

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.11 - phase: 0
(331 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
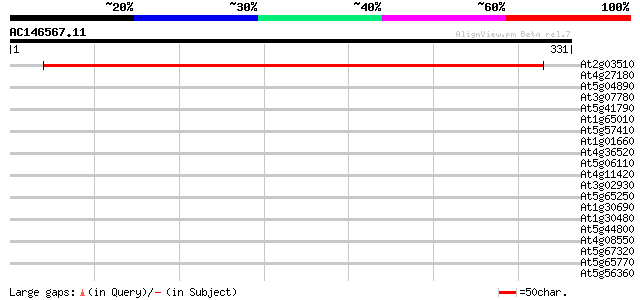
Score E
Sequences producing significant alignments: (bits) Value
At2g03510 unknown protein 511 e-145
At4g27180 heavy chain polypeptide of kinesin-like protein (katB) 37 0.011
At5g04890 unknown protein 37 0.015
At3g07780 unknown protein 35 0.043
At5g41790 myosin heavy chain-like protein 35 0.057
At1g65010 hypothetical protein 35 0.074
At5g57410 unknown protein 34 0.097
At1g01660 hypothetical protein 34 0.13
At4g36520 trichohyalin like protein 33 0.16
At5g06110 cell division related protein-like 33 0.22
At4g11420 putative protein 33 0.22
At3g02930 unknown protein 33 0.22
At5g65250 unknown protein 33 0.28
At1g30690 putative phosphatidyl-inositol-transfer protein 33 0.28
At1g30480 DNA damage repair protein, putative 33 0.28
At5g44800 helicase-like protein 32 0.37
At4g08550 hypothetical protein 32 0.37
At5g67320 unknown protein 32 0.48
At5g65770 nuclear matrix constituent protein 1 (NMCP1)-like 32 0.48
At5g56360 unknown protein 32 0.48
>At2g03510 unknown protein
Length = 356
Score = 511 bits (1315), Expect = e-145
Identities = 244/295 (82%), Positives = 274/295 (92%)
Query: 21 AIVHQVPEGHVGVYWRGGALLKTITEPGFHMKMPFLTQFEPVQVTLQTDEVTDIPCGTKG 80
++VHQVPEGHVG YWRGGALL ITEPGFH+K+PF+T +EPVQVTLQTD+V DIPCGTKG
Sbjct: 45 SLVHQVPEGHVGAYWRGGALLNIITEPGFHLKLPFITNYEPVQVTLQTDQVRDIPCGTKG 104
Query: 81 GVMIVFGKIEVVNRLHKESVYETLLNYGVQYDKTWIYDKIHHEINQFCSSHSLQQVYIDV 140
GV+I F KIEVVNRL K+ VY+TLLNYGV YD TWIYDKIHHEINQFCSSHSLQQVYID+
Sbjct: 105 GVLITFEKIEVVNRLRKDFVYDTLLNYGVNYDNTWIYDKIHHEINQFCSSHSLQQVYIDI 164
Query: 141 FDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQ 200
FDQIDE+MKDALQ DCTRYAPGIEI+ VRVTKP IPES+R NFEQMEEERTKVLIAIEKQ
Sbjct: 165 FDQIDERMKDALQADCTRYAPGIEILSVRVTKPKIPESVRRNFEQMEEERTKVLIAIEKQ 224
Query: 201 KVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADA 260
+V+EKEAET K MAISEAEKNANVSKILM+QKL+EKDS+RR+ +IEN MYL R+KSLADA
Sbjct: 225 RVAEKEAETKKIMAISEAEKNANVSKILMQQKLTEKDSSRREADIENQMYLDRQKSLADA 284
Query: 261 DFYRVIKEAEANRLKLTPEFLELKFIESIANNTKIFFGDKIPNMILDQRLLGNFL 315
D+YRV++EAEAN+LKLTPEFLELKFI++IA NTKIFFGDK+PNM+LDQRLLGNFL
Sbjct: 285 DYYRVLREAEANKLKLTPEFLELKFIDAIARNTKIFFGDKVPNMVLDQRLLGNFL 339
>At4g27180 heavy chain polypeptide of kinesin-like protein (katB)
Length = 745
Score = 37.4 bits (85), Expect = 0.011
Identities = 50/220 (22%), Positives = 102/220 (45%), Gaps = 21/220 (9%)
Query: 77 GTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYDKTWIYDK-IHHEINQFCSSHSLQQ 135
G++ G + F + +V LH+ Y++ NY + + T Y K + I F Q+
Sbjct: 25 GSEYGGPVEFTREDVETLLHERIKYKSKYNYKERCENTMDYVKRLRLCIRWF------QE 78
Query: 136 VYID-VFDQIDEKMKDALQVDCTRYAPGIEI-IGVRVTKPN-IPESIRHNFE----QMEE 188
+ +D F+Q EK+K+A++++ ++ +E+ + V+ + N + + +R NF Q+ +
Sbjct: 79 LELDYAFEQ--EKLKNAMEMN-EKHCADLEVNLKVKEEELNMVIDELRKNFASVQVQLAK 135
Query: 189 ERTKVLIAIE---KQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEI 245
E+T+ L A E K++ + E+++ E K + ++ + D + +E
Sbjct: 136 EQTEKLAANESLGKEREARIAVESLQAAITEELAKTQGELQTANQRIQAVNDMYKLLQEY 195
Query: 246 ENAMYLAREKSLADAD-FYRVIKEAEANRLKLTPEFLELK 284
+++ L K D D + IK E R + LK
Sbjct: 196 NSSLQLYNSKLQGDLDEAHENIKRGEKERTGIVESIGNLK 235
>At5g04890 unknown protein
Length = 366
Score = 37.0 bits (84), Expect = 0.015
Identities = 31/97 (31%), Positives = 52/97 (52%), Gaps = 9/97 (9%)
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARR-Q 242
++ EEE K + +KQ + EKEA K ++A++ A + K+ E K E+ +A++ Q
Sbjct: 141 KEKEEEEAKQM---KKQLLEEKEALIRKLQEEAKAKEEAEMRKLQEEAKAKEEAAAKKLQ 197
Query: 243 EEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPE 279
EEIE A E+ L + R ++E + +KL E
Sbjct: 198 EEIE-AKEKLEERKLEE----RRLEERKLEDMKLAEE 229
Score = 34.3 bits (77), Expect = 0.097
Identities = 36/171 (21%), Positives = 72/171 (42%), Gaps = 22/171 (12%)
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
E + +Q+ EE+ ++ ++++ +++EAE K ++A++ A K+ E + EK
Sbjct: 146 EEAKQMKKQLLEEKEALIRKLQEEAKAKEEAEMRKLQEEAKAKEEAAAKKLQEEIEAKEK 205
Query: 237 DSARRQEEIE------NAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKF----- 285
R+ EE M LA E L + + E+ L PE + K
Sbjct: 206 LEERKLEERRLEERKLEDMKLAEEAKLKKIQERKSVDESGEKEKILKPEVVYTKSGHVAT 265
Query: 286 -IESIANNTKIFFG----------DKIPNMILDQRLLGNFLVEEVPRGAAT 325
+ K FG +K+ N++ ++ +G ++E++ R T
Sbjct: 266 PKPESGSGLKSGFGGVGEVVKSAEEKLGNLVEKEKKMGKGIMEKIRRKEIT 316
Score = 28.5 bits (62), Expect = 5.3
Identities = 23/76 (30%), Positives = 36/76 (47%), Gaps = 6/76 (7%)
Query: 185 QMEEERTKVLIAIEKQKVSEKEAETMKKMAIS-EAEKNANVSKILMEQKLSEKDSARRQE 243
++EE+R +E+ + EKE E K+M EK A + K+ E K E+ R+ +
Sbjct: 128 KLEEKRL-----LEESRRKEKEEEEAKQMKKQLLEEKEALIRKLQEEAKAKEEAEMRKLQ 182
Query: 244 EIENAMYLAREKSLAD 259
E A A K L +
Sbjct: 183 EEAKAKEEAAAKKLQE 198
>At3g07780 unknown protein
Length = 566
Score = 35.4 bits (80), Expect = 0.043
Identities = 27/120 (22%), Positives = 58/120 (47%), Gaps = 9/120 (7%)
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGV-RVTKPNIPES----IRHNFEQMEEERTKVLIAI 197
+++EK K ++ R +I V R+ + E+ ++ N ++E ER + ++
Sbjct: 426 EVEEKAKQVAELQMERQKKKQQIEEVERIVRLKQAEAEMFQLKANEAKVEAERLERIVKA 485
Query: 198 EKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSL 257
+K+K E+ A K+ +SEAE K + +K+ E++S E A+ ++ + +
Sbjct: 486 KKEKTEEEYASNYLKLRLSEAE----AEKEYLFEKIKEQESGGNGGEASQAVMYSKIREM 541
Score = 30.0 bits (66), Expect = 1.8
Identities = 22/98 (22%), Positives = 44/98 (44%), Gaps = 4/98 (4%)
Query: 201 KVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADA 260
+++E ET++KM I EK K M + E++ + +++ ++K
Sbjct: 390 RIAEVVKETLRKMEIVGEEKTRMYKKARMGLEECEREVEEKAKQVAELQMERQKKKQQIE 449
Query: 261 DFYRVIK----EAEANRLKLTPEFLELKFIESIANNTK 294
+ R+++ EAE +LK +E + +E I K
Sbjct: 450 EVERIVRLKQAEAEMFQLKANEAKVEAERLERIVKAKK 487
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 35.0 bits (79), Expect = 0.057
Identities = 33/117 (28%), Positives = 57/117 (48%), Gaps = 10/117 (8%)
Query: 185 QMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQ-E 243
Q+E +VL E K +E+E+ TM ISE ++I++++ ++ + Q
Sbjct: 669 QLESSEHRVLELSESLKAAEEESRTM-STKISETSDELERTQIMVQELTADSSKLKEQLA 727
Query: 244 EIENAMYLAREKSLADADFYRVIKEAEAN--RLKLTPEFLELKFIE---SIANNTKI 295
E E+ ++L EK D+ IKE EA L+L E + + I+ IA+ T +
Sbjct: 728 EKESKLFLLTEK---DSKSQVQIKELEATVATLELELESVRARIIDLETEIASKTTV 781
>At1g65010 hypothetical protein
Length = 1318
Score = 34.7 bits (78), Expect = 0.074
Identities = 47/204 (23%), Positives = 91/204 (44%), Gaps = 46/204 (22%)
Query: 119 KIHHEINQFCSSHSLQQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTK-PNIPE 177
+++H +++ S ++ Q+ I++ EK +A + D Y + I +K N+ E
Sbjct: 319 ELNHVLHETKSDNAAQKEKIELL----EKTIEAQRTDLEEYGRQVCIAKEEASKLENLVE 374
Query: 178 SIRHNFEQMEEERTKVL-------------------IAIEKQKVSEKEAETMKKM----- 213
SI+ E +EE+T+ L ++IE ++ +E ++ K M
Sbjct: 375 SIKSELEISQEEKTRALDNEKAATSNIQNLLDQRTELSIELERCKVEEEKSKKDMESLTL 434
Query: 214 AISEAEKNANVSK---ILMEQKL----SEKDSARRQEEIENAMYLAREKSLADADFYRVI 266
A+ EA ++ +K ++ +++L S+ DS + + N Y EK L DA
Sbjct: 435 ALQEASTESSEAKATLLVCQEELKNCESQVDSLKLASKETNEKY---EKMLEDA------ 485
Query: 267 KEAEANRLKLTPEFLELKFIESIA 290
E + LK T + ++ +F S A
Sbjct: 486 -RNEIDSLKSTVDSIQNEFENSKA 508
>At5g57410 unknown protein
Length = 373
Score = 34.3 bits (77), Expect = 0.097
Identities = 31/128 (24%), Positives = 59/128 (45%), Gaps = 10/128 (7%)
Query: 164 EIIGVRVTKPNIPESIRHNFEQMEEERTKVL-IAIEKQKVSEKEAETMKKMAISEAEKNA 222
EI + T+ +++ E++++ER + + I Q+V ++ MKK +
Sbjct: 115 EIATITRTEAKNTAALKSQIEKLQQERDEFQRMVIGNQQVKAQQIHEMKKKEKDYIKLQE 174
Query: 223 NVSKILMEQKLSEKDSARRQEEIENAMYLAREKSL-----ADADFYRVIKEA-EANRLKL 276
++++LME+K K+S E + R++ D DFY+ I +A EA +L
Sbjct: 175 RLNQVLMEKK---KESRSGMEIMNLLQKEGRQRGTWNGKKTDTDFYKKIVDAYEAKNQEL 231
Query: 277 TPEFLELK 284
E L+
Sbjct: 232 MAENTSLR 239
>At1g01660 hypothetical protein
Length = 568
Score = 33.9 bits (76), Expect = 0.13
Identities = 31/105 (29%), Positives = 49/105 (46%), Gaps = 8/105 (7%)
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
E +RH ++MEE + +EK K ++EA + K + E ++K +E+
Sbjct: 357 EQLRHR-KEMEESMKRQEEELEKTKKEKEEACMISKNLMQLYEDEVR------QRKEAEE 409
Query: 237 DSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFL 281
RR+EE+E E +F R+ +E EA R K T E L
Sbjct: 410 LVKRRREELEKVKKEKEEACSVGQNFMRLYEE-EARRRKGTEEEL 453
>At4g36520 trichohyalin like protein
Length = 1432
Score = 33.5 bits (75), Expect = 0.16
Identities = 35/118 (29%), Positives = 54/118 (45%), Gaps = 14/118 (11%)
Query: 179 IRHNFEQMEEERT-KVLIAIEKQK---VSEKEAETMKKMAISEAEKNANVS---KILMEQ 231
++ FE+ EE R + A+E++K + E + + I EA + A + K +EQ
Sbjct: 713 LKEAFEKEEENRRMREAFALEQEKERRIKEAREKEENERRIKEAREKAELEQRLKATLEQ 772
Query: 232 KLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEA-----EANRLKLTPEFLELK 284
+ E+ RQE EN ++ L A+ R +KEA RLK T E E K
Sbjct: 773 EEKERQIKERQEREENER--RAKEVLEQAENERKLKEALEQKENERRLKETREKEENK 828
Score = 30.8 bits (68), Expect = 1.1
Identities = 23/64 (35%), Positives = 31/64 (47%), Gaps = 8/64 (12%)
Query: 204 EKEAETMKKMAISEAEKNANVSKILMEQKLSEKDS--------ARRQEEIENAMYLAREK 255
EKEAE +K+ E E+ V + ++ EKD A +E +E A AREK
Sbjct: 1132 EKEAERLKRERDLEMEQLRKVEEEREREREREKDRMAFDQRALADARERLEKACAEAREK 1191
Query: 256 SLAD 259
SL D
Sbjct: 1192 SLPD 1195
Score = 30.0 bits (66), Expect = 1.8
Identities = 28/101 (27%), Positives = 48/101 (46%), Gaps = 6/101 (5%)
Query: 179 IRHNFEQMEEERTKVLI---AIEKQKVSEK-EAETMKKMAISEAEKNANVSKI--LMEQK 232
I+ E+ E ER V A +++K+ E+ E E K A + E+N + + L ++K
Sbjct: 677 IKEAREKAENERRAVEAREKAEQERKMKEQQELELQLKEAFEKEEENRRMREAFALEQEK 736
Query: 233 LSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANR 273
AR +EE E + AREK+ + +++ E R
Sbjct: 737 ERRIKEAREKEENERRIKEAREKAELEQRLKATLEQEEKER 777
Score = 28.5 bits (62), Expect = 5.3
Identities = 25/96 (26%), Positives = 43/96 (44%), Gaps = 14/96 (14%)
Query: 172 KPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQ 231
K + + ++ EQ E+ER K++ +E E K + +AE + + L EQ
Sbjct: 759 KAELEQRLKATLEQEEKERQI------KERQEREENERRAKEVLEQAENERKLKEAL-EQ 811
Query: 232 KLSEKDSARRQEEIENAMYL-------AREKSLADA 260
K +E+ +E+ EN L +EK L +A
Sbjct: 812 KENERRLKETREKEENKKKLREAIELEEKEKRLIEA 847
Score = 28.1 bits (61), Expect = 6.9
Identities = 26/93 (27%), Positives = 45/93 (47%), Gaps = 5/93 (5%)
Query: 201 KVSEKEAETMKKMAISEA---EKNANVSKILMEQKLSEK--DSARRQEEIENAMYLAREK 255
K++E ++ I EA E+N ++ +E+ +EK +A QEE E + AREK
Sbjct: 624 KLNEPLKRMEEETRIKEARLREENDRRERVAVEKAENEKRLKAALEQEEKERKIKEAREK 683
Query: 256 SLADADFYRVIKEAEANRLKLTPEFLELKFIES 288
+ + ++AE R + LEL+ E+
Sbjct: 684 AENERRAVEAREKAEQERKMKEQQELELQLKEA 716
>At5g06110 cell division related protein-like
Length = 663
Score = 33.1 bits (74), Expect = 0.22
Identities = 36/147 (24%), Positives = 65/147 (43%), Gaps = 18/147 (12%)
Query: 176 PESIRHNFEQMEEERTKVLIAIEK----QKVSEKEAETMKKMAISEAEKNANVSKILMEQ 231
PE H+ EQ E K + E QK ++E ++ + + +K+ + K ++
Sbjct: 246 PEEEEHDIEQAESREEKRWMERENARKTQKARKEEYARIRTLVDNAYKKDIRIQKRKDDE 305
Query: 232 K---LSEKDS---ARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKF 285
K L +K++ A+RQ+E A + EK + + R AEA +L + E K
Sbjct: 306 KAKKLQKKEAKVMAKRQQEEAAAAAIEEEKRRKEEEAKRA---AEAAQLHKRAKEREKKL 362
Query: 286 IESIANNTKIFFGDKIPNMILDQRLLG 312
+ + ++ + +L QRLLG
Sbjct: 363 LRKERSRLRV-----LSAPVLSQRLLG 384
>At4g11420 putative protein
Length = 987
Score = 33.1 bits (74), Expect = 0.22
Identities = 23/63 (36%), Positives = 38/63 (59%), Gaps = 7/63 (11%)
Query: 185 QMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSE---KDSARR 241
++EEER + L E+ + E EAE +KK+ EAE+ AN+ K +Q+ E ++ +RR
Sbjct: 798 KIEEERIRKLQEEEEARKQE-EAERLKKV---EAERKANLDKAFEKQRQREIELEEKSRR 853
Query: 242 QEE 244
+ E
Sbjct: 854 ERE 856
>At3g02930 unknown protein
Length = 806
Score = 33.1 bits (74), Expect = 0.22
Identities = 32/130 (24%), Positives = 59/130 (44%), Gaps = 24/130 (18%)
Query: 128 CSSHSLQQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQME 187
C+S SL V + ++ ++ + T IE++ + V + +E
Sbjct: 331 CASVSLVSVTKQL--EVSNSRLHDMESEITDLKEKIELLEMTVASQKV---------DLE 379
Query: 188 EERTKVLIAIEKQKVSEKEAETMK-----------KMAISEAEKNANVSKILMEQK--LS 234
+ K+ IA E+ SEKEAE +K + E + ++V ++L E+K LS
Sbjct: 380 KSEQKLGIAEEESSKSEKEAEKLKNELETVNEEKTQALKKEQDATSSVQRLLEEKKKILS 439
Query: 235 EKDSARRQEE 244
E +S++ +EE
Sbjct: 440 ELESSKEEEE 449
>At5g65250 unknown protein
Length = 300
Score = 32.7 bits (73), Expect = 0.28
Identities = 27/102 (26%), Positives = 50/102 (48%), Gaps = 10/102 (9%)
Query: 163 IEIIGVR--VTKPNIPESIRHN--FEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEA 218
+E +G+R VT+ + E I Q E T+VL+A +Q++ EKE ++K+ ++
Sbjct: 193 LEKLGIRFRVTRKALKEPISETAALAQKNSEATRVLVA--QQEILEKELGEIQKVLLAMQ 250
Query: 219 EKNANVSKILM----EQKLSEKDSARRQEEIENAMYLAREKS 256
E+ ++++ KL E ++ +Q E A E S
Sbjct: 251 EQQRKQLELILTIAKSSKLFESSTSSKQSPSEQRKNKAEEPS 292
>At1g30690 putative phosphatidyl-inositol-transfer protein
Length = 540
Score = 32.7 bits (73), Expect = 0.28
Identities = 26/93 (27%), Positives = 45/93 (47%), Gaps = 11/93 (11%)
Query: 201 KVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADA 260
KV EK+ E+ +A + + V ++ E K+ E +S + E +E + E +
Sbjct: 6 KVEEKQVESEVVIAPAVVPEETTVKAVVEETKVEEDES--KPEGVEKSASFKEE-----S 58
Query: 261 DFYRVIKEAEANRLKLTPEFLELKFIESIANNT 293
DF+ +KE+E L L+ K E+I +NT
Sbjct: 59 DFFADLKESEKKAL----SDLKSKLEEAIVDNT 87
>At1g30480 DNA damage repair protein, putative
Length = 387
Score = 32.7 bits (73), Expect = 0.28
Identities = 16/59 (27%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIP--ESIRHNFE-QMEEERTKVLIAIE 198
Q+D++++D + +C +Y ++ +T+PN P E++R + EE TK L+ ++
Sbjct: 295 QVDDELEDEVGGECGKYGTVTRVLIFEITEPNFPVHEAVRIFVQFSRPEETTKALVDLD 353
>At5g44800 helicase-like protein
Length = 2228
Score = 32.3 bits (72), Expect = 0.37
Identities = 34/115 (29%), Positives = 50/115 (42%), Gaps = 26/115 (22%)
Query: 160 APGIEIIGVRVTKPNI--------PESIRHNF-------EQMEEERTKVLIAIEKQKVSE 204
APG + G+ T +I PE+I + E++E ERT L + EKQ+ E
Sbjct: 2124 APGPSLEGITGTTKSISLESQSSEPETINQDGDLDPETDEKVESERTP-LHSDEKQEEQE 2182
Query: 205 KEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLAD 259
E K+ EAE Q + ++ A QEE E +M + SL+D
Sbjct: 2183 SENALNKQCEPIEAES----------QNTNAEEEAEAQEEDEESMKMVTGNSLSD 2227
>At4g08550 hypothetical protein
Length = 587
Score = 32.3 bits (72), Expect = 0.37
Identities = 21/72 (29%), Positives = 37/72 (51%), Gaps = 4/72 (5%)
Query: 178 SIRHNFEQMEEERT-KVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
S +HN E+ EEE T K ++ IE++ E+E E++ E + N ++ +I+ +
Sbjct: 46 SPKHNVEKKEEEFTRKPVVEIEEE---EEEMESIDIHEEEEGDNNVSLDEIMSVDSSDDD 102
Query: 237 DSARRQEEIENA 248
D + EI A
Sbjct: 103 DDSESSAEITKA 114
>At5g67320 unknown protein
Length = 613
Score = 32.0 bits (71), Expect = 0.48
Identities = 38/128 (29%), Positives = 59/128 (45%), Gaps = 16/128 (12%)
Query: 186 MEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEI 245
ME+ER + + E K E+E E + A E +++ + E++ E++ R +E+I
Sbjct: 108 MEKERDR---SKENDKGVEREHEGDRNRA-KEKDRHEKQKEREREREKLEREKEREREKI 163
Query: 246 ENAMYLAREKSLADADFYRVIKEAEANRLKLTPE-----FLELKFIESIANNTKIFFGDK 300
E REK R I E E +RLKL E E + IE ++ K GD
Sbjct: 164 EREKEREREK------MEREIFEREKDRLKLEKEREIEREREREKIEREKSHEK-QLGDA 216
Query: 301 IPNMILDQ 308
M++DQ
Sbjct: 217 DREMVIDQ 224
Score = 29.3 bits (64), Expect = 3.1
Identities = 27/98 (27%), Positives = 42/98 (42%), Gaps = 14/98 (14%)
Query: 187 EEERTKVLIAIEKQKVSEKEAETMKKMAISEAEK---NANVSKILMEQKLSEKDSARRQE 243
+ R K EKQK E+E E +++ E EK + ME+++ E++ R +
Sbjct: 129 DRNRAKEKDRHEKQKEREREREKLEREKEREREKIEREKEREREKMEREIFEREKDRLKL 188
Query: 244 EIENAMYLAR-----------EKSLADADFYRVIKEAE 270
E E + R EK L DAD VI + +
Sbjct: 189 EKEREIEREREREKIEREKSHEKQLGDADREMVIDQTD 226
>At5g65770 nuclear matrix constituent protein 1 (NMCP1)-like
Length = 1042
Score = 32.0 bits (71), Expect = 0.48
Identities = 31/134 (23%), Positives = 64/134 (47%), Gaps = 16/134 (11%)
Query: 165 IIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEA-ETMKKMAISEAEK--- 220
++G+ + K + I + E++E A E++K E+E +++K+MA E E
Sbjct: 645 LLGIEMQKRELEYCIENKREELENSSRDREKAFEQEKKLEEERIQSLKEMAEKELEHVQV 704
Query: 221 -----NANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLK 275
+A +I ++++ E++ A ++ +E + + REK R A R +
Sbjct: 705 ELKRLDAERLEIKLDRERREREWAELKDSVEE-LKVQREKLETQRHMLR------AERDE 757
Query: 276 LTPEFLELKFIESI 289
+ E ELK +E++
Sbjct: 758 IRHEIEELKKLENL 771
>At5g56360 unknown protein
Length = 647
Score = 32.0 bits (71), Expect = 0.48
Identities = 24/101 (23%), Positives = 48/101 (46%), Gaps = 8/101 (7%)
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQK---- 232
+ ++ EQ+E+ K + EK++ +KEAE + +AE+ + S+ + E
Sbjct: 191 DQLKDRKEQIEKVEEKERLQKEKEEKEKKEAELAAQQGKGDAEEKTDDSEKVEESSHDEG 250
Query: 233 ---LSEKDSARRQEEIENAM-YLAREKSLADADFYRVIKEA 269
+S+ D +EI N Y + E+ A+ + ++ EA
Sbjct: 251 TPAVSQHDETTHHDEIGNYKDYPSDEEPAAEGEPTSILDEA 291
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,956,375
Number of Sequences: 26719
Number of extensions: 290250
Number of successful extensions: 1595
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 1393
Number of HSP's gapped (non-prelim): 247
length of query: 331
length of database: 11,318,596
effective HSP length: 100
effective length of query: 231
effective length of database: 8,646,696
effective search space: 1997386776
effective search space used: 1997386776
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146567.11


