

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146551.3 - phase: 1 /pseudo
(379 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
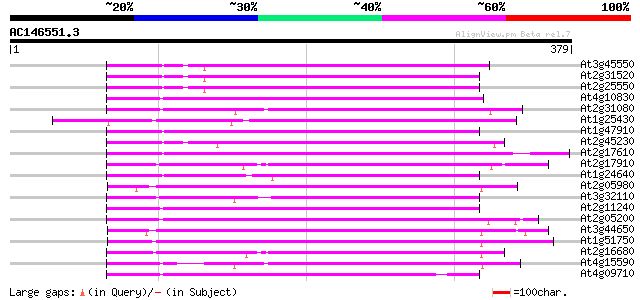
Score E
Sequences producing significant alignments: (bits) Value
At3g45550 putative protein 167 9e-42
At2g31520 putative non-LTR retroelement reverse transcriptase 165 4e-41
At2g25550 putative non-LTR retroelement reverse transcriptase 164 6e-41
At4g10830 putative protein 162 3e-40
At2g31080 putative non-LTR retroelement reverse transcriptase 151 6e-37
At1g25430 hypothetical protein 147 7e-36
At1g47910 reverse transcriptase, putative 147 9e-36
At2g45230 putative non-LTR retroelement reverse transcriptase 147 1e-35
At2g17610 putative non-LTR retroelement reverse transcriptase 145 4e-35
At2g17910 putative non-LTR retroelement reverse transcriptase 141 5e-34
At1g24640 hypothetical protein 141 5e-34
At2g05980 putative non-LTR retroelement reverse transcriptase 137 7e-33
At3g32110 non-LTR reverse transcriptase, putative 137 1e-32
At2g11240 pseudogene 137 1e-32
At2g05200 putative non-LTR retroelement reverse transcriptase 135 5e-32
At3g44650 putative protein 134 8e-32
At1g51750 hypothetical protein 133 1e-31
At2g16680 putative non-LTR retroelement reverse transcriptase 132 2e-31
At4g15590 reverse transcriptase like protein 131 7e-31
At4g09710 RNA-directed DNA polymerase -like protein 130 2e-30
>At3g45550 putative protein
Length = 851
Score = 167 bits (423), Expect = 9e-42
Identities = 98/263 (37%), Positives = 143/263 (54%), Gaps = 8/263 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L + + L+ + +GNG+ +THLQFADD+L C NV
Sbjct: 259 PLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPA-ITHLQFADDSLFF---CQANV 314
Query: 126 RTMMA---VLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGV 181
R A V ++E SG K+N KS++T G + S L+N YLG+
Sbjct: 315 RNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGGKYLGL 374
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P +K + +IDR+ R +SW+ K LS G+ +LLK V ++PVY +S FK P G
Sbjct: 375 PEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCFKLPQG 434
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
I+S IES+ +F+W R I W+ W + SK EGGLG R + FN +LL K WR+
Sbjct: 435 IVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRI 494
Query: 302 LTERDELWYRVLKAKYGEEGGRI 324
+ + L+ RV+KA+Y ++ I
Sbjct: 495 IQYPNSLFARVMKARYFKDNSII 517
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 165 bits (417), Expect = 4e-41
Identities = 95/256 (37%), Positives = 142/256 (55%), Gaps = 8/256 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L + + L+N + +GNG+ +THLQFADD+L C NV
Sbjct: 794 PLSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPA-ITHLQFADDSLFF---CQANV 849
Query: 126 RTMMA---VLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGV 181
R A V ++E SG K+N KSM+T G + S + ++ YLG+
Sbjct: 850 RNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGGKYLGL 909
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P +K ++ +IDR+ R S+W+ + LS G+ ++LK V ++PVY +S FK P G
Sbjct: 910 PEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKG 969
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
I+S IES+ +F+W R I W+ W + SK EGGLG R + FN +LL K WR+
Sbjct: 970 IVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRL 1029
Query: 302 LTERDELWYRVLKAKY 317
+ + L+ RV+KA+Y
Sbjct: 1030 IQYPNSLFARVMKARY 1045
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 164 bits (416), Expect = 6e-41
Identities = 95/256 (37%), Positives = 141/256 (54%), Gaps = 8/256 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L + + L+N + +GNG+ +THLQFADD+L C NV
Sbjct: 1020 PLSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPA-ITHLQFADDSLFF---CQANV 1075
Query: 126 RTMMA---VLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGV 181
R A V ++E SG K+N KSM+T G + S ++ YLG+
Sbjct: 1076 RNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSRLKQILEIPNQGGGGKYLGL 1135
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P +K ++ +IDR+ R S+W+ + LS G+ ++LK V ++PVY +S FK P G
Sbjct: 1136 PEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKG 1195
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
I+S IES+ +F+W R I W+ W + SK EGGLG R + FN +LL K WR+
Sbjct: 1196 IVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRL 1255
Query: 302 LTERDELWYRVLKAKY 317
+ + L+ RV+KA+Y
Sbjct: 1256 IQYPNSLFARVMKARY 1271
>At4g10830 putative protein
Length = 1294
Score = 162 bits (410), Expect = 3e-40
Identities = 94/256 (36%), Positives = 139/256 (53%), Gaps = 2/256 (0%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L A+ L+ V +GNG VTHLQFADD+L + N
Sbjct: 1000 PLSPYLFILCADILNHLIKNRVAEGDIRGIRIGNGVP-GVTHLQFADDSLFFCQSNVRNC 1058
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
+ + V ++E SG K+N KSM+T G + + ++ YLG+P
Sbjct: 1059 QALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQ 1118
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
+K + +I+R+ R SSW+ K LS G+ ++LK V S+PVY +S FK P I+S
Sbjct: 1119 FGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVS 1178
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
IE++ +F+W + R+I WI W + SK EGGLG R + FN +LL K WRM+
Sbjct: 1179 EIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINN 1238
Query: 305 RDELWYRVLKAKYGEE 320
+ L+ R++KA+Y E
Sbjct: 1239 PNSLFARIMKARYFRE 1254
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 151 bits (381), Expect = 6e-37
Identities = 93/288 (32%), Positives = 148/288 (51%), Gaps = 10/288 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L E L+ A VG + V G +++H+ FADD ++ E +
Sbjct: 501 PLSPYLFVLCLERLCHLIEASVGKREWKPIAVSCGGS-KLSHVCFADDLILFAEASVAQI 559
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT---GVNISDSWLVEASALMNCCRGNFTFVYLGVP 182
R + VL F E SG KV+ KS + V+ L+ + + C + YLG+P
Sbjct: 560 RIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLISEESGIGCTKE--LGKYLGMP 617
Query: 183 IGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGI 242
I + V++R+ RL+ W + LS+ GR+ L K VLSS+PV+ +S P
Sbjct: 618 ILQKRMNKETFGEVLERVSARLAGWKGRSLSLAGRITLTKAVLSSIPVHVMSAILLPVST 677
Query: 243 ISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRML 302
+ +++ ++F WG E +K + W IC KAEGG+G+R N +L+ K WR+L
Sbjct: 678 LDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKVGWRLL 737
Query: 303 TERDELWYRVLKAKYGEEGGR----IREGGRLSSAWWKMVCHIREGLV 346
+++ LW RV++ KY G + ++ R SS W + +RE +V
Sbjct: 738 QDKESLWARVVRKKYKVGGVQDTSWLKPQPRWSSTWRSVAVGLREVVV 785
>At1g25430 hypothetical protein
Length = 1213
Score = 147 bits (372), Expect = 7e-36
Identities = 104/322 (32%), Positives = 160/322 (49%), Gaps = 14/322 (4%)
Query: 30 LAEMDK*VCWDSNGID-PCKWESYGGISD*AWSSPRG-----PLSPFLFLLAAEGFTVLM 83
LA +K + W S I P S G + + S +G PLSP+LF+LA E F+ L+
Sbjct: 613 LAIPEKFINWISQCISTPTFTVSINGGNGGFFKSTKGLRQGDPLSPYLFVLAMEAFSNLL 672
Query: 84 NAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNVRTMMAVLLLFEEISGLKV 143
++ + + H S + ++HL FADD +I + ++ + L F SGLKV
Sbjct: 673 HSRYESGLIHYHP--KASNLSISHLMFADDVMIFFDGGSFSLHGICETLDDFASWSGLKV 730
Query: 144 NFHKS--MLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRIV 201
N KS L G+N +S A+A G YLG+P+ +++ ++P++++I
Sbjct: 731 NKDKSHLYLAGLNQLES---NANAAYGFPIGTLPIRYLGLPLMNRKLRIAEYEPLLEKIT 787
Query: 202 GRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEEN 261
R SW NK LS GR+ L+ V+ +++S F P G I IES+ F W G E
Sbjct: 788 ARFRSWVNKCLSFAGRIQLISSVIFGSINFWMSTFLLPKGCIKRIESLCSRFLWSGNIEQ 847
Query: 262 RKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEG 321
K + W +CL K+EGGLG+RR+ +N +L + WR+ +D LW + G
Sbjct: 848 AKGIKVSWAALCLPKSEGGLGLRRLLEWNKTLSMRLIWRLFVAKDSLWADWQHLHHLSRG 907
Query: 322 G-RIREGGRLSSAWWKMVCHIR 342
EGG+ S WK + +R
Sbjct: 908 SFWAVEGGQSDSWTWKRLLSLR 929
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 147 bits (371), Expect = 9e-36
Identities = 83/253 (32%), Positives = 132/253 (51%), Gaps = 2/253 (0%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L E + ++ V S V+HL FADD+L +
Sbjct: 422 PLSPYLFILCTEVLIANIRKAERQNLITGIKVATPSPA-VSHLLFADDSLFFCKANKEQC 480
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
++ +L +E +SG ++NF KS + G + DS + ++ YLG+P
Sbjct: 481 GIILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADIKLILGIHNLGGMGSYLGLPES 540
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
K + V DR+ R++ W+ K LS GG+ V++K V ++LP Y +S F+ P I S
Sbjct: 541 LGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAATLPRYVMSCFRLPKAITS 600
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
+ S F+W ++R + W+ WD +C SK++GGLG R + FN +LL K WR++T
Sbjct: 601 KLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAKQLWRLITA 660
Query: 305 RDELWYRVLKAKY 317
D L+ +V K +Y
Sbjct: 661 PDSLFAKVFKGRY 673
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 147 bits (370), Expect = 1e-35
Identities = 91/275 (33%), Positives = 142/275 (51%), Gaps = 10/275 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF++ E ++ + + V G+ ++HL FADD++ C +N
Sbjct: 638 PLSPYLFVICTEMLVKMLQSAEQKNQITGLKVARGAPP-ISHLLFADDSMFY---CKVND 693
Query: 126 RTMMAVLLLFEEIS---GLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGV 181
+ ++ + EE S G +VN+ KS + G +IS+ + R VYLG+
Sbjct: 694 EALGQIIRIIEEYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGL 753
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P K++ + DR+ ++ W + LS GG+ +LLK V +LP Y +S FK P
Sbjct: 754 PESFQGSKVATLSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKT 813
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
I IES+ F+W +E R + W W + KA GGLG + + AFN++LLGK WRM
Sbjct: 814 ICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRM 873
Query: 302 LTERDELWYRVLKAKYGEEGGRIRE--GGRLSSAW 334
+TE+D L +V K++Y + + G R S AW
Sbjct: 874 ITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAW 908
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 145 bits (366), Expect = 4e-35
Identities = 93/315 (29%), Positives = 153/315 (48%), Gaps = 14/315 (4%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
P+SP+L+LL EG + L+ A + H F ++HL FA D+L+ +
Sbjct: 93 PISPYLYLLCTEGLSALIQASIKAKQLHGFKASRNGPA-ISHLLFAHDSLVFCKATLEEC 151
Query: 126 RTMMAVLLLFEEISGLKVNFHKS-MLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
T++ VL L+E+ SG VNF KS +L G + + S L+ + YLG+P
Sbjct: 152 MTLVNVLKLYEKASGQAVNFQKSAILFGKGLDFRTSEQLSQLLGIYKTEGFGRYLGLPEF 211
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
K + + + + ++ +W NKLLS G+ VL+K +++++P Y +S F P +I
Sbjct: 212 VGRNKTNAFSFIAQTMDQKMDNWYNKLLSPAGKEVLIKSIVTAIPTYSMSCFLLPMRLIH 271
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
I S + F+W + KI W+ W + K GGL +R + FN++LL K WR+L +
Sbjct: 272 QITSAMRWFWWSNTKVKHKIPWVAWSKLNDPKKMGGLAIRDLKDFNIALLAKQSWRILQQ 331
Query: 305 RDELWYRVLKAKYGEEGGRIREGGRLSSAW-WKMVCHIREGLVDWGFTVTEEVSSSFQFD 363
L RV KAKY + + S++ WK + H T+ +S ++
Sbjct: 332 PFSLMARVFKAKYFPKERLLDAKATSQSSYAWKSILH-----------GTKLISRGLKYI 380
Query: 364 DGKGEYGEGYGEAWL 378
G G + + + WL
Sbjct: 381 AGNGNNIQLWKDNWL 395
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 141 bits (356), Expect = 5e-34
Identities = 95/305 (31%), Positives = 149/305 (48%), Gaps = 11/305 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP LF+L E ++N + +V V HL FADDTL++ +
Sbjct: 616 PLSPALFVLCTEALIHILNKAEQAGKITGIQFQD-KKVSVNHLLFADDTLLMCKATKQEC 674
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT-GVNIS---DSWLVEASALMNCCRGNFTFVYLGV 181
+M L + ++SG +N +KS +T G N+ W+ S + G T YLG+
Sbjct: 675 EELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSRSGIS--LEGG-TGKYLGL 731
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P K + + +++ RL+ W K LS GG+ VLLK + +LPVY +S FK P
Sbjct: 732 PECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIALALPVYVMSCFKLPKN 791
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
+ + ++ F+W ++ RKI W+ W + L K +GG G + + FN +LL K WR+
Sbjct: 792 LCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAWRV 851
Query: 302 LTERDELWYRVLKAKYGEEGGRI--REGGRLSSAWWKMVCHIREGLVDWGFTVTEEVSSS 359
L E+ L+ RV +++Y + G R S A W+ + RE L+ TV +
Sbjct: 852 LQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYA-WRSILFGRELLMQGLRTVIGNGQKT 910
Query: 360 FQFDD 364
F + D
Sbjct: 911 FVWTD 915
>At1g24640 hypothetical protein
Length = 1270
Score = 141 bits (356), Expect = 5e-34
Identities = 89/256 (34%), Positives = 129/256 (49%), Gaps = 8/256 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSPFLF+L EG T L+N + V HL FADD+L + +
Sbjct: 601 PLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPM-VHHLLFADDSLFLCKASREQS 659
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTF----VYLGV 181
+ +L ++ +G +N +KS +T D L + C G FT YLG+
Sbjct: 660 LVLQKILKVYGNATGQTINLNKSSITFGEKVDEQL---KGTIRTCLGIFTEGGAGTYLGL 716
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P K+ + DR+ +L W + LS GG+ VLLK V ++PV+ +S FK P
Sbjct: 717 PECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPIT 776
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
++ES SF+W + +RKI W W+ +CL K GGLG R + +FN +LL K WR+
Sbjct: 777 TCENLESAMASFWWDSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRL 836
Query: 302 LTERDELWYRVLKAKY 317
L D L R+LK++Y
Sbjct: 837 LHFPDCLLSRLLKSRY 852
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 137 bits (346), Expect = 7e-33
Identities = 85/281 (30%), Positives = 140/281 (49%), Gaps = 8/281 (2%)
Query: 67 LSPFLFLLAAEGFTVLMN--AVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLN 124
LSP+L+++ + +++ AV YH + +THL FADD ++ + +
Sbjct: 809 LSPYLYVICMNVLSCMLDKAAVEKKISYHP----RCRNMNLTHLCFADDIMVFSDGTSKS 864
Query: 125 VRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
++ +A+ F +S LK++ KS + IS + G YLG+P+
Sbjct: 865 IQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTSILQQFPFELGTLPVKYLGLPLL 924
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
S + P++++I R++SW N+ LS GRL L+K VLSS+ ++LS F+ P +
Sbjct: 925 TKRMTQSDYLPLVEKIRARITSWTNRFLSFAGRLQLIKSVLSSITNFWLSVFRLPKACLQ 984
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
IE +F +F W G + N K A I W +C K EGGLG++ + N L K WR+L+
Sbjct: 985 EIEKMFSAFLWSGPDLNTKKAKIAWSEVCKLKEEGGLGLKPLKEANEVSLLKLIWRILSA 1044
Query: 305 RDELWYRVLKAKY--GEEGGRIREGGRLSSAWWKMVCHIRE 343
RD LW + + E ++E L S W+ + R+
Sbjct: 1045 RDSLWVKWVNKHLIRKETFWSVKENTGLGSWLWRKILKQRD 1085
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 137 bits (344), Expect = 1e-32
Identities = 83/261 (31%), Positives = 131/261 (49%), Gaps = 18/261 (6%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L E L+ + + + + S R++H+ FADD ++ E +
Sbjct: 1177 PLSPYLFVLCIERLCHLIESSIAAKKWKPIKISQ-SGPRLSHICFADDLILFAEASIDQI 1235
Query: 126 RTMMAVLLLFEEISGLKVNFHKSML---------TGVNISDSWLVEASALMNCCRGNFTF 176
R + VL F SG KV+ KS + G IS+ ++A+ +
Sbjct: 1236 RVLRGVLEKFCGASGQKVSLEKSKIYFSKNVLRDLGKRISEESGMKATKDLG-------- 1287
Query: 177 VYLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFF 236
YLGVPI + VI R RL+ W ++LS GRL L K VL+S+ V+ +S
Sbjct: 1288 KYLGVPILQKRINKETFGEVIKRFSSRLAGWKGRMLSFAGRLTLTKAVLTSILVHTMSTI 1347
Query: 237 KAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGK 296
K P + ++ + ++F WG E +K + W +CL + EGGLG+R A N +L+ K
Sbjct: 1348 KLPQSTLDGLDKVSRAFLWGSTLEKKKQHLVAWTRVCLPRREGGLGIRSATAMNKALIAK 1407
Query: 297 WCWRMLTERDELWYRVLKAKY 317
WR+L + LW +V+++KY
Sbjct: 1408 VGWRVLNDGSSLWAQVVRSKY 1428
>At2g11240 pseudogene
Length = 1044
Score = 137 bits (344), Expect = 1e-32
Identities = 82/253 (32%), Positives = 125/253 (48%), Gaps = 2/253 (0%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSPFLF++ +E + L V G+ RV HL FADDT+ +
Sbjct: 464 PLSPFLFIICSEVLSGLCRKAQLDGSLLGLRVSKGNP-RVNHLLFADDTIFFCRSDLKSC 522
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
+T + +L +EE SG +N KS +T D EA ++ YLG+P
Sbjct: 523 KTFLCILKKYEEASGQMINKSKSAITFSRKTPDHIKTEAQQILGIQLVGGLGKYLGLPKM 582
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
+K + ++DRI R SW+++ LS G+ +LK VL+S+P Y +S FK +
Sbjct: 583 FGRKKRDLFNQIVDRIRQRSLSWSSRFLSTAGKTTMLKSVLASMPTYTMSCFKLLVSLCK 642
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
I+S F+W + +K+ WI W + +K EGGLG + + FN +LL K WR++
Sbjct: 643 RIQSALTHFWWDSSADKKKMCWIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIVQS 702
Query: 305 RDELWYRVLKAKY 317
+ R+L KY
Sbjct: 703 PSCVLVRILLGKY 715
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 135 bits (339), Expect = 5e-32
Identities = 95/299 (31%), Positives = 143/299 (47%), Gaps = 10/299 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVG-NGSEVRVTHLQFADDTLIIGEKCWLN 124
PLSP LF+L E + L V NG RV HL FADDT+ + +
Sbjct: 533 PLSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSINGP--RVNHLLFADDTMFFSKSDPES 590
Query: 125 VRTMMAVLLLFEEISGLKVNFHKSMLT-GVNISDSWLVEASALMNCCRGNFTFVYLGVPI 183
+ +L + + SG +NFHKS +T S + ++ + T YLG+P
Sbjct: 591 CNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPE 650
Query: 184 GGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGII 243
RK + +ID+I + SW ++ LS G+ V+LK VL+S+P+Y +S FK P+ +
Sbjct: 651 HFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALC 710
Query: 244 SSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLT 303
I+S+ F+W + RK +W+ W + K GGLG R + N SLL K WR+L
Sbjct: 711 RKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLN 770
Query: 304 ERDELWYRVLKAKYGEEGG--RIREGGRLSSAWWKMVCH---IREGLVDWGFTVTEEVS 357
+ L R+L KY + + S W ++ ++EGL W T E+VS
Sbjct: 771 SPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSIIAGREILKEGL-GWLITNGEKVS 828
>At3g44650 putative protein
Length = 762
Score = 134 bits (337), Expect = 8e-32
Identities = 92/304 (30%), Positives = 149/304 (48%), Gaps = 11/304 (3%)
Query: 67 LSPFLFLLAAEGFTVLMNAVVGTDM--YHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLN 124
LSP+LF++ + + L++ VVG YH + + +THL FADD +I+ + +
Sbjct: 215 LSPYLFVICMDVLSKLLDKVVGIGRIGYHP----HCKRMGLTHLSFADDLMILTDGQCRS 270
Query: 125 VRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
+ ++ V LF + SGLK++ KS + +S + + G YLG+P+
Sbjct: 271 IEGIIEVFDLFSKWSGLKISMEKSTIFSAGLSSTSRAQLHTHFPFEVGELPIRYLGLPLV 330
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
+ P+I++I R+ SW+++ LS GR L+ ++ S ++LS F+ P I
Sbjct: 331 TKRLSSVDYAPLIEQIRKRIGSWSSRFLSFAGRFNLISSIIWSSCNFWLSAFQLPRACIQ 390
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
IE + SF W G N K A I W+ +C K+EGGLG+R + N K WR+++
Sbjct: 391 EIEKLCSSFLWSGTNLNSKKAKISWNQVCKPKSEGGLGLRSLKEANDVCCLKLVWRIISH 450
Query: 305 RDELWYRVLKAKY--GEEGGRIREGGRLSSAWWKMVCHIREGLVD--WGFTVTEEVSSSF 360
D LW + ++ E ++E L S WK + R G+ V S+SF
Sbjct: 451 GDSLWVKWVEHNLLKREIFWIVKENANLGSWIWKKILKYR-GVAKRFCKAEVGNGESTSF 509
Query: 361 QFDD 364
FDD
Sbjct: 510 WFDD 513
>At1g51750 hypothetical protein
Length = 629
Score = 133 bits (335), Expect = 1e-31
Identities = 90/304 (29%), Positives = 145/304 (47%), Gaps = 5/304 (1%)
Query: 67 LSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNVR 126
LSP+LF+++ E + +++ G + + +THL FADD +I+ + +V
Sbjct: 67 LSPYLFVISMEVLSKMLDQAAGGKRFGFHP--KCKNLGLTHLCFADDLMILTDGKVRSVD 124
Query: 127 TMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGD 186
++ V+ LF + SGL++N K+ L +SD + G YLG+P+
Sbjct: 125 GIVEVMNLFAKRSGLQINMEKTTLYTAGVSDHNRYMMISRYPFGLGQLPVRYLGLPLVTK 184
Query: 187 PRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSI 246
P+ ++I R+ +W ++ LS GRL L+ VL S +++S F+ P+ + I
Sbjct: 185 RLTKEDLSPLFEQIRNRIGTWTSRYLSFAGRLNLISSVLWSTMNFWMSAFRLPSACLKEI 244
Query: 247 ESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERD 306
SI +F W G E +R+ A + WD IC K EGGLG+R + NV + K WR+ + D
Sbjct: 245 NSICSAFLWSGPELHRRKAKVSWDDICKPKQEGGLGLRSLTEANVVSVLKLIWRVTSNDD 304
Query: 307 ELWYRVLKAKY--GEEGGRIREGGRLSSAWWKMVCHIREGLVDWG-FTVTEEVSSSFQFD 363
LW + K E + L S WK + RE + V +SF FD
Sbjct: 305 SLWVKWSKMNLLKQESFWSLTPNSSLGSWMWKKMLKYRETAKPFSRVEVNNGARTSFWFD 364
Query: 364 DGKG 367
+ G
Sbjct: 365 NWSG 368
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 132 bits (333), Expect = 2e-31
Identities = 84/275 (30%), Positives = 134/275 (48%), Gaps = 10/275 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
P+SPFLF+L E ++ + NGS V HL F DDT ++ +
Sbjct: 590 PISPFLFVLCTEALIHILQQAENSKKVSGIQF-NGSGPSVNHLLFVDDTQLVCRATKSDC 648
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT-GVNISDS---WLVEASALMNCCRGNFTFVYLGV 181
MM L + ISG +N KS +T GV + + W+ S + G T YLG+
Sbjct: 649 EQMMLCLSQYGHISGQLINVEKSSITFGVKVDEDTKRWIKNRSGIH--LEGG-TGKYLGL 705
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P K + + +++ LS W +K LS GG+ +LLK + +LPVY ++ F+ P G
Sbjct: 706 PENLSGSKQDLFGYIKEKLQSHLSGWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKG 765
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
+ + + S+ F+W E + KI WI + L K+ GG G + + FN +LL K WR+
Sbjct: 766 LCTKLTSVMMDFWWNSMEFSNKIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRL 825
Query: 302 LTERDELWYRVLKAKY--GEEGGRIREGGRLSSAW 334
++ + ++ K++Y + R+G R S W
Sbjct: 826 FSDSKSIVSQIFKSRYFMNTDFLNARQGTRPSYTW 860
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 131 bits (329), Expect = 7e-31
Identities = 88/288 (30%), Positives = 137/288 (47%), Gaps = 29/288 (10%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
P+SP+LF+L E + VG + S + G +V+H+ FADD
Sbjct: 453 PISPYLFVLCIERLCHQIETAVGRGDWKSISISQGGP-KVSHVCFADD------------ 499
Query: 126 RTMMAVLLLFEEIS-GLKVNFHKSML---TGVNISDSWLVEASALMNCCRGNFTFVYLGV 181
L+LF E S KV+ KS + V+ L+ A + R YLG+
Sbjct: 500 ------LILFAEASVAQKVSLEKSKIFFSNNVSRDLEGLITAETGIGSTRE--LGKYLGM 551
Query: 182 PIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAG 241
P+ + V++R+ RLS W ++ LS+ GR+ L K VL S+P++ +S PA
Sbjct: 552 PVLQKRINKDTFGEVLERVSSRLSGWKSRSLSLAGRITLTKAVLMSIPIHTMSSILLPAS 611
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRM 301
++ ++ + ++F WG E RK + W +C KA GGLG+R N +LL K WR+
Sbjct: 612 LLEQLDKVSRNFLWGSTVEKRKQHLLSWKKVCRPKAAGGLGLRASKDMNRALLAKVGWRL 671
Query: 302 LTERDELWYRVLKAKYG----EEGGRIREGGRLSSAWWKMVCHIREGL 345
L ++ LW RVL+ KY + + SS W + +REG+
Sbjct: 672 LNDKVSLWARVLRRKYKVTDVHDSSWLVPKATWSSTWRSIGVGLREGV 719
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 130 bits (326), Expect = 2e-30
Identities = 86/253 (33%), Positives = 122/253 (47%), Gaps = 9/253 (3%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L E + L + V GS +V HL FADDT+ +
Sbjct: 579 PLSPYLFILCTEVLSGLCRKAQEKGVMVGIRVARGSP-QVNHLLFADDTMFFCKTNPTCC 637
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASAL-MNCCRGNFTFVYLGVPIG 184
+ +L +E SG +N KS +T + + + L + YLG+P
Sbjct: 638 GALSNILKKYELASGQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEH 697
Query: 185 GDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIIS 244
RK + ++DRI R SW+ + LS G+ +LLK VLSS+P Y + FK PA +
Sbjct: 698 FGRRKRDIFSSIVDRIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCK 757
Query: 245 SIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
I+S+ F+W + RK+AW+ WD + L EGGLG R + A K WR+L E
Sbjct: 758 QIQSVLTRFWWDSKPDKRKMAWVSWDKLTLPINEGGLGFREIEA-------KLSWRILKE 810
Query: 305 RDELWYRVLKAKY 317
L RVL KY
Sbjct: 811 PHSLLSRVLLGKY 823
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.331 0.146 0.494
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,967,025
Number of Sequences: 26719
Number of extensions: 398946
Number of successful extensions: 1183
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1044
Number of HSP's gapped (non-prelim): 80
length of query: 379
length of database: 11,318,596
effective HSP length: 101
effective length of query: 278
effective length of database: 8,619,977
effective search space: 2396353606
effective search space used: 2396353606
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146551.3


