

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.3 - phase: 0
(670 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
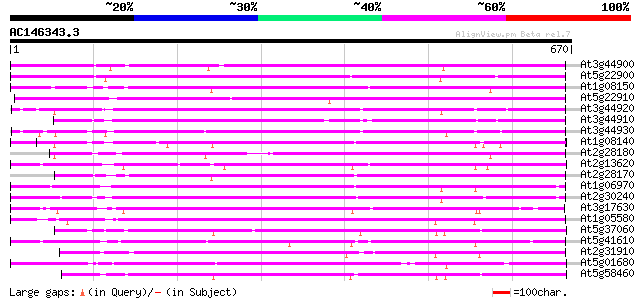
Score E
Sequences producing significant alignments: (bits) Value
At3g44900 putative protein 432 e-121
At5g22900 Na+/H+ antiporter-like protein 412 e-115
At1g08150 hypothetical protein 369 e-102
At5g22910 Na+/H+ antiporter-like protein 348 4e-96
At3g44920 putative protein 348 4e-96
At3g44910 putative protein 348 6e-96
At3g44930 putative protein 345 5e-95
At1g08140 hypothetical protein 345 5e-95
At2g28180 hypothetical protein 344 8e-95
At2g13620 putative Na/H antiporter 333 2e-91
At2g28170 hypothetical protein 332 5e-91
At1g06970 hypothetical protein 301 1e-81
At2g30240 putative Na/H antiporter 298 7e-81
At3g17630 unknown protein 259 3e-69
At1g05580 unknown protein 249 3e-66
At5g37060 putative transporter protein 246 4e-65
At5g41610 Na+/H+ antiporter-like protein 239 5e-63
At2g31910 putative Na+/H+ antiporter 237 1e-62
At5g01680 putative transporter protein 234 9e-62
At5g58460 putative protein 231 8e-61
>At3g44900 putative protein
Length = 817
Score = 432 bits (1111), Expect = e-121
Identities = 260/686 (37%), Positives = 389/686 (55%), Gaps = 31/686 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD ++I TGRKA I S ++ + ++ + + G +I I
Sbjct: 137 MDLSLIRSTGRKAVAIGLSSVLLSITVCALIFFLILRDVGTKKGEPVMSFFEIIFIYLIQ 196
Query: 61 CY--FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGD 118
C F VI +LL +L + NSELGRLA+S+A++ D SI++ + F+ +K D G
Sbjct: 197 CLSSFPVIGNLLFELRLQNSELGRLAMSSAVISDFSTSILSAV-LVFLKELKDDKSRLGS 255
Query: 119 --------GKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFI 170
G +K V ++CF + + RP++ + +K+TP GRP+KK Y+Y + I
Sbjct: 256 VFIGDVIVGNRPMKRAGTVVLFVCFAIY---IFRPLMFFIIKRTPSGRPVKKFYIYAIII 312
Query: 171 IALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMK 230
+ +L KQS+ G I+GL VP GPPLG+ ++++ E P FV + A +
Sbjct: 313 LVFGSAILADWCKQSIFIGPFILGLAVPHGPPLGSAILQKFESVVFGTFLPFFVATSAEE 372
Query: 231 IDLSVY-----VKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKG 285
ID S+ +KS I V + IV K +T + MP D + L+L++S KG
Sbjct: 373 IDTSILQSWIDLKSIVILVSVSFIV-----KFALTTLPAFLYGMPAKDCIALSLIMSFKG 427
Query: 286 VVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSL 345
+ +F + + + +VLS+ +L+ + +K +YDPSR YAGY+KRN+L +
Sbjct: 428 IFEFGAYGYAYQRGTIRPVTFTVLSLYILLNSAVIPPLLKRIYDPSRMYAGYEKRNMLHM 487
Query: 346 KPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ 405
KPNSEL+I+SCI K I P+ N+L+ P+ NP+ ++LHL+ELVG+++PV ISHRLQ
Sbjct: 488 KPNSELRILSCIYKTDDIRPMINLLEATCPSRENPVATYVLHLMELVGQANPVLISHRLQ 547
Query: 406 ERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIII 465
R + SE V+V+F+ F +D G+ VSTYTA+S + MH DIC LAL+ S+II
Sbjct: 548 TRKSENMSYNSENVVVSFEQFHNDFFGSVFVSTYTALSVPKMMHGDICMLALNNTTSLII 607
Query: 466 LPFHLRWSEDGSVESADE-TTRSLNTKVLERAPCSVAILVNRGHSSPFNHNE-----NSK 519
LPFH WS DGS +D R LN VL+ +PCSV I V R + E +S
Sbjct: 608 LPFHQTWSADGSAIVSDSLMIRQLNKSVLDLSPCSVGIFVYRSSNGRRTIKETAANFSSY 667
Query: 520 QIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKN-DEFTSWDVMLDDELLKGV 578
Q+ M+FLGG DDREAL LAKR ++ + V L+S+ + ++ T WD MLD ELL+ V
Sbjct: 668 QVCMLFLGGKDDREALSLAKRMARDSRITITVVSLISSEQRANQATDWDRMLDLELLRDV 727
Query: 579 KGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYP 638
K + ++ + + V + + T++ + IA ++D IVGR G KS T+ L W+E+
Sbjct: 728 KSNVLAGADIVFSEEVVNDANQTSQLLKSIANEYDLFIVGREKGRKSVFTEGLEEWSEFE 787
Query: 639 ELGVLGDLLASPDTNTKASILVVQQQ 664
ELG++GDLL S D N +AS+LV+QQQ
Sbjct: 788 ELGIIGDLLTSQDLNCQASVLVIQQQ 813
>At5g22900 Na+/H+ antiporter-like protein
Length = 822
Score = 412 bits (1060), Expect = e-115
Identities = 248/685 (36%), Positives = 383/685 (55%), Gaps = 25/685 (3%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD +I TGRKA TI S ++ V+ + + + +L +VI
Sbjct: 138 MDTGLIRTTGRKAITIGLSSVLLSTLVCSVIFFGNLRDVGTKNSDHTLNSLEYVVIYSIQ 197
Query: 61 CY--FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTD-----S 113
C F V+ +LL +L + NSELGRLA+S+A++ D SI+ + F+ +K + S
Sbjct: 198 CLSSFPVVGNLLFELRLQNSELGRLAISSAVISDFSTSILASV-LIFMKELKDEQTRLGS 256
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIAL 173
GD + + + F+ + V RP++ + +K+TP GRP+K +Y+ + ++
Sbjct: 257 VFIGDVIAGNRPLMRAGIVVLFVCIAIYVFRPLMFYIIKQTPSGRPVKAIYLSTIIVMVS 316
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
+L KQS+ G I+GL VP GPPLG+ +I++ E P F+ S + +ID+
Sbjct: 317 GSAILANWCKQSIFMGPFILGLAVPHGPPLGSAIIQKYESAIFGTFLPFFIASSSTEIDI 376
Query: 234 SVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNV 293
S + + + I+V + K + T + MPM D L+L++S KG+ +
Sbjct: 377 SALFGWEGLNGIILIMVTSFVVKFIFTTVPALFYGMPMEDCFALSLIMSFKGIFELGAYA 436
Query: 294 FLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKI 353
+ + E +V + + + + ++YLYDPSR YAGY+KRN+ LKPNSEL+I
Sbjct: 437 LAYQRGSVRPETFTVACLYITLNSAIIPPILRYLYDPSRMYAGYEKRNMQHLKPNSELRI 496
Query: 354 VSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSH 413
+SCI + I P+ N+L+ P+ +P+ ++LHL+ELVG+++P+FISH+LQ R +
Sbjct: 497 LSCIYRTDDISPMINLLEAICPSRESPVATYVLHLMELVGQANPIFISHKLQTR-RTEET 555
Query: 414 TFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
++S V+V+F+ F D G+ VSTYTA+S MH DIC LAL+ S+I+LPFH WS
Sbjct: 556 SYSNNVLVSFEKFRKDFYGSVFVSTYTALSMPDTMHGDICMLALNNTTSLILLPFHQTWS 615
Query: 474 EDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFN------------HNENSKQ 520
DGS + S + R+LN VL+ APCSV + V R S N N +S
Sbjct: 616 ADGSALISNNNMIRNLNKSVLDVAPCSVGVFVYRSSSGRKNISSGRKTINGTVPNLSSYN 675
Query: 521 IAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVST-IKNDEFTSWDVMLDDELLKGVK 579
I MIFLGG DDREA+ LA R ++ ++ + L++T K E T WD MLDDELL+ VK
Sbjct: 676 ICMIFLGGKDDREAVTLATRMARDPRINITIVRLITTDEKARENTVWDKMLDDELLRDVK 735
Query: 580 GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
++ ++ Y + +E+ ++T+ + + D IVGR NG S T+ L W+E+ E
Sbjct: 736 S--NTLVDIFYSEKAIEDAAETSSLLRSMVSDFDMFIVGRGNGRTSVFTEGLEEWSEFKE 793
Query: 640 LGVLGDLLASPDTNTKASILVVQQQ 664
LG++GDLL S D N +AS+LV+QQQ
Sbjct: 794 LGIIGDLLTSQDFNCQASVLVIQQQ 818
>At1g08150 hypothetical protein
Length = 815
Score = 369 bits (946), Expect = e-102
Identities = 230/677 (33%), Positives = 366/677 (53%), Gaps = 31/677 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD M+ +TG K + ++P+ +V + +E + E + I+ QS
Sbjct: 151 MDVGMVRKTGTKVIVTGIATVILPIIAANMVFGKLRETGGKYLTGMEYRT---ILFMQSI 207
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I+ LL DL I +SE GR+ +STAMV D TG G + + D +
Sbjct: 208 SAFTGISRLLRDLRINHSEFGRIVISTAMVADG-----TGFGVNLFALVAWM-----DWR 257
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
+ + + Y+ FMV V+RP + W +K+TP+ RP+K+ ++YI+ I+A
Sbjct: 258 VSALQGVGIIGYVIFMV---WVVRPAMFWVIKRTPQERPVKECFIYIILILAFGGYYFLK 314
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS- 239
G ++GL VP GPPLG++++++ E F + L P+F+ ++ID
Sbjct: 315 EIHMFPAVGPFLLGLCVPHGPPLGSQLVEKFESFNTGILLPLFLFFSMLQIDGPWLANQI 374
Query: 240 -------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNN 292
+Y L II+ V + K++ ++ MP+ D +AL+LS KG+V+ C
Sbjct: 375 GQLRHFDGQLYEALTIIIVVFVAKIIFSMIPALLAKMPLTDSFVMALILSNKGIVELCYF 434
Query: 293 VFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELK 352
++ +S +L ++ ++++ +LV T++ + + YLYD S+++ +QKRN++SLK SELK
Sbjct: 435 LYGVESNVLHVKSFTIMATMILVSSTISPVLIHYLYDSSKRFISFQKRNLMSLKLGSELK 494
Query: 353 IVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS 412
+ CI K HI + N+L P + + +++HL+ELVG +PVFISH++Q + +
Sbjct: 495 FLVCIHKADHISGMINLLAQSFPLHESTISCYVIHLVELVGLDNPVFISHQMQ-KAEPGN 553
Query: 413 HTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRW 472
++S V++ FD F+H + S+ +T IS R+MH +I LALDK AS ++LPFH+ W
Sbjct: 554 RSYSNNVLIAFDNFKH-YWKSISLELFTCISNPRYMHQEIYSLALDKQASFLMLPFHIIW 612
Query: 473 SED-GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDD 531
S D +V S D R+ N VL +APCSV I V+R + S ++ IF+GG DD
Sbjct: 613 SLDQTTVVSDDVMRRNANLNVLRQAPCSVGIFVHRQKLLSAQKSSPSFEVCAIFVGGKDD 672
Query: 532 REALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDD----ELLKGVKGVYGSVDN 587
REAL L ++ ++ +L V L+ + T WD MLD E+L+ G
Sbjct: 673 REALALGRQMMRNPNVNLTVLKLIPAKMDGMTTGWDQMLDSAEVKEVLRNNNNTVGQHSF 732
Query: 588 VTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLL 647
V Y + V + SDT+ + IA D +VGR G+ + AL+ WTE+ ELGV+GDLL
Sbjct: 733 VEYVEETVNDGSDTSTLLLSIANSFDLFVVGRSAGVGTDVVSALSEWTEFDELGVIGDLL 792
Query: 648 ASPDTNTKASILVVQQQ 664
S D + S+LVVQQQ
Sbjct: 793 VSQDFPRRGSVLVVQQQ 809
>At5g22910 Na+/H+ antiporter-like protein
Length = 800
Score = 348 bits (894), Expect = 4e-96
Identities = 224/671 (33%), Positives = 354/671 (52%), Gaps = 23/671 (3%)
Query: 6 ITRTGRKAWTIAFCSFVIPMFFGLVVC-YRFQEFWKLEMG-NFEAKNLPVIVIGQSGCYF 63
+ + G++ IA SF + M GL +R + L M VIV Q+
Sbjct: 137 VYQIGKRPIVIAVSSFFVTMISGLAFRNFRLDKVDPLYMPLRLAPTERSVIVSIQAVTLL 196
Query: 64 AVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGTL 123
VI L+ +L++ NSELGR+A+STA V D F +T + +++ + + S +
Sbjct: 197 PVITHLVYELKMSNSELGRIAISTAAVSD-FLGFLTLVCISYVGTYRYVSPGIANR---- 251
Query: 124 KAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGLLTK 183
++ + ++V + +P+ + V TPEG+P+ KVY+Y+ + A+A + +
Sbjct: 252 ----DIVALIILVLVILFIFKPMAQRIVDMTPEGKPVPKVYLYVTILTAIAASIYLSVFN 307
Query: 184 QSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL--SVYVKSDY 241
Q + G +VGL +P+GPPLG+ + + E + FPI + AMK D+ ++Y D
Sbjct: 308 QMYILGALLVGLAIPDGPPLGSALEARFESLVTNIFFPISIAVMAMKADVVRALYSFDDI 367
Query: 242 IY--VWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
+ + LG+ V V V I +C +P + + +A +++ KG VD C
Sbjct: 368 SFNILLLGLTVVVKWTASFVPCLI--FCELPTRESVIIATIMNYKGFVDLCFFDVALRRR 425
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILK 359
LS +V+ I VL+ + +K LYDP RKY GY KR+I+ LK NS+LKI++C+ K
Sbjct: 426 NLSRATYTVMIIYVLLNAGILPTIIKALYDPKRKYIGYVKRDIMHLKTNSDLKILTCLHK 485
Query: 360 PSHIIPIKNVLDICSPTSSNP------LVIHILHLLELVGRSSPVFISHRLQERVGSSSH 413
P +I ++L++ S +N + + LHL++L GR+ P+ I H + + +
Sbjct: 486 PDNISGAISLLELLSSPLNNDNKDRGVIAVTALHLVKLAGRTFPILIPHDKRSKARLLQN 545
Query: 414 TFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
++ + +++ F F+ +N + +VS++TA S M DIC LALD L S+II+P +WS
Sbjct: 546 SYIQTMMLAFTEFQQENWESTTVSSFTAYSHENLMDQDICNLALDHLTSMIIVPSGRKWS 605
Query: 474 EDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDRE 533
DG ES D R +N +L+ APCSV IL RG++ + + +IF+GG DDRE
Sbjct: 606 PDGEYESDDIMIRRVNESLLDLAPCSVGILNYRGYNKGKKKTNSIINVGVIFIGGKDDRE 665
Query: 534 ALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYEKV 593
AL LAK + L V +S + D+ +WD ++DDE+L +K Y +N Y +
Sbjct: 666 ALSLAKWMGQNSRVCLTVIRFLSGQELDKSKNWDYLVDDEVLNDLKATYSLANNFNYMEK 725
Query: 594 EVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTN 653
V + +A HD +IVGR + S LA W E PELGV+GDLLAS D
Sbjct: 726 VVNGGPAVATTVRLVAEDHDLMIVGRDHEDYSLDLTGLAQWMELPELGVIGDLLASKDLR 785
Query: 654 TKASILVVQQQ 664
+ S+LVVQQQ
Sbjct: 786 ARVSVLVVQQQ 796
>At3g44920 putative protein
Length = 671
Score = 348 bits (894), Expect = 4e-96
Identities = 218/683 (31%), Positives = 366/683 (52%), Gaps = 40/683 (5%)
Query: 3 FTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLP---------- 52
F M RT R+ +AF S +P+ G+V F + L N N+
Sbjct: 4 FLMTVRTSRR---VAFHSGKLPVVIGIVSF--FAPLFSLSFLNLFTDNIDPHYMSLDKAL 58
Query: 53 ----VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISS 108
VIVI QS +L +L+I+NSELGRLALS + + D + T +
Sbjct: 59 AERTVIVITQSQILLPSTTYILLELKIINSELGRLALSASAINDMLGIFAMIVATTQATY 118
Query: 109 IKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIV 168
I SH A+ ++ + F ++ V +P+++W + +TPE +P++ +Y++ V
Sbjct: 119 IHV-SH--------AIAYRDLVAVIIFFLIVFFVFKPMVQWIIDRTPEDKPVEDIYIHAV 169
Query: 169 FIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCA 228
+ A A + + G I+G+I+PEGPPLG+ + + E PI +T A
Sbjct: 170 ILTAFASAAYFVFFNMKYVLGPLIIGIIIPEGPPLGSALEAKFERLTMNVFLPISITFSA 229
Query: 229 MKID-LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVV 287
M+ D L + + IY + + + + + K++ + +C Y +P ++ L ++L+LS K V
Sbjct: 230 MRCDGLRILSQFTDIYFNIFLTLLILVIKLVACLTLCLYYKLPRSESLAVSLILSYKSFV 289
Query: 288 DFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP 347
+F + + +S + L + L+ + + V+ +YDP RKY YQKR+IL L+
Sbjct: 290 EFVLYEAVLEEKFISQATYAFLILYSLLSAGIVPMVVRSMYDPKRKYVNYQKRDILHLEA 349
Query: 348 NSELKIVSCILKPSHIIPIKNVLDI-CSPTSSNPLVIHILHLLELVGRSSPVFISH-RLQ 405
NS L+I++C+ KP ++ L + SP P+ + +LHL++LVG+ +P+ +SH +
Sbjct: 350 NSGLRILTCLHKPENVSETIAFLQLFSSPIHDFPIAVTVLHLVKLVGQINPIIVSHDKKL 409
Query: 406 ERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIII 465
+R+ +S+ + + F F ++ + +V+T+TA S MH+DIC LALD+ S+I+
Sbjct: 410 KRLHKNSYIHTANL--AFRQFMQESLESVTVTTFTAFSHENLMHEDICTLALDRTTSMIV 467
Query: 466 LPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH----NENSKQI 521
+P +W+ DG ES D R LN +L+RAPCS+ ILV+RG S ++ N + +
Sbjct: 468 VPSGRKWTVDGMFESDDLAARQLNQSLLDRAPCSIGILVDRGQFSRKSYVTSKNRYNIDV 527
Query: 522 AMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGV 581
++F+GG DDREAL L KR + V L+ ++ + WD +LD+E LK +K
Sbjct: 528 GVLFIGGKDDREALSLVKRMKYNPRVRVTVIRLI--FDHEIESEWDYILDNEGLKDLKST 585
Query: 582 YGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELG 641
+ D + E++ V + + + + +A ++D ++VGR + + S L W E PELG
Sbjct: 586 ESNEDILYTERI-VTSVVEVVKAVQLLAEEYDLMVVGRDHDMTSQDLSGLTEWVELPELG 644
Query: 642 VLGDLLASPDTNTKASILVVQQQ 664
V+GDLLA+ D N+K S+LVVQQQ
Sbjct: 645 VIGDLLAARDLNSKVSVLVVQQQ 667
>At3g44910 putative protein
Length = 705
Score = 348 bits (893), Expect = 6e-96
Identities = 205/614 (33%), Positives = 340/614 (54%), Gaps = 24/614 (3%)
Query: 53 VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTD 112
V++ QS + LS+L+ILNSELGRL LS +++ D F S V+ I + + K
Sbjct: 111 VVISSQSSILLPTVVHFLSELKILNSELGRLVLSASLINDIFASTVS-IFAYLVGTYKNI 169
Query: 113 SHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIA 172
S + A+ ++ + ++V VLRP+++W V++TPEG+P+ VY++ V +
Sbjct: 170 S--------PMTAYRDLIAVIILILVAFCVLRPVVEWIVERTPEGKPVADVYVHAVVLSV 221
Query: 173 LAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKID 232
+A L G ++G+I+PEGPP+G+ + + E L PI +T M+ D
Sbjct: 222 IASAAYSSFFNMKYLLGPFLLGIIIPEGPPIGSALEAKYEALTMNVLIPISITFSTMRCD 281
Query: 233 -LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCN 291
+ + + D I+ + ++ KM + C YC +P + + +L+L K +
Sbjct: 282 VMKIVYQYDDIWYNIFLMTFTGFLKMATGMVPCLYCKIPFKEAIAASLLLCSKSFSEIFL 341
Query: 292 NVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSEL 351
+D +S + L L+ + + LYDP RKY GYQK+NI++LKP+S+L
Sbjct: 342 YESTYDDSYISQATYTFLITCALINSGIIPTALAGLYDPKRKYVGYQKKNIMNLKPDSDL 401
Query: 352 KIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ-ERVGS 410
+I++CI +P +I + L T +V+ +LHL++LVG++ PV ISH Q RV +
Sbjct: 402 RILTCIHRPENISAAISFLQFLPST----IVVTVLHLVKLVGKTVPVLISHNKQINRVVT 457
Query: 411 SSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHL 470
+S+ + + F E + +++ +TAI+ MHD+IC +AL++ SIII+P
Sbjct: 458 NSYIHTANL--AFSQLE-----SVTMTMFTAITHENLMHDEICKVALEQATSIIIVPSGR 510
Query: 471 RWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSD 530
+W+ DG+ ES DE R LN +L+ A CS+ ILV+RG S + + + +IF+GG D
Sbjct: 511 KWTVDGAFESEDEAIRRLNESLLKSASCSIGILVDRGQLSLKGTRKFNIDVGVIFIGGKD 570
Query: 531 DREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTY 590
DREAL L K+ + + V L+S + E T+WD +LD E+L+ +K + +++ Y
Sbjct: 571 DREALSLVKKMKQNPRVKITVIRLISD-RETESTNWDYILDHEVLEDLKDT-EATNSIAY 628
Query: 591 EKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASP 650
+ V + + ++ +D ++VGR +G+ SP L W E PELGV+GDLLAS
Sbjct: 629 TERIVTGGPEVATTVRSLSEDYDLMVVGRDHGMASPDFDGLMEWVELPELGVIGDLLASR 688
Query: 651 DTNTKASILVVQQQ 664
+ +++ S+LVVQQQ
Sbjct: 689 ELDSRVSVLVVQQQ 702
>At3g44930 putative protein
Length = 731
Score = 345 bits (885), Expect = 5e-95
Identities = 222/689 (32%), Positives = 371/689 (53%), Gaps = 52/689 (7%)
Query: 3 FTMITRTGRKAWTIAFCSFVIPMFFGLVVCYR------FQEFWKLEMGNFEAKNLPV--- 53
F M RT R+ +AF S +P+ G+V + FQ F+ N + +P+
Sbjct: 64 FLMTVRTSRR---VAFHSGKLPVVIGIVSFFAPLFGLGFQNFFS---DNIDPHYMPLTKA 117
Query: 54 ------IVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFIS 107
IVI QS +L +L+I+NSELGRLALS ++ D + GI + ++
Sbjct: 118 LGERTAIVITQSSILLPSTTYILLELKIINSELGRLALSACVIND-----ILGIFSMIVA 172
Query: 108 SIKTD----SHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKV 163
SI+ SH A+ + + F +V LV +P+++W + +TPE +P++ +
Sbjct: 173 SIQATYIHVSHAT--------AYRDTVAVIIFFLVVFLVFKPMVQWVIDRTPEDKPVEDM 224
Query: 164 YMYIVFIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIF 223
Y++ V I ALA + + G ++G+I+PEGPPLG+ + + E PI
Sbjct: 225 YIHAVIITALASAAYFVFFNMKYILGPLMIGIIIPEGPPLGSALEAKFERLTMNVFLPIS 284
Query: 224 VTSCAMKIDLSVYVKSDYIYVWLGIIVA--VHLFKMLVTIGICWYCNMPMADGLCLALML 281
+T AM+ D + S + ++ I + + + K++ + C Y +P+++ L ++ +L
Sbjct: 285 ITFSAMRCD-GARILSQFNDIFFNIFLTFLILVIKLVACLAPCLYYKLPLSESLAVSFIL 343
Query: 282 SCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRN 341
S K DF + D +S S L + L+ + ++ +YDP RKY YQKR+
Sbjct: 344 SYKSFADFVLYEAVLDDTYISQATYSFLILYSLLNAGIVPTVLRRMYDPRRKYVNYQKRD 403
Query: 342 ILSLKPNSELKIVSCILKPSHIIPIKNVLDICS-PTSSNPLVIHILHLLELVGRSSPVFI 400
IL L+ NS+L+I++C+ KP ++ L + S P P+ + +LHL++LVG+ +P+ +
Sbjct: 404 ILHLERNSDLRILTCLHKPENVSETIAFLQLLSSPNLDFPIAVTVLHLVKLVGQINPIIV 463
Query: 401 SH-RLQERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDK 459
SH + +R+ S+ + + F F ++ + +V+T+TA S MH+DIC LALDK
Sbjct: 464 SHDKKLKRLNKDSYIHTANL--AFRQFVLESLESVTVTTFTAFSHENLMHEDICTLALDK 521
Query: 460 LASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSK 519
S+I++P +W+ DG ES + R LN +L+RAPCS+ ILV+RG S + + K
Sbjct: 522 TTSMIVVPSGRKWTVDGLFESDNTAIRHLNQSLLDRAPCSIGILVDRGQFSRKSIVTSKK 581
Query: 520 Q----IAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELL 575
+ + ++F+GG DDREAL L KR + V LV ++ + WD +LD+E L
Sbjct: 582 RYIIDVGVLFIGGKDDREALSLVKRMKNNPRIRVTVIRLV--FDHEIESDWDYILDNEGL 639
Query: 576 KGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWT 635
K +K + D + Y + V ++ + + + +A ++D ++VGR + + S L W
Sbjct: 640 KDLKSTEDNKD-IDYIERIVTSSVEVVKAVQLLAEEYDLMVVGRDHDMTSQDLSGLMEWV 698
Query: 636 EYPELGVLGDLLASPDTNTKASILVVQQQ 664
E PELGV+GDLLA+ D ++K S+LVVQQQ
Sbjct: 699 ELPELGVIGDLLAARDLSSKVSVLVVQQQ 727
>At1g08140 hypothetical protein
Length = 1536
Score = 345 bits (885), Expect = 5e-95
Identities = 228/660 (34%), Positives = 355/660 (53%), Gaps = 48/660 (7%)
Query: 33 YRFQEFWKLEMGNFEAKNLP------VIVIGQSGCYFAVIASLLSDLEILNSELGRLALS 86
+ F +W L+ +A+ LP +I+ QS F I +LL DL+I +SE GR+ALS
Sbjct: 130 FGFVLYWFLKGVTMDAE-LPFRTEKRLIIFLQSISAFTSIDTLLKDLQIKHSEFGRIALS 188
Query: 87 TAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPI 146
AMV D + G F ++I + L F+ + F+VV V+RP
Sbjct: 189 GAMVTD-----MLAFGVTFFNAIYYEK---------LYGFMQTVGFCLFVVVMICVVRPA 234
Query: 147 LKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGLLTKQSVL---GGICIVGLIVPEGPPL 203
+ W +K+TPEGRP+K Y+Y +F IA A K L G + GL VP G PL
Sbjct: 235 MYWVIKQTPEGRPVKDFYLYSIFGIAFAC--FTFFNKVIHLFGPAGSFVFGLTVPNGYPL 292
Query: 204 GTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSD--------YIYVWLGIIVAVHLF 255
GT +I++ E F + P+F + M++DL K IY + I+ V+
Sbjct: 293 GTTLIQKFESFNLGSILPLFGSLTMMQVDLLRLFKESGDLIRMEGQIYEVISFILLVNTT 352
Query: 256 KMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLV 315
K +VT + MP+ D LAL+LS KG+ + + + L+ E ++L+ L+
Sbjct: 353 KFVVTTITAYAFKMPLRDSFALALVLSNKGIFELAYYTYAVELKLIRPEVFTILAAYTLL 412
Query: 316 IGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKNVLDICSP 375
+ ++ ++DP++++ Y+KRN+ LK + L+ + C+ +P HI + ++L+ SP
Sbjct: 413 NSIFIPMLLELVHDPTKRFRCYRKRNLGILKDGAALQCLMCVYRPDHITSMTDLLETFSP 472
Query: 376 TSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEAVIVTFDLFEHDNAGTAS 435
+ +P+ +ILHL+ELVG+++P+FISH+LQ+ S+ + S+ VI++F F+ S
Sbjct: 473 SQDSPMACNILHLVELVGQANPMFISHQLQKPEPGST-SLSDNVIISFRGFQRQFFEYTS 531
Query: 436 VSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGS-VESADETTRSLNTKVLE 494
+ +T++S + MH+DIC+LAL + S+I+LPFH WS D S V S D+ R LN VL
Sbjct: 532 LDIFTSVSVSQHMHEDICWLALSRSLSLIVLPFHRTWSVDRSTVISNDDNLRMLNVNVLR 591
Query: 495 RAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAKR-TIKEDTYHLVVYH 553
RAPCSV I V R + ++ +I +IF GG DDREAL + R + E L +
Sbjct: 592 RAPCSVGIFVYRKPIVESHMAKSHSKICLIFNGGKDDREALAITNRMRLTEKRTRLTIIR 651
Query: 554 LV---STIKNDEFTSWDVMLDDELLKGVKGVYGS-----VDNVTYEKVEVENTSDTTEFI 605
+ S + NDE W+ L + V + GS VTY V + S+T+ +
Sbjct: 652 FIPKSSEMDNDE---WEQQQSINLKESVTSIVGSNIKENDAKVTYIDKAVSDGSETSRIL 708
Query: 606 SDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTNTKASILVVQQQV 665
+A +D IVG +GI + T ++ WTE+ ELG +GDLLAS + + AS+LVVQ+QV
Sbjct: 709 RAMANDYDLFIVGSGSGIGTEATSGISEWTEFNELGPIGDLLASHEYPSSASVLVVQKQV 768
Score = 285 bits (728), Expect = 8e-77
Identities = 201/671 (29%), Positives = 339/671 (49%), Gaps = 64/671 (9%)
Query: 2 DFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSGC 61
D ++ ++G K+ I S +IP G ++ ++ L M E V+ S
Sbjct: 921 DVGIMKKSGTKSVVIGITSMIIPWQIGKLLYSSREKSSILTMTEME---YTVMTFTMSMT 977
Query: 62 YFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKG 121
F + LL+DL+I++++ G++A S MV D +T +A++S +T G
Sbjct: 978 PFTCVNMLLTDLKIVHTDFGQIAQSAGMVTDLLAFFLTV--SAYVSRDETQGVKMG---- 1031
Query: 122 TLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIA-LAVGMLGL 180
AF+ F ++ ++R + W ++ TPEG P+K VY+YI ++A L+
Sbjct: 1032 --LAFMAFFIFV-------YLVRQFMLWVIRHTPEGAPVKNVYLYIGLLLAYLSYLYWSR 1082
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSD 240
LG + GL VP GPPLG+E L
Sbjct: 1083 FLFFGPLGAFAL-GLAVPNGPPLGSEFGNGRHLH-------------------------G 1116
Query: 241 YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSML 300
++Y + V++ K + +P+ D + L +++ K + + +
Sbjct: 1117 HMYECFSFLPIVYIAKFATSFLAALATKIPLRDSIILGVIMGTKSSFELGYVLTAFEKDR 1176
Query: 301 LSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILKP 360
+S E LS+L + +LV L + + +LYD S+++ Y +RN LK E++ + CI KP
Sbjct: 1177 ISLEVLSLLGVYILVNSLLTPMAIHFLYDRSKRFVCYGRRN---LKEKPEMQTLVCINKP 1233
Query: 361 SHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEAVI 420
+I + ++L SP+ +P+ +LHL+EL+G+++P FISH+LQ + S ++SE VI
Sbjct: 1234 DNITSMISLLRATSPSKDSPMECCVLHLIELLGQATPTFISHQLQ-KPKPGSRSYSENVI 1292
Query: 421 VTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGSVE- 479
+F LF+ +AS++ +T+++ + MH+ IC+ AL + +++I+L FH W +G+V
Sbjct: 1293 SSFQLFQEVYWDSASINMFTSLTSAKEMHEQICWFALSQGSNLILLSFHRTWEPNGNVII 1352
Query: 480 SADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAK 539
S D+T RSLN VL+RAPCSV I V R E+ ++ +I++GG+DD+EAL LA
Sbjct: 1353 SDDQTLRSLNLNVLKRAPCSVGIFVYRKPIWQTKALESPCRVCLIYVGGNDDKEALALAD 1412
Query: 540 RTIKEDTYHLVVYHLVSTIKNDEFT------SWDVMLDDELLKGVKGVYGSVDNVTYEKV 593
L V L+ T DE + D+ ++ G D T
Sbjct: 1413 HMRGNQQVILTVLRLIPTSYADESSLRIHSQMVDMNRHEDQRPG--------DKSTIIDW 1464
Query: 594 EVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTN 653
V + ++T++ + ++ +D IVGRR+G+ + T+ L W E+ ELGV+GDLLAS
Sbjct: 1465 TVGDGTETSKILHSVSYDYDLFIVGRRSGVGTTVTRGLGDWMEFEELGVIGDLLASEYFP 1524
Query: 654 TKASILVVQQQ 664
++AS+LVVQQQ
Sbjct: 1525 SRASVLVVQQQ 1535
>At2g28180 hypothetical protein
Length = 822
Score = 344 bits (883), Expect = 8e-95
Identities = 215/637 (33%), Positives = 346/637 (53%), Gaps = 57/637 (8%)
Query: 48 AKNLP-------VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTG 100
AKN P V+++ +S F+ IA LL DL + +S +GR+ALS+A+V D ++
Sbjct: 216 AKNRPLTFQEYDVMLLMESITSFSGIARLLRDLGMNHSSIGRVALSSALVSDIVGLLL-- 273
Query: 101 IGTAFISSIKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPM 160
I+++ S DG L F+V+ V+RPI+ +K+ EGRP+
Sbjct: 274 ----LIANVSRSSATLADGLAILTEIT------LFLVIAFAVVRPIMFKIIKRKGEGRPI 323
Query: 161 KKVYMYIVFIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLF 220
+ Y++ V ++ M Q G +GL +P GPP+G+ ++++LE F +
Sbjct: 324 EDKYIHGVLVLVCLSCMYWEDLSQFPPLGAFFLGLAIPNGPPIGSALVERLESFNFGIIL 383
Query: 221 PIFVTSCAMKID-------LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMAD 273
P+F+T+ ++ D L+ + D + +++ + L K+ V++ + + MP+ D
Sbjct: 384 PLFLTAVMLRTDTTAWKGALTFFSGDDKKFAVASLVLLIFLLKLSVSVIVPYLYKMPLRD 443
Query: 274 GLCLALMLSCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRK 333
+ LAL++S K VLSI ++ L + + +LYDPS++
Sbjct: 444 SIILALIMSHK-----------------------VLSI--VLNSLLIPMAIGFLYDPSKQ 478
Query: 334 YAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVG 393
+ YQKRN+ S+K ELK + CI +P HI + N+L+ + +PL ++LHL+EL G
Sbjct: 479 FICYQKRNLASMKNMGELKTLVCIHRPDHISSMINLLEASYQSEDSPLTCYVLHLVELRG 538
Query: 394 RSSPVFISHRLQERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDIC 453
+ P ISH++Q+ + + +SE VI++F+ F + S+ T+T I+ M DDIC
Sbjct: 539 QDVPTLISHKVQKLGVGAGNKYSENVILSFEHFHRSVCSSISIDTFTCIANANHMQDDIC 598
Query: 454 YLALDKLASIIILPFHLRWSED-GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPF 512
+LALDK ++IILPFH WS D S+ S E R LN VL++APCSV IL+ R +
Sbjct: 599 WLALDKAVTLIILPFHRTWSLDRTSIVSDVEAIRFLNVNVLKQAPCSVGILIERHLVNKK 658
Query: 513 NHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDD 572
S ++ +IF+GG DDREAL AKR +++ L V L+++ K+ + T WD MLD
Sbjct: 659 QEPHESLKVCVIFVGGKDDREALAFAKRMARQENVTLTVLRLLASGKSKDATGWDQMLDT 718
Query: 573 ----ELLKGVK-GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQ 627
EL+K G+ + Y + E+ + +DT+ + +A +D +VGR G
Sbjct: 719 VELRELIKSNNAGMVKEETSTIYLEQEILDGADTSMLLRSMAFDYDLFVVGRTCGENHEA 778
Query: 628 TQALASWTEYPELGVLGDLLASPDTNTKASILVVQQQ 664
T+ + +W E+ ELGV+GD LASPD +K S+LVVQQQ
Sbjct: 779 TKGIENWCEFEELGVIGDFLASPDFPSKTSVLVVQQQ 815
>At2g13620 putative Na/H antiporter
Length = 821
Score = 333 bits (853), Expect = 2e-91
Identities = 214/693 (30%), Positives = 354/693 (50%), Gaps = 49/693 (7%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD ++ +TG++A TIA V+P G + + E + + + + S
Sbjct: 119 MDIMVVRKTGKRALTIAIGGMVLPFLIGAAFSFSMH---RSEDHLGQGTYILFLGVALSV 175
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L++L+++N+E+GR+++S A+V D F I+ + A S KT
Sbjct: 176 TAFPVLARILAELKLINTEIGRISMSAALVNDMFAWILLALAIALAESDKT--------- 226
Query: 121 GTLKAFLNVFYYLC---FMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGM 177
+F +++ + F+ V V+RP + W ++KTPEG + ++ ++ + G
Sbjct: 227 ----SFASLWVMISSAVFIAVCVFVVRPGIAWIIRKTPEGENFSEFHICLILTGVMISGF 282
Query: 178 LGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
+ + G + GL++P GP LG +I++LE F S L P+F +K +++
Sbjct: 283 ITDAIGTHSVFGAFVFGLVIPNGP-LGLTLIEKLEDFVSGLLLPLFFAISGLKTNIAAIQ 341
Query: 238 KSDYIYVWLGIIVAVHLF---KMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVF 294
WL + + + L K++ T+ + ++ MP+ +G+ L L+L+ KG+V+
Sbjct: 342 GPA---TWLTLFLVIFLACAGKVIGTVIVAFFHGMPVREGITLGLLLNTKGLVEMIVLNV 398
Query: 295 LHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIV 354
D +L E + + + LV+ + V LY P +K Y++R I KP+SEL+++
Sbjct: 399 GKDQKVLDDETFATMVLVALVMTGVITPIVTILYKPVKKSVSYKRRTIQQTKPDSELRVL 458
Query: 355 SCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQER---VGSS 411
C+ P ++ I N+L+ PT +P+ I++LHL+EL GR+S + I H ++ +
Sbjct: 459 VCVHTPRNVPTIINLLEASHPTKRSPICIYVLHLVELTGRASAMLIVHNTRKSGRPALNR 518
Query: 412 SHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLR 471
+ S+ +I F+ +E +A +V TAISP MH+D+C LA DK S II+PFH +
Sbjct: 519 TQAQSDHIINAFENYEQ-HAAFVAVQPLTAISPYSTMHEDVCSLAEDKRVSFIIIPFHKQ 577
Query: 472 WSEDGSVESADETTRSLNTKVLERAPCSVAILVNRG--HSSPFNHNENSKQIAMIFLGGS 529
+ DG +ES + R +N +LE +PCSV ILV+RG ++ N N S Q+A++F GG
Sbjct: 578 QTVDGGMESTNPAYRLVNQNLLENSPCSVGILVDRGLNGATRLNSNTVSLQVAVLFFGGP 637
Query: 530 DDREALCLAKRTIKEDTYHLVVYHLV----------STIKNDEFTSWDVM-------LDD 572
DDREAL A R + L V + + ND M LDD
Sbjct: 638 DDREALAYAWRMAQHPGITLTVLRFIHDEDEADTASTRATNDSDLKIPKMDHRKQRQLDD 697
Query: 573 ELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALA 632
+ + + +++ Y + V N +T + + HD IVGR G+ SP T L
Sbjct: 698 DYINLFRAENAEYESIVYIEKLVSNGEETVAAVRSMDSSHDLFIVGRGEGMSSPLTAGLT 757
Query: 633 SWTEYPELGVLGDLLASPDTNTKASILVVQQQV 665
W+E PELG +GDLLAS D S+LVVQQ V
Sbjct: 758 DWSECPELGAIGDLLASSDFAATVSVLVVQQYV 790
>At2g28170 hypothetical protein
Length = 617
Score = 332 bits (850), Expect = 5e-91
Identities = 205/624 (32%), Positives = 346/624 (54%), Gaps = 30/624 (4%)
Query: 54 IVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDS 113
I++ QS FA + LL+DL+I +SE GR+ S A V D I+ GT + K
Sbjct: 9 IILLQSLSSFAGVNGLLTDLKINHSEFGRMVQSCAAVTDLVIFIMVS-GTVLLKGQKGLP 67
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIAL 173
H + + + F+V ++ P++ W +K+TPEGR +K VY+Y+V A
Sbjct: 68 HG-----------IVIVLVIGFLVY---IVWPVMLWIIKQTPEGRLVKDVYIYLVMATAY 113
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
V M L Q G I+GL P GPPLG+ +I++ E F L P+F + ++D+
Sbjct: 114 FVYMFWLNFFQFSTYGWFIIGLATPAGPPLGSALIQRFECFNVGVLLPLFGSLSMEQLDI 173
Query: 234 SVYVKS--------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKG 285
S ++ + Y + +I+ V + K +VT + +P D + LA++LS +
Sbjct: 174 SWLMREILNLKHMEGFAYEAISVILIVTVVKFVVTAITAFAVRIPYRDSIVLAMVLSNRS 233
Query: 286 VVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSL 345
+ + ++ + + +++ ++ +++VLV L I ++++Y+P ++ Y+ RN+L+L
Sbjct: 234 IFELGYLGYIVELKMFDNKSFTIAALSVLVSSLLTPIAIEFMYEPQHIFSSYRDRNMLTL 293
Query: 346 KPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ 405
K +S+LK + CI KP HI + N +++ +PT + L ++LHL+EL+G++ P FISH++Q
Sbjct: 294 KHDSKLKTLVCIHKPDHITSMVNFVELFNPTQESKLECNVLHLVELIGQAIPTFISHKMQ 353
Query: 406 E-RVGSSSHTFSEAVIVTF-DLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASI 463
+ +VG+ S S VI F L H S+ +T+ S V MH+D+C+LALDK ++
Sbjct: 354 KPKVGTRS--CSRNVITAFLSLRRHLTKEAISIDIFTSASLVEHMHEDLCWLALDKNVAL 411
Query: 464 IILPFHLRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIA 522
++LPFH WS D S + S D+ ++LN KVL+RA CSV I V R + + ++
Sbjct: 412 VVLPFHRSWSVDRSTIVSDDKAMQNLNHKVLKRASCSVGIFVYRKPLWESQMHGSCYKVC 471
Query: 523 MIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVY 582
I +GG DD+EAL R + + + HL+ + +E LD + +K +
Sbjct: 472 AIVVGGKDDKEALAFTNRMRRNKQTSVTILHLIPQLTTEESEDSVQKLDYDDIKEIMKTE 531
Query: 583 GSVDNVTYEKVE--VENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPEL 640
S +N ++ +E V+ ++T+ + IA +D IVGR +G+ S T+ L WTE+ EL
Sbjct: 532 DSNENDSWICIEKSVKEGAETSVILRSIAYDYDLFIVGRSSGMNSAVTKGLNEWTEFEEL 591
Query: 641 GVLGDLLASPDTNTKASILVVQQQ 664
G LGD++AS + ++AS+LV+QQQ
Sbjct: 592 GALGDVIASKEFPSRASVLVLQQQ 615
>At1g06970 hypothetical protein
Length = 829
Score = 301 bits (770), Expect = 1e-81
Identities = 185/672 (27%), Positives = 345/672 (50%), Gaps = 26/672 (3%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D ++I + G KA I S+ +P G + + + L + ++ +
Sbjct: 132 IDASIIRKAGSKAILIGTASYALPFSLGNLTVLFLKNTYNLPPDVVHC--ISTVISLNAM 189
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V ++L++L ILNS+LGRLA + ++V ++F+ IV + F+
Sbjct: 190 TSFPVTTTVLAELNILNSDLGRLATNCSIVCEAFSWIVALVFRMFLRD------------ 237
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMK-KVYMYIVFIIALAVGMLG 179
GTL + + + ++V V RP + W ++ ++ + + ++ L + +
Sbjct: 238 GTLASVWSFVWVTALILVIFFVCRPAIIWLTERRSISIDKAGEIPFFPIIMVLLTISLTS 297
Query: 180 LLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS 239
+ G +G+ +P+GPPLGT + +LE+F + + P F++ ++ + + +S
Sbjct: 298 EVLGVHAAFGAFWLGVSLPDGPPLGTGLTTKLEMFATSLMLPCFISISGLQTNFFIIGES 357
Query: 240 DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
++ + +I+ + K L T YCN+ + D LAL++ C+GV++ V D
Sbjct: 358 -HVKIIEAVILITYGCKFLGTAAASAYCNIQIGDAFSLALLMCCQGVIEIYTCVMWKDEK 416
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP-NSELKIVSCIL 358
+L++E ++L I +L++ ++R V LYDPS++Y KR IL + N + +++ C+
Sbjct: 417 VLNTECFNLLIITLLLVTGISRFLVVCLYDPSKRYRSKSKRTILDTRQRNLQFRLLLCVY 476
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEA 418
++ + N+L+ P+ +P+ + LHL+EL GR+ V + H ++ ++ S
Sbjct: 477 NVENVPSMVNLLEASYPSRFSPISVFTLHLVELKGRAHAVLVPHHQMNKLDPNT-VQSTH 535
Query: 419 VIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGSV 478
++ F FE N GT +TA +P ++DDIC LALDK A++I++PFH +++ DG+V
Sbjct: 536 IVNGFQRFEQQNQGTLMAQHFTAAAPFSSINDDICTLALDKKATLIVIPFHKQYAIDGTV 595
Query: 479 ESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH---NENSKQIAMIFLGGSDDREAL 535
+ + + R++N VLE+APCSV I ++RG + + + +A+IF+ G DD EAL
Sbjct: 596 DHVNPSIRNINLNVLEKAPCSVGIFIDRGETEGRRSVLMSYTWRNVAVIFIEGRDDAEAL 655
Query: 536 CLAKRTIKEDTYHLVVYHL---VSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYEK 592
+ R + + + H S +N + + L+ K S ++Y +
Sbjct: 656 AFSMRIAEHPEVSVTMIHFRHKSSLQQNHVVDVESELAESYLINDFKNFAMSKPKISYRE 715
Query: 593 VEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDT 652
V + +TT+ IS + D ++VGR + ++S L W+E PELGV+GD+ AS D
Sbjct: 716 EIVRDGVETTQVISSLGDSFDLVVVGRDHDLESSVLYGLTDWSECPELGVIGDMFASSDF 775
Query: 653 NTKASILVVQQQ 664
+ S+LV+ QQ
Sbjct: 776 H--FSVLVIHQQ 785
>At2g30240 putative Na/H antiporter
Length = 831
Score = 298 bits (763), Expect = 7e-81
Identities = 191/673 (28%), Positives = 343/673 (50%), Gaps = 29/673 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D ++I + G KA I S+ P G + + L + + + +
Sbjct: 134 IDGSIIRKAGSKAILIGTASYAFPFSLGNLTIMFISKTMGLPSDVISCTSSAISLSSMTS 193
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V ++L++L ILNSELGRLA +MV + + V A ++ T
Sbjct: 194 --FPVTTTVLAELNILNSELGRLATHCSMVCEVCSWFV-----ALAFNLYTRDR------ 240
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML-G 179
T+ + + + ++V V RPI+ W ++ + K V + ++ L++ L G
Sbjct: 241 -TMTSLYALSMIIGLLLVIYFVFRPIIVWLTQRKTKSMDKKDVVPFFPVLLLLSIASLSG 299
Query: 180 LLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS 239
G +G+ +P+GPPLGTE+ +LE+F S P F+ ++ + +S
Sbjct: 300 EAMGVHAAFGAFWLGVSLPDGPPLGTELAAKLEMFASNLFLPCFIAISGLQTNFFEITES 359
Query: 240 -DYIYVWLGIIVAV-HLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHD 297
++ V + II+ + + K L T YC + D LCLA ++ C+G+++ + D
Sbjct: 360 HEHHVVMIEIILLITYGCKFLGTAAASAYCQTQIGDALCLAFLMCCQGIIEVYTTIVWKD 419
Query: 298 SMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP-NSELKIVSC 356
+ ++ +E +++ I +L + ++R V YLYDPS++Y KR IL+ + N +L+++
Sbjct: 420 AQVVDTECFNLVIITILFVTGISRFLVVYLYDPSKRYKSKSKRTILNTRQHNLQLRLLLG 479
Query: 357 ILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFS 416
+ ++ + N+L+ PT NP+ LHL+EL GR+ + H ++ ++ S
Sbjct: 480 LYNVENVPSMVNLLEATYPTRFNPISFFTLHLVELKGRAHALLTPHHQMNKLDPNTAQ-S 538
Query: 417 EAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDG 476
++ F FE G +TA +P +++DIC LALDK A++I++PFH +++ DG
Sbjct: 539 THIVNAFQRFEQKYQGALMAQHFTAAAPYSSINNDICTLALDKKATLIVIPFHKQYAIDG 598
Query: 477 SVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH---NENSKQIAMIFLGGSDDRE 533
+V + R++N VL+ APCSVAI ++RG + + +AM+F+GG DD E
Sbjct: 599 TVGQVNGPIRTINLNVLDAAPCSVAIFIDRGETEGRRSVLMTNTWQNVAMLFIGGKDDAE 658
Query: 534 ALCLAKRTIKEDTYHLVVYHL--VSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYE 591
AL L R ++ ++ + H S +++++++ M + L+ K + + Y
Sbjct: 659 ALALCMRMAEKPDLNVTMIHFRHKSALQDEDYSD---MSEYNLISDFKSYAANKGKIHYV 715
Query: 592 KVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPD 651
+ V + +TT+ IS + +D ++VGR + ++S L W+E PELGV+GD+L SPD
Sbjct: 716 EEIVRDGVETTQVISSLGDAYDMVLVGRDHDLESSVLYGLTDWSECPELGVIGDMLTSPD 775
Query: 652 TNTKASILVVQQQ 664
+ S+LVV QQ
Sbjct: 776 FH--FSVLVVHQQ 786
>At3g17630 unknown protein
Length = 857
Score = 259 bits (662), Expect = 3e-69
Identities = 186/683 (27%), Positives = 343/683 (49%), Gaps = 47/683 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIV---IG 57
+DF I +TG+K+ IA +P G+ + + G LP IV +
Sbjct: 171 LDFAAIKKTGKKSLLIAIAGISLPFIVGVGTSFVLSA--TISKG---VDQLPFIVFMGVA 225
Query: 58 QSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNG 117
S F V+A +L++L++L +++GR+A+S A V D I+ + A +G
Sbjct: 226 LSITAFPVLARILAELKLLTTDIGRMAMSAAGVNDVAAWILLALAIAL----------SG 275
Query: 118 DGKGTLKAFLNVFYYLC---FMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALA 174
DG L ++V+ LC F++ + ++P+L + ++ PEG P+K++Y+ + + LA
Sbjct: 276 DGTSPL---VSVWVLLCGTGFVIFAVVAIKPLLAYMARRCPEGEPVKELYVCVTLTVVLA 332
Query: 175 VGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLS 234
+ L G +VG++ P+ P + +++E S L P++ + +K D++
Sbjct: 333 ASFVTDTIGIHALFGAFVVGIVAPKEGPFCRILTEKIEDLVSGLLLPLYFAASGLKTDVT 392
Query: 235 VYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVF 294
+ + + +I+ K++ T+G C +P + + L +++ KG+V+
Sbjct: 393 TIRGAQSWGLLVLVILTTCFGKIVGTVGSSMLCKVPFREAVTLGFLMNTKGLVELIVLNI 452
Query: 295 LHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIV 354
D +L+ +A ++L + L + V +Y P+RK A Y+ R I +SEL+I+
Sbjct: 453 GKDRKVLNDQAFAILVLMALFTTFITTPIVMLIYKPARKGAPYKHRTIQRKDHDSELRIL 512
Query: 355 SCILKPSHIIPIKNVLDICSPTSSNP-LVIHILHLLELVGRSSPVFISHRLQER---VGS 410
+C +I + N+++ T L ++ +HL+EL RSS + + H+ + + +
Sbjct: 513 ACFHSTRNIPTLINLIESSRGTGKKGRLCVYAMHLMELSERSSAIAMVHKARNNGLPIWN 572
Query: 411 SSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHL 470
++ +++ F+ ++H A +V TAIS + +H+DIC A K ++I+LPFH
Sbjct: 573 KIERSTDQMVIAFEAYQHLRA--VAVRPMTAISGLSSIHEDICTSAHQKRVAMILLPFHK 630
Query: 471 RWSEDGSVESADETTRSLNTKVLERAPCSVAILVNR--GHSSPFNHNENSKQIAMIFLGG 528
DG++ES +N +VL+RAPCSV ILV+R G +S +E + ++ + F GG
Sbjct: 631 HQRMDGAMESIGHRFHEVNQRVLQRAPCSVGILVDRGLGGTSQVVASEVAYKVVIPFFGG 690
Query: 529 SDDREALCLAKRTIKEDTYHLVVYHLVS---TIK------NDEFTSWDVMLDDELLKGVK 579
DDREAL + ++ L VY V+ T+K +DE + D+E ++ +
Sbjct: 691 LDDREALAYGMKMVEHPGITLTVYKFVAARGTLKRFEKSEHDEKEKKEKETDEEFVRELM 750
Query: 580 GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
+++ YE+ VE+ D + ++ + + +VGR + S L T+ PE
Sbjct: 751 NDPRGNESLAYEERVVESKDDIIATLKSMS-KCNLFVVGRNAAVAS-----LVKSTDCPE 804
Query: 640 LGVLGDLLASPDTNTKASILVVQ 662
LG +G LL+S + +T AS+LVVQ
Sbjct: 805 LGPVGRLLSSSEFSTTASVLVVQ 827
>At1g05580 unknown protein
Length = 867
Score = 249 bits (637), Expect = 3e-66
Identities = 185/690 (26%), Positives = 323/690 (46%), Gaps = 51/690 (7%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD M+ T K IAF ++ + G + Y P +I SG
Sbjct: 133 MDLRMVRITELKPVIIAFTGLLVALPVGAFLYY------------LPGNGHPDKII--SG 178
Query: 61 CYFAVIA----------SLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIK 110
C F +A +L+DL++L S++GR A+ A+V D ++ G A S
Sbjct: 179 CVFWSVALACTNFPDLARILADLKLLRSDMGRTAMCAAIVTDLCTWVLLVFGFASFSKSG 238
Query: 111 TDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFI 170
T + + F+ + F+++ V+RP + W KT + + +++ +
Sbjct: 239 TWNK--------MMPFV-IITTAIFVLLCIFVIRPGIAWIFAKTVKAGHVGDTHVWFILG 289
Query: 171 IALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMK 230
+ G++ + G + GL +P + + ++L F S L P+F C ++
Sbjct: 290 GVVLCGLITDACGVHSITGAFLFGLSIPHDHIIRNMIEEKLHDFLSGILMPLFYIICGLR 349
Query: 231 IDLSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFC 290
D+ ++ ++ + +I + L K++ T+ + ++PM D + +++ KG +
Sbjct: 350 ADIGFMLQFTDKFMMVVVICSSFLVKIVTTVITSLFMHIPMRDAFAIGALMNTKGTLSLV 409
Query: 291 NNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSE 350
D+ L S + ++I +LV+ + + + Y P +K A Y+ R + +K +E
Sbjct: 410 VLNAGRDTKALDSPMYTHMTIALLVMSLVVEPLLAFAYKPKKKLAHYKHRTVQKIKGETE 469
Query: 351 LKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRS--SPVFISHRLQERV 408
L++++C+ ++ I N+L + + T +PL + +HL+EL GR+ S + ++ + +
Sbjct: 470 LRVLACVHVLPNVSGITNLLQVSNATKQSPLSVFAIHLVELTGRTTASLLIMNDECKPKA 529
Query: 409 GSSSHTFSEA--VIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIIL 466
S +E+ + TF+ E +N +V T TA+SP MH+DIC LA DK IIL
Sbjct: 530 NFSDRVRAESDQIAETFEAMEVNN-DAMTVQTITAVSPYATMHEDICVLAEDKRVCFIIL 588
Query: 467 PFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRG----HSSPFNHNENSKQIA 522
P+H + DG + + + +N VL APCSV ILV+RG S F +++A
Sbjct: 589 PYHKHLTPDGRMGEGNSSHAEINQNVLSHAPCSVGILVDRGMAMVRSESFRGESMKREVA 648
Query: 523 MIFLGGSDDREALCLAKRTIKEDTYHLVVYH-------LVSTIKNDEFTSWDVMLDDELL 575
M+F+GG DDREAL A R + + L V L+S+ K + +DDE +
Sbjct: 649 MLFVGGPDDREALSYAWRMVGQHVIKLTVVRFVPGREALISSGKVAAEYEREKQVDDECI 708
Query: 576 KGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIA--IQHDFIIVGRRNGIKSPQTQALAS 633
+ +V Y + V + DT I ++ +D +VGR SP T L
Sbjct: 709 YEFNFKTMNDSSVKYIEKVVNDGQDTIATIREMEDNNSYDLYVVGRGYNSDSPVTAGLND 768
Query: 634 WTEYPELGVLGDLLASPDTNTKASILVVQQ 663
W+ PELG +GD LAS + AS+LV+QQ
Sbjct: 769 WSSSPELGTIGDTLASSNFTMHASVLVIQQ 798
>At5g37060 putative transporter protein
Length = 859
Score = 246 bits (627), Expect = 4e-65
Identities = 187/646 (28%), Positives = 306/646 (46%), Gaps = 46/646 (7%)
Query: 54 IVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDS 113
+V S F VI ++L D+ +LNSE+G+ A+S A++ D + G+ I T +
Sbjct: 198 VVFALSFTSFPVIYTVLRDMNLLNSEVGKFAMSVALLGD-----MAGVYVIVIFEAMTHA 252
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIAL 173
D G ++V + FM+ LV+R W V +TPEG + + Y+ ++ + L
Sbjct: 253 -DVGGAYSVFWFLVSVVIFAAFML---LVVRRAFDWIVSQTPEGTLVNQNYIVMILMGVL 308
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
A L + S+ G +GL+VP GPPLG+ + + E F FL P ++
Sbjct: 309 ASCFLTDMFGLSIAVGPIWLGLLVPHGPPLGSTLAVRSETFIYEFLMPFTYALVGQGTNI 368
Query: 234 SVYVKSDY---IYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFC 290
+ + + V + K L T + +P + + L LM++ +G +D
Sbjct: 369 HFLRDETWRNQLSPLFYMTVVGFITKFLSTAFAALFFKVPARESITLGLMMNLRGQMDLL 428
Query: 291 NNVFLH--DSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPN 348
V+LH D ++ +V+ ++ +V+ + + + YDP+R Y + R I N
Sbjct: 429 --VYLHWIDKRIVGFPGYTVMVLHTVVVTAVTTPLINFFYDPTRPYRSSKHRTIQHTPQN 486
Query: 349 SELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERV 408
+E+ +V + + + LD PT S+PL I + L+EL GR++P+FI H ++
Sbjct: 487 TEMGLVLAVSDHETLSGLITFLDFAYPTKSSPLSIFAVQLVELAGRATPLFIDHEQRKEE 546
Query: 409 GSSSHTFSE------------AVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLA 456
+ E V F L+E ++ +YTA +P R M+ DIC LA
Sbjct: 547 EEEEYEEEEEEPERKQSGRIDQVQSAFKLYEEKRNECVTLRSYTAHAPKRLMYQDICELA 606
Query: 457 LDKLASIIILPFHLRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGH------- 508
L K + I+LP+ ED + E D S+N VLE PCSV I ++G
Sbjct: 607 LGKKTAFILLPYQKERLEDAAPTELRDSGMLSVNADVLEHTPCSVCIYFDKGRLKNAVVR 666
Query: 509 -SSPFNHNENS-------KQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKN 560
S H+ NS + ++FLGG+D+REAL LA R L V +S
Sbjct: 667 LSMDLQHSTNSIRMRQETYRFVVLFLGGADNREALHLADRMSTNPDVTLTVIRFLSYNHE 726
Query: 561 DEFTSWDVMLDDELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAI-QHDFIIVGR 619
E + LDD ++ S + V+Y++V V+N ++T I + + +D I GR
Sbjct: 727 GE-DEREKKLDDGVVTWFWVKNESNERVSYKEVVVKNGAETLAAIQAMNVNDYDLWITGR 785
Query: 620 RNGIKSPQTQALASWTEYPELGVLGDLLASPDTNTKASILVVQQQV 665
R GI + L++W+E +LGV+GD +A+ ++ S+LVVQQQV
Sbjct: 786 REGINPKILEGLSTWSEDHQLGVIGDTVAASVFASEGSVLVVQQQV 831
>At5g41610 Na+/H+ antiporter-like protein
Length = 810
Score = 239 bits (609), Expect = 5e-63
Identities = 181/691 (26%), Positives = 334/691 (48%), Gaps = 47/691 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D + RTG+KA IA +P G+ + + + G L + + S
Sbjct: 113 IDTKALRRTGKKALGIALAGITLPFALGIGSSFVLKA--TISKGVNSTAFLVFMGVALSI 170
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L++L++L +E+GRLA+S A V D I+ + A S N
Sbjct: 171 TAFPVLARILAELKLLTTEIGRLAMSAAAVNDVAAWILLALAIALSGS-------NTSPL 223
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
+L FL+ F++ ++ PI +W ++ EG P+++ Y+ + L G +
Sbjct: 224 VSLWVFLSG---CAFVIGASFIIPPIFRWISRRCHEGEPIEETYICATLAVVLVCGFITD 280
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSD 240
+ G +VG+++P+ P +++++E S P++ + +K +++ ++
Sbjct: 281 AIGIHSMFGAFVVGVLIPKEGPFAGALVEKVEDLVSGLFLPLYFVASGLKTNVAT-IQGA 339
Query: 241 YIYVWLGIIVAVHLF-KMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
+ L ++ A F K+L T+G+ +PM + + L +++ KG+V+ D
Sbjct: 340 QSWGLLVLVTATACFGKILGTLGVSLAFKIPMREAITLGFLMNTKGLVELIVLNIGKDRK 399
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSR---KYAGYQKRNILSLKPNSELKIVSC 356
+L+ + +++ + L + V +Y P+R K Y+ R + N++L+I++C
Sbjct: 400 VLNDQTFAIMVLMALFTTFITTPVVMAVYKPARRAKKEGEYKHRAVERENTNTQLRILTC 459
Query: 357 ILKPSHIIPIKNVLDICSPTSSNP-LVIHILHLLELVGRSSPVFISHRLQE-------RV 408
I + N+L+ L ++ LHL EL RSS + + H++++ R
Sbjct: 460 FHGAGSIPSMINLLEASRGIEKGEGLCVYALHLRELSERSSAILMVHKVRKNGMPFWNRR 519
Query: 409 GSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPF 468
G ++ ++ V+V F F+ +V TAIS + +H+DIC A+ K A+I+ILPF
Sbjct: 520 GVNAD--ADQVVVAFQAFQQ--LSRVNVRPMTAISSMSDIHEDICTTAVRKKAAIVILPF 575
Query: 469 HLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNR--GHSSPFNHNENSKQIAMIFL 526
H DGS+E+ R +N +VL +APCSV I V+R G SS + + S + ++F
Sbjct: 576 HKHQQLDGSLETTRGDYRWVNRRVLLQAPCSVGIFVDRGLGGSSQVSAQDVSYSVVVLFF 635
Query: 527 GGSDDREALCLAKRTIKEDTYHLVVYHL--------------VSTIKNDEFTSWDVMLDD 572
GG DDREAL R + L V+ VS N+ + ++ D+
Sbjct: 636 GGPDDREALAYGLRMAEHPGIVLTVFRFVVSPERVGEIVNVEVSNNNNENQSVKNLKSDE 695
Query: 573 ELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALA 632
E++ ++ + ++V + + ++EN + + + + +VGR G A+
Sbjct: 696 EIMSEIRKISSVDESVKFVEKQIENAAVDVRSAIEEVRRSNLFLVGRMPG--GEIALAIR 753
Query: 633 SWTEYPELGVLGDLLASPDTNTKASILVVQQ 663
+E PELG +G LL SP+++TKAS+LV+QQ
Sbjct: 754 ENSECPELGPVGSLLISPESSTKASVLVIQQ 784
>At2g31910 putative Na+/H+ antiporter
Length = 735
Score = 237 bits (605), Expect = 1e-62
Identities = 177/629 (28%), Positives = 294/629 (46%), Gaps = 40/629 (6%)
Query: 60 GCY-FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGD 118
GC F +A +L+DL++L +++G A+ A+V D I+ G A S S +
Sbjct: 76 GCTNFPDLARILADLKLLRTDMGHTAMCAAVVTDLCTWILFIFGMAIFSK----SGVRNE 131
Query: 119 GKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML 178
A F LC+ V+ P V W T EG + +++ + ++
Sbjct: 132 MLPYSLASTIAFVLLCYFVIQPGVA-----WIFNNTVEGGQVGDTHVWYTLAGVIICSLI 186
Query: 179 GLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVK 238
+ + G + GL +P + + ++L F S L P+F C ++ D+ +
Sbjct: 187 TEVCGVHSITGAFLFGLSIPHDHIIRKMIEEKLHDFLSGMLMPLFYIICGLRADIGYMNR 246
Query: 239 SDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDS 298
+ + + + A + K+L T+ + +P+ DGL + +++ KG + D+
Sbjct: 247 TVSVGMMAVVTSASVMVKILSTMFCSIFLRIPLRDGLAIGALMNTKGTMALVILNAGRDT 306
Query: 299 MLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCIL 358
L + L++ LV+ + + + Y P +K Y+ R I K SEL +++C+
Sbjct: 307 KALDVIMYTHLTLAFLVMSMVVQPLLAIAYKPKKKLIFYKNRTIQKHKGESELCVLTCVH 366
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFI--------SHRLQERVGS 410
++ I N+L + +PT +PL + +HL+EL GR++ + +RV +
Sbjct: 367 VLPNVSGITNLLQLSNPTKKSPLNVFAIHLVELTGRTTASLLIMNDEAKPKANFADRVRA 426
Query: 411 SSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHL 470
S +E F E +N G V T TA+SP M +DIC LA DK A I+LP+H
Sbjct: 427 ESDQIAEM----FTALEVNNDGVM-VQTITAVSPYATMDEDICLLAEDKQACFILLPYHK 481
Query: 471 RWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRG------HSSPFNHNENSKQIAMI 524
+ DG + + +N V+ APCSV ILV+RG S F K+IAM+
Sbjct: 482 NMTSDGRLNEGNAVHAEINQNVMSHAPCSVGILVDRGMTTVRFESFMFQGETTKKEIAML 541
Query: 525 FLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIK--------NDEFTSWDVMLDDELLK 576
FLGG DDREAL A R + ++ L V V + + DE+ D +D+E +
Sbjct: 542 FLGGRDDREALAYAWRMVGQEMVQLTVVRFVPSQEALVSAGEAADEYEK-DKHVDEESIY 600
Query: 577 GVKGVYGSVDNVTYEKVEVENTSDTTEFISDIA--IQHDFIIVGRRNGIKSPQTQALASW 634
+ +VTY + V+N +T I ++ +D IVGR +++P T L W
Sbjct: 601 EFNFKTMNDPSVTYVEKVVKNGQETITAILELEDNNSYDLYIVGRGYQVETPVTSGLTDW 660
Query: 635 TEYPELGVLGDLLASPDTNTKASILVVQQ 663
P+LG++GD L S + +AS+LVVQQ
Sbjct: 661 NSTPDLGIIGDTLISSNFTMQASVLVVQQ 689
>At5g01680 putative transporter protein
Length = 780
Score = 234 bits (598), Expect = 9e-62
Identities = 192/678 (28%), Positives = 312/678 (45%), Gaps = 36/678 (5%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D +I R G+ A F+ P G + C + + + L ++ QS
Sbjct: 119 VDMGVIKRGGKLAIINGLSLFLFPYVVGAIACTVITSNIRGTVAKNNPEQLHNLLTNQSV 178
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
YF V S+LS+L++LNSE GRLALS+ MV + F G G F+ I DS + +
Sbjct: 179 VYFQVAYSVLSNLKMLNSEPGRLALSSIMVANCF-----GWGF-FLLLITFDSFLHQNYS 232
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
T +L F + +V +V RPI W VK+TPEG+ +K ++ + ++ L
Sbjct: 233 KT--TYLPTFTKVLLLVGIVVVCRPIFNWIVKRTPEGKKLKASHLCTICVMLCTATFLSE 290
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSD 240
+ G +GL+ P+ PP GT + ++ FC L P +V K+D + D
Sbjct: 291 TVGFPYVVGSVALGLVTPKTPPFGTGLTDKIGSFCYAVLMPCYVIGIGNKVDFFSFNLRD 350
Query: 241 YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSML 300
I + +I + K + Y +P++ + + ++ +G+ D L +
Sbjct: 351 IISLEF-LIFTISAAKFASIVLPSLYFQVPISHAVIVGFIVCIQGIYDVQIFKQLLNYKN 409
Query: 301 LSSEALSVLSINVLVIGTLARIGVKYLYD-PSRKYAGYQKRNILSLKPNSELKIVSCILK 359
+S EA ++ I+ +V T+ VK LY RK+ Y+++ + +PN LKI++C
Sbjct: 410 ISHEAFGIMVISAMVHSTIFTAIVKNLYGWVQRKHITYRRQTVQHYEPNKPLKILTCFYH 469
Query: 360 PSHIIPIKNVLDICS-PTSSNPLVIHILHLLELVGRSSPVFISHRL-QERVGSSSHTFSE 417
+ PI VL++ + P+S++ I ++L EL + P+ I H S+S + +
Sbjct: 470 RETVPPILTVLELSTCPSSASSHSIVSVNLEELEQNNVPLLIQHHPGHNDESSTSSSRRD 529
Query: 418 AVIVTFDLFE--HDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSED 475
+ F+ F HD SV +TA++P + MH+D+C LA +K +II D
Sbjct: 530 QISKAFEKFRSGHDLQENVSVECFTAVAPSKTMHEDVCALAFEKETDLIIFGM-----AD 584
Query: 476 GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQ-------IAMIFLGG 528
G+ R L V +P SVA+L+++G F + + + I IFLGG
Sbjct: 585 GTA-----AERRLCRNVRNASPSSVAVLMDQGRLPDFKNMGTAMKNGSMRINICSIFLGG 639
Query: 529 SDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTS-WDVMLDDELLKGVKGVYGSVDN 587
+DDRE L A R + +L V LV + LD ++ + + N
Sbjct: 640 ADDRETLAFAVRMTNQPYVNLTVLKLVDGENVSHLNDVVEKRLDFRTIEKFRQDTMNKHN 699
Query: 588 VTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLL 647
V E SD + + +D I+VG R+ Q L+ W+E ELG +GDLL
Sbjct: 700 VALR----EEASDLVNLLREEGNNYDLIMVGIRHEKSFEVLQGLSVWSEIEELGEIGDLL 755
Query: 648 ASPDTNTKASILVVQQQV 665
S D AS+L VQQQ+
Sbjct: 756 VSRDLKLSASVLAVQQQL 773
>At5g58460 putative protein
Length = 857
Score = 231 bits (590), Expect = 8e-61
Identities = 176/635 (27%), Positives = 301/635 (46%), Gaps = 44/635 (6%)
Query: 63 FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGT 122
F VI ++L D+ +LNSE+G+ A+S ++ D V + A + D G G +
Sbjct: 207 FPVIYTVLRDMNLLNSEIGKFAMSVTLLGDMVGVYVLVLFEAMAQA------DGGGGAYS 260
Query: 123 LKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGLLT 182
+ FL + ++ LV++ +W V KTPEG + + Y+ + + L L +
Sbjct: 261 VIWFLISAAIMAACLL--LVVKRSFEWIVAKTPEGGLVNQNYIVNILMGVLVSCFLTDMF 318
Query: 183 KQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSDY- 241
++ G +GL+VP GPPLG+ + + E F + FL P K ++++ K +
Sbjct: 319 GMAIAVGPIWLGLVVPHGPPLGSTLAIRSETFVNEFLMPFSFALVGQKTNVNLISKETWP 378
Query: 242 --IYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
I + + + + K + + G + +P D L L LM++ +G +D + D
Sbjct: 379 KQISPLIYMSIVGFVTKFVSSTGAALFFKVPTRDSLTLGLMMNLRGQIDILLYLHWIDKQ 438
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILK 359
++ SV+ + +V+ + + +LYDP+R Y ++R I N+E +V +
Sbjct: 439 MVGLPGYSVMVLYAIVVTGVTAPLISFLYDPTRPYRSSKRRTIQHTPQNTETGLVLAVTD 498
Query: 360 PSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ------------ER 407
+ LD PT ++P + + L+EL GR+ P+FI+H + ER
Sbjct: 499 HDTFSGLITFLDFAYPTKTSPFSVFAIQLVELEGRAQPLFIAHDKKREEEYEEEEEPAER 558
Query: 408 VGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILP 467
+GS + V F L++ + ++ YTA + M+ +IC LAL K + I+LP
Sbjct: 559 MGSRR---VDQVQSAFKLYQEKRSECVTMHAYTAHASKHNMYQNICELALTKKTAFILLP 615
Query: 468 FHLRWSEDGSV-ESADETTRSLNTKVLERAPCSVAILVNRGH--------SSPFNHNENS 518
+ +D ++ E D S+N VL PCSV I +G S H NS
Sbjct: 616 YQKERLQDAALTELRDSGMLSVNADVLAHTPCSVCIYYEKGRLKNAMVRSSMDPQHTTNS 675
Query: 519 K-------QIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLD 571
+ ++FLGG+D+REAL LA R + +L V ++ E + LD
Sbjct: 676 SHMRQEMYRFVVLFLGGADNREALHLADRMTENPFINLTVIRFLAHNHEGE-DEREKKLD 734
Query: 572 DELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAI-QHDFIIVGRRNGIKSPQTQA 630
D ++ S V+Y++V V+N ++T I + + +D I GRR GI +
Sbjct: 735 DGVVTWFWVKNESNARVSYKEVVVKNGAETLAAIQAMNVNDYDLWITGRREGINPKILEG 794
Query: 631 LASWTEYPELGVLGDLLASPDTNTKASILVVQQQV 665
L++W+E +LGV+GD +A ++ S+LVVQQQV
Sbjct: 795 LSTWSEDHQLGVIGDTVAGSVFASEGSVLVVQQQV 829
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,640,029
Number of Sequences: 26719
Number of extensions: 621630
Number of successful extensions: 1939
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1730
Number of HSP's gapped (non-prelim): 45
length of query: 670
length of database: 11,318,596
effective HSP length: 106
effective length of query: 564
effective length of database: 8,486,382
effective search space: 4786319448
effective search space used: 4786319448
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146343.3


