

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.12 + phase: 0 /pseudo
(176 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
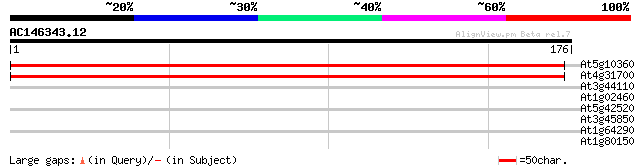
At5g10360 40S ribsomal protein S6 328 1e-90
At4g31700 ribosomal protein S6 - like 327 2e-90
At3g44110 dnaJ protein homolog atj3 28 1.9
At1g02460 polygalacturonase PG1, putative 28 3.3
At5g42520 unknown protein 27 5.7
At3g45850 kinesin-related protein - like 27 5.7
At1g64290 hypothetical protein 27 7.4
At1g80150 unknown protein 26 9.6
>At5g10360 40S ribsomal protein S6
Length = 249
Score = 328 bits (840), Expect = 1e-90
Identities = 158/174 (90%), Positives = 165/174 (94%)
Query: 1 FNVANPTTGCQKKLEIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKITGGCDKQGFP 60
FNVANPTTGCQKKLEIDDDQKLRAF+DKR+SQEV+GDALGEEFKGYVFKI GGCDKQGFP
Sbjct: 3 FNVANPTTGCQKKLEIDDDQKLRAFFDKRLSQEVSGDALGEEFKGYVFKIMGGCDKQGFP 62
Query: 61 MKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGEND 120
MKQGVLTPGRVRLLLHRGTPCFRGHGRR GERRRKSVRGCIVSPDLSVLNLVIVKKG +D
Sbjct: 63 MKQGVLTPGRVRLLLHRGTPCFRGHGRRTGERRRKSVRGCIVSPDLSVLNLVIVKKGVSD 122
Query: 121 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRRTFTTKDGAKLPR 174
LPGLTD EKPRMRGPKRASKIRKLFNL K+DDVRKYVNTYRRTFT K G K+ +
Sbjct: 123 LPGLTDTEKPRMRGPKRASKIRKLFNLGKEDDVRKYVNTYRRTFTNKKGKKVSK 176
>At4g31700 ribosomal protein S6 - like
Length = 250
Score = 327 bits (839), Expect = 2e-90
Identities = 158/174 (90%), Positives = 164/174 (93%)
Query: 1 FNVANPTTGCQKKLEIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKITGGCDKQGFP 60
FNVANPTTGCQKKLEIDDDQKLRAF+DKRISQEV+GDALGEEFKGYVFKI GGCDKQGFP
Sbjct: 3 FNVANPTTGCQKKLEIDDDQKLRAFYDKRISQEVSGDALGEEFKGYVFKIKGGCDKQGFP 62
Query: 61 MKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGEND 120
MKQGVLTPGRVRLLLHRGTPCFRGHGRR GERRRKSVRGCIVSPDLSVLNLVIVKKGEND
Sbjct: 63 MKQGVLTPGRVRLLLHRGTPCFRGHGRRTGERRRKSVRGCIVSPDLSVLNLVIVKKGEND 122
Query: 121 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRRTFTTKDGAKLPR 174
LPGLTD EKPRMRGPKRASKIRKLFNL K+DDVR YVNTYRR FT K G ++ +
Sbjct: 123 LPGLTDTEKPRMRGPKRASKIRKLFNLKKEDDVRTYVNTYRRKFTNKKGKEVSK 176
>At3g44110 dnaJ protein homolog atj3
Length = 420
Score = 28.5 bits (62), Expect = 1.9
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 3/67 (4%)
Query: 52 GGCDKQGFPMKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRR-KSVRGCIVSPDLSVLN 110
GGC G + L PG ++ + H C +G G +R R +G V P+ VL
Sbjct: 165 GGCQGSGMKVSIRQLGPGMIQQMQHACNEC-KGTGETINDRDRCPQCKGDKVIPEKKVLE 223
Query: 111 LVIVKKG 117
V V+KG
Sbjct: 224 -VNVEKG 229
>At1g02460 polygalacturonase PG1, putative
Length = 491
Score = 27.7 bits (60), Expect = 3.3
Identities = 19/60 (31%), Positives = 27/60 (44%), Gaps = 6/60 (10%)
Query: 15 EIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKI-----TGGC-DKQGFPMKQGVLTP 68
E DD + + WD + E N D++ GY F I TG C Q F + ++TP
Sbjct: 101 ETDDTEAFKTAWDSSCNNENNTDSVLLVPYGYTFMIQSTIFTGPCRSYQFFQVDGTIVTP 160
>At5g42520 unknown protein
Length = 342
Score = 26.9 bits (58), Expect = 5.7
Identities = 21/70 (30%), Positives = 31/70 (44%), Gaps = 7/70 (10%)
Query: 104 PDLSVLNLVIVKKGENDLPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRK--YVNTYR 161
P S L K+G+ P T RGPK K++K +DD+ K +V T
Sbjct: 156 PAGSTLESAKPKRGKRVNPKATTQTAANKRGPKNQRKVKK----ESEDDLNKIMFVKT-T 210
Query: 162 RTFTTKDGAK 171
+T +D +K
Sbjct: 211 HDYTDEDSSK 220
>At3g45850 kinesin-related protein - like
Length = 1058
Score = 26.9 bits (58), Expect = 5.7
Identities = 10/20 (50%), Positives = 17/20 (85%)
Query: 19 DQKLRAFWDKRISQEVNGDA 38
++ LRAF D+++S++ NGDA
Sbjct: 1006 EELLRAFRDEKLSKQANGDA 1025
>At1g64290 hypothetical protein
Length = 364
Score = 26.6 bits (57), Expect = 7.4
Identities = 16/46 (34%), Positives = 23/46 (49%), Gaps = 5/46 (10%)
Query: 94 RKSVRGCIVSPDLSVLNLVIVKKGEND-----LPGLTDVEKPRMRG 134
R S RGC+ + + LNLV KK +D P +D E+ + G
Sbjct: 312 RSSDRGCLYNINAEKLNLVYAKKEGSDCSFVCFPFCSDYERVDLNG 357
>At1g80150 unknown protein
Length = 397
Score = 26.2 bits (56), Expect = 9.6
Identities = 16/61 (26%), Positives = 25/61 (40%)
Query: 104 PDLSVLNLVIVKKGENDLPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRRT 163
P + + V N + GL +E+P + K KL+NL K + + V R
Sbjct: 14 PSSAATGIAYVSTESNQIQGLKPLEEPALVKLKSERDPEKLYNLFKANATNRLVIENRFA 73
Query: 164 F 164
F
Sbjct: 74 F 74
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,332,878
Number of Sequences: 26719
Number of extensions: 188155
Number of successful extensions: 375
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 370
Number of HSP's gapped (non-prelim): 8
length of query: 176
length of database: 11,318,596
effective HSP length: 93
effective length of query: 83
effective length of database: 8,833,729
effective search space: 733199507
effective search space used: 733199507
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146343.12


