

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.10 + phase: 0
(393 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
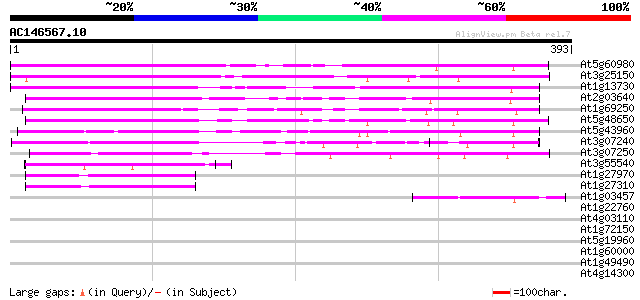
Score E
Sequences producing significant alignments: (bits) Value
At5g60980 ras-GTPase-activating protein SH3-domain binding prote... 253 1e-67
At3g25150 RNA binding like protein 244 5e-65
At1g13730 unknown protein 226 1e-59
At2g03640 unknown protein 213 2e-55
At1g69250 RNA-binding protein like 195 4e-50
At5g48650 NTF2-containing RNA-binding protein-like 156 2e-38
At5g43960 unknown protein 121 6e-28
At3g07240 putative RNA-binding protein 104 1e-22
At3g07250 putative RNA-binding protein 97 2e-20
At3g55540 putative protein 81 1e-15
At1g27970 unknown protein 52 6e-07
At1g27310 similar to nuclear transport factor 2 49 5e-06
At1g03457 ribonucleoprotein like 45 9e-05
At1g22760 putative polyA-binding protein, PAB3 40 0.002
At4g03110 putative ribonucleoprotein 40 0.002
At1g72150 putative cytosolic factor protein 40 0.003
At5g19960 glycine-rich RNA-binding protein - like 39 0.004
At1g60000 unknown protein 39 0.005
At1g49490 hypothetical protein 38 0.008
At4g14300 ribonucleoprotein like protein 38 0.011
>At5g60980 ras-GTPase-activating protein SH3-domain binding protein
- like
Length = 460
Score = 253 bits (647), Expect = 1e-67
Identities = 161/386 (41%), Positives = 222/386 (56%), Gaps = 37/386 (9%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA EA +P ++VG AFVEQYY ILH+ P VHRFY DSS ++RP+ G +TTVTT
Sbjct: 1 MAQQEASPSPGAEVVGRAFVEQYYHILHQSPGLVHRFYQDSSFLTRPDVTGAVTTVTTMQ 60
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+ KI SL+Y + E+ +ADAQ S+ GV+V+VTG LTG DN+++KF+QSFFLAPQDK
Sbjct: 61 AINDKILSLKYEDYTAEIETADAQESHERGVIVLVTGRLTGNDNVRKKFSQSFFLAPQDK 120
Query: 121 GFYVLNDVFRYVDAYK-SIDIESVPANDADESAPSEAIITPEPEPV-HVPEVIPPTQTVI 178
G++VLNDVFR+++ + + SVP N +A I PE V H PEV P
Sbjct: 121 GYFVLNDVFRFLEEKEVTAQARSVPINGTTRDV--QAPIEPERVVVSHEPEVEPE----- 173
Query: 179 PTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPT 238
P A I + + EV P + + V +V P+ EP
Sbjct: 174 PVAS---------IEEEDLDNVAEVYDPSDKDE-GVVVDVEPI-----------EPPTQI 212
Query: 239 SIEKVASNTQEDTPKKSFASIVNALKDNSAP-FHLRASPAKPAVHPPRVHSVPAPEAPTP 297
S ++ S Q D PK S+ASI+ +K + AP H+ + +PA ++ + PA A P
Sbjct: 213 SHNEILSVPQGDAPKHSYASILKQMKSSPAPTTHVARNKPRPAPVNQKLTAPPAEPAARP 272
Query: 298 ----NMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK-- 351
+ ++P + + H+I+V NLP +T QL+ FK FG IK +GIQVRSNK
Sbjct: 273 EASAHENVPNSSHVDVEDDGHSIYVRNLPFDSTPTQLEEVFKNFGAIKHEGIQVRSNKQQ 332
Query: 352 GSCFGFVEFESAASMQSALEVWSISI 377
G CFGFVEFE+++ QSALE ++I
Sbjct: 333 GFCFGFVEFETSSGKQSALEASPVTI 358
>At3g25150 RNA binding like protein
Length = 488
Score = 244 bits (624), Expect = 5e-65
Identities = 148/394 (37%), Positives = 214/394 (53%), Gaps = 34/394 (8%)
Query: 1 MAVPEAVQTP----TPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTV 56
MA+ A Q P TP MVGNAFV QYY ILH+ P+ VHRFY + S + RPEE+G M+
Sbjct: 1 MAMLGAQQVPAAACTPDMVGNAFVPQYYHILHQSPEHVHRFYQEISKLGRPEENGLMSIT 60
Query: 57 TTTAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLA 116
+T IDKKI +L Y E+ + D Q S+ G +V+VTG LTG D+++R F+Q+FFLA
Sbjct: 61 STLQAIDKKIMALGYGVISAEIATVDTQESHGGGYIVLVTGYLTGKDSVRRTFSQTFFLA 120
Query: 117 PQDKGFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQT 176
PQ+ G++VLND+FR++D + +P N+ AP + + IP
Sbjct: 121 PQETGYFVLNDMFRFIDEGTVVHGNQIPVNNV--QAPVNTY-----QDTAAAKEIPDDFV 173
Query: 177 VIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQ 236
Q QT + ++V P E+ ++S E + V E+
Sbjct: 174 QEKYVQENHAVKQTEVLSKSINEPEKVFTPSEDEQVSAAEEALVTETVNEA--------- 224
Query: 237 PTSIEKVASNTQE--DTPKKSFASIVNALKDNSAPFHLRASPAK--------PAVHPPRV 286
P ++KV + + PK+S+ASIV +K+N+AP +P K A+H P
Sbjct: 225 PIEVQKVGESDSRTGEIPKRSYASIVKVMKENAAPMSASRTPTKVEPKKQEDQAIHIPL- 283
Query: 287 HSVPAPEAPTPNMDIPLEKNNENAGRA--HAIFVANLPMSATVEQLDRAFKKFGPIKRDG 344
P E ++ + +NN+ RA +I++ LP+ AT L+ F+KFG I+ +G
Sbjct: 284 -PTPLSEKSDSGANVAVNENNQENERALGPSIYLKGLPLDATPALLENEFQKFGLIRTNG 342
Query: 345 IQVRSNKGSCFGFVEFESAASMQSALEVWSISIN 378
IQVRS KG CFGFVEFESA+SMQSA+E + +N
Sbjct: 343 IQVRSQKGFCFGFVEFESASSMQSAIEASPVMLN 376
>At1g13730 unknown protein
Length = 428
Score = 226 bits (577), Expect = 1e-59
Identities = 144/374 (38%), Positives = 200/374 (52%), Gaps = 41/374 (10%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA+ P +GN+FVE+YY++L++ P QVH+FY D SV+ RP DG M +V +
Sbjct: 1 MALESNAPVVDPNTIGNSFVEKYYNLLYKSPSQVHQFYLDDSVLGRPGSDGEMVSVKSLK 60
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+++I S +Y ++++L+AD+Q SY NGV+ +VTG LT + + +F+QSFFL P +
Sbjct: 61 AINEQIMSFDYEISKIQILTADSQASYMNGVVTLVTGLLTVKEGQRMRFSQSFFLVPLNG 120
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
++VLNDVFRYV ADE EA E E V +P+V+ PT+ V
Sbjct: 121 SYFVLNDVFRYV---------------ADEIVEPEA-NKKEVEEV-IPQVVQPTEQVDEV 163
Query: 181 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSI 240
A+ V PTQ A TEN + ++ H K E
Sbjct: 164 AEPVTIPTQQPEAK------------------QTTENTVKKPERAVANGHPKTQEDNVVN 205
Query: 241 EKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP-PRVHSVPAPEAPTPNM 299
+K + D PKKSFA IV L N A F+ +ASPAKP P + + +AP P
Sbjct: 206 DK---SNGVDAPKKSFAHIVQDLAQNGATFNAKASPAKPKSKPVTKPSAARESKAPAPVS 262
Query: 300 DIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSN--KGSCFGF 357
+ + + IFVANL M AT EQL+ FK FG I +DGIQVRS KG+CFGF
Sbjct: 263 EHSSAATIDQQAEGYTIFVANLLMDATPEQLNETFKGFGAITKDGIQVRSYRLKGNCFGF 322
Query: 358 VEFESAASMQSALE 371
V F SA +++ L+
Sbjct: 323 VTFASAEAVKLVLQ 336
>At2g03640 unknown protein
Length = 422
Score = 213 bits (541), Expect = 2e-55
Identities = 136/365 (37%), Positives = 202/365 (55%), Gaps = 47/365 (12%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
PQ VGN FV++YY+ L+ +VH+FY + S++SRP DG + T+ + I+ +I S++Y
Sbjct: 12 PQFVGNGFVQEYYNHLYDSTSEVHKFYLEDSMISRPGLDGEIVTIKSLKGINDQIMSIDY 71
Query: 72 TSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRY 131
S R+E+L+AD+Q + NGV+ +VTG + G D +RKF+QSFFL ++ ++VLND FRY
Sbjct: 72 KSSRIEILTADSQSTLKNGVVTLVTGLVIGNDGGRRKFSQSFFLVSRNGSYFVLNDTFRY 131
Query: 132 V-DAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
V D + +E + +ES + AI P E V + VI PT
Sbjct: 132 VSDEF----VEPEATKEVEESQSTNAITEPANESV----------------EAVIVPT-- 169
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQED 250
+ +T ++K S + NG V E + V E+S K E + QE+
Sbjct: 170 ---EAKTTVTKPAS-AIPNGHAKVPEEKV----VNENSSLPKAAE---------AKLQEE 212
Query: 251 TPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPE--APTPNMDIPLEKNNE 308
PKKSFA IV +L ++ ++ASP K P V APE AP+P ++ +
Sbjct: 213 VPKKSFALIVQSLAQSAGTLQVKASPVK---RKPVEKPVAAPERKAPSPIRKQASAESIK 269
Query: 309 NAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRS--NKGSCFGFVEFESAASM 366
+ +IFVANLPM AT+EQL FK FG I++DGIQVRS K +C GFV FE+ ++
Sbjct: 270 PQAQGSSIFVANLPMDATIEQLYETFKSFGAIRKDGIQVRSYPEKKNCIGFVAFENGEAV 329
Query: 367 QSALE 371
++ +
Sbjct: 330 KNVFQ 334
>At1g69250 RNA-binding protein like
Length = 427
Score = 195 bits (495), Expect = 4e-50
Identities = 132/375 (35%), Positives = 203/375 (53%), Gaps = 54/375 (14%)
Query: 10 PTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSL 69
P+ Q + FV QYY +L + P + R Y D+SV+SRP+ GTM + T+ I+K I S
Sbjct: 8 PSAQDIAAEFVRQYYHVLGQLPHEARRLYVDASVVSRPDVTGTMMSFTSVEAINKHILSC 67
Query: 70 EYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVF 129
++ + + EVLS D+Q S +G+ ++V G +TG DN +RKF+Q F+LA Q+ VLND+
Sbjct: 68 DFENTKFEVLSVDSQNSLEDGIFIMVIGFMTGKDNQRRKFSQMFYLARQNT-LVVLNDML 126
Query: 130 RYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQ 189
RYV D ++S+ +E P V E++ P + +T + Q
Sbjct: 127 RYV--------------DQEDSSTTETPCEP------VTEIVRPADGLKKAEKTEL--KQ 164
Query: 190 TVIADTETIISKEV---SLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASN 246
+A E ++ V + PL+NGK+ +E + + V EP+ +
Sbjct: 165 KNVASVEKSVNAAVEKNAAPLDNGKMKQSEKAV-------ITQKVTEPD---------AA 208
Query: 247 TQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVP----APEAPTPNMDIP 302
Q D K+SFA IV ++ N+APF ++ SP + V P+ P AP+ P +
Sbjct: 209 PQPDGAKRSFADIVGSMAKNAAPFQVK-SPVQAPVQKPKYVGQPRAAAAPQKPA-YVSKS 266
Query: 303 LEKNNENAGR--AHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS----CFG 356
++KN++ +IFVANLP++A QL FK FGPIK +GIQVRS++G+ CFG
Sbjct: 267 IKKNDQKVIEVPGTSIFVANLPLNAMPPQLFELFKDFGPIKENGIQVRSSRGNANPVCFG 326
Query: 357 FVEFESAASMQSALE 371
F+ FE+ AS+QS L+
Sbjct: 327 FISFETVASVQSVLQ 341
>At5g48650 NTF2-containing RNA-binding protein-like
Length = 458
Score = 156 bits (395), Expect = 2e-38
Identities = 121/404 (29%), Positives = 190/404 (46%), Gaps = 70/404 (17%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P VG+AFV QYY I P+ + RFY + S + R +DG M +T I ++++ L Y
Sbjct: 13 PLTVGSAFVNQYYYIFCNMPEHLPRFYQEISRVGRVGQDGVMRDFSTFQGISEELKRLTY 72
Query: 72 TSFR-VEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
E+ S D Q S+N G ++ VTG T + +RKF Q+FFLAPQ+KGF+VLND+ R
Sbjct: 73 GDCNSAEITSYDTQESHNGGFLLFVTGYFTLNERSRRKFTQTFFLAPQEKGFFVLNDILR 132
Query: 131 YVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
+V+ +DA ++ P EV+ + PT + ++
Sbjct: 133 FVN------------DDAKDNVPETI----------DGEVVSGINSTTPTIINGMKGSEQ 170
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE- 249
+ + KEVS PL+N + +NV+ V E ++ V E + ++VA ++Q+
Sbjct: 171 AACVSVNPVCKEVSKPLDNE--NAKDNVL----VPEIANEVARTE--ITCKEVADDSQKN 222
Query: 250 --------DTPKKSFASIVNALKDN-SAPFHLRASPAKPAVHPPRVHSVPAP-------- 292
D PKKS+AS++ KD P SP K + + H P+
Sbjct: 223 YDPDDGLADAPKKSYASVLKVTKDKFGVPAVSLPSPKK--IPKDQEHQAPSDPSTGQILK 280
Query: 293 --------------EAPTPNMDIPLEKNNEN---AGRAHAIFVANLPMSATVEQLDRAFK 335
E+ T + + +N N +I+V +LP +A ++ L+ FK
Sbjct: 281 DQGQQASSDPSQVIESDTVSESVDASENGHNQEAVAEGTSIYVRHLPFNANIDMLEAEFK 340
Query: 336 KFGPIKRDGIQVRSNK--GSCFGFVEFESAASMQSALEVWSISI 377
+FG I GIQV + + G +GFVEFE A + A+E + I
Sbjct: 341 QFGAITNGGIQVINQRGLGYPYGFVEFEEADAAHRAIEASPVKI 384
>At5g43960 unknown protein
Length = 450
Score = 121 bits (304), Expect = 6e-28
Identities = 107/400 (26%), Positives = 186/400 (45%), Gaps = 59/400 (14%)
Query: 6 AVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKK 65
A P VG+ FV QYY +L + PD +H+FY + S R + D T T + I
Sbjct: 2 ATPYPGATQVGSYFVGQYYQVLQQQPDLIHQFYSEPSRAIRIDGDST-ETANSLLHIHNM 60
Query: 66 IQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKGFYV 124
+ SL +T+ +EV + ++ S+ GV+VVV+G + T + +R F Q+FFLAPQ+KG++V
Sbjct: 61 VMSLNFTA--IEVKTINSVESWEGGVLVVVSGSVKTKEFSNRRSFVQTFFLAPQEKGYFV 118
Query: 125 LNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTV 184
L+DVF +VD PS E H ++ PPT+ P
Sbjct: 119 LSDVFLFVD-----------EGTVYYHQPSYL-----SEIKHEAQLNPPTRHPDPQVSDY 162
Query: 185 IPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVK-ESSHHVKE-PEQPTSIEK 242
+ + ++ + + ++ L + K S+ E+ H E ++E P + +++
Sbjct: 163 VLEEEA----SDYVNAVQIKDDLVD-KYSLQEDQHQPQHEDYEDEVAIEETPREEVAVDV 217
Query: 243 V----ASNTQE---DTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAP-EA 294
V A+ +E + K S+ASI+ K+ +A + A+ ++ P
Sbjct: 218 VHEHRAAPVEEPVGEKSKMSYASILKVAKE-AATVPVAATQPSYNKSSQDINEWDQPMRT 276
Query: 295 PTPNMDIPLEKNNENAGRAH--------------------AIFVANLPMSATVEQLDRAF 334
P+P + PL ++ + +++V NLP + +++ F
Sbjct: 277 PSPQLAAPLAPIQQSNSSTYVSDYGAEAEDGSGFEDFEFKSVYVRNLPSDISASEIEEEF 336
Query: 335 KKFGPIKRDGIQVRSNK---GSCFGFVEFESAASMQSALE 371
K FG IK DG+ +R+ K G C+ FVEFE S+++A++
Sbjct: 337 KNFGTIKPDGVFLRTRKDVMGVCYAFVEFEDMTSVENAIK 376
>At3g07240 putative RNA-binding protein
Length = 369
Score = 104 bits (259), Expect = 1e-22
Identities = 107/399 (26%), Positives = 172/399 (42%), Gaps = 90/399 (22%)
Query: 2 AVPEAVQTPTPQ-MVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
A P A T Q ++G+ F E YY+ L P+ + +Y D S ++RP DGTM + +T
Sbjct: 5 ARPSAKAVYTIQTIIGDGFAEHYYNNLQNSPEILPGYYKDVSKITRPGLDGTMRS-STLP 63
Query: 61 EIDKKIQSLEYTSF-RVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQD 119
+I + + L F VEV S +Q S++ G+ V V G T + R F Q+F APQ+
Sbjct: 64 DIIEDLDMLSPGGFDSVEVTSVMSQDSHDKGIRVAVDGYFTFNERPARNFTQNFTFAPQE 123
Query: 120 KGFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIP 179
KG +V D+F++V IP
Sbjct: 124 KGLFVSTDMFKFVG--------------------------------------------IP 139
Query: 180 TAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENV-----------IPVNHVKESS 228
A IPP A+ I K +LPL N +++ EN + V + + S
Sbjct: 140 EANATIPP-----ANNAAICVK--NLPL-NATIALVENAFKQFGEIRRGGVEVRNKRSFS 191
Query: 229 HHVKEPEQPTSIEK-----------VASNTQEDTPKKSFASIVNALKDNSAPFHLRASPA 277
+ E ++ + ++ V N + +P + + + P ++R
Sbjct: 192 YGFVEFKEENAAQRAIKNCLIGFDNVGMNLMQASPVT--IDLRSVYVEKKRPDYIRYWDT 249
Query: 278 KPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVA----NLPMSATVEQLDRA 333
P+ P ++ E + D + NEN A++ A NLP + T + ++ A
Sbjct: 250 -PSTGPGIIY---RSEGMSVTKDYGNKGGNENNQEPRALYAAVHVKNLPPNVTTDWVENA 305
Query: 334 FKKFGPIKRDGIQVRSNK--GSCFGFVEFESAASMQSAL 370
FK+FGPIKR G+QV SN+ G+ FG V+F AA+ + A+
Sbjct: 306 FKQFGPIKRGGVQV-SNRGVGNWFGNVKFVHAAAAERAV 343
Score = 58.2 bits (139), Expect = 8e-09
Identities = 30/77 (38%), Positives = 46/77 (58%), Gaps = 7/77 (9%)
Query: 295 PTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSC 354
P N IP N AI V NLP++AT+ ++ AFK+FG I+R G++VR+ +
Sbjct: 139 PEANATIPPANNA-------AICVKNLPLNATIALVENAFKQFGEIRRGGVEVRNKRSFS 191
Query: 355 FGFVEFESAASMQSALE 371
+GFVEF+ + Q A++
Sbjct: 192 YGFVEFKEENAAQRAIK 208
>At3g07250 putative RNA-binding protein
Length = 946
Score = 96.7 bits (239), Expect = 2e-20
Identities = 101/385 (26%), Positives = 155/385 (40%), Gaps = 78/385 (20%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
VG+ F QYY+ L P+ +++ Y D S +SRP DGTM T + K ++ SF
Sbjct: 281 VGDEFARQYYNTLQNAPENLYKLYKDKSTISRPGLDGTMRVFT----LSKDLKWRSPGSF 336
Query: 75 -RVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRYVD 133
V++ S +Q S G++VVV G LT + R F Q FFL PQ+KG+ V D+
Sbjct: 337 DSVKITSVTSQDSLKQGILVVVYGYLTFNERPARHFTQVFFLVPQEKGYIVCTDM----- 391
Query: 134 AYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIA 193
++ +DI P A IPP
Sbjct: 392 -FRFVDI--------------------------------------PEANAAIPPAND--- 409
Query: 194 DTETIISKEVSLPLENGKLSVTENVIPVNH-----VKESSHHVKE-PEQPTSIEKVASNT 247
+ N SV E +P V E +H P+ + E A
Sbjct: 410 ------GSKCLRSWFNANGSVIEEKVPETEGAALRVSEPNHGFDNVPKLSCASEGAAICA 463
Query: 248 QEDTPKKSFASIVNALKD----NSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNM--DI 301
++ + A + NA K +R + A P V A N+ D
Sbjct: 464 KKLPLDATIAFVENAFKQFGEIRRGGVEVRINWASPISIDGYRTYVEKKYAYNKNIKADA 523
Query: 302 PLEKNNENAGRAHAIF------VANLPMSATVEQLDRAFKKFGPIKRDGIQV--RSNKGS 353
+ N N+ + AI V +LP +ATV ++ FK+FGPIK+ I+V +N
Sbjct: 524 GADTGNGNSQDSQAITEDAHIRVKDLPPNATVALVESVFKQFGPIKKGRIRVINPANSNY 583
Query: 354 CFGFVEFESAASMQSALEVWSISIN 378
+ FVEFE A + + A++ ++++
Sbjct: 584 WYAFVEFEEADAAKRAIQASPLNVD 608
Score = 37.4 bits (85), Expect = 0.014
Identities = 23/69 (33%), Positives = 36/69 (51%), Gaps = 6/69 (8%)
Query: 315 AIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESA-----ASMQSA 369
++F+ LP+ T ++ A KKFGPI I ++S F FV+FE A A M S
Sbjct: 738 SVFLKRLPLDVTFALIEDALKKFGPINAISI-IKSGPLYKFAFVDFEKADVANRAIMASP 796
Query: 370 LEVWSISIN 378
+ + ++N
Sbjct: 797 VRICEKNVN 805
>At3g55540 putative protein
Length = 334
Score = 80.9 bits (198), Expect = 1e-15
Identities = 51/140 (36%), Positives = 81/140 (57%), Gaps = 8/140 (5%)
Query: 11 TPQMVGNAFVEQYYSILHRDPDQVHR-FYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSL 69
T + + N F++ Y+ L P+ V +Y D SVM+RP DGTM + T+ I ++I S
Sbjct: 28 TSEALANCFLQSYFLNLGVYPEVVQMMWYADDSVMTRPGPDGTMMSFTSPEAIQEQIVSC 87
Query: 70 EYTSFRVEVLSADAQ---PSYNNGVMVVVTGCLTGTD-NIKRKFAQSFFLA-PQDKGFYV 124
+Y +V+S AQ S +G ++VTG LT D ++R+F QS +LA QD+ + +
Sbjct: 88 DYEGASFDVMSFAAQSCNTSSEDGAFIMVTGFLTCKDKQVRRRFVQSLYLARRQDRSYAI 147
Query: 125 LNDVFRYVDAYKSIDIESVP 144
+ND RY+D+ + + SVP
Sbjct: 148 VNDFLRYIDSIPA--LPSVP 165
Score = 55.1 bits (131), Expect = 7e-08
Identities = 44/149 (29%), Positives = 72/149 (47%), Gaps = 9/149 (6%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDG--TMTTVTTTAEIDKKIQSL 69
P+ G FV+ YY + R+ ++ Y + S+MSRP TM + I+K++ +
Sbjct: 165 PESAG--FVKVYYELPMRE--ELGLMYVNESIMSRPTSTSGRTMVEMPGLDAINKRVSNE 220
Query: 70 EYTSFRVEVLSADAQ--PSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVL-N 126
+ + S D Q S+ + + ++V G +T D +RKF Q F++ G YV+ N
Sbjct: 221 HKRASNFILNSVDYQISRSFKDRMFIMVCGFVTLDDKTERKFLQFFYVTRCHNGSYVIYN 280
Query: 127 DVFRYVDAYKSIDIESVPANDADESAPSE 155
D+ RYVD +ES + A S E
Sbjct: 281 DILRYVDVTPRDTLESSSQSAAKPSTDVE 309
>At1g27970 unknown protein
Length = 126
Score = 52.0 bits (123), Expect = 6e-07
Identities = 38/122 (31%), Positives = 60/122 (49%), Gaps = 8/122 (6%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P V AFVE YYS + + Y ++S+++ + + I K+ SL +
Sbjct: 6 PDAVSKAFVEHYYSTFDTNRVGLAGLYQEASMLTFEGQ-----KIQGVQSIVAKLTSLPF 60
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + + D QPS +G++V V+G L + KF+Q F L P +G FYV ND+
Sbjct: 61 QQCKHHISTVDCQPSGPASGMLVFVSGNLQLAGEEHALKFSQMFHLMPTPQGSFYVFNDI 120
Query: 129 FR 130
FR
Sbjct: 121 FR 122
>At1g27310 similar to nuclear transport factor 2
Length = 122
Score = 48.9 bits (115), Expect = 5e-06
Identities = 34/121 (28%), Positives = 55/121 (45%), Gaps = 7/121 (5%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P V AFVE YYS + + Y + S+++ + + + I K+ L +
Sbjct: 3 PDAVAKAFVEHYYSTFDANRPGLVSLYQEGSMLTFEGQK-----IQGSQNIVAKLTGLPF 57
Query: 72 TSFRVEVLSADAQPSYN-NGVMVVVTGCLT-GTDNIKRKFAQSFFLAPQDKGFYVLNDVF 129
+ + + D QPS G++V V+G L + KF+Q F L +YV ND+F
Sbjct: 58 QQCKHNITTVDCQPSGPAGGMLVFVSGNLQLAGEQHALKFSQMFHLISNQGNYYVFNDIF 117
Query: 130 R 130
R
Sbjct: 118 R 118
>At1g03457 ribonucleoprotein like
Length = 429
Score = 44.7 bits (104), Expect = 9e-05
Identities = 31/110 (28%), Positives = 51/110 (46%), Gaps = 8/110 (7%)
Query: 283 PPRVH-SVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIK 341
PPR+ + +P IP E A+ +F+ N+P ++L F+ FG +
Sbjct: 299 PPRLPLTTVSPGISNNGTSIPSSLQTEGPAGAN-LFIYNIPREFEDQELAATFQPFGKVL 357
Query: 342 RDGIQVRSNKG--SCFGFVEFESAASMQSALEVWSISINSCYQQGCKLNV 389
+ V G CFGF+ ++S A+ Q+A+ ++N C G KL V
Sbjct: 358 SAKVFVDKATGISKCFGFISYDSQAAAQNAIN----TMNGCQLSGKKLKV 403
>At1g22760 putative polyA-binding protein, PAB3
Length = 660
Score = 40.4 bits (93), Expect = 0.002
Identities = 22/58 (37%), Positives = 32/58 (54%), Gaps = 3/58 (5%)
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS--CFGFVEFESAASMQSALE 371
++V NLP ++L + F KFG I + +R G+ CFGFV FE + SA+E
Sbjct: 231 VYVKNLPKEIGEDELRKTFGKFGVIS-SAVVMRDQSGNSRCFGFVNFECTEAAASAVE 287
Score = 31.6 bits (70), Expect = 0.79
Identities = 24/83 (28%), Positives = 38/83 (44%), Gaps = 7/83 (8%)
Query: 296 TPNMDIPLE---KNNENAGRAHA---IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVR- 348
TP D P+ N + + R IF+ NL S + L F FG I + +
Sbjct: 112 TPLFDRPIRIMLSNRDPSTRLSGKGNIFIKNLDASIDNKALFETFSSFGTILSCKVAMDV 171
Query: 349 SNKGSCFGFVEFESAASMQSALE 371
+ + +GFV+FE S Q+A++
Sbjct: 172 TGRSKGYGFVQFEKEESAQAAID 194
>At4g03110 putative ribonucleoprotein
Length = 441
Score = 40.0 bits (92), Expect = 0.002
Identities = 29/106 (27%), Positives = 51/106 (47%), Gaps = 11/106 (10%)
Query: 291 APEAPTPNMDIPLEKNNENAGRAHA-----IFVANLPMSATVEQLDRAFKKFGPIKRDGI 345
+P + +P M PL + + +F+ N+P ++L AF+ FG + +
Sbjct: 321 SPGSISPGMSTPLGIGLSSVVQTEGPEGANLFIYNIPREFGDQELAAAFQSFGIVLSAKV 380
Query: 346 QVRSNKG--SCFGFVEFESAASMQSALEVWSISINSCYQQGCKLNV 389
V G CFGFV ++S A+ Q+A+++ +N + G KL V
Sbjct: 381 FVDKATGVSKCFGFVSYDSQAAAQNAIDM----MNGRHLGGKKLKV 422
Score = 28.5 bits (62), Expect = 6.7
Identities = 18/63 (28%), Positives = 34/63 (53%), Gaps = 9/63 (14%)
Query: 314 HAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQV-----RSNKGSCFGFVEFESAASMQS 368
H +FV LP + + ++ F K+G IK +Q+ +++KG F+++E+ S
Sbjct: 106 HKLFVGMLPKNVSEAEVQSLFSKYGTIK--DLQILRGAQQTSKGC--AFLKYETKEQAVS 161
Query: 369 ALE 371
A+E
Sbjct: 162 AME 164
>At1g72150 putative cytosolic factor protein
Length = 573
Score = 39.7 bits (91), Expect = 0.003
Identities = 49/207 (23%), Positives = 72/207 (34%), Gaps = 36/207 (17%)
Query: 141 ESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIADTETIIS 200
E V A +DE A E +TPE E P + +TV+ + V+ E +
Sbjct: 38 EEVAAPVSDEKAVPEKEVTPEKE---APAAEAEKSVSVKEEETVVVAEKVVVLTAEEVQK 94
Query: 201 K--EVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQEDTPKKSFAS 258
K E L L+ E PV VKE K+ E+ T E+ +E+T +
Sbjct: 95 KALEEFKELVREALNKREFTAPVTPVKEEKTEEKKTEEETKEEEKTEEKKEETTTE---- 150
Query: 259 IVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFV 318
++ KPA VPA E + P+E +E A
Sbjct: 151 -------------VKVEEEKPA--------VPAAEEEKSSEAAPVETKSEEKPEEKAEVT 189
Query: 319 ANLPMSA------TVEQLDRAFKKFGP 339
SA TVE ++ + P
Sbjct: 190 TEKASSAEEDGTKTVEAIEESIVSVSP 216
>At5g19960 glycine-rich RNA-binding protein - like
Length = 337
Score = 39.3 bits (90), Expect = 0.004
Identities = 24/86 (27%), Positives = 41/86 (46%), Gaps = 9/86 (10%)
Query: 314 HAIFVANLPMSATVEQLDRAFKKFGPIKRDGI-QVRSNKGSCFGFVEFESAASMQSALEV 372
++++V LP T E + R F +G + I RS +G C+GFV F + S A+E
Sbjct: 7 NSVYVGGLPYDITEEAVRRVFSIYGSVLTVKIVNDRSVRGKCYGFVTFSNRRSADDAIED 66
Query: 373 W--------SISINSCYQQGCKLNVG 390
++ +N +G ++N G
Sbjct: 67 MDGKSIGGRAVRVNDVTTRGGRMNPG 92
>At1g60000 unknown protein
Length = 258
Score = 38.9 bits (89), Expect = 0.005
Identities = 29/89 (32%), Positives = 42/89 (46%), Gaps = 8/89 (8%)
Query: 285 RVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDG 344
R V + P PN + PL E H +FV NL + T E L AF++ G +
Sbjct: 154 RALKVNFADKPKPNKE-PLYPETE-----HKLFVGNLSWTVTSESLAGAFRECGDVVGAR 207
Query: 345 IQVRSNKGSC--FGFVEFESAASMQSALE 371
+ + G +GFV + S A M++ALE
Sbjct: 208 VVFDGDTGRSRGYGFVCYSSKAEMETALE 236
>At1g49490 hypothetical protein
Length = 847
Score = 38.1 bits (87), Expect = 0.008
Identities = 37/150 (24%), Positives = 64/150 (42%), Gaps = 16/150 (10%)
Query: 159 TPEPEPVHVPEVIP-----PTQTVIPTAQTVIPPTQTVIADTETIISK-EVSLPLENGKL 212
T P P H P PTQ P+++T PT + +D I+S + P+++
Sbjct: 636 TFSPPPTHNTNQPPMGAPTPTQAPTPSSETTQVPTPSSESDQSQILSPVQAPTPVQSSTP 695
Query: 213 SVTENVIPVNHVKES--SHHVKEPEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPF 270
S +P ES + ++ + PT ++ A T +T + S + + AP
Sbjct: 696 SSEPTQVPTPSSSESYQAPNLSPVQAPTPVQ--APTTSSETSQVPTPSSESNQSPSQAP- 752
Query: 271 HLRASPAKPAVHPPRVHSVPAPEAPTPNMD 300
+P VH P +S P ++PTP+ +
Sbjct: 753 ----TPILEPVHAPTPNSKPV-QSPTPSSE 777
Score = 30.8 bits (68), Expect = 1.3
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 10/90 (11%)
Query: 154 SEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIADTE--TIISKEVSLPLENGK 211
SE+ P PV P + T T+Q P +++ + ++ T I + V P N K
Sbjct: 708 SESYQAPNLSPVQAPTPVQAPTTSSETSQVPTPSSESNQSPSQAPTPILEPVHAPTPNSK 767
Query: 212 LSVTENVIPVNHVKESSHHVKEPEQPTSIE 241
PV SS V PEQ +E
Sbjct: 768 --------PVQSPTPSSEPVSSPEQSEEVE 789
Score = 29.6 bits (65), Expect = 3.0
Identities = 33/108 (30%), Positives = 47/108 (42%), Gaps = 26/108 (24%)
Query: 144 PANDADESAPSEAIITPEPEPVHVPEV-IPPTQTVIPTAQTVIPPTQTVIADTETIISKE 202
P++++++S PS+A TP EPVH P P Q+ P+++ V P Q S+E
Sbjct: 740 PSSESNQS-PSQAP-TPILEPVHAPTPNSKPVQSPTPSSEPVSSPEQ----------SEE 787
Query: 203 VSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQED 250
V P PVN SS P TSI +N +D
Sbjct: 788 VEAP----------EPTPVN---PSSVPSSSPSTDTSIPPPENNDDDD 822
Score = 28.1 bits (61), Expect = 8.7
Identities = 21/75 (28%), Positives = 31/75 (41%), Gaps = 11/75 (14%)
Query: 234 PEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP-PRVHSVPAP 292
P P + E S + PK S D+S P +P +P+ P P H P P
Sbjct: 396 PNPPRTSEPKPSKPEPVMPKPS---------DSSKP-ETPKTPEQPSPKPQPPKHESPKP 445
Query: 293 EAPTPNMDIPLEKNN 307
E P ++P +K +
Sbjct: 446 EEPENKHELPKQKES 460
>At4g14300 ribonucleoprotein like protein
Length = 406
Score = 37.7 bits (86), Expect = 0.011
Identities = 22/77 (28%), Positives = 37/77 (47%), Gaps = 2/77 (2%)
Query: 296 TPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQV--RSNKGS 353
T N++ + + IFV LP + T E+ + F+ +GP+ I +N+
Sbjct: 87 TGNLNTSRSSGGDAYNKTKKIFVGGLPPTLTDEEFRQYFEVYGPVTDVAIMYDQATNRPR 146
Query: 354 CFGFVEFESAASMQSAL 370
FGFV F+S ++ S L
Sbjct: 147 GFGFVSFDSEDAVDSVL 163
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,245,675
Number of Sequences: 26719
Number of extensions: 418174
Number of successful extensions: 1951
Number of sequences better than 10.0: 161
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 126
Number of HSP's that attempted gapping in prelim test: 1606
Number of HSP's gapped (non-prelim): 330
length of query: 393
length of database: 11,318,596
effective HSP length: 101
effective length of query: 292
effective length of database: 8,619,977
effective search space: 2517033284
effective search space used: 2517033284
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146567.10


