 122/524 (23%), Positives = 227/524 (43%), Gaps = 52/524 (9%)
Query: 26 GAMAIADLVKTTLGPKGMDKILQST-GRGREVTVTNDGATILKSLHIDNPA----AKVLV 80
G +ADLV TLGPKG + +L+S G R + NDG T+ + + +++P AK++
Sbjct: 70 GVNKLADLVGVTLGPKGRNVVLESKYGSPR---IVNDGVTVAREVELEDPVENIGAKLVR 126
Query: 81 DISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVEKV 140
+ +D GDGTT+ VVLA L+ E K+VAA +P+ I G E ALV ++
Sbjct: 127 QAASKTNDLAGDGTTTSVVLAQGLIAEGVKVVAAGANPVLITRGI----EKTTKALVAEL 182
Query: 141 VDNKGDAEKFRSDLMNIARTTLSS--KILSQDKEHFAKLAVDAVMRL-KGSTNLESVQII 197
K E S+L ++A + + ++ + E AK+ V+ L +G + S+ ++
Sbjct: 183 --KKMSKEVEDSELADVAAVSAGNNYEVGNMIAEAMAKVGRKGVVTLEEGKSAENSLYVV 240
Query: 198 KKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMDT--DKVKIYGARVRVDS 255
+ G ++ F+ D + + EN K+ + + + D + I ++
Sbjct: 241 E--GMQFDRGYISPYFVTDSEKMCAE---YENCKLFLVDKKITNARDIISILEDAIK-GG 294
Query: 256 MAKVAEIEGAEKEKMKE-KVNKIIG----HGINCFVNRQLIYNFPEELFADAGILAIEHA 310
+ E E+E + VNK+ G + + + +++ A G I
Sbjct: 295 YPLLIIAEDIEQEPLATLVVNKLRGTIKVAALKAPGFGERKSQYLDDIAALTGATVIREE 354
Query: 311 DFDGIERL---ALVTGGEIASTFDNPESVKLGQCE------------LIE--EIMIGEDK 353
+E++ L G++ T D V G E LIE E ++K
Sbjct: 355 VGLQLEKVGPEVLGNAGKVVLTKDTTTIVGDGSTEEVVKKRVEQIKNLIEAAEQDYEKEK 414
Query: 354 LIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMA 413
L + G I + + L E + + DAL V + +++GGG + +A
Sbjct: 415 LNERIAKLSGGVAVIQVGAQTETELKEKKLRVEDALNATKAAVEEG-IVVGGGCTLLRLA 473
Query: 414 KEIDALARKTPG-KKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAG 472
++DA+ ++ + + +AL IA NAG++ + + ++ + ++ G
Sbjct: 474 SKVDAIKETLANDEEKVGADIVKKALSYPLKLIAKNAGVNGSVVSEKVLS---SDNPKHG 530
Query: 473 IDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
+ +G D++ GI + KV + L A+ A+ L D ++
Sbjct: 531 YNAATGKYEDLMAAGIIDPTKVVRCCLEHASSVAKTFLMSDCVV 574
>At3g13470 chaperonin 60 beta - like protein
Length = 596
Score = 84.0 bits (206), Expect = 2e-16
Identities = 117/530 (22%), Positives = 223/530 (42%), Gaps = 56/530 (10%)
Query: 26 GAMAIADLVKTTLGPKGMDKILQST-GRGREVTVTNDGATILKSLHIDNPA----AKVLV 80
G +ADLV TLGPKG + +L+S G R + NDG T+ + + +++P AK++
Sbjct: 70 GVNKLADLVGVTLGPKGRNVVLESKYGSPR---IVNDGVTVAREVELEDPVENIGAKLVR 126
Query: 81 DISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVEKV 140
+ +D GDGTT+ VVLA + E K+VAA +P+ I G A+ ALV ++
Sbjct: 127 QAAAKTNDLAGDGTTTSVVLAQGFIAEGVKVVAAGANPVLITRGIEKTAK----ALVNEL 182
Query: 141 VDNKGDAEKFRSDLMNIARTTLSS--KILSQDKEHFAKLAVDAVMRLKGSTNLESVQIIK 198
+ E S+L ++A + + ++ S E +K+ V+ L+ + E+ +
Sbjct: 183 KLMSKEVED--SELADVAAVSAGNNHEVGSMIAEAMSKVGRKGVVTLEEGKSAEN-NLYV 239
Query: 199 KPGGSLIDSFLDEGFILD-KKIG--------LGQPKRIENAKILVA---NTAMDTDKVKI 246
G ++ F+ D +K+ L K++ NA+ LV + + I
Sbjct: 240 VEGMQFDRGYISPYFVTDSEKMSVEYDNCKLLLVDKKVTNARDLVGVLEDAIRGGYPILI 299
Query: 247 YGARVRVDSMA-----------KVAEIE----GAEKEKMKEKVNKIIGHGINCFVNRQLI 291
+ +++A K+A ++ G K + + + + G + +
Sbjct: 300 IAEDIEQEALATLVVNKLRGTLKIAALKAPGFGERKSQYLDDIAILTGATVIREEVGLSL 359
Query: 292 YNFPEELFADAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGE 351
+E+ +A + + + + + G N V++ E +
Sbjct: 360 DKAGKEVLGNASKVVL-------TKEMTTIVGDGTTQEAVNKRVVQIRNLIEQAEQDYEK 412
Query: 352 DKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMV 411
+KL + G I + + L E + + DAL V + +++GGG +
Sbjct: 413 EKLNERIAKLSGGVAVIQVGAQTETELKEKKLRVEDALNATKAAVEEG-IVVGGGCTLLR 471
Query: 412 MAKEIDALARKTPG-KKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCT 470
+A ++DA+ ++ + E RAL IA NAG++ + + ++ A N+
Sbjct: 472 LASKVDAIKDTLENDEEKVGAEIVKRALSYPLKLIAKNAGVNGSVVSEKVLA---NDNVK 528
Query: 471 AGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAP 520
G + +G D++ GI + KV + L A A+ L D ++ P
Sbjct: 529 FGYNAATGKYEDLMAAGIIDPTKVVRCCLEHAASVAKTFLMSDCVVVEIP 578
>At1g55490 Rubisco subunit binding-protein beta subunit
Length = 600
Score = 83.6 bits (205), Expect = 3e-16
Identities = 120/522 (22%), Positives = 219/522 (40%), Gaps = 48/522 (9%)
Query: 26 GAMAIADLVKTTLGPKGMDKILQST-GRGREVTVTNDGATILKSLHIDNPA----AKVLV 80
G +ADLV TLGPKG + +L+S G R + NDG T+ + + +++P AK++
Sbjct: 74 GVNKLADLVGVTLGPKGRNVVLESKYGSPR---IVNDGVTVAREVELEDPVENIGAKLVR 130
Query: 81 DISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVEKV 140
+ +D GDGTT+ VVLA + E K+VAA +P+ I G A+ ALV ++
Sbjct: 131 QAAAKTNDLAGDGTTTSVVLAQGFIAEGVKVVAAGANPVLITRGIEKTAK----ALVTEL 186
Query: 141 VDNKGDAEKFRSDLMNIARTTL--SSKILSQDKEHFAKLAVDAVMRLKGSTNLESVQIIK 198
K E S+L ++A + + +I + E +K+ V+ L+ + E+ +
Sbjct: 187 --KKMSKEVEDSELADVAAVSAGNNDEIGNMIAEAMSKVGRKGVVTLEEGKSAEN-NLYV 243
Query: 199 KPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMDT--DKVKIYGARVRVDSM 256
G ++ F+ D + +N K+L+ + + D V + +R
Sbjct: 244 VEGMQFDRGYISPYFVTDSE---KMSVEFDNCKLLLVDKKITNARDLVGVLEDAIRGGYP 300
Query: 257 AKVAEIEGAEKEKMKEKVNKIIGH------GINCFVNRQLIYNFPEELFADAGILAIE-H 309
+ + ++ VNK+ G F R+ Y + A ++ E
Sbjct: 301 ILIIAEDIEQEALATLVVNKLRGTLKIAALRAPGFGERKSQYLDDIAILTGATVIREEVG 360
Query: 310 ADFDGIERLALVTGGEIASTFDNPESVKLGQCE------------LIE--EIMIGEDKLI 355
D + L ++ T + V G + LIE E ++KL
Sbjct: 361 LSLDKAGKEVLGNASKVVLTKETSTIVGDGSTQDAVKKRVTQIKNLIEQAEQDYEKEKLN 420
Query: 356 KFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKE 415
+ G I + + L E + + DAL V + +++GGG + +A +
Sbjct: 421 ERIAKLSGGVAVIQVGAQTETELKEKKLRVEDALNATKAAVEEG-IVVGGGCTLLRLASK 479
Query: 416 IDAL-ARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGID 474
+DA+ A ++ + + RAL IA NAG++ + + ++ + N+ G +
Sbjct: 480 VDAIKATLDNDEEKVGADIVKRALSYPLKLIAKNAGVNGSVVSEKVLS---NDNVKFGYN 536
Query: 475 VISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
+G D++ GI + KV + L A A+ L D ++
Sbjct: 537 AATGKYEDLMAAGIIDPTKVVRCCLEHAASVAKTFLMSDCVV 578
>At1g26230 chaperonin precursor, putative
Length = 611
Score = 75.5 bits (184), Expect = 7e-14
Identities = 115/526 (21%), Positives = 224/526 (41%), Gaps = 56/526 (10%)
Query: 26 GAMAIADLVKTTLGPKGMDKILQST-GRGREVTVTNDGATILKSLHIDNPAAKVLVDISK 84
GA +A L+ TLGPKG + +LQ+ G R + NDG T+LK + +++P V V + +
Sbjct: 58 GADMVAKLLGVTLGPKGRNVVLQNKYGPPR---IVNDGETVLKEIELEDPLENVGVKLVR 114
Query: 85 VQ----DDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVEKV 140
+D GDG+T+ ++LA L+ E K+++A +P+ + G E ALV ++
Sbjct: 115 QAGAKTNDLAGDGSTTSIILAHGLITEGIKVISAGTNPIQVARGI----EKTTKALVLEL 170
Query: 141 VDNKGDAEKFRSDLMNIARTTLSS--KILSQDKEHFAKLAVDAVMRL-KGSTNLESVQII 197
+ E +L ++A + + ++ + F ++ V+ + KG + +++I+
Sbjct: 171 KSMSREIED--HELAHVAAVSAGNDYEVGNMISNAFQQVGRTGVVTIEKGKYLVNNLEIV 228
Query: 198 KKPGGSLIDSFLDEGFILDKKIGLGQ---------PKRIENAKIL--VANTAMDTD-KVK 245
+ G +L F+ D++ + K+I N K + + ++A+ + V
Sbjct: 229 E--GMQFNRGYLSPYFVTDRRKREAEFHDCKLLLVDKKITNPKDMFKILDSAVKEEFPVL 286
Query: 246 IYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHG---INCFVNRQLIYNFPEELFADA 302
I + D++A V I K +K K G +C + + + D
Sbjct: 287 IVAEDIEQDALAPV--IRNKLKGNLKVAAIKAPAFGERKSHCLDDLAIFTG--ATVIRDE 342
Query: 303 GILAIEHADFDGI----------ERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGED 352
L++E A + + + +VT G D S E EE +
Sbjct: 343 MGLSLEKAGKEVLGTAKRVLVTKDSTLIVTNGFTQKAVDERVSQIKNLIENTEENF--QK 400
Query: 353 KLIKFSGVAMGQACTIVLRGASHHV-LDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMV 411
K++ + I+ GA V L + + + DAL + + +++GGG +
Sbjct: 401 KILNERVARLSGGIAIIQVGALTQVELKDKQLKVEDALNATKSAIEEG-IVVGGGCALLR 459
Query: 412 MAKEIDALARKTPG-KKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCT 470
+A ++D + ++ + E F +AL IA NA + +I ++ + N+
Sbjct: 460 LATKVDRIKETLDNTEQKIGAEIFKKALSYPIRLIAKNADTNGNIVIEKVLS---NKNTM 516
Query: 471 AGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
G + D++ GI + KV + L A+ A+ L D ++
Sbjct: 517 YGYNAAKNQYEDLMLAGIIDPTKVVRCCLEHASSVAQTFLTSDCVV 562
>At4g33240 putative protein
Length = 1757
Score = 46.2 bits (108), Expect = 4e-05
Identities = 43/201 (21%), Positives = 81/201 (39%), Gaps = 25/201 (12%)
Query: 200 PGGSLIDSFLDEGFILDKKIGLGQ-PKRIENAKILVANTAMDTDKVKIYGARVRVDSMAK 258
P G +S + +G + K + + +IE ++L+ A++ ++ + ++
Sbjct: 434 PCGRRSESMVVKGVVCKKNVAHRRMTSKIEKPRLLILGGALEYQRIS--------NQLSS 485
Query: 259 VAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHADFDGIERL 318
+ E + +K V KI H + + + + F +E I + + +ER+
Sbjct: 486 FDTLLQQEMDHLKMAVAKIDSHNPDILLVEKSVSRFAQEYLLAKDISLVLNIKRSLLERI 545
Query: 319 ALVTGGEIASTFDNPESVKLGQCELIEEIMIGE-------------DKLIKFSGVAMGQA 365
+ TG +I + D S KLG C+L E L+ F G
Sbjct: 546 SRCTGAQIVPSIDQLTSPKLGYCDLFHVEKFVETHVSPCQVAKKMAKTLMFFDGCPKPLG 605
Query: 366 CTIVLRGASHHVLDEAERSLH 386
CTI+L+GA DE ++ H
Sbjct: 606 CTILLKGAHE---DELKKVKH 623
>At3g14270 unknown protein
Length = 1791
Score = 41.2 bits (95), Expect = 0.001
Identities = 44/195 (22%), Positives = 80/195 (40%), Gaps = 25/195 (12%)
Query: 206 DSFLDEGFILDKKI-GLGQPKRIENAKILVANTAMDTDKVKIYGARVRVDSMAKVAEIEG 264
DS + +G + K + +IE A++L+ ++ +V + ++ +
Sbjct: 452 DSMVVKGVVCKKNVVNRRMSTKIEKARLLILGGGLEYQRVS--------NQLSSFDTLLQ 503
Query: 265 AEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHADFDGIERLALVTGG 324
EK+ +K V KI N + + + F +E I + + ++R+A TG
Sbjct: 504 QEKDHLKMAVAKIHAERPNILLVEKSVSRFAQEYLLAKDISLVLNIKRPLLDRIARCTGA 563
Query: 325 EIASTFDNPESVKLGQCELIEEIMIGED---------KLIK----FSGVAMGQACTIVLR 371
+I + D+ S KLG CE E+ K++K F TI+LR
Sbjct: 564 QIIPSVDHLSSQKLGYCENFRVDRYPEEHGSTGQVGKKVVKTLMYFEHCPKPLGFTILLR 623
Query: 372 GASHHVLDEAERSLH 386
GA+ DE ++ H
Sbjct: 624 GANE---DELKKVKH 635
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,254,542
Number of Sequences: 26719
Number of extensions: 411303
Number of successful extensions: 1236
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1140
Number of HSP's gapped (non-prelim): 52
length of query: 527
length of database: 11,318,596
effective HSP length: 104
effective length of query: 423
effective length of database: 8,539,820
effective search space: 3612343860
effective search space used: 3612343860
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146571.2


