

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.2 + phase: 2 /partial
(358 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
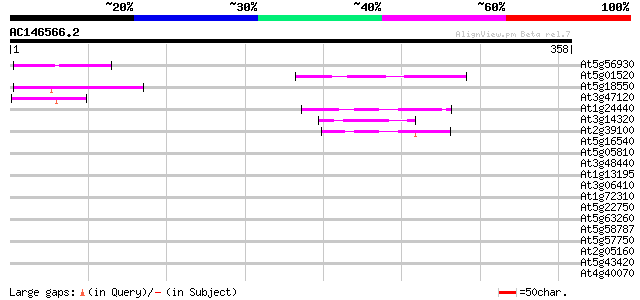
Score E
Sequences producing significant alignments: (bits) Value
At5g56930 unknown protein 53 2e-07
At5g01520 C3HC4 type Zinc RING finger like protein 47 1e-05
At5g18550 zinc finger -like protein 43 2e-04
At3g47120 putative RNA-binding protein 43 2e-04
At1g24440 unknown protein 43 2e-04
At3g14320 putative zinc finger protein 42 5e-04
At2g39100 RING zinc finger like protein 42 7e-04
At5g16540 zinc finger protein 3 (ZFN3) 41 0.001
At5g05810 putative protein 40 0.002
At3g48440 unknown protein 40 0.002
At1g13195 unknown protein 40 0.002
At3g06410 hypothetical protein 40 0.003
At1g72310 RING-H2 zinc finger protein (ATL3) 40 0.003
At5g22750 putative protein 39 0.004
At5g63260 unknown protein 39 0.006
At5g58787 unknown protein 39 0.006
At5g57750 putative protein 39 0.006
At2g05160 hypothetical protein 39 0.006
At5g43420 unknown protein 38 0.007
At4g40070 unknown protein 38 0.007
>At5g56930 unknown protein
Length = 675
Score = 53.1 bits (126), Expect = 2e-07
Identities = 24/63 (38%), Positives = 34/63 (53%), Gaps = 2/63 (3%)
Query: 3 CKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPSS 62
C +FA +C+KG+ C F HD P N F KG C G C + H ++ S TPS+
Sbjct: 385 CCYFATQSCMKGDDCPFDHDLSKYPCN--NFITKGFCYRGDSCLFSHKGTPQSASDTPSA 442
Query: 63 SIT 65
++T
Sbjct: 443 NVT 445
Score = 28.1 bits (61), Expect = 7.7
Identities = 10/26 (38%), Positives = 13/26 (49%)
Query: 24 KAPPNNICTFYQKGVCAYGSRCRYDH 49
K P C Y KG C G +C++ H
Sbjct: 349 KPKPIKYCRHYLKGRCHEGDKCKFSH 374
>At5g01520 C3HC4 type Zinc RING finger like protein
Length = 242
Score = 47.4 bits (111), Expect = 1e-05
Identities = 30/110 (27%), Positives = 47/110 (42%), Gaps = 21/110 (19%)
Query: 183 RNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPT 242
R + + + +E EC +CLE R +L C+H C++C RNWR+ +
Sbjct: 129 RTDKGKMSEIDLEREEECGICLEI---------RNKVVLPTCNHSMCINCYRNWRARS-- 177
Query: 243 LGMDVNSTLRACPICRKLSYFVVPSVIW-YATSEEKMEIIDTYKAKLKSI 291
++CP CR V +W Y S E ++ YK LK +
Sbjct: 178 ---------QSCPFCRGSLKRVNSGDLWIYTCSAEIADLPAIYKENLKRL 218
>At5g18550 zinc finger -like protein
Length = 456
Score = 43.1 bits (100), Expect = 2e-04
Identities = 29/101 (28%), Positives = 42/101 (40%), Gaps = 18/101 (17%)
Query: 3 CKFFAH-GACLKGEHCEFSHDWKA----------------PPNNICT-FYQKGVCAYGSR 44
C++F G C G C F H +A P CT F Q G+C +G
Sbjct: 297 CQYFMRTGDCKFGTSCRFHHPMEAASPEASTLSHIGLPLRPGAVPCTHFAQHGICKFGPA 356
Query: 45 CRYDHVKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSN 85
C++DH S + S +PS S P+ + LG +S+
Sbjct: 357 CKFDHSLGSSSLSYSPSPSSLTDMPVAPYPSSLGTLAPSSS 397
Score = 33.1 bits (74), Expect = 0.24
Identities = 13/25 (52%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDH 49
P C +Y + GVC YGSRCR++H
Sbjct: 43 PDEPDCIYYLRTGVCGYGSRCRFNH 67
Score = 32.7 bits (73), Expect = 0.31
Identities = 15/47 (31%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Query: 283 TYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLG 329
T+ + + +C++F G+C FGTSC + H E L H+G
Sbjct: 287 TFPQRPEQPECQYF-MRTGDCKFGTSCRFHHPMEAASPEASTLSHIG 332
Score = 29.6 bits (65), Expect = 2.7
Identities = 15/40 (37%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Query: 293 CKHFDFGEGNCPFGTSCFYKHAYRDG---RLEEVALRHLG 329
C+HF G C FG SC Y H + G + V+L ++G
Sbjct: 94 CQHF-MRTGTCKFGASCKYHHPRQGGGGDSVTPVSLNYMG 132
Score = 29.3 bits (64), Expect = 3.5
Identities = 16/65 (24%), Positives = 24/65 (36%), Gaps = 20/65 (30%)
Query: 5 FFAHGACLKGEHCEFSHDWKA-------------------PPNNICTFYQK-GVCAYGSR 44
F G C G C++ H + P C+++ + G C +GS
Sbjct: 97 FMRTGTCKFGASCKYHHPRQGGGGDSVTPVSLNYMGFPLRPGEKECSYFMRTGQCKFGST 156
Query: 45 CRYDH 49
CRY H
Sbjct: 157 CRYHH 161
Score = 29.3 bits (64), Expect = 3.5
Identities = 19/73 (26%), Positives = 26/73 (35%), Gaps = 19/73 (26%)
Query: 9 GACLKGEHCEFSHDWKAPP-----------------NNICT-FYQKGVCAYGSRCRYDHV 50
G C G C F+H P +C F + G C +G+ C+Y H
Sbjct: 55 GVCGYGSRCRFNHPRNRAPVLGGLRTEAGEFPERMGQPVCQHFMRTGTCKFGASCKYHHP 114
Query: 51 K-ASRAQSSTPSS 62
+ S TP S
Sbjct: 115 RQGGGGDSVTPVS 127
>At3g47120 putative RNA-binding protein
Length = 352
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/70 (27%), Positives = 29/70 (41%), Gaps = 22/70 (31%)
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAPPN----------------------NICTFYQKGVC 39
+C+ F G C +G+ C+FSHD K N +C +Q+G C
Sbjct: 135 VCRAFQRGECTRGDSCKFSHDEKRAANTGWGHEEDRSSKWDHDKNREGRGVCRAFQRGEC 194
Query: 40 AYGSRCRYDH 49
G C++ H
Sbjct: 195 TRGDSCKFSH 204
Score = 32.7 bits (73), Expect = 0.31
Identities = 11/23 (47%), Positives = 16/23 (68%)
Query: 2 LCKFFAHGACLKGEHCEFSHDWK 24
+C+ F G C +G+ C+FSHD K
Sbjct: 185 VCRAFQRGECTRGDSCKFSHDEK 207
>At1g24440 unknown protein
Length = 251
Score = 43.1 bits (100), Expect = 2e-04
Identities = 24/96 (25%), Positives = 40/96 (41%), Gaps = 22/96 (22%)
Query: 187 KHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMD 246
+++ ++ +E EC +CLE +L C H C+ C RNW
Sbjct: 142 RYMNSIDLEREDECGICLEPCTKM---------VLPNCCHAMCIKCYRNW---------- 182
Query: 247 VNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIID 282
N+ +CP CR V +W T +E +++D
Sbjct: 183 -NTKSESCPFCRGSIKRVNSEDLWVLTCDE--DVVD 215
>At3g14320 putative zinc finger protein
Length = 204
Score = 42.0 bits (97), Expect = 5e-04
Identities = 25/62 (40%), Positives = 29/62 (46%), Gaps = 16/62 (25%)
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC VCL + A K +L CDH F V CI +W SN T CPIC
Sbjct: 86 LECVVCLSEL-----ADGDKARVLPSCDHWFHVECIDSWLQSNST-----------CPIC 129
Query: 258 RK 259
RK
Sbjct: 130 RK 131
>At2g39100 RING zinc finger like protein
Length = 296
Score = 41.6 bits (96), Expect = 7e-04
Identities = 24/84 (28%), Positives = 42/84 (49%), Gaps = 18/84 (21%)
Query: 200 CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC-- 257
C +CLE + + +AA +++ C H +C++CIR W +S R CP+C
Sbjct: 39 CPICLENLTERRSAA-----VITVCKHGYCLACIRKW-----------SSFKRNCPLCNT 82
Query: 258 RKLSYFVVPSVIWYATSEEKMEII 281
R S+F+V +E++ I+
Sbjct: 83 RFDSWFIVSDFASRKYHKEQLPIL 106
>At5g16540 zinc finger protein 3 (ZFN3)
Length = 375
Score = 40.8 bits (94), Expect = 0.001
Identities = 30/107 (28%), Positives = 49/107 (45%), Gaps = 27/107 (25%)
Query: 3 CKFFAH-GACLKGEHCEFSH--DWKAPPNN---------------ICTFYQK-GVCAYGS 43
C+F+ G C G C+F H D + PP + +C FY + G+C +G
Sbjct: 249 CQFYMKTGDCKFGTVCKFHHPRDRQTPPPDCVLSSVGLPLRPGEPLCVFYSRYGICKFGP 308
Query: 44 RCRYDH-----VKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSN 85
C++DH + S +PSSS+ + + +E L N V+S+
Sbjct: 309 SCKFDHPMRVFTYNNNTASPSPSSSLHQETAITTE---LRNLLVSSS 352
>At5g05810 putative protein
Length = 407
Score = 40.0 bits (92), Expect = 0.002
Identities = 24/61 (39%), Positives = 32/61 (52%), Gaps = 16/61 (26%)
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC+VCL R +PT R LL +C H F V C+ W +D +ST CP+C
Sbjct: 144 LECAVCLARF--EPTEVLR---LLPKCKHAFHVECVDTW--------LDAHST---CPLC 187
Query: 258 R 258
R
Sbjct: 188 R 188
>At3g48440 unknown protein
Length = 448
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 3/48 (6%)
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDHVKASRAQSSTPSSSITEHQPLVS 72
P NICT+Y + G+C +G CR+DH + + ST SS P VS
Sbjct: 393 PDQNICTYYSRYGICKFGPACRFDH--SVQPPYSTESSQAIVEPPQVS 438
Score = 35.0 bits (79), Expect = 0.063
Identities = 21/66 (31%), Positives = 29/66 (43%), Gaps = 19/66 (28%)
Query: 3 CKF-FAHGACLKGEHCEFSHDW------KAPPNNI-----------CTFYQK-GVCAYGS 43
CK+ F G C GE C F+H AP N C +Y + G C YG+
Sbjct: 164 CKYYFRTGGCKYGETCRFNHTIPKSGLASAPELNFLGLPLRPGEVECPYYMRNGSCKYGA 223
Query: 44 RCRYDH 49
C+++H
Sbjct: 224 ECKFNH 229
Score = 32.0 bits (71), Expect = 0.53
Identities = 21/79 (26%), Positives = 31/79 (38%), Gaps = 21/79 (26%)
Query: 3 CKFFAH-GACLKGEHCEFSHDWK-----APPNNI--------------CTFY-QKGVCAY 41
C F+ G+C G C+F+H A N + C +Y + G C Y
Sbjct: 116 CSFYMRTGSCKFGSSCKFNHPLARKFQIARDNKVREKEDDGGKLGLIDCKYYFRTGGCKY 175
Query: 42 GSRCRYDHVKASRAQSSTP 60
G CR++H +S P
Sbjct: 176 GETCRFNHTIPKSGLASAP 194
Score = 30.0 bits (66), Expect = 2.0
Identities = 16/44 (36%), Positives = 22/44 (49%), Gaps = 2/44 (4%)
Query: 287 KLKSIDCKHFDFGEGNCPFGTSCFYKHAY-RDGRLEEVALRHLG 329
KL IDCK++ F G C +G +C + H + G L LG
Sbjct: 158 KLGLIDCKYY-FRTGGCKYGETCRFNHTIPKSGLASAPELNFLG 200
>At1g13195 unknown protein
Length = 260
Score = 40.0 bits (92), Expect = 0.002
Identities = 24/96 (25%), Positives = 37/96 (38%), Gaps = 22/96 (22%)
Query: 196 QEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACP 255
+E EC +CLE +L C H C+ C RNW N ++CP
Sbjct: 158 REEECGICLETCTKM---------VLPNCCHSMCIKCYRNW-----------NLKSQSCP 197
Query: 256 ICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKLKSI 291
CR V +W + +++DT A + +
Sbjct: 198 FCRGSMKRVNSEDLWVLAGDN--DVVDTRTASREDL 231
>At3g06410 hypothetical protein
Length = 437
Score = 39.7 bits (91), Expect = 0.003
Identities = 24/83 (28%), Positives = 35/83 (41%), Gaps = 19/83 (22%)
Query: 3 CKFFAH-GACLKGEHCEFSHDWKAPPNNI-----------------CT-FYQKGVCAYGS 43
C++F G C G C + H A P CT F Q G+C +G
Sbjct: 288 CQYFMRTGDCKFGSSCRYHHPVDAVPPKTGIVLSSIGLPLRPGVAQCTHFAQHGICKFGP 347
Query: 44 RCRYDHVKASRAQSSTPSSSITE 66
C++DH +S S +SS+T+
Sbjct: 348 ACKFDHSMSSSLSYSPSASSLTD 370
Score = 33.1 bits (74), Expect = 0.24
Identities = 13/25 (52%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDH 49
P C +Y + GVC YGSRCR++H
Sbjct: 30 PDEPDCIYYLRTGVCGYGSRCRFNH 54
Score = 30.0 bits (66), Expect = 2.0
Identities = 16/40 (40%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Query: 293 CKHFDFGEGNCPFGTSCFYKHAYR---DGRLEEVALRHLG 329
C+HF G C FG SC Y H + G + V+L +LG
Sbjct: 81 CQHF-MRTGTCKFGASCKYHHPRQGGGGGSVAPVSLSYLG 119
>At1g72310 RING-H2 zinc finger protein (ATL3)
Length = 324
Score = 39.7 bits (91), Expect = 0.003
Identities = 22/61 (36%), Positives = 29/61 (47%), Gaps = 16/61 (26%)
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+ECS+CL ++ A LL +C+H F V CI W S+ T CPIC
Sbjct: 125 LECSICLSELVKGDKAR-----LLPKCNHSFHVECIDMWFQSHST-----------CPIC 168
Query: 258 R 258
R
Sbjct: 169 R 169
>At5g22750 putative protein
Length = 1029
Score = 38.9 bits (89), Expect = 0.004
Identities = 22/74 (29%), Positives = 36/74 (47%), Gaps = 20/74 (27%)
Query: 186 QKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSC-IRNWRSSNPTLG 244
Q+ +E L++ ++ EC +CLE + + +L+ C H C C + +WR
Sbjct: 780 QEVVEELRKGEQGECPICLEAL---------EDAVLTPCAHRLCRECLLASWR------- 823
Query: 245 MDVNSTLRACPICR 258
NST CP+CR
Sbjct: 824 ---NSTSGLCPVCR 834
>At5g63260 unknown protein
Length = 435
Score = 38.5 bits (88), Expect = 0.006
Identities = 22/78 (28%), Positives = 33/78 (42%), Gaps = 20/78 (25%)
Query: 3 CKFFAH-GACLKGEHCEFSH------------------DWKAPPNNICTFY-QKGVCAYG 42
C F+ G+C G C+F+H D + P C +Y + G C YG
Sbjct: 107 CSFYMRTGSCKYGSSCKFNHPVRRKLQIGRERVRERDEDVENPKLMECKYYFRTGGCKYG 166
Query: 43 SRCRYDHVKASRAQSSTP 60
CR+ H+K + +S P
Sbjct: 167 ESCRFSHMKEHNSPASVP 184
Score = 36.2 bits (82), Expect = 0.028
Identities = 37/135 (27%), Positives = 49/135 (35%), Gaps = 29/135 (21%)
Query: 3 CKF-FAHGACLKGEHCEFSHDWK-----------------APPNNICTFYQK-GVCAYGS 43
CK+ F G C GE C FSH + P C FY + G C +GS
Sbjct: 154 CKYYFRTGGCKYGESCRFSHMKEHNSPASVPELNFLGLPIRPGEKECPFYMRNGSCKFGS 213
Query: 44 RCRYDHVKA-------SRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVATAAEF-SL 95
C+++H S S + P + S +TR NG TA S+
Sbjct: 214 DCKFNHPDPTAIGGVDSPLYRGNNGGSFSPKAPSQASSTSWSSTR-HMNGTGTAPFIPSM 272
Query: 96 F-STPFVLPSEQAWN 109
F + V P WN
Sbjct: 273 FPHSRGVTPQASDWN 287
Score = 35.0 bits (79), Expect = 0.063
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDHVKASRAQSSTPSSSIT 65
P ++CT Y + G+C +G CR+DH S + +PSSS T
Sbjct: 381 PDQSMCTHYSRYGICKFGPACRFDH---SIPPTFSPSSSQT 418
Score = 28.9 bits (63), Expect = 4.5
Identities = 15/43 (34%), Positives = 24/43 (54%), Gaps = 3/43 (6%)
Query: 274 SEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYR 316
SE++M ++ Y + S DC F G+C +G+SC + H R
Sbjct: 90 SEKRMMMV--YPVRPDSEDCS-FYMRTGSCKYGSSCKFNHPVR 129
>At5g58787 unknown protein
Length = 242
Score = 38.5 bits (88), Expect = 0.006
Identities = 23/87 (26%), Positives = 40/87 (45%), Gaps = 21/87 (24%)
Query: 173 EEREEHMMSCRNK-QKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVS 231
+++E M R K + + ++ +E EC +C+E + SK +L C H C+
Sbjct: 116 KQKEVCKMRYRKKDESEMSEIEIEREEECGICME-MNSKV--------VLPNCTHSLCIK 166
Query: 232 CIRNWRSSNPTLGMDVNSTLRACPICR 258
C R+WR + ++CP CR
Sbjct: 167 CYRDWRGRS-----------QSCPFCR 182
>At5g57750 putative protein
Length = 210
Score = 38.5 bits (88), Expect = 0.006
Identities = 21/60 (35%), Positives = 27/60 (45%), Gaps = 16/60 (26%)
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
+C+VCL AE + LL +C H F V CI W +N T CP+CR
Sbjct: 121 DCAVCLREF-----TAEDELRLLPKCSHAFHVECIDTWLLTNST-----------CPLCR 164
>At2g05160 hypothetical protein
Length = 536
Score = 38.5 bits (88), Expect = 0.006
Identities = 17/51 (33%), Positives = 30/51 (58%), Gaps = 5/51 (9%)
Query: 27 PNNICTFYQKGVCAYGSRCRYDH-----VKASRAQSSTPSSSITEHQPLVS 72
P IC ++ KG C +G+ CRY H + S AQ P++++++ + +VS
Sbjct: 158 PVKICHYFNKGFCKHGNNCRYFHGQIIPERESFAQMFNPNNNLSDEEHVVS 208
Score = 33.1 bits (74), Expect = 0.24
Identities = 41/175 (23%), Positives = 65/175 (36%), Gaps = 27/175 (15%)
Query: 11 CLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPSSSITEHQPL 70
C+K E SH + +PP+ DH+ + S T SS ++
Sbjct: 57 CVKYELARNSHHYHSPPS-------------------DHIPTPKFGSFTGSSPLSVSVSP 97
Query: 71 VSESAVLGNTRVTSNGVATAAEFSLFSTPFVLP--SEQAWNQESAQLDFLREDDVVQSVI 128
++ N+ + + +F F P P S ++QE L LR S+
Sbjct: 98 PMKTGFWENS-TEMDTLQNNLQFLNFEDPLTSPEFSNGFFSQERQCLP-LRTSRRSPSLP 155
Query: 129 TSPSELPICSFAAAGNCPRGEQCPHVHGDLCP--SCGRQCLHPFRPEEREEHMMS 181
P + IC + G C G C + HG + P Q +P EEH++S
Sbjct: 156 EFP--VKICHYFNKGFCKHGNNCRYFHGQIIPERESFAQMFNPNNNLSDEEHVVS 208
>At5g43420 unknown protein
Length = 375
Score = 38.1 bits (87), Expect = 0.007
Identities = 24/69 (34%), Positives = 29/69 (41%), Gaps = 18/69 (26%)
Query: 190 EALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNS 249
E KRSQE CSVCL E K ++ C H F + CI W +N
Sbjct: 130 EEEKRSQE--CSVCLSEFQD-----EEKLRIIPNCSHLFHIDCIDVWLQNNAN------- 175
Query: 250 TLRACPICR 258
CP+CR
Sbjct: 176 ----CPLCR 180
>At4g40070 unknown protein
Length = 322
Score = 38.1 bits (87), Expect = 0.007
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 16/64 (25%)
Query: 195 SQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRAC 254
S+++EC++CL + T LL C+H F + CI W S+ T C
Sbjct: 118 SKDLECAICLNELEDHETVR-----LLPICNHLFHIDCIDTWLYSHAT-----------C 161
Query: 255 PICR 258
P+CR
Sbjct: 162 PVCR 165
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,833,878
Number of Sequences: 26719
Number of extensions: 376313
Number of successful extensions: 1499
Number of sequences better than 10.0: 235
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 180
Number of HSP's that attempted gapping in prelim test: 1192
Number of HSP's gapped (non-prelim): 432
length of query: 358
length of database: 11,318,596
effective HSP length: 100
effective length of query: 258
effective length of database: 8,646,696
effective search space: 2230847568
effective search space used: 2230847568
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146566.2


