

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.6 + phase: 0 /pseudo
(799 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
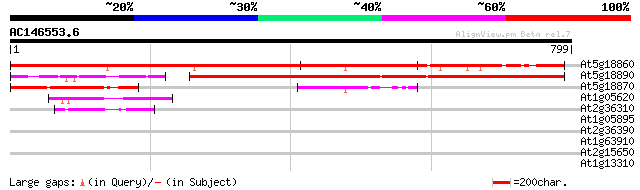
At5g18860 unknown protein 866 0.0
At5g18890 putative protein 503 e-142
At5g18870 putative protein 235 6e-62
At1g05620 unknown protein 58 2e-08
At2g36310 unknown protein 52 1e-06
At1g05895 putative protein 33 0.48
At2g36390 starch branching enzyme class II (sbe2-1) 32 1.8
At1g63910 putative MYB family transcription factor 31 3.1
At2g15650 putative retroelement pol polyprotein 30 7.0
At1g13310 hypothetical protein 29 9.1
>At5g18860 unknown protein
Length = 890
Score = 866 bits (2237), Expect = 0.0
Identities = 458/821 (55%), Positives = 576/821 (69%), Gaps = 50/821 (6%)
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLDI 60
MM RDDI VGVGGEGGI +GTI +VGGY PIIEQGMTT G CRYRQAIP G GG LDI
Sbjct: 87 MMDRDDIPVGVGGEGGISDDGTIHSDVGGYFPIIEQGMTTTGECRYRQAIPKGLGGLLDI 146
Query: 61 DTNLGIRKAFLPQGKRKYTPLRQPTTQQVLIDKISAGPITLIVTGAHTNLAIFLMNNPHL 120
D+N G RK FLPQG R+YTPL+QPT Q+V++DKIS GP T+I+ G+HTN A+FLM+NPHL
Sbjct: 147 DSNYGFRKQFLPQGNRRYTPLQQPTAQKVIVDKISEGPTTVILLGSHTNFALFLMSNPHL 206
Query: 121 KKNVEHVYIMGGVIRSKT---CCTKNAS-SSCIPSKCGDTGNVLTNYNANPYAEYNIFGD 176
K N++H+YIMGG +RS+ CC N++ + C P +CG+ GN+ T+Y +NPY+E+NIF D
Sbjct: 207 KHNIQHIYIMGGGVRSQNPTGCCPANSTVAECQPRQCGNRGNLFTDYTSNPYSEFNIFAD 266
Query: 177 PFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKS-QDTYEAQYCFKSLKMAHDTWF 235
PFAAY+V HSG+P+TLVPLDATNTIPI+++FF+ FE + Q TYEAQY F SLK+A DTWF
Sbjct: 267 PFAAYQVFHSGVPVTLVPLDATNTIPINQKFFETFENNYQRTYEAQYVFLSLKIARDTWF 326
Query: 236 DNQFYTSYFMWDSFTSGVAVSIMRNS---NRKKGENEFAEMEYINITVITSNKPYGISDG 292
D++FY SYFMWDSFT+GVAVSIMRNS N K GEN+FAEMEY+NITV+TSNKPYG SDG
Sbjct: 327 DDEFYKSYFMWDSFTAGVAVSIMRNSANKNNKNGENDFAEMEYMNITVVTSNKPYGRSDG 386
Query: 293 SNPLFDGLKVPKFNLKKGGVHSGHIQQGLTDPFCFVKNG--KGRCQDGYTKEVNGQESVK 350
SNP FD + PKFNL GGVHSGH+Q GL DP C K+G +G+C+DGYT+E++G +SV+
Sbjct: 387 SNPFFDNRRTPKFNLALGGVHSGHVQTGLRDPTCLPKSGIGRGKCKDGYTQEISGSDSVR 446
Query: 351 VLVATKAKPNKDIRSSLDREYFKSFLNVLKQPQQAGKFNFTTQFPYYEEVTHKPDFWNKI 410
VLVAT+AKPN +I+S LDRE++ FL VL +P++ G+FNF++QFPYY+E +PD
Sbjct: 447 VLVATRAKPNINIKSKLDREFYVDFLEVLNRPEETGRFNFSSQFPYYKEELFRPDLSKTR 506
Query: 411 LGKPVVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANAATIDVIYDLLHMMGR 470
GKPVVFDMDMSAGDFL+L YLLKVPV+ I+LKAIIVSPTGWANAATIDV+YDLLHMMGR
Sbjct: 507 PGKPVVFDMDMSAGDFLSLFYLLKVPVDKIDLKAIIVSPTGWANAATIDVVYDLLHMMGR 566
Query: 471 DDIKVGIGDFFAMNQSN--FSPVGDCKYVKAIPHGNGGFLDSDTLFGLARDLPRSPRRYT 528
DDI VG+GD A+NQS+ F PVG CKYVKAIP G GGFLDSDTL+GLARDLPRSPRRYT
Sbjct: 567 DDIPVGLGDMLALNQSDPIFPPVGGCKYVKAIPRGCGGFLDSDTLYGLARDLPRSPRRYT 626
Query: 529 AENSKKFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVTVLTNGPLTNLAKVVSI-KN 587
AENS GAPRDTD PELRQP A+E+W+++ ++ SK+TVLTNGPLTNLAK++S K
Sbjct: 627 AENSVTHGAPRDTDRPELRQPLAIEVWQNLTKSGNGVSKITVLTNGPLTNLAKIISSDKK 686
Query: 588 ISSIIQA*QFQNVSGGLCNGRTYQQ----E*Q*QRKCIFCSFQQVCRIQYVLGSFSSKDS 643
SS+I+ + GG N + F F + VL S +
Sbjct: 687 SSSLIKE---VYIVGGHINREKSDKGNIFTIPSNAYAEFNMFLDPLAAKTVLESALNITL 743
Query: 644 VSIRS*H----------HTHSTRYPAQSKFIFKYF----KLA*SDRKNTRSSVFQASSVK 689
V + + H +S+ +++F+ + L R+ T +F +
Sbjct: 744 VPLATQHKLSSFQTMLDRLYSSTKTPEARFVKRLLVRLQALHQKHRRYTHIDMFLGEVLG 803
Query: 690 ATPFEENPPQISPYGAVVLANGHSSLLDAKFELKSVKLLAEGIESTDGKMVVDEKYGKLV 749
A + + P + H ++ E + K+L I+ GK +
Sbjct: 804 AVLLGGDDASLKP----KMRAEHIKVIAEGDESRDGKIL---IDKLRGKQI--------- 847
Query: 750 RILRHVDAKTYHEIYAKRLGDPNQSAKVGSFKEQKRKWSHP 790
+IL VD + E +A RL D QSA +GSF+EQK+ WS P
Sbjct: 848 KILERVDLISISESFASRLDDKKQSAVIGSFEEQKKIWSTP 888
Score = 89.0 bits (219), Expect = 1e-17
Identities = 60/184 (32%), Positives = 88/184 (47%), Gaps = 38/184 (20%)
Query: 415 VVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANAA-TIDVIYDLLHMMGRDDI 473
++ D D+ D A+ YLLK+ +L I +S W NA ++ +YDLLHMM RDDI
Sbjct: 34 ILVDTDVDTDDLFAILYLLKLNKSEFDLVGITLSANAWTNAGHAVNQVYDLLHMMDRDDI 93
Query: 474 KVG----------------IGDFFAMNQSNFSPVGDCKYVKAIPHGNGGFLDSDTLFGLA 517
VG +G +F + + + G+C+Y +AIP G GG LD D+ +G
Sbjct: 94 PVGVGGEGGISDDGTIHSDVGGYFPIIEQGMTTTGECRYRQAIPKGLGGLLDIDSNYGFR 153
Query: 518 RD-LPRSPRRYTAENSKKFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVTVLTNGPL 576
+ LP+ RRYT L+QP A ++ I+ + G TV+ G
Sbjct: 154 KQFLPQGNRRYT----------------PLQQPTAQKV---IVDKISEG-PTTVILLGSH 193
Query: 577 TNLA 580
TN A
Sbjct: 194 TNFA 197
>At5g18890 putative protein
Length = 550
Score = 503 bits (1294), Expect = e-142
Identities = 268/539 (49%), Positives = 353/539 (64%), Gaps = 8/539 (1%)
Query: 256 SIMRNSNRKKGENEFAEMEYINITVITSNKPYGISDGSNPLFDGLKVPKFNLKKGGVHSG 315
S N N KG+N+FAEMEY+NITV+TSN+PYG+ D SNP F + PKFNL GGVHSG
Sbjct: 4 SASSNKNNNKGQNDFAEMEYMNITVVTSNEPYGLFDSSNPFFYKRRTPKFNLTLGGVHSG 63
Query: 316 HIQQGLTDPFCFVKNGKGRCQDGYTKEVNGQESVKVLVATKAKPNKDIRSSLDREYFKSF 375
H+Q+GL DP C +GKG C+DGYTKE +G +SV+VLVAT+AKP+K++ S LDRE++ F
Sbjct: 64 HVQRGLRDPICISTSGKGNCRDGYTKETSGPDSVRVLVATRAKPSKNLNSELDREFYDHF 123
Query: 376 LNVLKQPQQAGKFNFTTQFPYYEEVTHKPDFWNKIL-GKPVVFDMDMSAGDFLALSYLLK 434
L VL +P++ G+F+F+TQF YY E + N L GKPVVFDMDMSAGDFL+L YLLK
Sbjct: 124 LEVLNRPEETGRFHFSTQFLYYREELFIAELNNSRLGGKPVVFDMDMSAGDFLSLFYLLK 183
Query: 435 VPVEVINLKAIIVSPTGWANAATIDVIYDLLHMMGRDDIKVGIGDFFAMNQSN--FSPVG 492
VPVE+I+LKA+IVSPTGWAN ATIDV+YDLLHMMGRDDI VG+GD FA+NQS F G
Sbjct: 184 VPVEIIDLKAVIVSPTGWANTATIDVVYDLLHMMGRDDIPVGLGDMFAINQSEPVFPSAG 243
Query: 493 DCKYVKAIPHGNGGFLDSDTLFGLARDLPRSPRRYTAENSKKFGAPRDTDHPELRQPQAM 552
DCKY KA+P G GGFLDSDTL+GLARDLPRSPRRY ENS GAP DTD PELRQP A+
Sbjct: 244 DCKYAKAVPQGCGGFLDSDTLYGLARDLPRSPRRY--ENSVAHGAPSDTDRPELRQPLAL 301
Query: 553 EIWESILQTLKPGSKVTVLTNGPLTNLAKVVSI-KNISSIIQA*QFQNVSGGLCNGRTYQ 611
E+W+++ +++ SK+TVLTNGPLT+LAK++S KN SSII+ + V G + G++ +
Sbjct: 302 EVWQNLTKSVDEVSKITVLTNGPLTSLAKIISSDKNSSSIIK--EVYIVGGHISRGKSDK 359
Query: 612 QE*Q*QRKCIFCSFQQVCRIQYVLGSFSSKDSVSIRS*HHTHSTRYPAQSKFIFKYFKLA 671
+ F S ++++ + A ++ K
Sbjct: 360 GNIFTVPSNSYAEFNMFLDPLAAKTVLESGLNITLIPLATQREFSFQAMLNRLYSSTKTP 419
Query: 672 *SDRKNTRSSVFQASSVKATPFEENPPQISPYGAVVLANGHSSLLDAKFELKSVKLLAEG 731
+ + QA K + + + G +LL K + +K++AEG
Sbjct: 420 EARFVKRLLTRLQALHQKQRRYMHMDMFLGEILGAIFLGGDHALLKPKMRTEYIKVIAEG 479
Query: 732 IESTDGKMVVDEKYGKLVRILRHVDAKTYHEIYAKRLGDPNQSAKVGSFKEQKRKWSHP 790
ES DG +++D+ GK ++IL VD + +E +A RL D QSA +GSF+EQ+ KW+ P
Sbjct: 480 DESKDGHILIDKLRGKQIKILERVDLRGCYESFASRLDDKKQSAVIGSFEEQRMKWNTP 538
Score = 94.0 bits (232), Expect = 3e-19
Identities = 79/243 (32%), Positives = 116/243 (47%), Gaps = 56/243 (23%)
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLDI 60
MMGRDDI VG+G I + + P+ AG C+Y +A+P G GG LD
Sbjct: 216 MMGRDDIPVGLGDMFAINQSEPVFPS--------------AGDCKYAKAVPQGCGGFLDS 261
Query: 61 DTNLGIRKAFLPQGKRKYT--------------PLRQPTTQQVL------IDKISAGPIT 100
DT G+ + LP+ R+Y LRQP +V +D++S IT
Sbjct: 262 DTLYGLARD-LPRSPRRYENSVAHGAPSDTDRPELRQPLALEVWQNLTKSVDEVSK--IT 318
Query: 101 LIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVI-RSKTCCTKNASSSCIPSKCGDTGNV 159
++ G T+LA + ++ + ++ VYI+GG I R K+ D GN+
Sbjct: 319 VLTNGPLTSLAKIISSDKNSSSIIKEVYIVGGHISRGKS----------------DKGNI 362
Query: 160 LTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKSQDTYE 219
T +N YAE+N+F DP AA V+ SG+ ITL+PL AT + + S T E
Sbjct: 363 FT-VPSNSYAEFNMFLDPLAAKTVLESGLNITLIPL-ATQREFSFQAMLNRLYSSTKTPE 420
Query: 220 AQY 222
A++
Sbjct: 421 ARF 423
>At5g18870 putative protein
Length = 258
Score = 235 bits (600), Expect = 6e-62
Identities = 116/184 (63%), Positives = 142/184 (77%), Gaps = 17/184 (9%)
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLD- 59
MMGRDDI VGVGGEGGIL +GTILP+VG YLPIIEQGMTTAG CRYRQ+IP G R+
Sbjct: 84 MMGRDDITVGVGGEGGILEDGTILPDVGDYLPIIEQGMTTAGGCRYRQSIPKG---RIQK 140
Query: 60 IDTNLGIRKAFLPQGKRKYTPLRQPTTQQVLIDKISAGPITLIVTGAHTNLAIFLMNNPH 119
ID+N G RK FLPQG R+YTPL QPT Q+V++DK+S GPI++ V G+HTNLA+F+M+NPH
Sbjct: 141 IDSNYGFRKHFLPQGNRRYTPLEQPTAQKVIVDKVSEGPISIFVIGSHTNLALFMMSNPH 200
Query: 120 LKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAEYNIFGDPFA 179
LK N++H+Y+MGG +R C N G GN+ T+Y +NPYAE+NIF DPFA
Sbjct: 201 LKHNIQHIYVMGGSVR---CQNPN----------GFCGNLFTDYTSNPYAEFNIFTDPFA 247
Query: 180 AYKV 183
AY+V
Sbjct: 248 AYQV 251
Score = 77.8 bits (190), Expect = 2e-14
Identities = 59/191 (30%), Positives = 91/191 (46%), Gaps = 42/191 (21%)
Query: 410 ILGKP--VVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANAAT-IDVIYDLLH 466
+L P ++ D D+ DF+AL YLLK+ +L I +S W NA ++ IYD+L+
Sbjct: 24 VLNSPHRILLDTDVDTDDFIALLYLLKLNKTEFDLVGITLSANSWTNAGHGVNHIYDILY 83
Query: 467 MMGRDDIKVG----------------IGDFFAMNQSNFSPVGDCKYVKAIPHGNGGFLDS 510
MMGRDDI VG +GD+ + + + G C+Y ++IP G +DS
Sbjct: 84 MMGRDDITVGVGGEGGILEDGTILPDVGDYLPIIEQGMTTAGGCRYRQSIPKGRIQKIDS 143
Query: 511 DTLFGLARD-LPRSPRRYTAENSKKFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVT 569
+ +G + LP+ RRYT L QP A ++ I+ + G ++
Sbjct: 144 N--YGFRKHFLPQGNRRYT----------------PLEQPTAQKV---IVDKVSEG-PIS 181
Query: 570 VLTNGPLTNLA 580
+ G TNLA
Sbjct: 182 IFVIGSHTNLA 192
>At1g05620 unknown protein
Length = 322
Score = 58.2 bits (139), Expect = 2e-08
Identities = 52/198 (26%), Positives = 81/198 (40%), Gaps = 47/198 (23%)
Query: 56 GRLDIDTNLGIRKAFLPQ---------------GKRKYTPLR----QPTTQQVLID--KI 94
GR DI G K FL G + + P + + + + L++ K+
Sbjct: 61 GRTDIPVAEGTHKTFLNDTKLRIADFVHGKDGLGNQNFPPPKGKPIEKSGPEFLVEQAKL 120
Query: 95 SAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCG 154
G IT++ G TNLA+ + +P KNV + ++GG
Sbjct: 121 CPGEITVVALGPLTNLALAVQLDPEFSKNVGQIVLLGGAFA------------------- 161
Query: 155 DTGNVLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKS 214
N N NP +E NIFGDP AA V G I V ++ T+ + ++ + D+ S
Sbjct: 162 ------VNGNVNPASEANIFGDPEAADIVFTCGADIIAVGINVTHQVIMTADDKDKLASS 215
Query: 215 QDTYEAQYCFKSLKMAHD 232
+ AQY K L + +D
Sbjct: 216 KGKL-AQYLCKILDVYYD 232
>At2g36310 unknown protein
Length = 336
Score = 52.0 bits (123), Expect = 1e-06
Identities = 41/144 (28%), Positives = 65/144 (44%), Gaps = 30/144 (20%)
Query: 65 GIRKAFLPQGKRKYTPLRQPTTQQVLIDKISA--GPITLIVTGAHTNLAIFLMNNPHLKK 122
G+ LP RK + + + + L +K+ G +T++ G TNLA+ + +
Sbjct: 106 GLGDVSLPPPSRKKS---EKSAAEFLDEKVEEYPGEVTILALGPLTNLALAIKRDSSFAS 162
Query: 123 NVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAEYNIFGDPFAAYK 182
V+ + I+GG S GNV NP AE NI+GDP AA
Sbjct: 163 KVKKIVILGGAFFS-------------------LGNV------NPAAEANIYGDPEAADV 197
Query: 183 VIHSGIPITLVPLDATNTIPISEE 206
V SG IT+V ++ T + +S++
Sbjct: 198 VFTSGADITVVGINITTQLKLSDD 221
>At1g05895 putative protein
Length = 457
Score = 33.5 bits (75), Expect = 0.48
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 9/69 (13%)
Query: 523 SPRRYTAENSK-KFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVTVLTNGPLTNLAK 581
SPRR N+K G+ + +L Q + ++ LK GSK ++ NGPLTN AK
Sbjct: 286 SPRRKKLLNAKVSAGSSPRSMRAQLLQARKLD--------LKDGSKRKLVFNGPLTNAAK 337
Query: 582 VVSIKNISS 590
+ +K +S
Sbjct: 338 IPELKRNNS 346
>At2g36390 starch branching enzyme class II (sbe2-1)
Length = 858
Score = 31.6 bits (70), Expect = 1.8
Identities = 28/117 (23%), Positives = 47/117 (39%), Gaps = 20/117 (17%)
Query: 467 MMGRDDIKVGIGDFFAMNQSNFSPVGDCKYVKAIPHGNGGFLDSDTLFGLARDLPR---- 522
+M R+D G+ + F N ++ SP AIPHG+ + DT G+ +P
Sbjct: 241 VMARNDF--GVWEIFLPNNADGSP--------AIPHGSRVKIRMDTPSGIKDSIPAWIKY 290
Query: 523 --SPRRYTAENSKKFGAPRDT----DHPELRQPQAMEIWESILQTLKPGSKVTVLTN 573
P N + P + HP ++P ++ I+ES + K+ N
Sbjct: 291 SVQPPGEIPYNGVYYDPPEEDKYAFKHPRPKKPTSLRIYESHVGMSSTEPKINTYAN 347
>At1g63910 putative MYB family transcription factor
Length = 370
Score = 30.8 bits (68), Expect = 3.1
Identities = 10/31 (32%), Positives = 19/31 (61%)
Query: 223 CFKSLKMAHDTWFDNQFYTSYFMWDSFTSGV 253
C K ++++D W D+Q +++F WD+ V
Sbjct: 312 CLKRQELSYDQWDDSQQCSNFFFWDNLNINV 342
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 29.6 bits (65), Expect = 7.0
Identities = 24/96 (25%), Positives = 43/96 (44%), Gaps = 7/96 (7%)
Query: 212 EKSQDTYEAQYCFKSLKMAHDTWFDNQFYTSYFMWDSFTSGVAVSIMRNSNRKKGENEFA 271
EK Y+A Y LK A W++ SYF+ + F + + + + +KKGE+
Sbjct: 974 EKVLRLYKALY---GLKQAPRAWYER--IDSYFIQNGFARSMNDAALYS--KKKGEDVLI 1026
Query: 272 EMEYINITVITSNKPYGISDGSNPLFDGLKVPKFNL 307
Y++ +IT N + I+ + D ++ L
Sbjct: 1027 VSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGL 1062
>At1g13310 hypothetical protein
Length = 323
Score = 29.3 bits (64), Expect = 9.1
Identities = 28/94 (29%), Positives = 42/94 (43%), Gaps = 17/94 (18%)
Query: 694 EENPPQISPYGAVVLANGHS--SLLDAKFELKSVKLLAEGIESTDGKMVVDEKYGKLVRI 751
E PP S + LA G + + A ELKS K LAE +ES D + G V +
Sbjct: 189 ELTPPLCSALSLLCLAEGQEYVTTMGALHELKSQKYLAELLESED-------RVGDAVGV 241
Query: 752 LRHVDAKTYHEIYAKRLGDPNQSAK-VGSFKEQK 784
LR + A + P++ K + FK+++
Sbjct: 242 LRRA-------LAAAKKSTPSKDDKWIAIFKKER 268
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,405,831
Number of Sequences: 26719
Number of extensions: 827994
Number of successful extensions: 1688
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1648
Number of HSP's gapped (non-prelim): 19
length of query: 799
length of database: 11,318,596
effective HSP length: 107
effective length of query: 692
effective length of database: 8,459,663
effective search space: 5854086796
effective search space used: 5854086796
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146553.6


