

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146565.8 - phase: 0 /pseudo
(1145 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
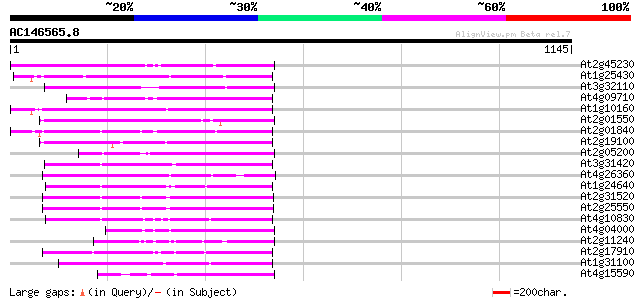
Score E
Sequences producing significant alignments: (bits) Value
At2g45230 putative non-LTR retroelement reverse transcriptase 215 1e-55
At1g25430 hypothetical protein 204 2e-52
At3g32110 non-LTR reverse transcriptase, putative 197 4e-50
At4g09710 RNA-directed DNA polymerase -like protein 192 9e-49
At1g10160 putative reverse transcriptase 189 6e-48
At2g01550 putative non-LTR retroelement reverse transcriptase 189 8e-48
At2g01840 putative non-LTR retroelement reverse transcriptase 183 6e-46
At2g19100 putative non-LTR retroelement reverse transcriptase 182 7e-46
At2g05200 putative non-LTR retroelement reverse transcriptase 181 2e-45
At3g31420 hypothetical protein 179 6e-45
At4g26360 putative protein 176 7e-44
At1g24640 hypothetical protein 174 2e-43
At2g31520 putative non-LTR retroelement reverse transcriptase 174 3e-43
At2g25550 putative non-LTR retroelement reverse transcriptase 174 3e-43
At4g10830 putative protein 171 2e-42
At4g04000 putative reverse transcriptase 166 5e-41
At2g11240 pseudogene 163 5e-40
At2g17910 putative non-LTR retroelement reverse transcriptase 156 7e-38
At1g31100 hypothetical protein 149 1e-35
At4g15590 reverse transcriptase like protein 144 4e-34
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 215 bits (548), Expect = 1e-55
Identities = 156/542 (28%), Positives = 244/542 (44%), Gaps = 19/542 (3%)
Query: 3 MIVLYWNVRGFGNLDTKIALQNFYLSNKPTFIFLAEPMISFSYVPSWYWHSIGVKKCCLN 62
M +L WN +G GN T L+ P IFL E +Y+ + H +G
Sbjct: 1 MRILSWNCQGVGNTPTVRHLREIRGLYFPEVIFLCETKKRRNYLENVVGH-LGFFDLHTV 59
Query: 63 NRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWAD 122
+ L +W D V + V+ + + I + WQ +L IY R LW
Sbjct: 60 EPIGKSGGLALMWKDSVQIKVLQSDKRLIDALLIWQDKEFYLTCIYGEPVQAERGELWER 119
Query: 123 LTHLQGCFLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLL 182
LT L GPW+ GDFN ++ EK G +SC +F N+ L + SG
Sbjct: 120 LTRLGLSRSGPWMLTGDFNELVDPSEKIGGPARKESSCLEFRQMLNSCGLWEVNHSGYQF 179
Query: 183 TWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAI 242
+W R V RLDR++ N+ W+ + L + SDH PL+ + + +
Sbjct: 180 SWYGNR-NDELVQCRLDRTVANQAWMELFPQAKATYLQKICSDHSPLINNLVGDNWRKWA 238
Query: 243 SFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQV 302
FK+ K W + L+ W++ + ++ K+ + +W R V
Sbjct: 239 GFKYDKRWVQREGFKDLLCNFWSQQSTKTNALMME-KIASCRREISKWKR--VSKPSSAV 295
Query: 303 RLATDEVSRIQQIIDSVGFSDDLYRQDL-EAQLLLTNALNVQDQFWKEKSRHQSFIHGDR 361
R + +Q +D+ R++L + L+ N ++QFW+EKSR +GDR
Sbjct: 296 R-----IQELQFKLDAATKQIPFDRRELARLKKELSQEYNNEEQFWQEKSRIMWMRNGDR 350
Query: 362 NTAYFHRVARIKASSKNISLLYD--GATVLSDAGDIAAHILNYFQGIF--DVVNNCIPND 417
NT YFH + + + I L D G SD D+ YF+ +F + V +
Sbjct: 351 NTKYFHAATKNRRAQNRIQKLIDEEGREWTSDE-DLGRVAEAYFKKLFASEDVGYTVEE- 408
Query: 418 LVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
++ LV++ NN L+ ++E++ A F +N PGPDG G YQ FW+ +G
Sbjct: 409 --LENLTPLVSDQMNNNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQ 466
Query: 478 VVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLA 537
+ VQ FF GS + +N + LIPKI A+ M DFRPI+L N +K++ K++A+RL
Sbjct: 467 ITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLK 526
Query: 538 SI 539
I
Sbjct: 527 KI 528
>At1g25430 hypothetical protein
Length = 1213
Score = 204 bits (519), Expect = 2e-52
Identities = 163/557 (29%), Positives = 264/557 (47%), Gaps = 48/557 (8%)
Query: 8 WNVRGFGNLDTKIALQNFYLSNKPTFIFLAEPMISF--------SYVPSWYWHSIGVKKC 59
WN+RGF N+ + + + +NKP F + E + + +P W + V+
Sbjct: 8 WNIRGFNNVSHRSGFKKWVKANKPIFGGVIETHVKQPKDRKFINALLPGWSF----VENY 63
Query: 60 CLNNRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSS--VFLAAIYANTSYLFRR 117
++ G +W +W+ V VVV+ S Q I EV S + ++ +YA R+
Sbjct: 64 AFSDLG----KIWVMWDPSVQVVVVAKSLQMITCEVLLPGSPSWIIVSVVYAANEVASRK 119
Query: 118 LLWADLTHLQ-GCFLG--PWLFIGDFNAVMGAHEKRGRWPPTAASCS----DFGSWSNAN 170
LW ++ ++ +G PWL +GDFN V+ E P + + DF A
Sbjct: 120 ELWIEIVNMVVSGIIGDRPWLVLGDFNQVLNPQEHSN---PVSLNVDINMRDFRDCLLAA 176
Query: 171 LLTHLPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLL 230
L+ L G TW N + + VA ++DR + N+ W A + + SDH
Sbjct: 177 ELSDLRYKGNTFTWWN-KSHTTPVAKKIDRILVNDSWNALFPSSLGIFGSLDFSDHVSCG 235
Query: 231 LSADVASVKNAISFKFYKAWTSHVDCRRLVLENW-AKNVRGVGMVRLQGKLRILKTVFKQ 289
+ + S+K FKF+ ++D LV +NW NV G M R+ KL+ LK K
Sbjct: 236 VVLEETSIKAKRPFKFFNYLLKNLDFLNLVRDNWFTLNVVGSSMFRVSKKLKALKKPIKD 295
Query: 290 WNRSVFGDVDRQVRLATDEVSRIQQ--IIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFW 347
++R + +++++ + A D + Q + D + +LEA+ ++ F+
Sbjct: 296 FSRLNYSELEKRTKEAHDFLIGCQDRTLADPTPINASF---ELEAERKWHILTAAEESFF 352
Query: 348 KEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLYDGATVLSDAGD-IAAHILNYFQGI 406
++KSR F GD NT YFHR+A + SS +IS LYDG L D+ + I +YF +
Sbjct: 353 RQKSRISWFAEGDGNTKYFHRMADARNSSNSISALYDGNGKLVDSQEGILDLCASYFGSL 412
Query: 407 F-DVVNNCI--PND----LVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGP 459
D V+ + ND L R A V E E+ ++I++A+F L + + GP
Sbjct: 413 LGDEVDPYLMEQNDMNLLLSYRCSPAQVCELESTFS-----NEDIRAALFSLPRNKSCGP 467
Query: 460 DGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIA 519
DGF F+ W IVG++V ++++FF G N+ +VLIPKI DFRPI+
Sbjct: 468 DGFTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATTIVLIPKIVNPTCTSDFRPIS 527
Query: 520 LANFQFKIVTKILADRL 536
N +K++ ++L DRL
Sbjct: 528 CLNTLYKVIARLLTDRL 544
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 197 bits (500), Expect = 4e-50
Identities = 141/475 (29%), Positives = 220/475 (45%), Gaps = 45/475 (9%)
Query: 71 LWALWNDEV-LVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHLQGC 129
LW LW + V V+ ++DQ I + +V L +YA + R LW L +
Sbjct: 633 LWLLWRTGIGEVSVVDSTDQFIHAKDVNGKDNVNLVVVYAAPTASRRSGLWDRLGDVIRS 692
Query: 130 FLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRL 189
GP + GDFN ++ E+ G ++ FG W N + L L G TW GR
Sbjct: 693 MDGPVVIGGDFNTIVRLDERSGGNGRLSSDSLAFGEWINDHSLIDLGFKGNKFTWKRGRE 752
Query: 190 GSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLL--LSADVASVKNAISFKFY 247
+VA RLDR +C ++ S L SDH PL L+ +V+ + F+F
Sbjct: 753 ERFFVAKRLDRVLCCAHARLKWQEASVLHLPFLASDHAPLYVQLTPEVSGNRGRRPFRFE 812
Query: 248 KAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATD 307
AW SH + L+L +W K++ +T
Sbjct: 813 AAWLSHPGFKELLLTSWNKDI------------------------------------STP 836
Query: 308 EVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFH 367
E ++Q+++D + SDDL +++ E L ++ W +KSR + F+HGDRNT +FH
Sbjct: 837 EALKVQELLD-LHQSDDLLKKEEELLKDFDVVLEQEEVVWMQKSREKWFVHGDRNTKFFH 895
Query: 368 RVARIKASSKNISLLYDG-ATVLSDAGDIAAHILNYFQGIFDVVN-NCIPNDLVARSIHA 425
I+ I +L D LS+A ++ H ++Y++ ++ + + + + L A
Sbjct: 896 TSTIIRRRRNQIEMLQDNDGRWLSNAQELETHAIDYYKRLYSLDDLDAVVEQLPQEGFTA 955
Query: 426 LVTEDENNMLIRL-PLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQD 484
L D +++ PL E++ A+ + APGPDGF FYQ W++VG V V D
Sbjct: 956 LSEADFSSLTKPFSPL--EVEGAIRSMGKYKAPGPDGFQPVFYQQGWEVVGESVTKFVMD 1013
Query: 485 FFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
FF GSF N L+VLI K+ + + FRPI+L N FK +TK++ RL +
Sbjct: 1014 FFSSGSFPQETNDVLVVLIAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGV 1068
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 192 bits (488), Expect = 9e-49
Identities = 131/427 (30%), Positives = 208/427 (48%), Gaps = 23/427 (5%)
Query: 116 RRLLWADLTHLQGCFLGPWLFIGDFNAVMGAHEKRG---RWPPTAASCSDFGSWSNANLL 172
R + W ++ L WL GDFN ++ EK+G RW + F S+ + N L
Sbjct: 57 RSVFWDKISSLGAQRSSAWLLTGDFNDILDNSEKQGGPLRWEGFFLA---FRSFVSQNGL 113
Query: 173 THLPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLS 232
+ +G L+W R S ++ RLDR++ N W + ++ C L SDH PL+
Sbjct: 114 WDINHTGNSLSWRGTRY-SHFIKSRLDRALGNCSWSELFPMSKCEYLRFEGSDHRPLVTY 172
Query: 233 ADVASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNR 292
+K + F+F + + R LV E W + + ++ R +++ K W +
Sbjct: 173 FGAPPLKRSKPFRFDRRLREKEEIRALVKEVWELARQDSVLYKIS---RCRQSIIK-WTK 228
Query: 293 SVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLL--LTNALNVQDQFWKEK 350
Q + + + QQ ++S S D+ L + L A ++ FWK+
Sbjct: 229 E-------QNSNSAKAIKKAQQALESA-LSADIPDPSLIGSITQELEAAYRQEELFWKQW 280
Query: 351 SRHQSFIHGDRNTAYFHRVARIKASSKNISLLYDGA-TVLSDAGDIAAHILNYFQGIFDV 409
SR Q GDRN YFH R + N+S++ DG+ + IA+ I +YFQ IF
Sbjct: 281 SRVQWLNSGDRNKGYFHATTRTRRMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIFTT 340
Query: 410 VNNCIPNDLVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQS 469
NN +V ++ +++ N LI++ EIK A+F ++ D APGPDGF F+ +
Sbjct: 341 SNNS-DLQVVQEALSPIISSHCNEELIKISSLLEIKEALFSISADKAPGPDGFSASFFHA 399
Query: 470 FWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVT 529
+WDI+ +DV ++ FF+ + +N + LIPKI + + D+RPIAL N Q+KIV
Sbjct: 400 YWDIIEADVSRDIRSFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVA 459
Query: 530 KILADRL 536
KIL RL
Sbjct: 460 KILTRRL 466
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 189 bits (481), Expect = 6e-48
Identities = 143/555 (25%), Positives = 250/555 (44%), Gaps = 32/555 (5%)
Query: 3 MIVLYWNVRGFGNLDTKIALQNFYLSNKPTFIFLAEPMIS--------FSYVPSWYWHSI 54
M V WN+RG + + + ++++ SN E ++ S +P W S
Sbjct: 1 MKVFCWNIRGLNSRNRQRVVRSWIASNNLLVGCFLETHVAQENANSVLASTLPGWRMDS- 59
Query: 55 GVKKCCLNNRGPRLPNLWALWNDEVLVVVIFNSDQCI--ALEVTWQHSSVFLAAIYANTS 112
CC L +W +W+ + V+V +DQ + ++++ S +A +Y S
Sbjct: 60 --NYCC-----SELGRIWIVWDPSISVLVFKRTDQIMFCSIKIPSLLQSFAVAFVYGRNS 112
Query: 113 YLFRRLLWAD---LTHLQGCFLGPWLFIGDFNAVMGA--HEKRGRWPPTAASCSDFGSWS 167
L RR LW D L+ + PWL +GDFN + A H + D
Sbjct: 113 ELDRRSLWEDILVLSRTSPLSVTPWLLLGDFNQIAAASEHYSINQSLLNLRGMEDLQCCL 172
Query: 168 NANLLTHLPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHH 227
+ L+ LPS G TWSN + + + +LDR++ N EW A + SDH
Sbjct: 173 RDSQLSDLPSRGVFFTWSNHQQDNP-ILRKLDRALANGEWFAVFPSALAVFDPPGDSDHA 231
Query: 228 PLLLSADVASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVR-GVGMVRLQGKLRILKTV 286
P ++ D + SFK++ +SH + W +N G M L+ L++ K
Sbjct: 232 PCIILIDNQPPPSKKSFKYFSFLSSHPSYLAALSTAWEENTLVGSHMFSLRQHLKVAKLC 291
Query: 287 FKQWNRSVFGDVDRQVRLATDEVSRIQ-QIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQ 345
+ NR F ++ ++ + + IQ +++ S SD L+R++ A+ +
Sbjct: 292 CRTLNRLRFSNIQQRTAQSLTRLEDIQVELLTSP--SDTLFRREHVARKQWIFFAAALES 349
Query: 346 FWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLY-DGATVLSDAGDIAAHILNYFQ 404
F+++KSR + GD NT +FHR ++ I L D + + I ++ Y+
Sbjct: 350 FFRQKSRIRWLHEGDANTRFFHRAVIAHQATNLIKFLRGDDGFRVENVDQIKGMLIAYYS 409
Query: 405 GIFDVVNNCIPNDLVARSIHALVTEDEN---NMLIRLPLRDEIKSAVFDLNGDGAPGPDG 461
+ + + + V + L ++ + L +P +EI +F + + APGPDG
Sbjct: 410 HLLGIPSENVTPFSVEKIKGLLPFRCDSFLASQLTTIPSEEEITQVLFSMPRNKAPGPDG 469
Query: 462 FGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALA 521
F F+ W IV S VV ++++FF+ G+ N+ + LIPK+ GA + FRP+A
Sbjct: 470 FPVEFFIEAWAIVKSSVVAAIREFFISGNLPRGFNATAITLIPKVTGADRLTQFRPVACC 529
Query: 522 NFQFKIVTKILADRL 536
+K++T+I++ RL
Sbjct: 530 TTIYKVITRIISRRL 544
>At2g01550 putative non-LTR retroelement reverse transcriptase
Length = 1449
Score = 189 bits (480), Expect = 8e-48
Identities = 144/501 (28%), Positives = 232/501 (45%), Gaps = 34/501 (6%)
Query: 62 NNRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTW--QHSSVFLAAIYANTSYLFRRLL 119
N RG LW +W + V + SDQ I V Q F + +YA+ R++L
Sbjct: 479 NRRG----RLWVVWRENVRFTPFYKSDQLITCSVKLESQEEEFFYSFVYASNFAEERKIL 534
Query: 120 WADLT-HLQGCFLG--PWLFIGDFNAV--MGAHEKRGRWPPTAASCSDFGSWSNANLLTH 174
W DL H+ + PW+ GDFN + M H + P + DF S N +
Sbjct: 535 WNDLRDHMDSPIIRDKPWIIFGDFNEILDMDEHSRMEDHPAVTSGMRDFQSLVNYCSFSD 594
Query: 175 LPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSAD 234
L S GPL TW N R + +LDR + NE W Y + SDH ++ +
Sbjct: 595 LASHGPLFTWCNKRDNDP-IWKKLDRVMVNEAWKMVYPQSYNVFEAGGCSDHLRCRINLN 653
Query: 235 V---ASVKNAISFKFYKAWTSHVDCRRLVLENWAK----NVRGVGMVRLQGKLRILKTVF 287
+ A V+ FKF A + + LV W + ++ + R KL+ LK
Sbjct: 654 MNSGAQVRGNKPFKFVNAVADMEEFKPLVENFWRETEPIHMSTSSLFRFTKKLKALKPKL 713
Query: 288 KQWNRSVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFW 347
+ + G++ ++ R A + + QQ +S S + EA + ++++++
Sbjct: 714 RGLAKEKMGNLVKRTREAYLSLCQAQQS-NSQNPSQRAMEIESEAYVRWDRIASIEEKYL 772
Query: 348 KEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLY--DGATVLSDAGDIAAHILNYFQG 405
K+ S+ GD+N FHR A +A+ +I + DG+T + DI +FQ
Sbjct: 773 KQVSKLHWLKVGDKNNKTFHRAATARAAQNSIREIQKEDGSTATTK-DDIKNETERFFQE 831
Query: 406 IFDVVNNCIPNDLVARSIHALVT-------EDENNMLIRLPLRDEIKSAVFDLNGDGAPG 458
+ IPND ++ L + E +ML EI+ A+F + D +PG
Sbjct: 832 FLQL----IPNDYEGITVEKLTSLLPYHCSPAEKDMLTASVSAKEIRGALFSMPNDKSPG 887
Query: 459 PDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPI 518
PDG+ FY+ WDI+G++ VL+V+ FF +G +N+ +L LIPK A+ M D+RPI
Sbjct: 888 PDGYTSEFYKRAWDIIGAEFVLAVKSFFEKGFLPKGVNTTILALIPKKLEAKEMKDYRPI 947
Query: 519 ALANFQFKIVTKILADRLASI 539
+ N +K+++KI+A+RL +
Sbjct: 948 SCCNVIYKVISKIIANRLKHV 968
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 183 bits (464), Expect = 6e-46
Identities = 143/543 (26%), Positives = 234/543 (42%), Gaps = 23/543 (4%)
Query: 1 SSMIVLYWNVRGFGNLDTKIALQNFYLSNKPTFIFLAEPMISFSYVPSWYWHSIGVKK-- 58
S M V +WN +G G T L+ +FL E +Y +GVK
Sbjct: 361 SPMRVGFWNCQGLGQPLTVRRLEEVQRVYFLDMLFLIETKQQDNYT-----RDLGVKMGF 415
Query: 59 ---CCLNNRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLF 115
C ++ RG L W + + VI + + + L V +++ + +L+ IY +
Sbjct: 416 EDMCIISPRGLS-GGLVVYWKKHLSIQVISHDVRLVDLYVEYKNFNFYLSCIYGHPIPSE 474
Query: 116 RRLLWADLTHLQGCFLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHL 175
R LW L + GPW+ GDFN ++ +EK+G + S +F + N + L
Sbjct: 475 RHHLWEKLQRVSAHRSGPWMMCGDFNEILNLNEKKGGRRRSIGSLQNFTNMINCCNMKDL 534
Query: 176 PSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADV 235
S G +W G+ + + LDR N +W A + L SDH P+++
Sbjct: 535 KSKGNPYSWV-GKRQNETIESCLDRVFINSDWQASFPAFETEFLPIAGSDHAPVIIDIAE 593
Query: 236 ASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVF 295
F++ + D V W + R KL + +W R
Sbjct: 594 EVCTKRGQFRYDRRHFQFEDFVDSVQRGWNRG-RSDSHGGYYEKLHCCRQELAKWKR--- 649
Query: 296 GDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQD-LEAQLLLTNALNVQDQFWKEKSRHQ 354
R +++ ++ +D+ L Q L + L A ++ +W KSR++
Sbjct: 650 ----RTKTNTAEKIETLKYRVDAAERDHTLPHQTILRLRQDLNQAYRDEELYWHLKSRNR 705
Query: 355 SFIHGDRNTAYFHRVARIKASSKNISLLYDGATVLSDAGDIAAHIL-NYFQGIFDVVNNC 413
+ GDRNT +F+ +++ S I + D + + D + NYF +F
Sbjct: 706 WMLLGDRNTMFFYASTKLRKSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQTS 765
Query: 414 IPNDLVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDI 473
++++ I VTE N+ L++ E++ AVF + D APG DGF FY WD+
Sbjct: 766 DWEEIIS-GIAPKVTEQMNHELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWDL 824
Query: 474 VGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILA 533
+G+DV L V+ FF + IN + LIPKI + M D+RPI+L +KI++KIL
Sbjct: 825 IGNDVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILI 884
Query: 534 DRL 536
RL
Sbjct: 885 KRL 887
>At2g19100 putative non-LTR retroelement reverse transcriptase
Length = 1447
Score = 182 bits (463), Expect = 7e-46
Identities = 135/498 (27%), Positives = 238/498 (47%), Gaps = 34/498 (6%)
Query: 62 NNRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTWQH--SSVFLAAIYANTSYLFRRLL 119
N+RG +W +W V + + S+Q I V ++ F + +YA+ R++L
Sbjct: 472 NSRG----RIWVVWRRNVRLSPFYKSEQLITCSVKLENRDDEFFCSFVYASNFRDDRKVL 527
Query: 120 WADLT-HLQGCFLG--PWLFIGDFNAVMGA--HEKRGRWPPTAASCSDFGSWSNANLLTH 174
W +L H + PW+ GDFN + H K P + DF S N LT
Sbjct: 528 WNELQDHYDSPIIKKKPWIIFGDFNETLELEEHSKVEDNPVVSMGMRDFRSMVNYCSLTD 587
Query: 175 LPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWL-------AFYRVTSCCTLIRHQSDHH 227
+ GPL TWSN R +A +LDR + N+ W + + C +R + +
Sbjct: 588 MAHHGPLYTWSNKREHDL-IAKKLDRVMVNDVWTQSFPQSYSVFEAGGCLDHLRGRIN-- 644
Query: 228 PLLLSADVASVKNAISFKFYKAWTSHVDCRRLVLENWAKN----VRGVGMVRLQGKLRIL 283
L + V+ FKF T D + V W + + + R KL+ L
Sbjct: 645 --LNDGPGSIVRGKRPFKFVNVLTEMEDFKPTVDSYWKETEPIFLSTSSLFRFSKKLKSL 702
Query: 284 KTVFKQWNRSVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQ 343
K + + + G++ ++ R A D + + Q+ + + + ++++EA + ++
Sbjct: 703 KPLLRNLAKERLGNLVKKTREAYDTLCKKQESTLN-NPTPNAMKEEVEAHDRWEHVAGLE 761
Query: 344 DQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLY--DGATVLSDAGDIAAHILN 401
++F K+KS+ GD+N FHR + + +IS + DG+ V + +I A+
Sbjct: 762 EKFLKKKSKLHWLDGGDKNNKAFHRAVVTREAQNSISEIQCQDGS-VTAKGDEIKAYAER 820
Query: 402 YFQGIFDVVNNCIPNDLVARSIHAL---VTEDENNMLIRLPLRDEIKSAVFDLNGDGAPG 458
+F+ ++ N +A L +E E+ +L R+ +EIK +F + D +PG
Sbjct: 821 FFREFLQLIPNEYEGVTMADLQDLLPFRCSETEHELLTRVVTAEEIKKVLFSMPNDKSPG 880
Query: 459 PDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPI 518
PDGF F+++ W+I+G++ +L++Q FF +G IN+ +L LIPK A+ M D+RPI
Sbjct: 881 PDGFTSEFFKATWEILGNEFILAIQSFFAKGFLPKGINTTILALIPKKKEAKEMKDYRPI 940
Query: 519 ALANFQFKIVTKILADRL 536
+ N +K+++KI+A+RL
Sbjct: 941 SCCNVIYKVISKIIANRL 958
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 181 bits (459), Expect = 2e-45
Identities = 125/401 (31%), Positives = 191/401 (47%), Gaps = 15/401 (3%)
Query: 141 NAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRLGSAYVALRLDR 200
N ++ EKRG P S DF S+ + N L L SG +W R +V RLDR
Sbjct: 36 NEILDNSEKRGGPPRDQGSFIDFRSFISKNGLWDLKYSGNPFSWRGMRY-DWFVRQRLDR 94
Query: 201 SICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVDCRRLV 260
++ N WL + L SDH PL++ D A VK F+F + L+
Sbjct: 95 AMSNNSWLESFPSGRSEYLRFEGSDHRPLVVFVDEARVKRRGQFRFDNRLRDNDVVNALI 154
Query: 261 LENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRIQQIIDSVG 320
E W G +L T Q R + Q + + + + Q+ ++
Sbjct: 155 QETWTN----------AGDASVL-TKMNQCRREIINWTRLQNLNSAELIEKTQKALEEAL 203
Query: 321 FSDDLYRQDLEA-QLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNI 379
+D + A L +A +++QFWK++SR GDRNT YFH V R + + +
Sbjct: 204 TADPPNPTTIGALTATLEHAYKLEEQFWKQRSRVLWLHSGDRNTGYFHAVTRNRRTQNRL 263
Query: 380 SLLYD-GATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIHALVTEDENNMLIRL 438
+++ D + I+ I YFQ IF ++ +V +I +V++ +N+ L R+
Sbjct: 264 TVMEDINGVAQHEEHQISQIISGYFQQIFTSESDG-DFSVVDEAIEPMVSQGDNDFLTRI 322
Query: 439 PLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSN 498
P +E+K AVF +N APGPDGF FY S+W I+ +DV ++ FF +F +N
Sbjct: 323 PNDEEVKDAVFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNET 382
Query: 499 LLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
+ LIPK G + + D+RPIAL N +KIV KI+ R+ I
Sbjct: 383 HIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLI 423
>At3g31420 hypothetical protein
Length = 1491
Score = 179 bits (455), Expect = 6e-45
Identities = 126/470 (26%), Positives = 204/470 (42%), Gaps = 11/470 (2%)
Query: 71 LWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHLQGCF 130
L WN V + + F++ + L V S +L+ +Y + + R LW L L
Sbjct: 248 LVVFWNSSVDIFLCFSNSNLVDLHVKSNEGSFYLSFVYGHPNPSHRHHLWERLERLNTTR 307
Query: 131 LGP-WLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRL 189
G W +GDFN ++ EKRG + DF + T L S G +W+ G+
Sbjct: 308 QGTAWFIMGDFNEILSNREKRGGRLRPERTFQDFRNMVRGCNFTDLKSVGDRFSWA-GKR 366
Query: 190 GSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKA 249
G +V LDR++ N EW + + + +SDH PL+ + + F +
Sbjct: 367 GDHHVTCSLDRTMANNEWHTLFPESETVFMEYGESDHRPLVTNISAQKEERRGFFSYDSR 426
Query: 250 WTSHVDCRRLVLENW-AKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDE 308
T +++VL W +N G L KL + +W ++ + + ++++ +
Sbjct: 427 LTHKEGFKQVVLNQWHRRNGSFEGDSSLNRKLVECRQAISRWKKNNRVNAEERIKIIRHK 486
Query: 309 VSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHR 368
+ R + RQDL A ++ +W+ KSR + GDRNT YFH
Sbjct: 487 LDRAIATGTATPRERRQMRQDLN------QAYADEEIYWQTKSRSRWLNAGDRNTRYFHS 540
Query: 369 VARIKASSKNISLLYDG-ATVLSDAGDIAAHILNYFQGIF-DVVNNCIPNDLVARSIHAL 426
+ + + + D + +IA +NYF ++ N + V +
Sbjct: 541 TTKTRRCRNRLLSVQDSDGDICRGDENIAKVAINYFDDLYKSTPNTSLRYADVFQGFQQK 600
Query: 427 VTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFF 486
+T++ N LIR EI+ +VF + P PDGF FYQ FW + V+ V FF
Sbjct: 601 ITDEINEDLIRPVTELEIEESVFSVAPSRTPDPDGFTADFYQQFWPDIKQKVIDEVTRFF 660
Query: 487 LQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
+ + N L LIPK+ + FRPIAL N +KI++KIL +RL
Sbjct: 661 ERSELDERHNHTNLCLIPKVETPTTIAKFRPIALCNVSYKIISKILVNRL 710
>At4g26360 putative protein
Length = 1141
Score = 176 bits (446), Expect = 7e-44
Identities = 139/488 (28%), Positives = 227/488 (46%), Gaps = 32/488 (6%)
Query: 68 LPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSS--VFLAAIYANTSYLFRRLLWADLTH 125
L +W LW+ V VV++ S Q I EV + +S + ++ +YA R+ LW ++T
Sbjct: 60 LGKIWVLWDPSVEVVIVAKSLQMITCEVLFPNSRTWIVISVVYAANEDDKRKELWREITA 119
Query: 126 LQGC---FLGPWLFIGDFNAVMGAHE-KRGRWPPTAASCSDFGSWSNANLLTHLPSSGPL 181
L F PW+ +GDFN V+ HE R DF L+ L G
Sbjct: 120 LVASPVTFNRPWILLGDFNQVLHPHEHSRHVSLNVDRRIRDFRECLLDAELSDLVYKGSS 179
Query: 182 LTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNA 241
TW N + + VA ++DR + NE W + + SDH + ++ +K
Sbjct: 180 FTWWN-KSKTRPVAKKIDRILVNESWSNLFPSSFGLFGPPDFSDHASCGVVLELDPIKAK 238
Query: 242 ISFKFYKAWTSHVDCRRLVLENW-AKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDR 300
FKF+ + + LV + W + NV G M R+ KL+ LK K ++R + ++++
Sbjct: 239 RPFKFFNFLLKNPEFLNLVWDVWYSTNVVGSSMFRVSKKLKALKKPIKDFSRLNYSNLEK 298
Query: 301 QVRLATDEVSRIQQI-IDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHG 359
+ A + + Q + +D+ + + +LEAQ ++ F++++SR F G
Sbjct: 299 RTEEAHETLLSFQNLTLDNPSLENAAH--ELEAQRKWQILATAEESFFRQRSRVTWFAEG 356
Query: 360 DRNTAYFHRVARIKASSKNISLLYDGA-TVLSDAGDIAAHILNYFQGIFDVVNNCIPNDL 418
D NT YFHR+A + S I+ L D + T + IA H YF+ + N+ P L
Sbjct: 357 DGNTRYFHRMADSRKSVNTITTLVDDSGTQIDSQQGIADHCALYFENLLSDDND--PYSL 414
Query: 419 VARSIHALVTE----DENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIV 474
++ L+T + L + ++IK+A F L + A GPDGF
Sbjct: 415 EQDDMNLLLTYRCPYSQVADLEAMFSDEDIKAAFFGLPSNKACGPDGF------------ 462
Query: 475 GSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILAD 534
V +V++FF+ G+ N+ +VLIPK P A DFRPI+ N +K++ ++L D
Sbjct: 463 --PVTAAVREFFISGNLLKQWNATTIVLIPKFPNASCTSDFRPISCMNTLYKVIARLLTD 520
Query: 535 RLASIAMC 542
RL + C
Sbjct: 521 RLQKLLSC 528
>At1g24640 hypothetical protein
Length = 1270
Score = 174 bits (442), Expect = 2e-43
Identities = 132/470 (28%), Positives = 206/470 (43%), Gaps = 22/470 (4%)
Query: 74 LWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHLQGCFLGP 133
LW V V + F + +V + + ++ +Y + R W ++ +
Sbjct: 34 LWKSSVQVDLKFVDKNLMDAQVQFGAVNFCVSCVYGDPDRSKRSQAWERISRIGVGRRDK 93
Query: 134 WLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRLGSAY 193
W GDFN ++ EK G + C F L +P+ G TW+ GR G +
Sbjct: 94 WCMFGDFNDILHNGEKNGGPRRSDLDCKAFNEMIKGCDLVEMPAHGNGFTWA-GRRGDHW 152
Query: 194 VALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSH 253
+ RLDR+ N+EW F+ V++ L SDH P+L+ + F+F K +
Sbjct: 153 IQCRLDRAFGNKEWFCFFPVSNQTFLDFRGSDHRPVLIKLMSSQDSYRGQFRFDKRFLFK 212
Query: 254 VDCRRLVLENWAKNVRGVGMV---RLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVS 310
D + ++ W++ G + RL+ + L + KQ N + +++ E S
Sbjct: 213 EDVKEAIIRTWSRGKHGTNISVADRLRACRKSLSSWKKQNNLNSLDKINQLEAALEKEQS 272
Query: 311 RIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVA 370
+ I V + ++DL A ++ +WK+KSR + G+RN+ YFH
Sbjct: 273 LVWPIFQRVS----VLKKDL------AKAYREEEAYWKQKSRQKWLRSGNRNSKYFHAAV 322
Query: 371 RIKASSKNISLLYD--GATVLSDA--GDIAAHILNYFQGIFDVVNNCIPNDLVARSIHAL 426
+ K I L D G S+A G++AA YF +F N D + +
Sbjct: 323 KQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAA---YFGNLFKSSNPSGFTDWFSGLVPR- 378
Query: 427 VTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFF 486
V+E N L+ EIK AVF + APGPDG F+Q +W VG+ V V+ FF
Sbjct: 379 VSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFF 438
Query: 487 LQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
G N L LIPK M D RPI+L + +KI++KI+A RL
Sbjct: 439 ADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRL 488
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 174 bits (441), Expect = 3e-43
Identities = 124/474 (26%), Positives = 214/474 (44%), Gaps = 11/474 (2%)
Query: 67 RLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHL 126
R L +W + V + +I ++ I VT+ + S +L+ +Y + + R LW L H+
Sbjct: 219 RSGGLALMWKNNVSLSLISQDERLIDSHVTFNNKSFYLSCVYGHPTQSERHQLWQTLEHI 278
Query: 127 QGCFLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSN 186
WL +GDFN ++ EK G + +F + + + + S G +W
Sbjct: 279 SDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIEDMRSKGDRFSWV- 337
Query: 187 GRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKF 246
G + V LDR N W A + L SDH P+L+ + + + + F+F
Sbjct: 338 GERHTHTVKCCLDRVFINSAWTATFPYAETEFLDFTGSDHKPVLVHFNESFPRRSKLFRF 397
Query: 247 YKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLAT 306
+R+V +W N R + ++ + + + + +++++
Sbjct: 398 DNRLIDIPTFKRIVQTSWRTN-RNSRSTPITERISSCRQAMARLKHASNLNSEQRIKKLQ 456
Query: 307 DEVSRIQQIIDSVGFSDDLYRQDL-EAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAY 365
++R + V RQ + + Q L A + ++ +WK+KSR+Q GD+NT Y
Sbjct: 457 SSLNRAMESTRRVD------RQLIPQLQESLAKAFSDEEIYWKQKSRNQWMKEGDQNTGY 510
Query: 366 FHRVARIKASSKNISLLYDG-ATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIH 424
FH + + S ++ + D + + +I H ++F IF N + +
Sbjct: 511 FHACTKTRYSQNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFST-NGIKVSPIDFADFK 569
Query: 425 ALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQD 484
+ VT N L + EI A+ + D APGPDG FY++ WDIVG DV+L V+
Sbjct: 570 STVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKK 629
Query: 485 FFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLAS 538
FF +IN + +IPKI + D+RPIAL N +K+++K L +RL S
Sbjct: 630 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKS 683
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 174 bits (441), Expect = 3e-43
Identities = 124/474 (26%), Positives = 214/474 (44%), Gaps = 11/474 (2%)
Query: 67 RLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHL 126
R L +W + V + +I ++ I VT+ + S +L+ +Y + + R LW L H+
Sbjct: 445 RSGGLALMWKNNVSLSLISQDERLIDSHVTFNNKSFYLSCVYGHPTQSERHQLWQTLEHI 504
Query: 127 QGCFLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSN 186
WL +GDFN ++ EK G + +F + + + + S G +W
Sbjct: 505 SDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIEDMRSKGDRFSWV- 563
Query: 187 GRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKF 246
G + V LDR N W A + L SDH P+L+ + + + + F+F
Sbjct: 564 GERHTHTVKCCLDRVFINSAWTATFPYAEIEFLDFTGSDHKPVLVHFNESFPRRSKLFRF 623
Query: 247 YKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLAT 306
+R+V +W N R + ++ + + + + +++++
Sbjct: 624 DNRLIDIPTFKRIVQTSWRTN-RNSRSTPITERISSCRQAMARLKHASNLNSEQRIKKLQ 682
Query: 307 DEVSRIQQIIDSVGFSDDLYRQDL-EAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAY 365
++R + V RQ + + Q L A + ++ +WK+KSR+Q GD+NT Y
Sbjct: 683 SSLNRAMESTRRVD------RQLIPQLQESLAKAFSDEEIYWKQKSRNQWMKEGDQNTGY 736
Query: 366 FHRVARIKASSKNISLLYDG-ATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIH 424
FH + + S ++ + D + + +I H ++F IF N + +
Sbjct: 737 FHACTKTRYSQNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFST-NGIKVSPIDFADFK 795
Query: 425 ALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQD 484
+ VT N L + EI A+ + D APGPDG FY++ WDIVG DV+L V+
Sbjct: 796 STVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKK 855
Query: 485 FFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLAS 538
FF +IN + +IPKI + D+RPIAL N +K+++K L +RL S
Sbjct: 856 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKS 909
>At4g10830 putative protein
Length = 1294
Score = 171 bits (434), Expect = 2e-42
Identities = 125/465 (26%), Positives = 214/465 (45%), Gaps = 11/465 (2%)
Query: 74 LWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHLQGCFLGP 133
LW D V + ++ D+ I + ++ + + +L+ +Y + R LW +L P
Sbjct: 432 LWKDSVRLSNLYQDDRHIDVHISINNINFYLSRVYGHPCQSERHSLWTHFENLSKTRNDP 491
Query: 134 WLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRLGSAY 193
W+ IGDFN ++ +EK G + F + + L + S G +W G S
Sbjct: 492 WILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTCDLKDIRSIGDRFSWV-GERHSHT 550
Query: 194 VALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSH 253
V LDR+ N E + L SDH PL LS + + F+F K
Sbjct: 551 VKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTETRKMRPFRFDKRLLEV 610
Query: 254 VDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRIQ 313
+ V W K + G L ++R + + +++ ++R+ + + +
Sbjct: 611 PHFKTYVKAGWNKAINGQRK-HLPDQVRTCRQAMAKLKHK--SNLNSRIRINQLQAA-LD 666
Query: 314 QIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIK 373
+ + SV ++ R Q LT A ++++W++KSR+Q GDRNT +FH + +
Sbjct: 667 KAMSSVNRTER--RTISHIQRELTVAYRDEERYWQQKSRNQWMKEGDRNTEFFHACTKTR 724
Query: 374 ASSKNISLLYDGATVLSDAG-DIAAHILNYFQGIFDVVNNCIPNDLVA-RSIHALVTEDE 431
S + + D ++ +I H +F +++ +N P ++ +VTE
Sbjct: 725 FSVNRLVTIKDEEGMIYRGDKEIGVHAQEFFTKVYE--SNGRPVSIIDFAGFKPIVTEQI 782
Query: 432 NNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSF 491
N+ L + EI +A+ + D APGPDG FY+S W+IVG DV+ V+ FF
Sbjct: 783 NDDLTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYM 842
Query: 492 ADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
+IN + +IPKI + + D+RPIAL N +KI++K L +RL
Sbjct: 843 KQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERL 887
>At4g04000 putative reverse transcriptase
Length = 1077
Score = 166 bits (421), Expect = 5e-41
Identities = 108/345 (31%), Positives = 168/345 (48%), Gaps = 12/345 (3%)
Query: 196 LRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVD 255
LRLDR++ N +W+ Y L SDH PL+ S D K F++ + + +
Sbjct: 171 LRLDRAMANSDWIIAYPSGRSEYLRFEGSDHRPLVTSFDHVQKKTKNIFRYDRTLKDNPE 230
Query: 256 CRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRIQQI 315
+L+ E W N ++ +L + + W++ ++ Q + + V + +
Sbjct: 231 ISKLMNETWKSNPSA----KVDYRLHLCRAALLNWSKE--HHLNSQKEILSLRVQLEEAL 284
Query: 316 IDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKAS 375
D D + DL Q LL A ++ FWK++SR+ GD+N+ +FH + +
Sbjct: 285 TDDAATQDSV---DLLNQNLLL-AYRKEETFWKQRSRNLWLALGDKNSCFFHAATNSRRA 340
Query: 376 SKNISLLYDGA-TVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIHALVTEDENNM 434
IS++ + A T + + +I I +YFQ IF +VA ++ +T D NN
Sbjct: 341 INTISVIENSAGTSVYEDSEIIETISDYFQDIFTSQEGD-RTSVVAEALSPCITTDMNNT 399
Query: 435 LIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADN 494
LI LP EIK A ++ D APG DGF F+QS W V ++ VQ FF+
Sbjct: 400 LISLPTVAEIKQACLSIHPDKAPGRDGFSASFFQSNWSTVSEEITAEVQAFFISEILPQK 459
Query: 495 INSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
IN + LIPKIP Q + D+RPIAL + +KI+ K+LA RL I
Sbjct: 460 INHTHVRLIPKIPAPQKVSDYRPIALCSVYYKIIAKLLAKRLQPI 504
>At2g11240 pseudogene
Length = 1044
Score = 163 bits (413), Expect = 5e-40
Identities = 113/370 (30%), Positives = 180/370 (48%), Gaps = 23/370 (6%)
Query: 172 LTHLPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLL 231
L L SG L+W G+ V RLDR++ N W Y + C L SDH PLL
Sbjct: 6 LYDLRHSGNFLSW-RGKRHDHVVHCRLDRALSNGAWAEDYPASRCIYLCFEGSDHRPLLT 64
Query: 232 SADVASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWN 291
D++ K F++ + ++ + LV E W N+ +V + K+ + +W+
Sbjct: 65 HFDLSKKKKKGVFRYDRRLKNNDEVTALVQEAW--NLYDTDIV--EEKISRCRLEIVKWS 120
Query: 292 RSVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKS 351
R+ +L + ++++ + S + +L + LLL A ++++WK++S
Sbjct: 121 RA---KQQSSQKLIEENRQKLEEAMSSQDHNQELL-STINTNLLL--AYKAEEEYWKQRS 174
Query: 352 RHQSFIHGDRNTAYFHRVARIKASSKNISLLY--DGATVLSDAGDIAAHILNYFQGIFDV 409
R GD+N+ YFH + R + S++ DG +AG I I YFQ +F
Sbjct: 175 RQLWLALGDKNSGYFHAITRGRTVINKFSVIEKEDGVPEYEEAG-ILNVISEYFQKLFSA 233
Query: 410 VNNCIPNDLVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQS 469
N + +I ++ ++N +EIKSA F ++ D APGPDGF F+QS
Sbjct: 234 -NEGARAATIKEAIKPFISPEQNP--------EEIKSACFSIHADKAPGPDGFSASFFQS 284
Query: 470 FWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVT 529
W VG ++VL +Q FF + IN + LIPKI + M D+RPIAL +KI++
Sbjct: 285 NWMTVGPNIVLEIQSFFSSSTLQPTINKTHITLIPKIQSLKRMVDYRPIALCTVFYKIIS 344
Query: 530 KILADRLASI 539
K+L+ RL I
Sbjct: 345 KLLSRRLQPI 354
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 156 bits (394), Expect = 7e-38
Identities = 128/479 (26%), Positives = 210/479 (43%), Gaps = 26/479 (5%)
Query: 67 RLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHL 126
R L W + + ++ + L+V+ ++ F++ +Y R LW L +
Sbjct: 42 RSGGLAIFWKSHLEIEFLYADKNLMDLQVSSRNKVWFISCVYGLPVTHMRPKLWEHLNSI 101
Query: 127 QGCFLGPWLFIGDFNAVMGAHEKRG--RWPPTAASCSDFGSWSNANLLTHLPSSGPLLTW 184
W IGDFN + EK G R P++ C + + + + L S+G TW
Sbjct: 102 GLKRAEAWCLIGDFNDIRSNDEKLGGPRRSPSSFQCFEHMLLNCS--MHELGSTGNSFTW 159
Query: 185 SNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISF 244
R +V +LDR N W + + L + SDH P+L+ + F
Sbjct: 160 GGNR-NDQWVQCKLDRCFGNPAWFSIFPNAHQWFLEKFGSDHRPVLVKFTNDNELFRGQF 218
Query: 245 KFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRL 304
++ K C ++ +W + G L + W S D + Q R
Sbjct: 219 RYDKRLDDDPYCIEVIHRSW-NSAMSQGTHSSFFSLIECRRAISVWKHS--SDTNAQSR- 274
Query: 305 ATDEVSRIQQIID---SVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDR 361
+ R+++ +D S+ + ++ QL L A ++ FW++KSR + GD+
Sbjct: 275 ----IKRLRKDLDAEKSIQIPCWPRIEYIKDQLSL--AYGDEELFWRQKSRQKWLAGGDK 328
Query: 362 NTAYFHRVARIKASSKNISLLYDGA----TVLSDAGDIAAHILNYFQGIFDVVNNCIPND 417
NT +FH + +S L D T SD G IA+ ++F+ +F N+
Sbjct: 329 NTGFFHATVHSERLKNELSFLLDENDQEFTRNSDKGKIAS---SFFENLFTSTYILTHNN 385
Query: 418 LVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
+ + A VT + N+ LI+ E+ +AVF +N + APGPDGF F+Q WD+V
Sbjct: 386 HL-EGLQAKVTSEMNHNLIQEVTELEVYNAVFSINKESAPGPDGFTALFFQQHWDLVKHQ 444
Query: 478 VVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
++ + FF G + N + LIPKI Q M D RPI+L + +KI++KIL RL
Sbjct: 445 ILTEIFGFFETGVLPQDWNHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRL 503
>At1g31100 hypothetical protein
Length = 1090
Score = 149 bits (375), Expect = 1e-35
Identities = 125/453 (27%), Positives = 199/453 (43%), Gaps = 24/453 (5%)
Query: 101 SVFLAAIYANTSYLFRRLLWADLTHLQGCFLG---PWLFIGDFNAVM-GAHEKRGRWPPT 156
SV ++ +YA + R+ LW +L L G PW+ +GDFN V+ A +
Sbjct: 14 SVVVSIVYAANEAITRKELWEELLLLSVSLSGNGKPWIMLGDFNQVLCPAEHSQATSLNV 73
Query: 157 AASCSDFGSWSNANLLTHLPSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSC 216
F L L G TW N + + VA +LDR + NE W + +
Sbjct: 74 NRRMKVFRDCLFEAELCDLVFKGNTFTWWN-KSATRPVAKKLDRILVNESWCSRFPSAYA 132
Query: 217 CTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVDCRRLVLENWAK-NVRGVGMVR 275
SDH + + + F+FY + D LV E W NV G M +
Sbjct: 133 VFGEPDFSDHASCGVIINPLMHREKRPFRFYNFLLQNPDFISLVGELWYSINVVGSSMFK 192
Query: 276 LQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQ---DLEA 332
+ KL+ LK + ++ F +++++V+ A + V Q + SD ++EA
Sbjct: 193 MSKKLKALKNPIRTFSMENFSNLEKRVKEAHNLVLYRQ----NKTLSDPTIPNAALEMEA 248
Query: 333 QLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLYDGATVLSDA 392
Q + ++ F+ ++SR GD NT+YFHR+A + + I ++ D V D
Sbjct: 249 QRKWLILVKAEESFFCQRSRVTWMGEGDSNTSYFHRMADSRKAVNTIHIIIDDNGVKIDT 308
Query: 393 G-DIAAHILNYFQGIFDVVNNCIPNDLVARSIHALV----TEDENNMLIRLPLRDEIKSA 447
I H + YF + P L+ L+ + D+ L R +IKSA
Sbjct: 309 QLGIKEHCIEYFSNLLG--GEVGPPMLIQEDFDLLLPFRCSHDQKKELAMSFSRQDIKSA 366
Query: 448 VFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIP 507
F + GPDGF F++ W ++G++V +V +FF N+ LVLIPKI
Sbjct: 367 FFSFPSNKTSGPDGFPVEFFKETWSVIGTEVTDAVSEFFTSSVLLKQWNATTLVLIPKIT 426
Query: 508 GAQAMGDFRPIALANF----QFKIVTKILADRL 536
A M DFRPI+ +F +K++ ++L +RL
Sbjct: 427 NASKMNDFRPISCNDFGPITLYKVIARLLTNRL 459
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 144 bits (362), Expect = 4e-34
Identities = 100/366 (27%), Positives = 169/366 (45%), Gaps = 31/366 (8%)
Query: 179 GPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASV 238
G TW G + S +VA RLDR + ++ C + P
Sbjct: 5 GNRFTWRRGLVESTFVAKRLDRVLFCAHARLKWQEALLCPAQNVDARRRP---------- 54
Query: 239 KNAISFKFYKAWTSHVDCRRLVLENWAKNVRG-VGMVRLQGKLRILKTVFKQWNRSVFGD 297
F+F AW SH + L+ +W + V + RL+ +L K+WN+ VFG+
Sbjct: 55 -----FRFEAAWLSHEGFKELLTASWDTGLSTPVALNRLRWQL-------KKWNKEVFGN 102
Query: 298 VDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDL---EAQLLLTNALNVQDQFWKEKSRHQ 354
+ + ++ +Q +++ V D L ++D E +LL + ++ W +KSR +
Sbjct: 103 IHVRKEKVVSDLKAVQDLLEVVQTDDLLMKEDTLLKEFDVLL----HQEETLWFQKSREK 158
Query: 355 SFIHGDRNTAYFHRVARIKASSKNISLLYDGATV-LSDAGDIAAHILNYFQGIFDVVNNC 413
GDRNT +FH I+ I +L D +++ + ++Y++ ++ + +
Sbjct: 159 LLALGDRNTTFFHTSTVIRRRRNRIEMLKDSEDRWVTEKEALEKLAMDYYRKLYSLEDVS 218
Query: 414 IPNDLVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDI 473
+ + +T +E N L R RDE+ AV + APGPDG+ FYQ W+
Sbjct: 219 VVRGTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPGPDGYQPVFYQQCWET 278
Query: 474 VGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILA 533
VG V V +FF G + N LLVL+ K+ + + FRP++L N FKI+TK++
Sbjct: 279 VGESVSKFVMEFFESGVLPKSTNDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMMV 338
Query: 534 DRLASI 539
RL ++
Sbjct: 339 IRLKNV 344
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.346 0.153 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,079,049
Number of Sequences: 26719
Number of extensions: 1060105
Number of successful extensions: 3923
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 3681
Number of HSP's gapped (non-prelim): 108
length of query: 1145
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1035
effective length of database: 8,379,506
effective search space: 8672788710
effective search space used: 8672788710
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146565.8


