

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.4 - phase: 0 /pseudo
(322 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
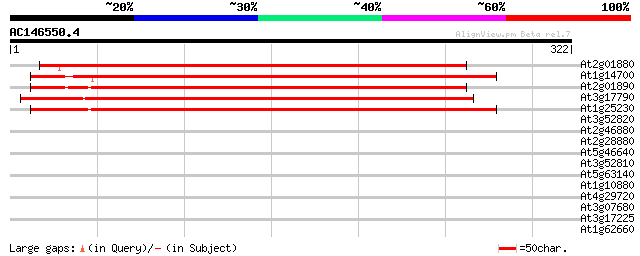
Score E
Sequences producing significant alignments: (bits) Value
At2g01880 purple acid phosphatase like protein 339 1e-93
At1g14700 purple acid phosphatase -like protein 338 2e-93
At2g01890 purple acid phosphatase like protein 333 1e-91
At3g17790 acid phosphatase type 5 320 7e-88
At1g25230 unknown protein 315 2e-86
At3g52820 purple acid phosphatase (PAP22) 32 0.36
At2g46880 unknown protein 30 1.3
At2g28880 para-aminobenzoate synthase and glutamine amidotransfe... 30 1.8
At5g46640 unknown protein 30 2.3
At3g52810 purple acid phosphatase-like protein 29 3.0
At5g63140 unknown protein 29 3.9
At1g10880 hypothetical protein 29 3.9
At4g29720 unknown protein 28 6.7
At3g07680 putative coated vesicle membrane protein 28 6.7
At3g17225 putative protein 28 8.7
At1g62660 beta-fructosidase (At1g62660) like protein 28 8.7
>At2g01880 purple acid phosphatase like protein
Length = 328
Score = 339 bits (870), Expect = 1e-93
Identities = 160/248 (64%), Positives = 194/248 (77%), Gaps = 3/248 (1%)
Query: 18 SVLLRLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQMGIV 74
SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQMG+V
Sbjct: 8 SVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQMGVV 67
Query: 75 GDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVD 134
G+ L+IDFVIS GDNFY DGL+GV+DP+F SF +IYT PSLQK WYSVLGNHDYRG+V+
Sbjct: 68 GEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRGNVE 127
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQLS +L QKD RW C RSF+L G+V+FFF DT PF+EKYFT+P++HTYDW VLPR
Sbjct: 128 AQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLPRNK 187
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
Y + LL ++DL + KS+A WK VVGHH IK+ G+HG TQEL QLLPIL+ N +D YING
Sbjct: 188 YISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLYING 247
Query: 255 HDHCLQHI 262
HDHCLQHI
Sbjct: 248 HDHCLQHI 255
>At1g14700 purple acid phosphatase -like protein
Length = 366
Score = 338 bits (867), Expect = 2e-93
Identities = 161/269 (59%), Positives = 207/269 (76%), Gaps = 6/269 (2%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQSLVAHQ 70
++F L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQS VA Q
Sbjct: 41 LVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQSQVALQ 96
Query: 71 MGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYR 130
MG +G+ L+IDFVISTGDNFY +GL + DP F +SF NIYTAPSLQK WYSVLGNHDYR
Sbjct: 97 MGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYSVLGNHDYR 156
Query: 131 GDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVL 190
GDV AQLS +LR D+RWVC+RSFI++ IV+ FFVDTTPF++KYF P +H YDW+GVL
Sbjct: 157 GDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVYDWSGVL 216
Query: 191 PRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDA 250
PR++Y LLK +D+AL +S AKWKIV+GHHTIKS GHHGNT ELE+ LLPIL++N +D
Sbjct: 217 PRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQANEVDL 276
Query: 251 YINGHDHCLQHIIDNERHGEAISSHGTRK 279
Y+NGHDHCL+HI + + + ++S G K
Sbjct: 277 YVNGHDHCLEHISSVDSNIQFMTSGGGSK 305
>At2g01890 purple acid phosphatase like protein
Length = 335
Score = 333 bits (853), Expect = 1e-91
Identities = 162/250 (64%), Positives = 196/250 (77%), Gaps = 2/250 (0%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+IF L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G +LNIDF+ISTGDNFY DG+ D F +SF NIYTA SLQK WY+VLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V AQLS ILR D RW+CLRS++++ IV+ FFVDTTPF+++YF +PK+H YDW GVLPR
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
Y LL +VD+AL +S AKWKIVVGHHTIKS GHHGNT ELE+QLLPIL++N +D YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLQHI 262
NGHDHCL+HI
Sbjct: 250 NGHDHCLEHI 259
>At3g17790 acid phosphatase type 5
Length = 338
Score = 320 bits (820), Expect = 7e-88
Identities = 146/261 (55%), Positives = 200/261 (75%), Gaps = 2/261 (0%)
Query: 7 QRMLSPVIFPASVLLRLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQS 65
+R L S+LL + + ++ +L RF K S++F+V+GDWGR+G++NQS
Sbjct: 5 RRSLMSATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQS 63
Query: 66 LVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLG 125
LVA+QMG +G+ +++DFV+STGDNFY +GL DP F +SF NIYTAPSLQK WYSVLG
Sbjct: 64 LVAYQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLQKQWYSVLG 123
Query: 126 NHDYRGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYD 185
NHDYRGD +AQLSS+LR+ DSRW+CLRSF++D +VE FFVDTTPF+++Y+T+ H+YD
Sbjct: 124 NHDYRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYD 183
Query: 186 WNGVLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKS 245
W V R SY LL++++++L SKA+WKIVVGHH ++S+GHHG+T+EL ++LLPILK
Sbjct: 184 WRAVPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKE 243
Query: 246 NNIDAYINGHDHCLQHIIDNE 266
N +D Y+NGHDHCLQH+ D +
Sbjct: 244 NGVDLYMNGHDHCLQHMSDED 264
>At1g25230 unknown protein
Length = 339
Score = 315 bits (808), Expect = 2e-86
Identities = 148/267 (55%), Positives = 193/267 (71%), Gaps = 1/267 (0%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
++ + LLS + +L +H P S++FLV+GDWGR G YNQS VA QMG
Sbjct: 12 IVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQVALQMG 70
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G+ ++I+FV+STGDN Y +G++ +DDP F SF NIYT+PSLQK WY VLGNHDYRGD
Sbjct: 71 RIGEEMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNHDYRGD 130
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V+AQLS ILR DSRW+C+RSFI+D I E FFVDTTPF++ YF P++ TYDW+GV PR
Sbjct: 131 VEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWSGVSPR 190
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
+SY +L +++ L +S AKWKIVVGHH IKS HGNT+ELE LLPIL++N +D Y+
Sbjct: 191 KSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANKVDLYM 250
Query: 253 NGHDHCLQHIIDNERHGEAISSHGTRK 279
NGHDHCLQHI ++ + ++S G K
Sbjct: 251 NGHDHCLQHISTSQSPIQFLTSGGGSK 277
>At3g52820 purple acid phosphatase (PAP22)
Length = 434
Score = 32.3 bits (72), Expect = 0.36
Identities = 46/239 (19%), Positives = 92/239 (38%), Gaps = 45/239 (18%)
Query: 35 PRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGDNFYKDG 94
P F P + F +VGD G+ + + ++H ++ + D + GD Y D
Sbjct: 128 PEFSFKTPPSTFPVEFAIVGDLGQT-EWTAATLSHI-----NSQDYDVFLLPGDLSYADT 181
Query: 95 LEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSF 154
+ + ++SF + + ++ W GNH+ + ++ + ++RW+
Sbjct: 182 HQPL-----WDSFGRLVEPLASKRPWMVTEGNHEIEFFPIIEHTTF-KSYNARWLM---- 231
Query: 155 ILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGV----------LPRESYRAELLKNVD 204
P E + T +++D GV ES + + L+ D
Sbjct: 232 ---------------PHTESFSTSNLYYSFDVAGVHTVMLGSYTDFDCESDQYQWLQ-AD 275
Query: 205 LALVKSKAK-WKIVVGHHTIKSVG--HHGNTQELEQQLLPILKSNNIDAYINGHDHCLQ 260
LA V K W +V+ H + H G + + + + +L + +D +GH H +
Sbjct: 276 LAKVDRKTTPWVVVLLHAPWYNTNEAHEGEGESMREAMESLLFNARVDVVFSGHVHAYE 334
>At2g46880 unknown protein
Length = 401
Score = 30.4 bits (67), Expect = 1.3
Identities = 15/55 (27%), Positives = 30/55 (54%), Gaps = 7/55 (12%)
Query: 81 DFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQK--IWYSVLGNHDYRGDV 133
D ++ +GDN Y G+ + + +++ AP+++ W ++LGNHD D+
Sbjct: 94 DLIVFSGDNVY-----GLCETSDVAKSMDMAFAPAIESGIPWVAILGNHDQESDM 143
>At2g28880 para-aminobenzoate synthase and glutamine
amidotransferase, a bifunctional enzyme like protein
Length = 919
Score = 30.0 bits (66), Expect = 1.8
Identities = 16/59 (27%), Positives = 30/59 (50%), Gaps = 10/59 (16%)
Query: 231 NTQELEQQLLPILKSNNI----------DAYINGHDHCLQHIIDNERHGEAISSHGTRK 279
+T++LE Q LP++ S+ + YIN C+++I D E + +++ RK
Sbjct: 617 STRKLEDQTLPVIDSSQSKTSFVPDKSREQYINDVQSCMKYIKDGESYELCLTTQNRRK 675
>At5g46640 unknown protein
Length = 386
Score = 29.6 bits (65), Expect = 2.3
Identities = 23/68 (33%), Positives = 33/68 (47%), Gaps = 1/68 (1%)
Query: 19 VLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNL 78
V+LR S+ S I RFE + LN+ V G R GN + +L GIVG ++
Sbjct: 220 VMLRQASHSSGIVTYEGRFEI-ITLSGSVLNYEVNGSTNRSGNLSVALAGPDGGIVGGSV 278
Query: 79 NIDFVIST 86
+ V +T
Sbjct: 279 VGNLVAAT 286
>At3g52810 purple acid phosphatase-like protein
Length = 437
Score = 29.3 bits (64), Expect = 3.0
Identities = 47/229 (20%), Positives = 81/229 (34%), Gaps = 35/229 (15%)
Query: 37 FEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLE 96
F P + + F V GD G+ ++L + + D + GD Y D +
Sbjct: 134 FSFKTPPSKFPIEFAVAGDLGQTDWTVRTLDQIR------KRDFDVFLLPGDLSYADTHQ 187
Query: 97 GVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSFIL 156
+ ++SF + + + W GNH+ S + ++RW+ + L
Sbjct: 188 PL-----WDSFGRLLETLASTRPWMVTEGNHEIESFPTNDHISF-KSYNARWLMPHAESL 241
Query: 157 DGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAEL------LKNVDLALVKS 210
+ + F D HT P ES+ + L+ VD +
Sbjct: 242 SHSNLYYSF-DVAGV----------HTVMLGSYTPYESHSDQYHWLQADLRKVD----RK 286
Query: 211 KAKWKIVVGHHTIKSVG--HHGNTQELEQQLLPILKSNNIDAYINGHDH 257
K W +VV H S H+G +++ L +L +D GH H
Sbjct: 287 KTPWLVVVMHTPWYSTNKAHYGEGEKMRSALESLLYRAQVDVVFAGHVH 335
>At5g63140 unknown protein
Length = 389
Score = 28.9 bits (63), Expect = 3.9
Identities = 19/50 (38%), Positives = 28/50 (56%), Gaps = 8/50 (16%)
Query: 81 DFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSL-QKI-WYSVLGNHD 128
D ++ TGDN + G D +S +N AP++ KI W ++LGNHD
Sbjct: 95 DLIVFTGDNIF-----GFDVKDALKS-INAAFAPAIASKIPWVAILGNHD 138
>At1g10880 hypothetical protein
Length = 651
Score = 28.9 bits (63), Expect = 3.9
Identities = 23/80 (28%), Positives = 36/80 (44%), Gaps = 16/80 (20%)
Query: 194 SYRAELLKNVDLALVKSKAKWKIVVGHHTI-----------KSVGHHGN-TQELEQQLLP 241
SY E ++ + K+ VGHH I KS+ + N +E++QQ+L
Sbjct: 352 SYEGEAMEVAKIGRKKTSRN----VGHHDIVGSKRNQYHYAKSIEENENMVKEMQQQMLQ 407
Query: 242 ILKSNNIDAYINGHDHCLQH 261
I K YI+G ++ L H
Sbjct: 408 IDKEIREKTYISGLEYNLNH 427
>At4g29720 unknown protein
Length = 533
Score = 28.1 bits (61), Expect = 6.7
Identities = 19/64 (29%), Positives = 30/64 (46%), Gaps = 5/64 (7%)
Query: 211 KAKW---KIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNER 267
K+KW + G ++ +VG G+ +L+ P+ K N +NGHD H +
Sbjct: 441 KSKWGSDPLFRGSYSYVAVGSSGD--DLDAMAEPLPKINKKVGQVNGHDQAKVHELQVMF 498
Query: 268 HGEA 271
GEA
Sbjct: 499 AGEA 502
>At3g07680 putative coated vesicle membrane protein
Length = 208
Score = 28.1 bits (61), Expect = 6.7
Identities = 22/106 (20%), Positives = 45/106 (41%), Gaps = 30/106 (28%)
Query: 68 AHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNH 127
+H+ GD L++ FV+ D+ + +GVD + P+ ++I H
Sbjct: 33 SHKAEYEGDTLHVSFVVIKSDSQWHFNEDGVD---------LVIHGPTGEQI-------H 76
Query: 128 DYRGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIE 173
D+R + A+ ++++K G+ F F + +P+ E
Sbjct: 77 DFREQISAKHDFVVQKK--------------GVYRFCFTNKSPYHE 108
>At3g17225 putative protein
Length = 185
Score = 27.7 bits (60), Expect = 8.7
Identities = 15/44 (34%), Positives = 23/44 (52%)
Query: 166 VDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVK 209
+DT I +DP +T D G+L + +LL N +LA +K
Sbjct: 49 IDTNFCITWLISDPTTYTLDLQGLLDLVFQKTQLLGNKNLAAMK 92
>At1g62660 beta-fructosidase (At1g62660) like protein
Length = 648
Score = 27.7 bits (60), Expect = 8.7
Identities = 31/147 (21%), Positives = 55/147 (37%), Gaps = 19/147 (12%)
Query: 92 KDGLEGVDDPTFYESFVNI-YTAPSLQKIWYSVLGNHDYRGDVD-------AQLSSILRQ 143
K+ + + P FY+ + + Y +W ++ H D+ A +
Sbjct: 115 KNWMNDPNGPLFYKGWYHFFYQYNPNAAVWGDIVWGHAVSKDLIHWLYLPIAMVPDQWYD 174
Query: 144 KDSRWVCLRSFILDGGIVEFFFVDTTPFIE----KYFTDPKEH------TYDWNGVL-PR 192
+ W +F+ DG IV + T F++ Y DP + + N VL P
Sbjct: 175 ANGVWTGSATFLDDGSIVMLYTGSTDEFVQVQNLAYPEDPSDPLLLKWVKFSGNPVLVPP 234
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVG 219
A+ ++ A S KW+I +G
Sbjct: 235 PGIGAKDFRDPTTAWKTSSGKWRITIG 261
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,599,538
Number of Sequences: 26719
Number of extensions: 330811
Number of successful extensions: 928
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 916
Number of HSP's gapped (non-prelim): 19
length of query: 322
length of database: 11,318,596
effective HSP length: 99
effective length of query: 223
effective length of database: 8,673,415
effective search space: 1934171545
effective search space used: 1934171545
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146550.4


