

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.9 + phase: 0
(80 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
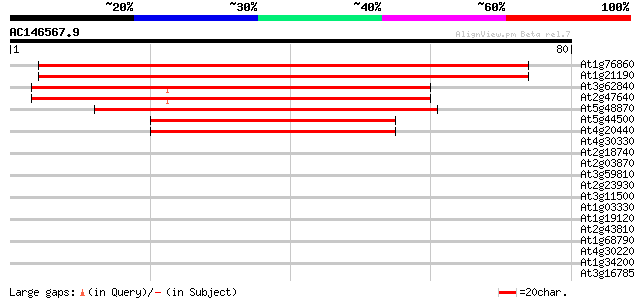
At1g76860 unknown protein 127 1e-30
At1g21190 putative protein 125 5e-30
At3g62840 small nuclear ribonucleoprotein-like protein 44 2e-05
At2g47640 putative small nuclear ribonucleoprotein D2 44 2e-05
At5g48870 Sm-like protein (SAD1) 39 4e-04
At5g44500 unknown protein 38 8e-04
At4g20440 putative snRNP protein 38 8e-04
At4g30330 small nuclear ribonucleoprotein homolog 35 0.007
At2g18740 small nuclear ribonucleoprotein E like protein 35 0.007
At2g03870 putative snRNP splicing factor 35 0.007
At3g59810 U6 snRNA-associated Sm-like protein 33 0.019
At2g23930 putative small nuclear ribonucleoprotein E 33 0.033
At3g11500 putative small nuclear ribonucleoprotein polypeptide G 32 0.057
At1g03330 unknown protein 32 0.057
At1g19120 unknown protein 32 0.074
At2g43810 putative small nuclear ribonucleoprotein polypeptide F 31 0.096
At1g68790 putative nuclear matrix constituent protein 1 (NMCP1) 30 0.21
At4g30220 snRNP Sm protein F - like 28 0.82
At1g34200 F23M19.12/F23M19.12 27 1.8
At3g16785 phospholipase D, putative, 5' partial 27 2.4
>At1g76860 unknown protein
Length = 98
Score = 127 bits (319), Expect = 1e-30
Identities = 62/70 (88%), Positives = 67/70 (95%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EEE+ V+EPLDLIRLSLDERIYVKLRSDRELRGKLHA+DQHLNMILGDVEE +TTVEIDD
Sbjct: 4 EEEATVREPLDLIRLSLDERIYVKLRSDRELRGKLHAFDQHLNMILGDVEETITTVEIDD 63
Query: 65 ETYEEIVRVS 74
ETYEEIVR +
Sbjct: 64 ETYEEIVRTT 73
>At1g21190 putative protein
Length = 97
Score = 125 bits (313), Expect = 5e-30
Identities = 58/70 (82%), Positives = 68/70 (96%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EE++ V+EPLDLIRLS++ERIYVKLRSDRELRGKLHA+DQHLNMILGDVEE++TT+EIDD
Sbjct: 4 EEDATVREPLDLIRLSIEERIYVKLRSDRELRGKLHAFDQHLNMILGDVEEVITTIEIDD 63
Query: 65 ETYEEIVRVS 74
ETYEEIVR +
Sbjct: 64 ETYEEIVRTT 73
>At3g62840 small nuclear ribonucleoprotein-like protein
Length = 109
Score = 43.5 bits (101), Expect = 2e-05
Identities = 21/59 (35%), Positives = 39/59 (65%), Gaps = 2/59 (3%)
Query: 4 TEEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
TEEE PL ++ +S+ + ++ + R++R+L G++ A+D+H NM+L +V E+ T V
Sbjct: 14 TEEEEFNTGPLSVLMMSVKNNTQVLINCRNNRKLLGRVRAFDRHCNMVLENVREMWTEV 72
>At2g47640 putative small nuclear ribonucleoprotein D2
Length = 108
Score = 43.5 bits (101), Expect = 2e-05
Identities = 21/59 (35%), Positives = 39/59 (65%), Gaps = 2/59 (3%)
Query: 4 TEEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
TEEE PL ++ +S+ + ++ + R++R+L G++ A+D+H NM+L +V E+ T V
Sbjct: 13 TEEEEFNTGPLSVLMMSVKNNTQVLINCRNNRKLLGRVRAFDRHCNMVLENVREMWTEV 71
>At5g48870 Sm-like protein (SAD1)
Length = 88
Score = 39.3 bits (90), Expect = 4e-04
Identities = 19/49 (38%), Positives = 30/49 (60%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
P +LI + +I+V ++ D+EL G L +D ++NM+L DV E T E
Sbjct: 10 PSELIDRCIGSKIWVIMKGDKELVGILKGFDVYVNMVLEDVTEYEITAE 58
>At5g44500 unknown protein
Length = 254
Score = 38.1 bits (87), Expect = 8e-04
Identities = 15/35 (42%), Positives = 26/35 (73%)
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEE 55
++ R+ V ++ R+L GK A+D+H+N++LGD EE
Sbjct: 13 INYRMRVTIQDGRQLIGKFMAFDRHMNLVLGDCEE 47
>At4g20440 putative snRNP protein
Length = 257
Score = 38.1 bits (87), Expect = 8e-04
Identities = 15/35 (42%), Positives = 26/35 (73%)
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEE 55
++ R+ V ++ R+L GK A+D+H+N++LGD EE
Sbjct: 13 INYRMRVTIQDGRQLVGKFMAFDRHMNLVLGDCEE 47
>At4g30330 small nuclear ribonucleoprotein homolog
Length = 88
Score = 35.0 bits (79), Expect = 0.007
Identities = 14/60 (23%), Positives = 36/60 (59%), Gaps = 4/60 (6%)
Query: 1 MAGTEEESAVKEPLDLIRLSLDERIYVKL----RSDRELRGKLHAYDQHLNMILGDVEEI 56
MA T+ + + +P++LI L + +++ + D + G++ +D+++N++L + EE+
Sbjct: 1 MASTKVQRIMTQPINLIFRFLQSKARIQIWLFEQKDLRIEGRITGFDEYMNLVLDEAEEV 60
>At2g18740 small nuclear ribonucleoprotein E like protein
Length = 88
Score = 35.0 bits (79), Expect = 0.007
Identities = 14/60 (23%), Positives = 36/60 (59%), Gaps = 4/60 (6%)
Query: 1 MAGTEEESAVKEPLDLIRLSLDERIYVKL----RSDRELRGKLHAYDQHLNMILGDVEEI 56
MA T+ + + +P++LI L + +++ + D + G++ +D+++N++L + EE+
Sbjct: 1 MASTKVQRIMTQPINLIFRFLQSKARIQIWLFEQKDLRIEGRITGFDEYMNLVLDEAEEV 60
>At2g03870 putative snRNP splicing factor
Length = 99
Score = 35.0 bits (79), Expect = 0.007
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Query: 14 LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
LDL + +D+ + VKL R++ G L YDQ LN++L + E V
Sbjct: 9 LDLAKF-VDKGVQVKLTGGRQVTGTLKGYDQLLNLVLDEAVEFV 51
>At3g59810 U6 snRNA-associated Sm-like protein
Length = 91
Score = 33.5 bits (75), Expect = 0.019
Identities = 19/59 (32%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Query: 1 MAGTEEE--SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
M+G EE+ K P D ++ + VKL S + RG L D ++N+ + EE V
Sbjct: 1 MSGVEEKVSGTTKTPADFLKSIRGRPVVVKLNSGVDYRGTLTCLDGYMNIAMEQTEEYV 59
>At2g23930 putative small nuclear ribonucleoprotein E
Length = 80
Score = 32.7 bits (73), Expect = 0.033
Identities = 13/45 (28%), Positives = 30/45 (65%), Gaps = 1/45 (2%)
Query: 12 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
+P DL + +D+++ +KL ++R + G L +DQ +N+++ + E+
Sbjct: 6 QPPDLKKY-MDKKLQIKLNANRMVTGTLRGFDQFMNLVVDNTVEV 49
>At3g11500 putative small nuclear ribonucleoprotein polypeptide G
Length = 79
Score = 32.0 bits (71), Expect = 0.057
Identities = 13/45 (28%), Positives = 30/45 (65%), Gaps = 1/45 (2%)
Query: 12 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
+P DL + +D+++ +KL ++R + G L +DQ +N+++ + E+
Sbjct: 6 QPPDLKKY-MDKKLQIKLNANRMVVGTLRGFDQFMNLVVDNTVEV 49
>At1g03330 unknown protein
Length = 93
Score = 32.0 bits (71), Expect = 0.057
Identities = 15/51 (29%), Positives = 30/51 (58%), Gaps = 6/51 (11%)
Query: 23 ERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIVRV 73
+ + V+L++D +RG LH+ DQ+LN+ ++ T +D + Y ++ V
Sbjct: 13 QEVTVELKNDLAIRGTLHSVDQYLNI------KLENTRVVDQDKYPHMLSV 57
>At1g19120 unknown protein
Length = 128
Score = 31.6 bits (70), Expect = 0.074
Identities = 15/37 (40%), Positives = 23/37 (61%)
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
LD+++ V LR R+L G L ++DQ N +L + E V
Sbjct: 19 LDKKLLVLLRDGRKLMGLLRSFDQFANAVLEEAYERV 55
>At2g43810 putative small nuclear ribonucleoprotein polypeptide F
Length = 91
Score = 31.2 bits (69), Expect = 0.096
Identities = 16/55 (29%), Positives = 26/55 (47%)
Query: 3 GTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
G + K P D ++ + + VKL S + RG L D ++N+ + EE V
Sbjct: 5 GEKASGTTKTPADFLKSIRGKPVVVKLNSGVDYRGILTCLDGYMNIAMEQTEEYV 59
>At1g68790 putative nuclear matrix constituent protein 1 (NMCP1)
Length = 1085
Score = 30.0 bits (66), Expect = 0.21
Identities = 17/67 (25%), Positives = 38/67 (56%), Gaps = 1/67 (1%)
Query: 4 TEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEID 63
T++ES ++E + +R++ +ER+ LR EL+ ++ Q ++L + EE+ E
Sbjct: 474 TKQESRIREEHESLRITKEERVEF-LRLQSELKQQIDKVKQEEELLLKEREELKQDKERF 532
Query: 64 DETYEEI 70
++ +E +
Sbjct: 533 EKEWEAL 539
>At4g30220 snRNP Sm protein F - like
Length = 88
Score = 28.1 bits (61), Expect = 0.82
Identities = 12/33 (36%), Positives = 21/33 (63%)
Query: 25 IYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
+ VKL+ E +G L + D ++N+ LG+ EE +
Sbjct: 20 VIVKLKWGMEYKGFLASVDSYMNLQLGNTEEYI 52
>At1g34200 F23M19.12/F23M19.12
Length = 352
Score = 26.9 bits (58), Expect = 1.8
Identities = 14/72 (19%), Positives = 39/72 (53%), Gaps = 1/72 (1%)
Query: 7 ESAVKEP-LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDE 65
ES +++P +D + + R++V+ + ++GK D+ + + + + ++IV E++
Sbjct: 65 ESLLEDPDVDAVYFPIPTRLHVEWATLAAIKGKHILLDKPVALNVAEFDQIVEACEVNGV 124
Query: 66 TYEEIVRVSHNP 77
+ + + H+P
Sbjct: 125 QFMDGTQWMHSP 136
>At3g16785 phospholipase D, putative, 5' partial
Length = 682
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/33 (33%), Positives = 21/33 (63%), Gaps = 2/33 (6%)
Query: 38 KLHAYDQHLNMILGDVEEIVTTVEIDDETYEEI 70
+L + +HL + G++++I+ V D TY+EI
Sbjct: 603 RLSLWSEHLGLRTGEIDQIIDPV--SDSTYKEI 633
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.134 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,759,150
Number of Sequences: 26719
Number of extensions: 61211
Number of successful extensions: 173
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 150
Number of HSP's gapped (non-prelim): 30
length of query: 80
length of database: 11,318,596
effective HSP length: 56
effective length of query: 24
effective length of database: 9,822,332
effective search space: 235735968
effective search space used: 235735968
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146567.9


