

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.14 - phase: 0 /pseudo
(312 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
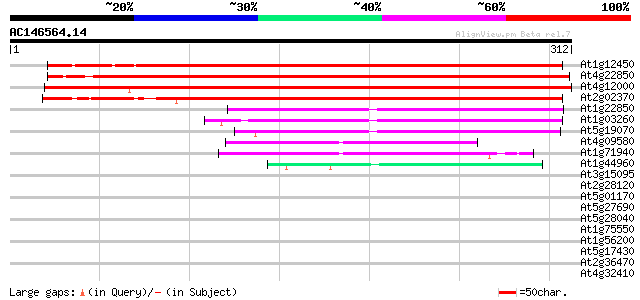
Sequences producing significant alignments: (bits) Value
At1g12450 hypothetical protein 358 2e-99
At4g22850 unknown protein 352 2e-97
At4g12000 unknown protein 327 5e-90
At2g02370 unknown protein 246 1e-65
At1g22850 unknown protein 71 7e-13
At1g03260 hypothetical protein 66 2e-11
At5g19070 unknown protein 65 4e-11
At4g09580 unknown protein 49 3e-06
At1g71940 unknown protein 47 2e-05
At1g44960 unknown protein 43 3e-04
At3g15095 unknown protein, 3' partial 32 0.34
At2g28120 nodulin-like protein 30 1.3
At5g01170 unknown protein 30 2.2
At5g27690 unknown protein (At5g27690) 29 3.8
At5g28040 unknown protein 28 4.9
At1g75550 hypothetical protein 28 4.9
At1g56200 unknown protein 28 4.9
At5g17430 ovule development protein aintegumenta-like protein 28 6.4
At2g36470 unknown protein 28 6.4
At4g32410 cellulose synthase catalytic subunit (RSW1) 28 8.4
>At1g12450 hypothetical protein
Length = 303
Score = 358 bits (919), Expect = 2e-99
Identities = 170/286 (59%), Positives = 229/286 (79%), Gaps = 3/286 (1%)
Query: 22 VSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMVEECSLPSRRSVVWYWVKMVLLFL 81
V ++ + +E D DN GDY+KL GS+E ++ E C + S SV W+WVK++ L +
Sbjct: 11 VPELKLRVEEDSDN-GDYLKLRGGSNEEDEGSSAES-SRCPIGSVTSV-WFWVKLISLVV 67
Query: 82 SLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSM 141
LG LA ++KWVGP+LI+KE+IP INW TFS PVL +LLFASVA+FP+ILLPS+PSM
Sbjct: 68 CLGSLAFVIIKWVGPFLIEKELIPFINWVRNTFSIPVLGLLLFASVALFPSILLPSSPSM 127
Query: 142 WVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNW 201
W+AG+T GYG GFLLI++AA+IGV+LPF+IG +F HK++ WL+KYPKKA+IL++AG G W
Sbjct: 128 WMAGLTFGYGKGFLLILSAASIGVTLPFLIGHLFLHKMQEWLKKYPKKAAILRAAGEGTW 187
Query: 202 FHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLA 261
FHQF+AV LIR+SPFPY+++NYCA+AT V YGPYI+GSLVGMVPEIFV+IYTGI+++TLA
Sbjct: 188 FHQFQAVTLIRVSPFPYIIYNYCALATGVHYGPYILGSLVGMVPEIFVSIYTGIMLRTLA 247
Query: 262 DASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEED 307
AS+ R +LS +I++N LGFC+T T++ T+YAK++L +Q ED
Sbjct: 248 VASDTRHTLSVVEIVVNVLGFCVTASATIVCTIYAKKKLSAMQSED 293
>At4g22850 unknown protein
Length = 296
Score = 352 bits (902), Expect = 2e-97
Identities = 173/291 (59%), Positives = 223/291 (76%), Gaps = 6/291 (2%)
Query: 22 VSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMVEECSLPSRRSVVWYWVKMVLLFL 81
V ++ + +E DDD KG Y+KL E L E+ + S W+WVK+ LLF
Sbjct: 10 VLELLVRVE-DDDEKGPYVKL----SEVLEKGQESEQEKEEEKTDSSRFWFWVKLSLLFA 64
Query: 82 SLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSM 141
L LAV W+GP ++DKE+IP+I WE TF+ PV +L+FASVAIFPTILLPSTPSM
Sbjct: 65 FLAALAVVGYIWIGPLIMDKELIPLIQWEIRTFTHPVCGLLVFASVAIFPTILLPSTPSM 124
Query: 142 WVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNW 201
W+AG+T GYG+GFLLII+AAA+GVSLP+ IG +F HKI+GWLE+YP +A++L++AG GNW
Sbjct: 125 WIAGMTFGYGYGFLLIISAAAVGVSLPYFIGQLFCHKIQGWLERYPDQAAVLRAAGEGNW 184
Query: 202 FHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLA 261
HQF V LIRISPFPY+++NYC+VAT VKYGPYI GSL+GMVPE+FVAIYTGIL++TLA
Sbjct: 185 LHQFLLVTLIRISPFPYILYNYCSVATRVKYGPYITGSLLGMVPEVFVAIYTGILVRTLA 244
Query: 262 DASN-ERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEEDKMLL 311
+AS+ E Q LS Q+ILN LGF TV TT++IT YAKR+L+ +++ED+ LL
Sbjct: 245 EASSAEEQGLSVTQVILNILGFLATVATTILITKYAKRQLETMKKEDEALL 295
>At4g12000 unknown protein
Length = 306
Score = 327 bits (838), Expect = 5e-90
Identities = 166/298 (55%), Positives = 218/298 (72%), Gaps = 5/298 (1%)
Query: 20 EVVSDVTITIEADD-DNKGDYIKLIPGSDECLPLTAVEMVEECSLPS---RRSVVWYWVK 75
+ VS+ + +E D D G Y+KL + T E S S ++ VW+W+K
Sbjct: 8 DTVSEFRVRVEEDGVDKLGHYVKLTEDFEVHRQETEQESSSSPSSSSSCGQKRSVWFWIK 67
Query: 76 MVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPTILL 135
+ L F L L +A KW+ P ++DKE+IP+I WE ETF+ PV IL+FASV++FP IL+
Sbjct: 68 LGLFFTFLTALGLAGYKWLYPLIMDKELIPLIKWEMETFTHPVCGILVFASVSLFPVILI 127
Query: 136 PSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKS 195
P+TPSMWVAG+T GY +G LL + A AIGVSLP+ I +F +KI+GWLE+YP +A++L++
Sbjct: 128 PTTPSMWVAGITFGYFYGLLLTLPAVAIGVSLPYFISYLFLNKIQGWLERYPDQAAMLRA 187
Query: 196 AGAGNWFHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGI 255
AG G+WFHQFRAV LIRISPFP+ V+NYCAVAT VK+GPY+ GSLVGM PEIFVAIYTGI
Sbjct: 188 AGGGSWFHQFRAVTLIRISPFPFAVYNYCAVATRVKFGPYMAGSLVGMAPEIFVAIYTGI 247
Query: 256 LIKTLADASN-ERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEEDKMLLL 312
LI+TLADAS E++ LS QI+LN GF TVVTTV+IT YAKR+L+ +++E + LLL
Sbjct: 248 LIRTLADASTAEQKGLSILQIVLNIFGFVATVVTTVLITKYAKRQLEVMKKEKEALLL 305
>At2g02370 unknown protein
Length = 320
Score = 246 bits (628), Expect = 1e-65
Identities = 121/295 (41%), Positives = 186/295 (63%), Gaps = 17/295 (5%)
Query: 19 KEVVSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMVEECSLPSRRSVVWYWVKMVL 78
KE D+ + DN +Y++L+ + E P V + + + S++ + +W+K
Sbjct: 6 KESREDIANSTPHMRDN--EYVRLVV-AHEASPAETVLSLSQSEVQSKKFM--WWLK--- 57
Query: 79 LFLSLGFLAVAVL------KWVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPT 132
+LG AVA+L KW P++ K +IPI+ WE F P+L I+L S+A+FP
Sbjct: 58 ---ALGICAVALLLTLVFGKWGVPFVFQKVLIPILQWEATAFGRPMLAIVLVVSLALFPV 114
Query: 133 ILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASI 192
L+PS PSMW+AG+ GYG GF++I+ IG+ LP++IG +F ++ WL+++P++A++
Sbjct: 115 FLIPSGPSMWLAGMIFGYGLGFVIIMVGTTIGMVLPYLIGLMFRDRLHQWLKRWPRQAAV 174
Query: 193 LKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIY 252
L+ A G+WFHQFR VA+ R+SPFPY +FNY V T++++ PY GS+ GM+PE F+ IY
Sbjct: 175 LRLAAEGSWFHQFRVVAIFRVSPFPYTIFNYAIVVTSMRFWPYFFGSIAGMIPEAFIYIY 234
Query: 253 TGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEED 307
+G LI+T AD Q L+ +I+ N + + VVTTV TVYAKR L+ELQ +
Sbjct: 235 SGRLIRTFADVQYGHQRLTTVEIVYNVISLVIAVVTTVAFTVYAKRALRELQNAE 289
>At1g22850 unknown protein
Length = 344
Score = 71.2 bits (173), Expect = 7e-13
Identities = 50/190 (26%), Positives = 92/190 (48%), Gaps = 7/190 (3%)
Query: 122 LLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIF-HHKIE 180
L A A + +P+ P AG+ G G +++ + + S+ F+I F +I
Sbjct: 155 LFIAVYAGLEILAIPALPLTMSAGLLFGPLIGTIIVSISGTMAASVAFLIARYFARERIL 214
Query: 181 GWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYIIGS 239
+E K +I K+ G FR V L+R+SP P+ + NY T+VK+ PY++GS
Sbjct: 215 KLVEDNKKFLAIDKAIGENG----FRVVTLLRLSPLLPFSLGNYLYGLTSVKFVPYVLGS 270
Query: 240 LVGMVPEIFVAIYTGILIKT-LADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKR 298
+GM+P + + G + + + SN Q++ +G +T + +T AK
Sbjct: 271 WLGMLPGSWAYVSAGAFGRAIIQEESNVGLPGGNGQLLTLGVGLLVTALAGTYVTSLAKD 330
Query: 299 RLKELQEEDK 308
+K++ +++K
Sbjct: 331 AIKDIDDDEK 340
>At1g03260 hypothetical protein
Length = 269
Score = 66.2 bits (160), Expect = 2e-11
Identities = 50/204 (24%), Positives = 94/204 (45%), Gaps = 12/204 (5%)
Query: 109 WETETFSP--PVLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVS 166
W E P P+ L + + I + +P++ G G GF+ A +G +
Sbjct: 36 WIKEDLGPFGPLALALAYIPLTI---VAVPASVLTLGGGYLFGLPVGFVADSLGATLGAT 92
Query: 167 LPFIIG-SIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYC 224
F++G +I + ++ YPK ++ + F+ V L+R+ P P+ + NY
Sbjct: 93 AAFLLGRTIGKSYVTSKIKHYPKFQAVSVAIQKSG----FKIVLLLRVVPILPFNMLNYL 148
Query: 225 AVATNVKYGPYIIGSLVGMV-PEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFC 283
T V+ G Y++ + +GM+ P F +Y G +K L+D ++ +S + ++ +G
Sbjct: 149 LSVTPVRLGEYMLATWLGMMQPITFALVYVGTTLKDLSDITHGWHEVSVFRWVIMMVGVA 208
Query: 284 LTVVTTVIITVYAKRRLKELQEED 307
L V+ + IT AK L + E+
Sbjct: 209 LAVILIICITRVAKSSLDKALAEN 232
>At5g19070 unknown protein
Length = 280
Score = 65.5 bits (158), Expect = 4e-11
Identities = 46/185 (24%), Positives = 85/185 (45%), Gaps = 8/185 (4%)
Query: 126 SVAIFPTILL--PSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIG-SIFHHKIEGW 182
+VA P +L P++ G G GF+ A +G F++G +I +
Sbjct: 54 AVAYIPLTVLAVPASVLTLGGGYLFGLPIGFVADSVGATLGSGAAFLLGRTIGKPFVVAK 113
Query: 183 LEKYPKKASILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYIIGSLV 241
L+ YP+ S+ + F+ L+R++P P+ + NY T ++ GPY++ S +
Sbjct: 114 LKDYPQFQSVALAIEKSG----FKICLLLRLAPLLPFSMLNYLLSVTPIRLGPYLLSSWL 169
Query: 242 GMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLK 301
GM+P +Y G +K L+D +++ S + ++V+ V +T AK L+
Sbjct: 170 GMMPITLALVYVGTTLKDLSDVTHKWSEFSPGRWAFLISSLVISVILMVCVTKVAKDALR 229
Query: 302 ELQEE 306
+ E
Sbjct: 230 KALAE 234
>At4g09580 unknown protein
Length = 287
Score = 49.3 bits (116), Expect = 3e-06
Identities = 34/143 (23%), Positives = 66/143 (45%), Gaps = 5/143 (3%)
Query: 121 ILLFASVAIF-PTILLPSTPSM-WVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHK 178
IL + S IF T ++P T M +AG G GF+L++ A G F + +
Sbjct: 112 ILGYCSTYIFMQTFMIPGTIFMSLLAGALFGVVRGFVLVVLNATAGACSCFFLSKLVGRP 171
Query: 179 IEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYGPYII 237
+ WL +P+K ++ A + +RI+P P + N + ++ + + +
Sbjct: 172 LVNWL--WPEKLRFFQAEIAKRRDRLLNYMLFLRITPTLPNLFINLSSPIVDIPFHVFFL 229
Query: 238 GSLVGMVPEIFVAIYTGILIKTL 260
+LVG++P ++ + G+ + L
Sbjct: 230 ATLVGLMPASYITVRAGLALGDL 252
>At1g71940 unknown protein
Length = 272
Score = 46.6 bits (109), Expect = 2e-05
Identities = 38/180 (21%), Positives = 84/180 (46%), Gaps = 12/180 (6%)
Query: 117 PVLTILLFASVAIF-PTILLPSTPSM-WVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSI 174
P +L + + IF T ++P T M +AG G G +L++ A G + F + +
Sbjct: 93 PAQFVLGYCATYIFMQTFMIPGTIFMSLLAGALFGVFKGVVLVVFNATAGATSCFFLSKL 152
Query: 175 FHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYG 233
+ WL +P K ++ + + +RI+P P + N + +V +
Sbjct: 153 IGRPLITWL--WPDKLRFFQAEISKRRDKLLNYMLFLRITPTLPNLFINLASPIVDVPFH 210
Query: 234 PYIIGSLVGMVPEIFVAIYTGILIKTLADASN--ERQSLSAQQIILNALGFCLTVVTTVI 291
+ + +L+G++P ++ + G+ I L + + ++LS +L +GF ++++ T++
Sbjct: 211 VFFLATLIGLIPAAYITVRAGLAIGDLKSVKDLYDFKTLS----VLFLIGF-ISILPTIL 265
>At1g44960 unknown protein
Length = 261
Score = 42.7 bits (99), Expect = 3e-04
Identities = 36/159 (22%), Positives = 62/159 (38%), Gaps = 10/159 (6%)
Query: 144 AGVTLGYGF--GFLLIITAAAIGVSLPFIIGSIFHH---KIEGWLEKYPKKASILKSAGA 198
AG ++ +GF L + +A + S F IG + GW + +
Sbjct: 71 AGASMLFGFLPALLCVFSAKVLAASFSFWIGRFVFKSSTRATGWAHSNKYFNILSRGVER 130
Query: 199 GNWFHQFRAVALIRISPFPYMVFNYCAVATNVKY-GPYIIGSLVGMVPEIFVAIYTGILI 257
W + V L R SP P V NY AT V++ ++ +++G +P I G L
Sbjct: 131 DGW----KFVLLARFSPIPSYVINYALAATEVRFVADFLFPTVIGCLPMILQNASVGSLA 186
Query: 258 KTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYA 296
+ +Q + LG +V+ ++ I Y+
Sbjct: 187 GMAVASVAGKQKSQVWGYVFPVLGILSSVLISLRIKKYS 225
>At3g15095 unknown protein, 3' partial
Length = 417
Score = 32.3 bits (72), Expect = 0.34
Identities = 15/47 (31%), Positives = 23/47 (48%)
Query: 2 KDNTDDCGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDE 48
+ N D CGGGGGG V + + +E K I+L+ G ++
Sbjct: 208 ESNNDGCGGGGGGSNSCGAVFTRWFVAVEETSGGKRREIELVVGGED 254
>At2g28120 nodulin-like protein
Length = 577
Score = 30.4 bits (67), Expect = 1.3
Identities = 20/71 (28%), Positives = 34/71 (47%), Gaps = 9/71 (12%)
Query: 93 WVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGF 152
+V YL+ K +P P ++T++L S A I P S+++A + +G+ F
Sbjct: 391 FVSEYLLAKYKLP---------RPLMMTLVLLLSCAGHLLIAFPVPGSVYIASILMGFSF 441
Query: 153 GFLLIITAAAI 163
G L + A I
Sbjct: 442 GAQLPLLFAII 452
>At5g01170 unknown protein
Length = 568
Score = 29.6 bits (65), Expect = 2.2
Identities = 10/11 (90%), Positives = 11/11 (99%)
Query: 9 GGGGGGGGWKK 19
GGGGGGGGW+K
Sbjct: 538 GGGGGGGGWEK 548
>At5g27690 unknown protein (At5g27690)
Length = 352
Score = 28.9 bits (63), Expect = 3.8
Identities = 13/32 (40%), Positives = 16/32 (49%)
Query: 7 DCGGGGGGGGWKKEVVSDVTITIEADDDNKGD 38
D GGGGGGGG + V V + +GD
Sbjct: 137 DVGGGGGGGGGNFDQVKQVVTFVNGQLQPQGD 168
>At5g28040 unknown protein
Length = 427
Score = 28.5 bits (62), Expect = 4.9
Identities = 18/36 (50%), Positives = 22/36 (61%), Gaps = 3/36 (8%)
Query: 3 DNTDDCGGGGGG-GGWKKEVVSDVTI--TIEADDDN 35
D +D GGGGGG GG + E DV I EA+DD+
Sbjct: 16 DLEEDGGGGGGGRGGGETESDEDVVIPEPNEAEDDD 51
>At1g75550 hypothetical protein
Length = 167
Score = 28.5 bits (62), Expect = 4.9
Identities = 10/11 (90%), Positives = 10/11 (90%)
Query: 9 GGGGGGGGWKK 19
GGGGGGGGW K
Sbjct: 96 GGGGGGGGWYK 106
>At1g56200 unknown protein
Length = 154
Score = 28.5 bits (62), Expect = 4.9
Identities = 10/16 (62%), Positives = 12/16 (74%)
Query: 2 KDNTDDCGGGGGGGGW 17
+D+ D GGGGGGG W
Sbjct: 108 RDDGDGGGGGGGGGNW 123
>At5g17430 ovule development protein aintegumenta-like protein
Length = 581
Score = 28.1 bits (61), Expect = 6.4
Identities = 10/14 (71%), Positives = 10/14 (71%)
Query: 3 DNTDDCGGGGGGGG 16
D DCGGG GGGG
Sbjct: 118 DGNGDCGGGDGGGG 131
>At2g36470 unknown protein
Length = 327
Score = 28.1 bits (61), Expect = 6.4
Identities = 15/30 (50%), Positives = 18/30 (60%), Gaps = 7/30 (23%)
Query: 8 CGGGGGGGG-----WKKEVVSDVTITIEAD 32
CGGGGGGGG WK V S T+++ D
Sbjct: 183 CGGGGGGGGEEGYLWK--VKSPETMSVYVD 210
>At4g32410 cellulose synthase catalytic subunit (RSW1)
Length = 1081
Score = 27.7 bits (60), Expect = 8.4
Identities = 15/64 (23%), Positives = 29/64 (44%), Gaps = 18/64 (28%)
Query: 51 PLTAVEMVEECSLP--------------SRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGP 96
P+T++ ++ C LP S + +W+ +LLF+S+ + L+W G
Sbjct: 860 PITSIPLIAYCILPAFCLITDRFIIPEISNYASIWF----ILLFISIAVTGILELRWSGV 915
Query: 97 YLID 100
+ D
Sbjct: 916 SIED 919
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,328,687
Number of Sequences: 26719
Number of extensions: 318269
Number of successful extensions: 2321
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2246
Number of HSP's gapped (non-prelim): 54
length of query: 312
length of database: 11,318,596
effective HSP length: 99
effective length of query: 213
effective length of database: 8,673,415
effective search space: 1847437395
effective search space used: 1847437395
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146564.14


