

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.10 - phase: 0
(301 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
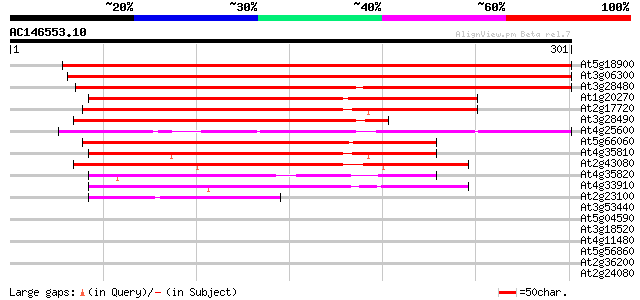
Sequences producing significant alignments: (bits) Value
At5g18900 unknown protein 454 e-128
At3g06300 unknown protein 435 e-122
At3g28480 prolyl 4-hydroxylase like protein 352 2e-97
At1g20270 unknown protein 240 6e-64
At2g17720 similar to prolyl 4-hydroxylase alpha subunit 237 5e-63
At3g28490 prolyl 4-hydroxylase, putative 216 9e-57
At4g25600 unknown protein 203 1e-52
At5g66060 prolyl 4-hydroxylase, alpha subunit-like protein 198 3e-51
At4g35810 putative protein 190 7e-49
At2g43080 unknown protein 184 5e-47
At4g35820 putative protein 159 2e-39
At4g33910 unknown protein 153 1e-37
At2g23100 hypothetical protein 72 5e-13
At3g53440 unknown protein 30 1.6
At5g04590 sulphite reductase 30 2.1
At3g18520 histone deacetylase, putative 29 2.7
At4g11480 serine/threonine kinase-like protein 29 3.6
At5g56860 unknown protein 28 6.1
At2g36200 putative kinesin-related cytokinesis protein 28 6.1
At2g24080 unknown protein 28 6.1
>At5g18900 unknown protein
Length = 298
Score = 454 bits (1167), Expect = e-128
Identities = 212/273 (77%), Positives = 240/273 (87%)
Query: 29 AGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLS 88
+ +S ++P+KVKQVS KPRAFVY+GFLT+LECDH++S+AK+ LKRSAVADN SGESK S
Sbjct: 26 SSSSVFVNPSKVKQVSSKPRAFVYEGFLTELECDHMVSLAKASLKRSAVADNDSGESKFS 85
Query: 89 EVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADK 148
EVRTSSG FISK KD IVSGIEDKIS+WTFLPKENGEDIQVLRYEHGQKYD H+DYF DK
Sbjct: 86 EVRTSSGTFISKGKDPIVSGIEDKISTWTFLPKENGEDIQVLRYEHGQKYDAHFDYFHDK 145
Query: 149 VNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRR 208
VNI RGGHR+AT+LMYL+NVTKGGETVFP+AE R LSE EDLS+C K+G+AV PR+
Sbjct: 146 VNIVRGGHRMATILMYLSNVTKGGETVFPDAEIPSRRVLSENKEDLSDCAKRGIAVKPRK 205
Query: 209 GDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESC 268
GDALLF +LHP+AIPD LSLH GCPVIEGEKWSATKWIHVDSFD+ V G+CTD +ESC
Sbjct: 206 GDALLFFNLHPDAIPDPLSLHGGCPVIEGEKWSATKWIHVDSFDRIVTPSGNCTDMNESC 265
Query: 269 ERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
ERWA LGECTKNPEYMVGT+ LPGYCR+SCK C
Sbjct: 266 ERWAVLGECTKNPEYMVGTTELPGYCRRSCKAC 298
>At3g06300 unknown protein
Length = 299
Score = 435 bits (1118), Expect = e-122
Identities = 204/270 (75%), Positives = 236/270 (86%)
Query: 32 SAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVR 91
S+II+P+KVKQVS KPRAFVY+GFLTDLECDHLIS+AK L+RSAVADN +GES++S+VR
Sbjct: 30 SSIINPSKVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVR 89
Query: 92 TSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNI 151
TSSG FISK KD IVSGIEDK+S+WTFLPKENGED+QVLRYEHGQKYD H+DYF DKVNI
Sbjct: 90 TSSGTFISKGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNI 149
Query: 152 ARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDA 211
ARGGHR+ATVL+YL+NVTKGGETVFP+A+E R LSE +DLS+C KKG+AV P++G+A
Sbjct: 150 ARGGHRIATVLLYLSNVTKGGETVFPDAQEFSRRSLSENKDDLSDCAKKGIAVKPKKGNA 209
Query: 212 LLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERW 271
LLF +L +AIPD SLH GCPVIEGEKWSATKWIHVDSFDK + G+CTD +ESCERW
Sbjct: 210 LLFFNLQQDAIPDPFSLHGGCPVIEGEKWSATKWIHVDSFDKILTHDGNCTDVNESCERW 269
Query: 272 AALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
A LGEC KNPEYMVGT +PG CR+SCK C
Sbjct: 270 AVLGECGKNPEYMVGTPEIPGNCRRSCKAC 299
>At3g28480 prolyl 4-hydroxylase like protein
Length = 316
Score = 352 bits (902), Expect = 2e-97
Identities = 162/266 (60%), Positives = 202/266 (75%), Gaps = 3/266 (1%)
Query: 36 DPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSG 95
DPT+V Q+SW PR F+Y+GFL+D ECDH I +AK +L++S VADN SGES SEVRTSSG
Sbjct: 52 DPTRVTQLSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSG 111
Query: 96 MFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGG 155
MF+SK +D IVS +E K+++WTFLP+ENGE +Q+L YE+GQKY+PH+DYF D+ N+ GG
Sbjct: 112 MFLSKRQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGG 171
Query: 156 HRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFC 215
HR+ATVLMYL+NV KGGETVFP + D+ +EC K+G AV PR+GDALLF
Sbjct: 172 HRIATVLMYLSNVEKGGETVFPMWKGKATQL---KDDSWTECAKQGYAVKPRKGDALLFF 228
Query: 216 SLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALG 275
+LHPNA D+ SLH CPV+EGEKWSAT+WIHV SF++ C D++ SCE+WA G
Sbjct: 229 NLHPNATTDSNSLHGSCPVVEGEKWSATRWIHVKSFERAFNKQSGCMDENVSCEKWAKAG 288
Query: 276 ECTKNPEYMVGTSGLPGYCRKSCKTC 301
EC KNP YMVG+ GYCRKSCK C
Sbjct: 289 ECQKNPTYMVGSDKDHGYCRKSCKAC 314
>At1g20270 unknown protein
Length = 287
Score = 240 bits (613), Expect = 6e-64
Identities = 114/209 (54%), Positives = 152/209 (72%), Gaps = 2/209 (0%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNK 102
+SW+PRAFVY FL+ EC++LIS+AK + +S V D+ +G+SK S VRTSSG F+ + +
Sbjct: 79 LSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRRGR 138
Query: 103 DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVL 162
D I+ IE +I+ +TF+P ++GE +QVL YE GQKY+PHYDYF D+ N GG R+AT+L
Sbjct: 139 DKIIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRMATML 198
Query: 163 MYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAI 222
MYL++V +GGETVFP A + +LSECGKKG++V PR GDALLF S+ P+A
Sbjct: 199 MYLSDVEEGGETVFPAA--NMNFSSVPWYNELSECGKKGLSVKPRMGDALLFWSMRPDAT 256
Query: 223 PDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
D SLH GCPVI G KWS+TKW+HV +
Sbjct: 257 LDPTSLHGGCPVIRGNKWSSTKWMHVGEY 285
>At2g17720 similar to prolyl 4-hydroxylase alpha subunit
Length = 291
Score = 237 bits (605), Expect = 5e-63
Identities = 116/214 (54%), Positives = 152/214 (70%), Gaps = 6/214 (2%)
Query: 40 VKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFIS 99
V+ +SW+PRA VY FLT+ EC+HLIS+AK + +S V D +G SK S VRTSSG F+
Sbjct: 80 VEVISWEPRAVVYHNFLTNEECEHLISLAKPSMVKSTVVDEKTGGSKDSRVRTSSGTFLR 139
Query: 100 KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVA 159
+ D +V IE +IS +TF+P ENGE +QVL Y+ GQKY+PHYDYF D+ N GG R+A
Sbjct: 140 RGHDEVVEVIEKRISDFTFIPVENGEGLQVLHYQVGQKYEPHYDYFLDEFNTKNGGQRIA 199
Query: 160 TVLMYLTNVTKGGETVFPNAEESPRHKLSETD--EDLSECGKKGVAVNPRRGDALLFCSL 217
TVLMYL++V GGETVFP A R +S +LS+CGK+G++V P++ DALLF ++
Sbjct: 200 TVLMYLSDVDDGGETVFPAA----RGNISAVPWWNELSKCGKEGLSVLPKKRDALLFWNM 255
Query: 218 HPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
P+A D SLH GCPV++G KWS+TKW HV F
Sbjct: 256 RPDASLDPSSLHGGCPVVKGNKWSSTKWFHVHEF 289
>At3g28490 prolyl 4-hydroxylase, putative
Length = 213
Score = 216 bits (551), Expect = 9e-57
Identities = 105/171 (61%), Positives = 139/171 (80%), Gaps = 6/171 (3%)
Query: 35 IDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRS-AVADNLSGESKLSEVRTS 93
+DPT++ Q+SW PRAF+YKGFL+D ECDHLI +AK +L++S VAD SGES+ SEVRTS
Sbjct: 27 VDPTRITQLSWTPRAFLYKGFLSDEECDHLIKLAKGKLEKSMVVADVDSGESEDSEVRTS 86
Query: 94 SGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIAR 153
SGMF++K +D IV+ +E K+++WTFLP+ENGE +Q+L YE+GQKYDPH+DYF DK +
Sbjct: 87 SGMFLTKRQDDIVANVEAKLAAWTFLPEENGEALQILHYENGQKYDPHFDYFYDKKALEL 146
Query: 154 GGHRVATVLMYLTNVTKGGETVFPNAE-ESPRHKLSETDEDLSECGKKGVA 203
GGHR+ATVLMYL+NVTKGGETVFPN + ++P+ K D+ S+C K+G A
Sbjct: 147 GGHRIATVLMYLSNVTKGGETVFPNWKGKTPQLK----DDSWSKCAKQGYA 193
>At4g25600 unknown protein
Length = 291
Score = 203 bits (516), Expect = 1e-52
Identities = 111/275 (40%), Positives = 160/275 (57%), Gaps = 29/275 (10%)
Query: 27 SYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESK 86
SY S +DPT+V Q+SW PR F+Y+GFL++ ECDHLIS+ K + +V + G+++
Sbjct: 46 SYVLGSKFVDPTRVLQLSWLPRVFLYRGFLSEEECDHLISLRKETTEVYSV--DADGKTQ 103
Query: 87 LSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFA 146
L D +V+GIE+K+S+WTFLP ENG I+V Y +K DYF
Sbjct: 104 L---------------DPVVAGIEEKVSAWTFLPGENGGSIKVRSYT-SEKSGKKLDYFG 147
Query: 147 DKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNP 206
++ + +ATV++YL+N T+GGE +FPN+E P++ C + G + P
Sbjct: 148 EEPSSVLHESLLATVVLYLSNTTQGGELLFPNSEMKPKNS----------CLEGGNILRP 197
Query: 207 RRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHE 266
+G+A+LF + NA D S H CPV++GE ATK I+ + G+C+D+ E
Sbjct: 198 VKGNAILFFTRLLNASLDGKSTHLRCPVVKGELLVATKLIYAKK-QARIEESGECSDEDE 256
Query: 267 SCERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
+C RWA LGEC KNP YM+G+ G CRKSC C
Sbjct: 257 NCGRWAKLGECKKNPVYMIGSPDYYGTCRKSCNAC 291
>At5g66060 prolyl 4-hydroxylase, alpha subunit-like protein
Length = 267
Score = 198 bits (504), Expect = 3e-51
Identities = 99/190 (52%), Positives = 134/190 (70%), Gaps = 2/190 (1%)
Query: 40 VKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFIS 99
V+ +SW+PRA VY FLT EC +LI +AK +++S V D +G+S S VRTSSG F++
Sbjct: 78 VEIISWEPRASVYHNFLTKEECKYLIELAKPHMEKSTVVDEKTGKSTDSRVRTSSGTFLA 137
Query: 100 KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVA 159
+ +D + IE +IS +TF+P E+GE +QVL YE GQKY+PHYDYF D+ N GG R+A
Sbjct: 138 RGRDKTIREIEKRISDFTFIPVEHGEGLQVLHYEIGQKYEPHYDYFMDEYNTRNGGQRIA 197
Query: 160 TVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHP 219
TVLMYL++V +GGETVFP A+ + + +LSECGK G++V P+ GDALLF S+ P
Sbjct: 198 TVLMYLSDVEEGGETVFPAAKGN--YSAVPWWNELSECGKGGLSVKPKMGDALLFWSMTP 255
Query: 220 NAIPDTLSLH 229
+A D SLH
Sbjct: 256 DATLDPSSLH 265
>At4g35810 putative protein
Length = 307
Score = 190 bits (483), Expect = 7e-49
Identities = 101/220 (45%), Positives = 141/220 (63%), Gaps = 37/220 (16%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGES----------------- 85
+SW+PRAFVY FLT+ EC+HLIS+AK + +S V D +G+S
Sbjct: 83 ISWEPRAFVYHNFLTNEECEHLISLAKPSMMKSKVVDVKTGKSIDSRFCTLTSVVVFTFQ 142
Query: 86 --------------KLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLR 131
L VRTSSG F+++ D IV IE++IS +TF+P ENGE +QVL
Sbjct: 143 LNLERFENSKFANPSLCRVRTSSGTFLNRGHDEIVEEIENRISDFTFIPPENGEGLQVLH 202
Query: 132 YEHGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETD 191
YE GQ+Y+PH+DYF D+ N+ +GG R+ATVLMYL++V +GGETVFP A + +S+
Sbjct: 203 YEVGQRYEPHHDYFFDEFNVRKGGQRIATVLMYLSDVDEGGETVFPAA----KGNVSDVP 258
Query: 192 --EDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLH 229
++LS+CGK+G++V P++ DALLF S+ P+A D SLH
Sbjct: 259 WWDELSQCGKEGLSVLPKKRDALLFWSMKPDASLDPSSLH 298
>At2g43080 unknown protein
Length = 283
Score = 184 bits (467), Expect = 5e-47
Identities = 103/216 (47%), Positives = 137/216 (62%), Gaps = 14/216 (6%)
Query: 35 IDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSS 94
I K + VSW PR V FL+ EC++L +IA+ L+ S V D +G+ S+VRTSS
Sbjct: 72 IGNVKPEVVSWSPRIIVLHDFLSPEECEYLKAIARPRLQVSTVVDVKTGKGVKSDVRTSS 131
Query: 95 GMFIS--KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIA 152
GMF++ + I+ IE +I+ ++ +P ENGE IQVLRYE Q Y PH+DYFAD N+
Sbjct: 132 GMFLTHVERSYPIIQAIEKRIAVFSQVPAENGELIQVLRYEPQQFYKPHHDYFADTFNLK 191
Query: 153 RGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGK--KGVAVNPRRGD 210
RGG RVAT+LMYLT+ +GGET FP A D D + GK KG++V P +GD
Sbjct: 192 RGGQRVATMLMYLTDDVEGGETYFPLA----------GDGDCTCGGKIMKGISVKPTKGD 241
Query: 211 ALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWI 246
A+LF S+ + D S+H GC V+ GEKWSATKW+
Sbjct: 242 AVLFWSMGLDGQSDPRSIHGGCEVLSGEKWSATKWM 277
>At4g35820 putative protein
Length = 272
Score = 159 bits (401), Expect = 2e-39
Identities = 91/195 (46%), Positives = 118/195 (59%), Gaps = 32/195 (16%)
Query: 43 VSWKPRAFVYKGFL--------TDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSS 94
++ +PRAFVY FL T+ ECDHLIS+AK + RS V + L+G + S RTSS
Sbjct: 91 ITKEPRAFVYHNFLALFFKICKTNEECDHLISLAKPSMARSKVRNALTGLGEESSSRTSS 150
Query: 95 GMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARG 154
G FI D IV IE +IS +TF+P+ENGE +QV+ YE GQK++PH+D G
Sbjct: 151 GTFIRSGHDKIVKEIEKRISEFTFIPQENGETLQVINYEVGQKFEPHFD----------G 200
Query: 155 GHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLF 214
R+ATVLMYL++V KGGETVFP A+ KKGV+V P++GDALLF
Sbjct: 201 FQRIATVLMYLSDVDKGGETVFPEAKGIK--------------SKKGVSVRPKKGDALLF 246
Query: 215 CSLHPNAIPDTLSLH 229
S+ P+ D S H
Sbjct: 247 WSMRPDGSRDPSSKH 261
>At4g33910 unknown protein
Length = 288
Score = 153 bits (387), Expect = 1e-37
Identities = 87/207 (42%), Positives = 121/207 (58%), Gaps = 9/207 (4%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSE-VRTSSGMFISKN 101
+SW+PRA + F T +C +I AK LK SA+A ++ ++ RTSSG FIS +
Sbjct: 81 LSWRPRAIYFPNFATAEQCQAIIERAKVNLKPSALALRKGETAENTKGTRTSSGTFISAS 140
Query: 102 KDAI--VSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVA 159
+++ + +E KI+ T +P+ +GE +LRYE GQKYD HYD F + R+A
Sbjct: 141 EESTGALDFVERKIARATMIPRSHGESFNILRYELGQKYDSHYDVFNPTEYGPQSSQRIA 200
Query: 160 TVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHP 219
+ L+YL++V +GGET+FP S D +C G+ V PR+GD LLF S+ P
Sbjct: 201 SFLLYLSDVEEGGETMFPFENGSN----MGIGYDYKQC--IGLKVKPRKGDGLLFYSVFP 254
Query: 220 NAIPDTLSLHAGCPVIEGEKWSATKWI 246
N D SLH CPV +GEKW ATKWI
Sbjct: 255 NGTIDQTSLHGSCPVTKGEKWVATKWI 281
>At2g23100 hypothetical protein
Length = 1036
Score = 71.6 bits (174), Expect = 5e-13
Identities = 39/104 (37%), Positives = 61/104 (58%), Gaps = 3/104 (2%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSS-GMFISKN 101
+SW PR F F T +C+ +I +AK +LK S +A L E+K +++ S ++
Sbjct: 800 LSWNPRVFYLPNFATKQQCEAVIDMAKPKLKPSTLA--LRKETKHFQMQYRSLHQHTDED 857
Query: 102 KDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYF 145
+ +++ IE+KI+ T PK+ E +LRY+ GQKYD HYD F
Sbjct: 858 ESGVLAAIEEKIALATRFPKDYYESFNILRYQLGQKYDSHYDAF 901
>At3g53440 unknown protein
Length = 512
Score = 30.0 bits (66), Expect = 1.6
Identities = 24/103 (23%), Positives = 40/103 (38%), Gaps = 11/103 (10%)
Query: 32 SAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVR 91
+ ++ P +KQV K L + DHL + KR A + L+G+ + +
Sbjct: 414 TTLVSPPVLKQVKNKA--------LHEKYIDHLHI---RDAKRKAESTRLAGKENIRPIE 462
Query: 92 TSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEH 134
+ KDA+ ++D I L EN Y+H
Sbjct: 463 IQKKDSVRAAKDALFFDVQDAIQKLKGLEAENSSSSSEFCYDH 505
>At5g04590 sulphite reductase
Length = 642
Score = 29.6 bits (65), Expect = 2.1
Identities = 40/193 (20%), Positives = 74/193 (37%), Gaps = 30/193 (15%)
Query: 30 GTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSE 89
GT+ + P + + PR F + D+ + + +++ V+D +GE +
Sbjct: 266 GTNFVDSPEPIYGTQFLPRKFKVA---VTVPTDNSVDLLTNDIGVVVVSDE-NGEPQGFN 321
Query: 90 VRTSSGMFISKNKDAIVSGIEDKISSWTFLPKEN----GEDIQVLRYEHGQKYDPHYDYF 145
+ GM + ++ + + + I ++PKE+ + I V + EHG++ D Y
Sbjct: 322 IYVGGGMGRTHRMESTFARLAEPIG---YVPKEDILYAVKAIVVTQREHGRRDDRKYSR- 377
Query: 146 ADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVN 205
+ YL ++ G F + E K E +L E K
Sbjct: 378 ----------------MKYL--ISSWGIEKFRDVVEQYYGKKFEPSRELPEWEFKSYLGW 419
Query: 206 PRRGDALLFCSLH 218
+GD FC LH
Sbjct: 420 HEQGDGAWFCGLH 432
>At3g18520 histone deacetylase, putative
Length = 552
Score = 29.3 bits (64), Expect = 2.7
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 1/51 (1%)
Query: 27 SYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAV 77
S AG ++D ++ W A Y L++LE LI K+++KR V
Sbjct: 476 SVAGLQTVLDVLNIQLEFWPSLAISYSKLLSELEA-RLIENKKNQMKRKVV 525
>At4g11480 serine/threonine kinase-like protein
Length = 656
Score = 28.9 bits (63), Expect = 3.6
Identities = 17/73 (23%), Positives = 34/73 (46%), Gaps = 3/73 (4%)
Query: 34 IIDPTKVKQVSWKPRAFVYKGF---LTDLECDHLISIAKSELKRSAVADNLSGESKLSEV 90
++DPTK Q+ WK R + G L L D ++I ++K S + + K+++
Sbjct: 414 LLDPTKKSQLDWKRRYNIIGGITRGLLYLHQDSRLTIIHRDIKASNILLDADMNPKIADF 473
Query: 91 RTSSGMFISKNKD 103
+ + + +D
Sbjct: 474 GMARNFRVDQTED 486
>At5g56860 unknown protein
Length = 398
Score = 28.1 bits (61), Expect = 6.1
Identities = 13/34 (38%), Positives = 22/34 (64%)
Query: 69 KSELKRSAVADNLSGESKLSEVRTSSGMFISKNK 102
K E ++ A+ ++G+S++S+ TSS IS NK
Sbjct: 324 KEEEEKEMEAETVAGDSEISKSTTSSNSSISSNK 357
>At2g36200 putative kinesin-related cytokinesis protein
Length = 1056
Score = 28.1 bits (61), Expect = 6.1
Identities = 14/41 (34%), Positives = 23/41 (55%)
Query: 70 SELKRSAVADNLSGESKLSEVRTSSGMFISKNKDAIVSGIE 110
S L RSA N ++++ RT++ ++KN D I+ IE
Sbjct: 853 SSLVRSACDSNEQHDAEVDSARTAAEKDVTKNSDDIIQQIE 893
>At2g24080 unknown protein
Length = 374
Score = 28.1 bits (61), Expect = 6.1
Identities = 20/85 (23%), Positives = 38/85 (44%), Gaps = 18/85 (21%)
Query: 122 ENGEDIQVLRYEHGQ------KYDPHYDYFADKVNIARGGHRVATVLM------------ 163
+NG+ + V ++E GQ K D HY +V++ R + V++
Sbjct: 138 KNGDFVVVWQFEQGQFLGFCKKGDLHYRDIPIRVDVRREFRGLQDVVLKGYNLYILVNRS 197
Query: 164 YLTNVTKGGETVFPNAEESPRHKLS 188
++ ++ G+ F N E+PR +S
Sbjct: 198 FIRHIDLSGQDGFKNVSETPRFPIS 222
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,502,690
Number of Sequences: 26719
Number of extensions: 324980
Number of successful extensions: 766
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 729
Number of HSP's gapped (non-prelim): 27
length of query: 301
length of database: 11,318,596
effective HSP length: 99
effective length of query: 202
effective length of database: 8,673,415
effective search space: 1752029830
effective search space used: 1752029830
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146553.10


