

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146569.8 - phase: 0
(166 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
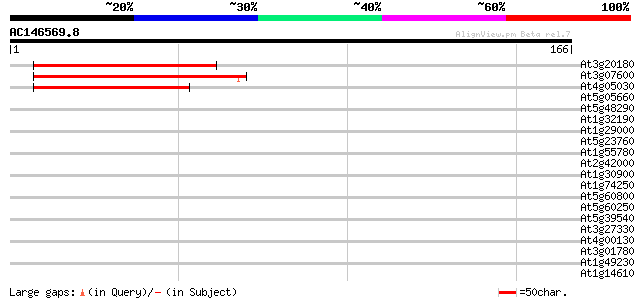
Score E
Sequences producing significant alignments: (bits) Value
At3g20180 hypothetical protein 50 4e-07
At3g07600 unknown protein 47 4e-06
At4g05030 45 2e-05
At5g05660 putative protein 37 0.004
At5g48290 ATFP5 37 0.005
At1g32190 unknown protein 37 0.006
At1g29000 hypothetical protein 36 0.008
At5g23760 unknown protein 36 0.011
At1g55780 hypothetical protein 35 0.014
At2g42000 metallothionein-like protein 34 0.041
At1g30900 F17F8.23 33 0.071
At1g74250 putative heat shock protein 33 0.092
At5g60800 unknown protein 32 0.16
At5g60250 unknown protein 32 0.16
At5g39540 putative protein 32 0.16
At3g27330 hypothetical protein 32 0.16
At4g00130 hypothetical protein 32 0.21
At3g01780 unknown protein 32 0.21
At1g49230 putative RING zinc finger protein gb|AAF16660.1; simil... 32 0.21
At1g14610 valyl tRNA synthetase (valRS) 32 0.21
>At3g20180 hypothetical protein
Length = 118
Score = 50.4 bits (119), Expect = 4e-07
Identities = 27/54 (50%), Positives = 35/54 (64%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEK 61
GVTSV++EG+ +D + V GD VD L L+KK +VT+ T EEVKK EK
Sbjct: 25 GVTSVAMEGEFQDELVVVGDGVDSASLIMALRKKACHVTLETLEEVKKPQVEEK 78
>At3g07600 unknown protein
Length = 157
Score = 47.4 bits (111), Expect = 4e-06
Identities = 24/66 (36%), Positives = 42/66 (63%), Gaps = 3/66 (4%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKE- 66
GV +V ++GD ++++ VTG VD++ L N L+KK +++ +V+ + +KK E+E
Sbjct: 29 GVNAVEIKGDHRNQIEVTGVEVDMIALINTLRKKVAFAELVSVAKVEPPKDGDKKPEEEK 88
Query: 67 --EEKK 70
EEKK
Sbjct: 89 KPEEKK 94
>At4g05030
Length = 110
Score = 44.7 bits (104), Expect = 2e-05
Identities = 20/46 (43%), Positives = 30/46 (64%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEV 53
GVT V +EG++KD+V V G+ VD CL +L+KK I++ +V
Sbjct: 62 GVTFVGIEGEEKDKVVVIGEGVDAACLVVRLRKKVGFADIISVTDV 107
>At5g05660 putative protein
Length = 820
Score = 37.4 bits (85), Expect = 0.004
Identities = 31/100 (31%), Positives = 38/100 (38%), Gaps = 19/100 (19%)
Query: 74 EACHAVLHGSC-KCNNFNCH-GKCNSCTKCESSK--CDGTNCVTIC---SFKCGN----- 121
E C L +C C CH G C SC K +K C G V C F CG+
Sbjct: 188 EVCERPLSNNCGHCCLLLCHPGPCASCPKLVKAKCFCGGVEDVRRCGHKQFSCGDVCERV 247
Query: 122 SKCDGNKCNG-CTPKVTSPCFQRCPQWCTCSK------CC 154
C+ + C C PC +R C+C K CC
Sbjct: 248 LDCNIHNCREICHDGECPPCRERAVYKCSCGKVKEEKDCC 287
Score = 28.9 bits (63), Expect = 1.3
Identities = 24/98 (24%), Positives = 31/98 (31%), Gaps = 12/98 (12%)
Query: 63 KEKEEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNS 122
K KEE+ + C V C N GK C + +C S CG
Sbjct: 279 KVKEEK-----DCCERVFRCEASCENMLNCGKHVCERGCHAGECGLCPYQGKRSCPCGKR 333
Query: 123 KCDGNKCNGCTPKVTSPC-------FQRCPQWCTCSKC 153
G C+ P C + RCP+ C C
Sbjct: 334 FYQGLSCDVVAPLCGGTCDKVLGCGYHRCPERCHRGPC 371
Score = 27.7 bits (60), Expect = 3.0
Identities = 20/88 (22%), Positives = 30/88 (33%), Gaps = 10/88 (11%)
Query: 75 ACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTP 134
AC + C N C C++ SS + C +C + TP
Sbjct: 502 ACDNLCGNPLPCGNHYCSYFCHALDIRSSSLDKRSESCEKCDLRCQKER---------TP 552
Query: 135 KVTSPCFQRC-PQWCTCSKCCAPYQPYC 161
+ PC +RC P+ C K +C
Sbjct: 553 RCQHPCPRRCHPEDCPPCKTLVKRSCHC 580
>At5g48290 ATFP5
Length = 181
Score = 37.0 bits (84), Expect = 0.005
Identities = 17/56 (30%), Positives = 36/56 (63%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKK 63
GV++V ++GD ++++ VTG VD++ L L+KK +++ +V+ + ++KK
Sbjct: 29 GVSAVEIKGDHRNQIEVTGVEVDMIPLIQILRKKVAFAELVSVTKVEPPKKEDEKK 84
>At1g32190 unknown protein
Length = 422
Score = 36.6 bits (83), Expect = 0.006
Identities = 21/66 (31%), Positives = 25/66 (37%), Gaps = 7/66 (10%)
Query: 91 CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQWCTC 150
C G C C +C +C S CG C KC+ T K CF C +
Sbjct: 297 CSGLCRPSCSCPKPRCPKPSC----SCGCGCGDCGCFKCSCPTLK---GCFSCCKKPSCV 349
Query: 151 SKCCAP 156
S CC P
Sbjct: 350 SSCCCP 355
Score = 32.7 bits (73), Expect = 0.092
Identities = 34/132 (25%), Positives = 47/132 (34%), Gaps = 37/132 (28%)
Query: 49 TTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNCH----------GKCNSC 98
TT + + KT ++ + ++E C + SC C C G C C
Sbjct: 273 TTTKSRLKTIWQEIRRRDESTGC---CCSGLCRPSCSCPKPRCPKPSCSCGCGCGDCG-C 328
Query: 99 TKCES-------SKCDGTNCVTIC---SFKCGNS-------KCDGNKCNGCTPKVTSPCF 141
KC S C +CV+ C +FKC + KC KC C P
Sbjct: 329 FKCSCPTLKGCFSCCKKPSCVSSCCCPTFKCSSCFGKPKCPKCSCWKCLKC------PDT 382
Query: 142 QRCPQWCTCSKC 153
+ C C CS C
Sbjct: 383 ECCRSSCCCSGC 394
>At1g29000 hypothetical protein
Length = 287
Score = 36.2 bits (82), Expect = 0.008
Identities = 21/58 (36%), Positives = 31/58 (53%), Gaps = 3/58 (5%)
Query: 17 DDKDRVCVTGDNVDIVCLANQLKKKFNN---VTILTTEEVKKKTEAEKKKEKEEEKKK 71
D K + ++ L +KKK + + TEE KKK E +KKK++EE+KKK
Sbjct: 146 DTKAQTLTVQGTIESAKLLAYIKKKVHKHAEIISSKTEEEKKKEEEDKKKKEEEDKKK 203
Score = 32.3 bits (72), Expect = 0.12
Identities = 15/21 (71%), Positives = 17/21 (80%)
Query: 51 EEVKKKTEAEKKKEKEEEKKK 71
EE KKK E E KK+KE+EKKK
Sbjct: 190 EEDKKKKEEEDKKKKEDEKKK 210
Score = 29.6 bits (65), Expect = 0.78
Identities = 16/33 (48%), Positives = 21/33 (63%)
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK EE KKK E EKKKE+E++K++
Sbjct: 186 KKKEEEDKKKKEEEDKKKKEDEKKKEEEKKKEE 218
Score = 27.7 bits (60), Expect = 3.0
Identities = 12/21 (57%), Positives = 17/21 (80%)
Query: 51 EEVKKKTEAEKKKEKEEEKKK 71
+E +KK E EKKKE+E +KK+
Sbjct: 204 KEDEKKKEEEKKKEEENKKKE 224
Score = 27.3 bits (59), Expect = 3.9
Identities = 12/22 (54%), Positives = 17/22 (76%)
Query: 51 EEVKKKTEAEKKKEKEEEKKKM 72
EE KKK E KKKE E++K+++
Sbjct: 211 EEEKKKEEENKKKEGEKKKEEV 232
>At5g23760 unknown protein
Length = 103
Score = 35.8 bits (81), Expect = 0.011
Identities = 20/59 (33%), Positives = 34/59 (56%), Gaps = 1/59 (1%)
Query: 13 SLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
S+ D KD+ +D V + +LKK V +++ K++ + EKK+EK+EEKK+
Sbjct: 33 SIAADMKDQKLTVIGLMDAVAVVKKLKK-VGKVDLISVGPAKEEKKEEKKEEKKEEKKE 90
>At1g55780 hypothetical protein
Length = 133
Score = 35.4 bits (80), Expect = 0.014
Identities = 23/93 (24%), Positives = 43/93 (45%), Gaps = 7/93 (7%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEE 67
GV SVS++G + D++ + G+ +D+ L +LKKK TI+T + + + +
Sbjct: 21 GVRSVSIQGQN-DQLVLLGEGIDLAELTRELKKKVCMTTIITVQAAPPQQPPQPHPMGQY 79
Query: 68 EKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTK 100
+ C +C+ N G C S ++
Sbjct: 80 NQMPPARRC------TCEIPNSGFCGFCRSMSQ 106
>At2g42000 metallothionein-like protein
Length = 115
Score = 33.9 bits (76), Expect = 0.041
Identities = 30/112 (26%), Positives = 41/112 (35%), Gaps = 8/112 (7%)
Query: 42 FNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKC 101
F L E + T K K + K + C+ SC C + G C
Sbjct: 10 FCTAQCLYISECGRGTLRLKTKMADTGKGSSVAGCN----DSCGCPSPCPGGNSCRCRMR 65
Query: 102 ESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQWCTCSKC 153
E+S D + V C CG + C+ K T TS C + CTC+ C
Sbjct: 66 EASAGDQGHMVCPCGEHCGCNPCNCPK----TQTQTSAKGCTCGEGCTCASC 113
>At1g30900 F17F8.23
Length = 649
Score = 33.1 bits (74), Expect = 0.071
Identities = 21/64 (32%), Positives = 28/64 (42%), Gaps = 8/64 (12%)
Query: 66 EEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC-SFKCGNSKC 124
E +K AC + C+C G KCE+ +CDG NC F+C KC
Sbjct: 507 ETKKGLTFSACSNLETSGCRCPP----GFKGDGLKCEACQCDGCNCKNKWGGFEC---KC 559
Query: 125 DGNK 128
GN+
Sbjct: 560 SGNR 563
Score = 26.6 bits (57), Expect = 6.6
Identities = 19/80 (23%), Positives = 29/80 (35%), Gaps = 10/80 (12%)
Query: 75 ACHAVLHGSCKCNNFNCHGKCN---SCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNG 131
+C C N C + + + C + + G C FK KC+ +C+G
Sbjct: 488 SCEPYGPARCSINQGGCWSETKKGLTFSACSNLETSGCRCPP--GFKGDGLKCEACQCDG 545
Query: 132 CTPKVTSPCFQRCPQWCTCS 151
C K F+ C CS
Sbjct: 546 CNCKNKWGGFE-----CKCS 560
>At1g74250 putative heat shock protein
Length = 630
Score = 32.7 bits (73), Expect = 0.092
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 3/41 (7%)
Query: 34 LANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIE 74
LA +KK+ V + VKK E EKKKE+E E+KK +E
Sbjct: 226 LAEFVKKRDKRVIDML---VKKNAEMEKKKEEERERKKKME 263
>At5g60800 unknown protein
Length = 283
Score = 32.0 bits (71), Expect = 0.16
Identities = 23/67 (34%), Positives = 36/67 (53%), Gaps = 2/67 (2%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEE 67
GV +V E ++ VTG +D V L +L++K L + + KK+ E E K + +E
Sbjct: 52 GVETVKSESAT-GKLTVTGA-LDPVKLREKLEEKTKKKVDLVSPQPKKEKEKENKNKNDE 109
Query: 68 EKKKMIE 74
+KKK E
Sbjct: 110 DKKKSEE 116
Score = 32.0 bits (71), Expect = 0.16
Identities = 28/90 (31%), Positives = 42/90 (46%), Gaps = 8/90 (8%)
Query: 25 TGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSC 84
T VD+V + Q KK+ +E KKK+E +KK + ++K K AVL
Sbjct: 84 TKKKVDLV--SPQPKKEKEKENKNKNDEDKKKSEEKKKPDNNDKKPKETPVTTAVLK--- 138
Query: 85 KCNNFNCHGKCNSCTKCESSKCDGTNCVTI 114
NF+C G C + +K G N +T+
Sbjct: 139 --LNFHCQG-CIGKIQKTVTKTKGVNGLTM 165
>At5g60250 unknown protein
Length = 655
Score = 32.0 bits (71), Expect = 0.16
Identities = 17/55 (30%), Positives = 22/55 (39%), Gaps = 13/55 (23%)
Query: 98 CTKCE-----SSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQW 147
C KC+ S C+ C +CG+ C C G K+ C RCP W
Sbjct: 470 CGKCQHMIELSQGCNHITC------RCGHEFC--YNCGGGWNKIMGTCLNRCPTW 516
>At5g39540 putative protein
Length = 196
Score = 32.0 bits (71), Expect = 0.16
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 2/58 (3%)
Query: 32 VCLANQLKKKFNNVTI-LTTEEVKKKTEAEKKKEKEEEKKKMIEACHA-VLHGSCKCN 87
V L++ K +N I LT + +KKK E E++K+K+++ + ++ H L CN
Sbjct: 87 VLLSHNSNKDYNFCKIYLTPQAIKKKKEVEEEKKKQKKGEAVVSVAHVEALEEQQPCN 144
>At3g27330 hypothetical protein
Length = 913
Score = 32.0 bits (71), Expect = 0.16
Identities = 23/93 (24%), Positives = 37/93 (39%), Gaps = 17/93 (18%)
Query: 54 KKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVT 113
K++ E E K+EK+ +K + + H VL C K C C+
Sbjct: 686 KREMETETKEEKKIDKDEKSLSDHVVL---------PCIDKLRDELSC-------AICLE 729
Query: 114 ICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQ 146
IC F+ + C + C C C ++CP+
Sbjct: 730 IC-FEPSTTTCGHSFCKKCLRSAADKCGRKCPK 761
>At4g00130 hypothetical protein
Length = 247
Score = 31.6 bits (70), Expect = 0.21
Identities = 22/53 (41%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Query: 20 DRVCVTGDNVDIVCL-ANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
D GD +I+ L A LKKKFNN + K +TE E + KE EKK+
Sbjct: 84 DFTSTKGDPYEILMLFAFMLKKKFNNA--VKNARKKGQTEDEVEYAKESEKKR 134
>At3g01780 unknown protein
Length = 1192
Score = 31.6 bits (70), Expect = 0.21
Identities = 14/21 (66%), Positives = 18/21 (85%)
Query: 51 EEVKKKTEAEKKKEKEEEKKK 71
EEVK+K E E+ K+KEE+KKK
Sbjct: 1131 EEVKEKKEKEEGKDKEEKKKK 1151
>At1g49230 putative RING zinc finger protein gb|AAF16660.1; similar
to gb|N37302.1, gb|T75752.1
Length = 219
Score = 31.6 bits (70), Expect = 0.21
Identities = 12/41 (29%), Positives = 18/41 (43%)
Query: 67 EEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCD 107
EE+ K++ CH H C + H C +C C C+
Sbjct: 141 EERVKLLPTCHHGFHVRCIDKWLSSHSSCPTCRHCLIQTCE 181
>At1g14610 valyl tRNA synthetase (valRS)
Length = 1108
Score = 31.6 bits (70), Expect = 0.21
Identities = 21/50 (42%), Positives = 29/50 (58%), Gaps = 9/50 (18%)
Query: 34 LANQLKKKF--------NNVTILTTEEV-KKKTEAEKKKEKEEEKKKMIE 74
L NQL + F + ILT EE+ +KK + EK KEKE +K+K +E
Sbjct: 31 LTNQLVRSFHGSRTMSESEKKILTEEELERKKKKEEKAKEKELKKQKALE 80
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.134 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,495,613
Number of Sequences: 26719
Number of extensions: 211274
Number of successful extensions: 3078
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 97
Number of HSP's successfully gapped in prelim test: 77
Number of HSP's that attempted gapping in prelim test: 2245
Number of HSP's gapped (non-prelim): 693
length of query: 166
length of database: 11,318,596
effective HSP length: 92
effective length of query: 74
effective length of database: 8,860,448
effective search space: 655673152
effective search space used: 655673152
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146569.8


