

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
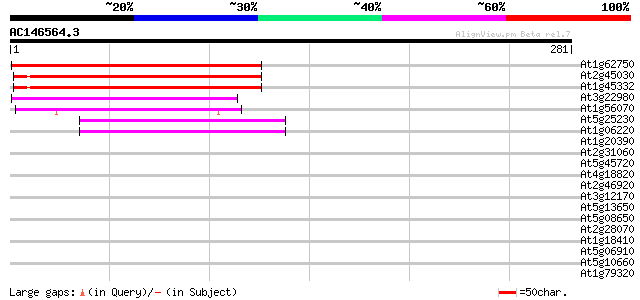
Score E
Sequences producing significant alignments: (bits) Value
At1g62750 elongation factor G, putative 180 7e-46
At2g45030 putative mitochondrial translation elongation factor G 92 4e-19
At1g45332 mitochondrial elongation factor, putative 92 4e-19
At3g22980 eukaryotic translation elongation factor 2, putative 59 4e-09
At1g56070 elongation factor like protein 52 4e-07
At5g25230 translation Elongation Factor 2 - like protein 49 3e-06
At1g06220 elongation factor like protein 49 3e-06
At1g20390 hypothetical protein 33 0.17
At2g31060 putative GTP-binding protein 32 0.29
At5g45720 putative protein 32 0.50
At4g18820 putative protein 30 1.5
At2g46920 unknown protein 30 1.9
At3g12170 hypothetical protein 29 2.5
At5g13650 GTP-binding protein typA (tyrosine phosphorylated prot... 28 4.2
At5g08650 GTP-binding protein LepA homolog 28 4.2
At2g28070 putative ABC transporter 28 4.2
At1g18410 kinesin-related protein, putative 28 4.2
At5g06910 DnaJ homologue (gb|AAB91418.1|) 28 5.5
At5g10660 vacuolar calcium binding protein - like 27 9.4
At1g79320 27 9.4
>At1g62750 elongation factor G, putative
Length = 783
Score = 180 bits (457), Expect = 7e-46
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 460 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 519
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 520 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 579
Query: 121 RYVFNK 126
+Y K
Sbjct: 580 KYTHKK 585
Score = 37.4 bits (85), Expect = 0.009
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 749 MTKGRASYTMQLAKFDVVPQHIQNQLSS 776
>At2g45030 putative mitochondrial translation elongation factor G
Length = 754
Score = 91.7 bits (226), Expect = 4e-19
Identities = 47/125 (37%), Positives = 77/125 (61%), Gaps = 2/125 (1%)
Query: 3 IVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAEE 62
IV + G+ + +G+T D + + M+ P V+ +A++P +K + L + +E
Sbjct: 430 IVAVFGI-ECASGDTFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGGQFSKALNRFQKE 488
Query: 63 DPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEVR 121
DP+FRV D E +TII G GELHL+ V+R++RE+KV+A VG P+ N+RE+I++ E
Sbjct: 489 DPTFRVGLDPESGQTIISGMGELHLDIYVERMRREYKVDATVGKPRVNFRETITQRAEFD 548
Query: 122 YVFNK 126
Y+ K
Sbjct: 549 YLHKK 553
>At1g45332 mitochondrial elongation factor, putative
Length = 754
Score = 91.7 bits (226), Expect = 4e-19
Identities = 47/125 (37%), Positives = 77/125 (61%), Gaps = 2/125 (1%)
Query: 3 IVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAEE 62
IV + G+ + +G+T D + + M+ P V+ +A++P +K + L + +E
Sbjct: 430 IVAVFGI-ECASGDTFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGGQFSKALNRFQKE 488
Query: 63 DPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEVR 121
DP+FRV D E +TII G GELHL+ V+R++RE+KV+A VG P+ N+RE+I++ E
Sbjct: 489 DPTFRVGLDPESGQTIISGMGELHLDIYVERMRREYKVDATVGKPRVNFRETITQRAEFD 548
Query: 122 YVFNK 126
Y+ K
Sbjct: 549 YLHKK 553
>At3g22980 eukaryotic translation elongation factor 2, putative
Length = 1015
Score = 58.5 bits (140), Expect = 4e-09
Identities = 39/116 (33%), Positives = 57/116 (48%), Gaps = 3/116 (2%)
Query: 2 DIVTLVGLKDTIA-GETLCDPKGPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
++V + GL I+ TL + + L M+F V +++AIEP ADM + GL L
Sbjct: 502 NVVAIRGLGPYISKSATLSSTRNCWPLASMEFQVSPTLRVAIEPSDPADMSALMKGLRLL 561
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREF-KVEANVGAPQANYRESI 114
DP ++ ++ GE+HLE V LK F KV V P +YRE+I
Sbjct: 562 NRADPFVEITVSARGEHVLAAAGEVHLERCVKDLKERFAKVNLEVSPPLVSYRETI 617
>At1g56070 elongation factor like protein
Length = 843
Score = 52.0 bits (123), Expect = 4e-07
Identities = 36/119 (30%), Positives = 62/119 (51%), Gaps = 6/119 (5%)
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V +VGL I TL + K + M F V V+++A++ K +D+ K+ GL +L
Sbjct: 452 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 511
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANV--GAPQANYRESISE 116
A+ DP + ++ I+ G GELHLE + L+ +F A + P ++RE++ +
Sbjct: 512 AKSDPMVVCTMEESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVCD 570
>At5g25230 translation Elongation Factor 2 - like protein
Length = 973
Score = 48.9 bits (115), Expect = 3e-06
Identities = 28/104 (26%), Positives = 54/104 (51%), Gaps = 1/104 (0%)
Query: 36 VIKIAIEPKTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKR 95
V+K A EP +++ KM GL K+++ P ++ I GTGEL+L++I+ L+
Sbjct: 587 VVKTATEPLNPSELPKMVEGLRKISKSYPLAITKVEESGEHTILGTGELYLDSIIKDLRE 646
Query: 96 EF-KVEANVGAPQANYRESISEVTEVRYVFNKPYILNPWTQVVD 138
+ +V+ V P ++ E++ E + ++ P N T + +
Sbjct: 647 LYSEVQVKVADPVVSFCETVVESSSMKCFAETPNKKNKLTMIAE 690
>At1g06220 elongation factor like protein
Length = 987
Score = 48.9 bits (115), Expect = 3e-06
Identities = 29/104 (27%), Positives = 54/104 (51%), Gaps = 1/104 (0%)
Query: 36 VIKIAIEPKTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKR 95
V+K A EP +++ KM GL K+++ P ++ I GTGEL+L++I+ L+
Sbjct: 601 VVKTATEPLNPSELPKMVEGLRKISKSYPLAITKVEESGEHTILGTGELYLDSIMKDLRE 660
Query: 96 EF-KVEANVGAPQANYRESISEVTEVRYVFNKPYILNPWTQVVD 138
+ +VE V P ++ E++ E + ++ P N T + +
Sbjct: 661 LYSEVEVKVADPVVSFCETVVESSSMKCFAETPNKKNKITMIAE 704
>At1g20390 hypothetical protein
Length = 1791
Score = 33.1 bits (74), Expect = 0.17
Identities = 26/77 (33%), Positives = 42/77 (53%), Gaps = 9/77 (11%)
Query: 59 LAEEDPSFRVSQDKEDRTIIQGT----GELHLETIVDRLKR-EFKVE---ANVGAPQANY 110
LAE++P R + D+E+R + GT G L L+ I+D L R E + E A + A QA+
Sbjct: 20 LAEQNPPVRRNTDQEERRVTDGTYVGPGGL-LQNILDHLSRQEARTEARFATLAAAQAHS 78
Query: 111 RESISEVTEVRYVFNKP 127
+ I+ + R ++P
Sbjct: 79 MDEINRLQTSRTSVHRP 95
>At2g31060 putative GTP-binding protein
Length = 664
Score = 32.3 bits (72), Expect = 0.29
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 17/133 (12%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLD---------QMDFPVHVIKIAIEPKTKADMEKM 52
DI+ + GL G T+ + L M F V+ +A + T ++
Sbjct: 330 DIICMAGLTAPSIGHTVASAEVTTALPTVELDPPTISMTFGVNDSPLAGQDGTHLTGGRI 389
Query: 53 EAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYR- 111
L+ AE + + V + +QG GEL L +++ ++RE E +V P+ Y+
Sbjct: 390 GDRLMAEAETNLAINVIPGLSESYEVQGRGELQLGILIENMRRE-GFELSVSPPKVMYKT 448
Query: 112 ------ESISEVT 118
E I EVT
Sbjct: 449 EKGQKLEPIEEVT 461
>At5g45720 putative protein
Length = 900
Score = 31.6 bits (70), Expect = 0.50
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 42 EPKTKADMEKMEAGLIKLAEEDPSFRVSQDK 72
+P +K DMEK++ L L+E + RVS DK
Sbjct: 647 QPLSKEDMEKLKQALKTLSESEKQLRVSNDK 677
>At4g18820 putative protein
Length = 1057
Score = 30.0 bits (66), Expect = 1.5
Identities = 14/31 (45%), Positives = 18/31 (57%)
Query: 42 EPKTKADMEKMEAGLIKLAEEDPSFRVSQDK 72
+P K DMEK+ L L+E + RVS DK
Sbjct: 752 QPLPKEDMEKLRQALKTLSEAEKQLRVSNDK 782
>At2g46920 unknown protein
Length = 814
Score = 29.6 bits (65), Expect = 1.9
Identities = 15/42 (35%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Query: 91 DRLKREFKVEANVGA---PQANYRESISEVTEVRYVFNKPYI 129
DR+K + KV GA + N+ E++ E+ +V Y+ PYI
Sbjct: 699 DRVKGQLKVTRAFGAGFLKKPNFNEALLEMFQVEYIGTDPYI 740
>At3g12170 hypothetical protein
Length = 262
Score = 29.3 bits (64), Expect = 2.5
Identities = 27/103 (26%), Positives = 45/103 (43%), Gaps = 15/103 (14%)
Query: 35 HVIKIAIEP-KTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELH----LETI 89
H + + + P K K D + E K + + D+E R + TG + +
Sbjct: 32 HKLALRLHPDKNKDDEDAKE----KFQQLQKVISILGDEEKRAVYDQTGSVDDADLSGDV 87
Query: 90 VDRLKREFKV------EANVGAPQANYRESISEVTEVRYVFNK 126
VD L+ FK E ++ +ANYR S SE ++ ++NK
Sbjct: 88 VDNLRDFFKAMYKKVTEEDIEEFEANYRGSESEKNDLIELYNK 130
>At5g13650 GTP-binding protein typA (tyrosine phosphorylated protein
A)
Length = 609
Score = 28.5 bits (62), Expect = 4.2
Identities = 25/116 (21%), Positives = 47/116 (39%), Gaps = 11/116 (9%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTK---------ADMEKM 52
DI + G+ + GET+ D L + +K++ T +
Sbjct: 283 DICAVCGIDNIQIGETIADKVHGKPLPTIKVEEPTVKMSFSVNTSPFSGREGKYVTSRNL 342
Query: 53 EAGLIKLAEEDPSFRVSQ-DKEDRTIIQGTGELHLETIVDRLKREFKVEANVGAPQ 107
L + E + + +V + D I+ G G LH+ +++ ++RE E VG P+
Sbjct: 343 RDRLNRELERNLAMKVEDGETADTFIVSGRGTLHITILIENMRRE-GYEFMVGPPK 397
>At5g08650 GTP-binding protein LepA homolog
Length = 675
Score = 28.5 bits (62), Expect = 4.2
Identities = 12/30 (40%), Positives = 18/30 (60%)
Query: 82 GELHLETIVDRLKREFKVEANVGAPQANYR 111
G LH+E + +RL+RE+ + AP YR
Sbjct: 419 GLLHMEIVQERLEREYNLNLITTAPSVVYR 448
>At2g28070 putative ABC transporter
Length = 705
Score = 28.5 bits (62), Expect = 4.2
Identities = 17/53 (32%), Positives = 26/53 (48%)
Query: 20 DPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAEEDPSFRVSQDK 72
D G F MD V + + K+ AD + +EA +IKL E + + S+ K
Sbjct: 384 DDNGDFSAVNMDTAVAIRTLEATYKSSADADSVEAMIIKLTEREGTQLKSKGK 436
>At1g18410 kinesin-related protein, putative
Length = 1081
Score = 28.5 bits (62), Expect = 4.2
Identities = 30/136 (22%), Positives = 60/136 (44%), Gaps = 7/136 (5%)
Query: 27 LDQMDFPVHVIKIAIEPKTKADMEKM--EAGLIKLAEEDPSFRVSQDKEDRTIIQGTGEL 84
L+QM V + A+E + + ++EKM EA +K+ E+ + Q +D TI T
Sbjct: 401 LEQMRKDASVARKALEERVR-ELEKMGKEADAVKMNLEE-KVKELQKYKDETITVTTSIE 458
Query: 85 HLETIVDRLKRE-FKVEANVGAPQANYRESISEVTEVRYVFNKPY--ILNPWTQVVDTNL 141
+++ K+E V ++ A ++I E V + + + N
Sbjct: 459 GKNRELEQFKQETMTVTTSLEAQNRELEQAIKETMTVNTSLEAKNRELEQSKKETMTVNT 518
Query: 142 RIKSKKKHFQKNSLYW 157
+K+K + ++N ++W
Sbjct: 519 SLKAKNRELEQNLVHW 534
>At5g06910 DnaJ homologue (gb|AAB91418.1|)
Length = 284
Score = 28.1 bits (61), Expect = 5.5
Identities = 26/103 (25%), Positives = 45/103 (43%), Gaps = 15/103 (14%)
Query: 35 HVIKIAIEP-KTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELH-------- 85
H + + + P K + D E + K + + D+E R + TG +
Sbjct: 50 HKLALKLHPDKNQDDKEAKD----KFQQLQKVISILGDEEKRAVYDQTGSIDDADIPGDA 105
Query: 86 LETIVDRLKREFKV--EANVGAPQANYRESISEVTEVRYVFNK 126
E + D + +K EA++ +ANYR S SE ++ +FNK
Sbjct: 106 FENLRDFFRDMYKKVNEADIEEFEANYRGSESEKKDLLELFNK 148
>At5g10660 vacuolar calcium binding protein - like
Length = 407
Score = 27.3 bits (59), Expect = 9.4
Identities = 20/78 (25%), Positives = 35/78 (44%), Gaps = 5/78 (6%)
Query: 42 EPKTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEA 101
EPK ++E+ + EE+ + S D+E+ + T +++ V+ K E K EA
Sbjct: 271 EPKEPENIEENNS-----EEEEEVKKKSDDEENSETVATTTDMNEAVNVEESKEEEKEEA 325
Query: 102 NVGAPQANYRESISEVTE 119
V + + E TE
Sbjct: 326 EVKEEEGESSAAKEETTE 343
>At1g79320
Length = 368
Score = 27.3 bits (59), Expect = 9.4
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Query: 31 DFPVHVIKIAIEPKTKADMEKMEAGLIKLAEEDP--SFRVSQDKEDRTIIQGTGELHLET 88
D P+ +I + D K + G ++D S ++++ E I G L LET
Sbjct: 129 DCPITIISDSCHSGGLIDEAKEQIGESTKKKKDSGDSSTINKETEAEIIEVGNRSLPLET 188
Query: 89 IVDRLKRE 96
++D LK+E
Sbjct: 189 LIDMLKQE 196
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,817,794
Number of Sequences: 26719
Number of extensions: 225741
Number of successful extensions: 798
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 782
Number of HSP's gapped (non-prelim): 22
length of query: 281
length of database: 11,318,596
effective HSP length: 98
effective length of query: 183
effective length of database: 8,700,134
effective search space: 1592124522
effective search space used: 1592124522
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146564.3


