

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
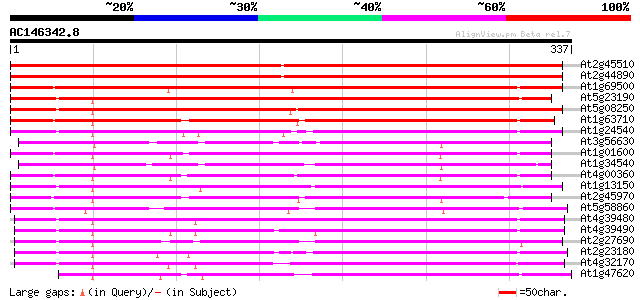
Query= AC146342.8 - phase: 0
(337 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g45510 putative cytochrome P450 446 e-126
At2g44890 putative cytochrome P450 446 e-125
At1g69500 unknown protein 296 1e-80
At5g23190 cytochrome P450-like protein 262 2e-70
At5g08250 cytochrome P450-like protein 261 5e-70
At1g63710 cytochrome P450 like protein 225 3e-59
At1g24540 putative cytochrome P450 221 4e-58
At3g56630 cytochrome P450-like protein 218 5e-57
At1g01600 unknown protein 214 7e-56
At1g34540 cytochrome p450, putative 213 2e-55
At4g00360 probable cytochrome P450 211 3e-55
At1g13150 putative cytochrome P450 monooxygenase 209 1e-54
At2g45970 putative cytochrome P450 207 5e-54
At5g58860 cytochrome P450 CYP86A1 207 7e-54
At4g39480 cytochrome P450 - like protein 207 9e-54
At4g39490 cytochrome P450 - like protein 205 2e-53
At2g27690 putative cytochrome P450 204 6e-53
At2g23180 putative cytochrome P450 202 2e-52
At4g32170 cytochrome p450 - like protein 201 6e-52
At1g47620 hypothetical protein 196 2e-50
>At2g45510 putative cytochrome P450
Length = 511
Score = 446 bits (1147), Expect = e-126
Identities = 216/332 (65%), Positives = 265/332 (79%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+CTLDSIFKVGFGVEL CL+ SKE FM+AF++ N R++DPLWKLK N GS
Sbjct: 178 MRCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGS 237
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
++ LKK+I ID FV+SLI T+R+ L+ +++ +EDILSRFL+ES+KD NM DKYLRD
Sbjct: 238 QSKLKKSIATIDKFVYSLITTKRKELAKEQNTVVREDILSRFLVESEKDPENMNDKYLRD 297
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILNFMIAGKDT+A LSWF YMLCKNPL+QEK+ QE+ +VT S E +++ FV +I+
Sbjct: 298 IILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTF-SHEKTTDVNGFVESIN 356
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ LD+MHYLHAAL+ETLRLYP VP+ R AE D+LPDG+ V+KG+ +YY++YAMGRM
Sbjct: 357 EEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMT 416
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
YIWG DA+EF PERWLKDG+FQPES FKFISFHAGPRICLGKDFAYRQMKIVSMAL+ FF
Sbjct: 417 YIWGQDAEEFKPERWLKDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLHFF 476
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFK+ +E + V Y+ M TLH+D GL L A PR
Sbjct: 477 RFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>At2g44890 putative cytochrome P450
Length = 490
Score = 446 bits (1146), Expect = e-125
Identities = 216/332 (65%), Positives = 264/332 (79%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
MKCTLDSIFKVGFGVEL CL+ SKE FMKAF++ N R DP WKLK LN GS
Sbjct: 157 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 216
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E+ LKK+I IID FV+SLI T+R+ LS +++ S +EDILS+FLLES+KD NM DKYLRD
Sbjct: 217 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTSVREDILSKFLLESEKDPENMNDKYLRD 276
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILN M+AGKDT+A +LSWF YMLCKNPL+QEK+ QE+ +VTS S E +++ F+ +++
Sbjct: 277 IILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTS-SHEKTTDVNGFIESVT 335
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ L +M YLHAAL+ET+RLYP VP R AE D+LPDG+ V+KG+ +YY+SYAMGRM
Sbjct: 336 EEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRMT 395
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
YIWG DA+EF PERWLKDG+FQPES FKFISFHAGPRIC+GKDFAYRQMKIVSMAL+ FF
Sbjct: 396 YIWGQDAEEFKPERWLKDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLHFF 455
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFK+ +E + V+Y+ M TLH+D GL L A PR
Sbjct: 456 RFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>At1g69500 unknown protein
Length = 524
Score = 296 bits (757), Expect = 1e-80
Identities = 168/358 (46%), Positives = 219/358 (60%), Gaps = 28/358 (7%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDSI KVGFGVE+ L E N F KAF+ +N V R++DPLWK+K+ LN GS
Sbjct: 168 MRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 226
Query: 61 EAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSD---------KEDILSRFLLESKKDS 110
EA L K+IK+++DF +S+I+ R+ ELL Q ++ K DILSRF+ S
Sbjct: 227 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 286
Query: 111 SNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQES-- 168
S T+K LRDI+LNF+IAG+DT+A TL+W YM+ N + EK+ E+ + S E+
Sbjct: 287 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 346
Query: 169 --------------NLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEH 214
N + EF ++ +L K+HYLHA +TETLRLYPAVP + E
Sbjct: 347 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 406
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+LP+G V G V Y+ Y+MGRM Y WG DA F PERWLKDG+FQ S FKF +F A
Sbjct: 407 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWLKDGVFQNASPFKFTAFQA 466
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
GPRICLGKD AY QMK+ L RF++F L + V YR M L + HGL + + R
Sbjct: 467 GPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRR 523
>At5g23190 cytochrome P450-like protein
Length = 559
Score = 262 bits (670), Expect = 2e-70
Identities = 133/328 (40%), Positives = 205/328 (61%), Gaps = 5/328 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL F KAF D+ R++ P +WK R L+
Sbjct: 208 LRLTFDNVCMIAFGVDPGCLGPDQPVIP-FAKAFEDATEAAVVRFVMPTCVWKFMRYLDI 266
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
G+E LK++IK +DDF +I+TR++ LS++ + + + D+L+ F+ + + +DK+L
Sbjct: 267 GTEKKLKESIKGVDDFADEVIRTRKKELSLEGETTKRSDLLTVFMGLRDEKGESFSDKFL 326
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQE-SNLNLDEFVS 177
RDI +NF++AG+DTS+ LSWFF++L KNP ++EK+ E+ + + N ++
Sbjct: 327 RDICVNFILAGRDTSSVALSWFFWLLEKNPEVEEKIMVEMCKILRQRDDHGNAEKSDYEP 386
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
+ KM YL AAL+E LRLYP+VP+ + +E D+ PDG ++ KG+ V Y YAMG
Sbjct: 387 VFGPEEIKKMDYLQAALSEALRLYPSVPVDHKEVQEDDVFPDGTMLKKGDKVIYAIYAMG 446
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
RM IWG D EF PERWL+DG F ES++KF +F+ GPR+CLGKDFAY QMK + A+V
Sbjct: 447 RMEAIWGKDCLEFRPERWLRDGRFMSESAYKFTAFNGGPRLCLGKDFAYYQMKSTAAAIV 506
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGL 325
++ K+ N + V + T+++ HGL
Sbjct: 507 YRYKVKVVN-GHKVEPKLALTMYMKHGL 533
>At5g08250 cytochrome P450-like protein
Length = 550
Score = 261 bits (666), Expect = 5e-70
Identities = 132/338 (39%), Positives = 212/338 (62%), Gaps = 8/338 (2%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL E F KAF D+ R++ P +WKL R LN
Sbjct: 205 LRLTFDNVCMIAFGVDPGCLSPKLPEIP-FAKAFEDATEATVVRFVMPKFVWKLMRSLNL 263
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
G+E LK++I +DDF +I+TR++ +S++ + + + D+L+ F+ ++ +DK+L
Sbjct: 264 GTEKKLKESINGVDDFAEEVIRTRKKEMSLETEIAKRPDLLTIFMGLRDENGQKFSDKFL 323
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQE---SNLNLDEF 175
RDI +NF++AG+DTS+ LSWFF+++ KNP ++EK+ + + + + N+ E+
Sbjct: 324 RDICVNFILAGRDTSSVALSWFFWLIEKNPEVEEKIMMGICKILEQRVDHGDTKKNM-EY 382
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
+ KM YL AAL+ETLRLYP+VP+ + E D+ PDG + KGE V Y YA
Sbjct: 383 EPVFRPEEIKKMDYLQAALSETLRLYPSVPVDHKEVLEDDVFPDGTKLKKGEKVIYAIYA 442
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
MGRM IWG D +EF PERWL+DG + ES++KF +F+ GPR+CLGKDFAY QM+ V+ A
Sbjct: 443 MGRMETIWGKDCREFKPERWLRDGRYMSESAYKFTAFNGGPRLCLGKDFAYYQMRYVAAA 502
Query: 296 LVRFFRFKLENE-TNDVTYRTMFTLHIDHGLPLYATPR 332
++ ++ +++++ + V + T+++ HGL + R
Sbjct: 503 IIYRYKVRVDDKGGHKVEPKMALTMYMKHGLKVNMVKR 540
>At1g63710 cytochrome P450 like protein
Length = 523
Score = 225 bits (573), Expect = 3e-59
Identities = 122/332 (36%), Positives = 204/332 (60%), Gaps = 14/332 (4%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D+I + FG + L E N F AF+ + +R++ P +WK+++ L
Sbjct: 180 LRLTFDNICGLTFGKDPRTLSPEFPE-NGFAVAFDGATEATLQRFIMPEFIWKIRKWLRL 238
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKY 117
G E + ++I +D+++ +I TR+ ELL Q+D S +D+LSRF+ K + +DKY
Sbjct: 239 GLEDDMSRSISHVDNYLSEIINTRKLELLGQQQDESRHDDLLSRFM----KKKESYSDKY 294
Query: 118 LRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLN--LDEF 175
L+ + LNF++AG+DTS+ +SWFF+++ NP ++EK+ E+ + ++++N++ DE
Sbjct: 295 LKYVALNFILAGRDTSSVAMSWFFWLVSLNPRVEEKIINEICTILIKTRDTNVSKWTDE- 353
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
++ +D++ YL AAL+ETLRLYP+VP + +D+LPDG V G V Y Y+
Sbjct: 354 --PLTFDEIDQLVYLKAALSETLRLYPSVPEDSKFVVANDVLPDGTFVPSGSNVTYSIYS 411
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
+GRM +IWG+D EF PERWL++ + + +KF++F+AGPRICLGKD AY QMK ++ +
Sbjct: 412 VGRMKFIWGEDCLEFKPERWLEESRDEKCNQYKFVAFNAGPRICLGKDLAYLQMKSITAS 471
Query: 296 LVRFFRFKLENETNDVTYRTMFTLHIDHGLPL 327
++ R + + V + TL + GL +
Sbjct: 472 ILLRHRLTVA-PGHRVEQKMSLTLFMKFGLKM 502
>At1g24540 putative cytochrome P450
Length = 522
Score = 221 bits (563), Expect = 4e-58
Identities = 124/342 (36%), Positives = 201/342 (58%), Gaps = 19/342 (5%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D+I FGV LE E F KAF D+ + R+L P +WK R L
Sbjct: 188 LRFTFDNICISAFGVYPGSLETGLPEIP-FAKAFEDATEYTLARFLIPPFVWKPMRFLGI 246
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFL---LESKKDSSN--- 112
G E L ++I+ F + ++ RR + + +D D+LSR + E ++D++
Sbjct: 247 GYERKLNNAVRIVHAFANKTVRERRNKMRKLGNLNDYADLLSRLMQREYEKEEDTTRGNY 306
Query: 113 MTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTS--TSQESNL 170
+DKY R+ +F+IAG+DT++ L WFF+++ K+P +++++ +E+ + T+QE+
Sbjct: 307 FSDKYFREFCTSFIIAGRDTTSVALVWFFWLVQKHPEVEKRILREIREIKRKLTTQETE- 365
Query: 171 NLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVY 230
D+F + +M YL AALTE+LRLYP+VPM + A E D+LPDG V KG ++
Sbjct: 366 --DQFEAE----DFREMVYLQAALTESLRLYPSVPMEMKQALEDDVLPDGTRVKKGARIH 419
Query: 231 YLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMK 290
Y Y+MGR+ IWG D +EF PERW+K+G E FK++ F+ GPR+C+GK FAY QMK
Sbjct: 420 YSVYSMGRIESIWGKDWEEFKPERWIKEGRIVSEDQFKYVVFNGGPRLCVGKKFAYTQMK 479
Query: 291 IVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
+V+ A++ + K+ + ++ + TL++ +G+ + PR
Sbjct: 480 MVAAAILMRYSVKVV-QGQEIVPKLTTTLYMKNGMNVMLQPR 520
>At3g56630 cytochrome P450-like protein
Length = 499
Score = 218 bits (554), Expect = 5e-57
Identities = 125/326 (38%), Positives = 193/326 (58%), Gaps = 20/326 (6%)
Query: 6 DSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPL---WKLKRVLNFGSEA 62
D+I K+ F V+ CL FM+AF + + +R+ + WK+K+ LN GSE
Sbjct: 180 DNICKLAFNVDSACLGDDGAAGVNFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSER 239
Query: 63 ALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD-KEDILSRFLLESKKDSSNMTDKYLRDI 121
L+++I I+ F +++ R E Q SD KED+LSRF+ + + +S + LRDI
Sbjct: 240 VLRESIMIVHKFADEIVRNRIE----QGKVSDHKEDLLSRFISKEEMNSPEI----LRDI 291
Query: 122 ILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISD 181
+++F++AG+DT+++ LSWFF++L +P +++K+ QE+ S + + + E V D
Sbjct: 292 VISFILAGRDTTSSALSWFFWLLSMHPEVKDKILQEL---NSIRERTGKRIGE-VYGFED 347
Query: 182 ATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPY 241
L M+YLHAA+TE+LRLYP VP+ + E ++LPDG + K + Y +YAMGRM
Sbjct: 348 LKL--MNYLHAAITESLRLYPPVPVDTMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMES 405
Query: 242 IWGDDAQEFLPERWLKD--GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRF 299
IWG D F PERW+ + G F+ E+ +KF +FHAGPR+CLGK+ AY QMK + A++
Sbjct: 406 IWGKDCDRFDPERWIDETNGGFRGENPYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLER 465
Query: 300 FRFKLENETNDVTYRTMFTLHIDHGL 325
F ++ + TL I GL
Sbjct: 466 FVVEVPGKKERPEILMSVTLRIRGGL 491
>At1g01600 unknown protein
Length = 554
Score = 214 bits (544), Expect = 7e-56
Identities = 119/332 (35%), Positives = 200/332 (59%), Gaps = 12/332 (3%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D+I + FG + E N F AF+ + +R++ P +WKLK+ L
Sbjct: 180 LRLTFDNICGLAFGKDTRTCAPGLPE-NGFASAFDRATEASLQRFIIPKFMWKLKKWLGL 238
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSDK--EDILSRFLLESKKDSSNMTD 115
G E +L +++ ID+++ ++I TR+ EL+S Q+ + + +D+LSRF++ K + + +D
Sbjct: 239 GLEVSLSRSLGEIDEYLAAVINTRKQELMSQQESGTHQRHDDLLSRFMM---KKTESYSD 295
Query: 116 KYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEF 175
+L+ + LNF++AG+DTS+ LSWFF+++ +P +++K+ +E+ +V ++ ++
Sbjct: 296 TFLQHVALNFILAGRDTSSVALSWFFWLITMHPTVEDKIVREICSVLIETRGTDDVASWT 355
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
+ +D++ YL AA++ETLRLYP+VP + E D+LPDG V G +V Y YA
Sbjct: 356 EEPLGFDEIDRLVYLKAAISETLRLYPSVPEDSKHVENDDVLPDGTFVPAGSSVTYSIYA 415
Query: 236 MGRMPYIWGDDAQEFLPERWLK--DGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVS 293
GRM WG+D EF PERW+ DG F ++F++F+AGPRICLGKD AY QMK ++
Sbjct: 416 AGRMKSTWGEDCLEFNPERWISPIDGKFINHDQYRFVAFNAGPRICLGKDLAYLQMKTIA 475
Query: 294 MALVRFFRFKLENETNDVTYRTMFTLHIDHGL 325
A++ R + + V + TL + +GL
Sbjct: 476 AAVLLRHRLTVV-PGHKVEQKMSLTLFMKNGL 506
>At1g34540 cytochrome p450, putative
Length = 498
Score = 213 bits (541), Expect = 2e-55
Identities = 118/325 (36%), Positives = 190/325 (58%), Gaps = 19/325 (5%)
Query: 6 DSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPL---WKLKRVLNFGSEA 62
D+I K+ F V+ CL FM+AF + + +R+ W++K+ LN GSE
Sbjct: 180 DNICKLAFNVDCACLGHDGAVGVNFMRAFETAATIISQRFRSVASCAWRIKKKLNIGSER 239
Query: 63 ALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRDII 122
L+++I + F +++ R + + KED+LSRF+ + + +S + LRDI+
Sbjct: 240 VLRESIATVHKFADEIVRNR---IDQGRSSDHKEDLLSRFISKEEMNSPEI----LRDIV 292
Query: 123 LNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDA 182
++F++AG+DT+++ LSWFF++L +P +++K+ QE+ ++ + + + + F
Sbjct: 293 ISFILAGRDTTSSALSWFFWLLSMHPEVEDKILQELNSIRARTGKRIGEVYGFEH----- 347
Query: 183 TLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYI 242
L M+YLHAA+TE+LRLYP VP+ ++ E ++LPDG V KG + Y +AMGRM I
Sbjct: 348 -LKMMNYLHAAITESLRLYPPVPVDIKSCAEDNVLPDGTFVGKGWAITYNIFAMGRMESI 406
Query: 243 WGDDAQEFLPERWLKD--GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
WG D F PERW+ + G F+ E KF +FHAGPR+C+GKD AY QMK + A++ F
Sbjct: 407 WGKDCDRFDPERWIDETNGCFRGEDPSKFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERF 466
Query: 301 RFKLENETNDVTYRTMFTLHIDHGL 325
++ + +M TL I GL
Sbjct: 467 VVEVPGKERPEILLSM-TLRIKGGL 490
>At4g00360 probable cytochrome P450
Length = 553
Score = 211 bits (538), Expect = 3e-55
Identities = 119/331 (35%), Positives = 197/331 (58%), Gaps = 12/331 (3%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D+I + FG + E N F AF+ + +R++ P LW+LK+ L
Sbjct: 180 LRLTFDNICGLAFGKDTRTCAPGLPE-NGFASAFDRATEASLQRFILPEFLWRLKKWLGL 238
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDK--EDILSRFLLESKKDSSNMTDK 116
G E +L +++ ID ++ ++I TR++ L Q++ + +D+LSRF+ KK + ++
Sbjct: 239 GLEVSLSRSLGEIDGYLDAVINTRKQELLSQRESGVQRHDDLLSRFM---KKKDQSYSET 295
Query: 117 YLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFV 176
+LR + LNF++AG+DTS+ LSWFF+++ +P +++K+ +E+ +V ++ ++++
Sbjct: 296 FLRHVALNFILAGRDTSSVALSWFFWLITTHPTVEDKIVREICSVLIETRGTDVS-SWTA 354
Query: 177 SNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAM 236
+ +D++ YL AAL+ETLRLYP+VP + DILPDG V G +V Y YA
Sbjct: 355 EPLEFDEVDRLVYLKAALSETLRLYPSVPEDSKHVVNDDILPDGTFVPAGSSVTYSIYAA 414
Query: 237 GRMPYIWGDDAQEFLPERWLK--DGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSM 294
GRM WG+D EF PERW+ DG F ++F++F+AGPRICLGKD AY QMK ++
Sbjct: 415 GRMKSTWGEDCLEFKPERWISPDDGKFVNHDQYRFVAFNAGPRICLGKDLAYLQMKTIAA 474
Query: 295 ALVRFFRFKLENETNDVTYRTMFTLHIDHGL 325
A++ R + + V + TL + +GL
Sbjct: 475 AVLLRHRLTVA-PGHKVEQKMSLTLFMKNGL 504
>At1g13150 putative cytochrome P450 monooxygenase
Length = 529
Score = 209 bits (533), Expect = 1e-54
Identities = 117/338 (34%), Positives = 194/338 (56%), Gaps = 10/338 (2%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D I G G + L + F KAF ++ R++ P +WK R L+
Sbjct: 184 LRLTFDIICLAGLGADPETLAVDLPQVP-FAKAFEEATESTLFRFMIPPFIWKPMRFLDT 242
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFL-LESKKDSSNM---T 114
G E L+ + ++ FV +I R L ++ ++ D+LSR + +ES K + + T
Sbjct: 243 GYEKGLRIAVGVVHGFVDKMIVDRICELKEEETLDNRSDVLSRIIQIESHKRENEIDPST 302
Query: 115 DKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDE 174
++ R +F++AG+DTS+ LSWF +++ K+P ++ K+ E+ + +S + +E
Sbjct: 303 IRFFRQFCTSFILAGRDTSSVALSWFCWVIQKHPEVENKIICEIREILRQRGDSPTSKNE 362
Query: 175 FVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSY 234
+ + + L+ M YL AAL+ETLRL+P +PM + A E D+LPDG V KG VY+ Y
Sbjct: 363 SLFTVKE--LNNMVYLQAALSETLRLFPPIPMEMKQAIEDDVLPDGTFVRKGSRVYFSIY 420
Query: 235 AMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSM 294
AMGRM IWG D + F PERW++ G F + FK++ F+AGPR+C+GK FAY QMK+++
Sbjct: 421 AMGRMESIWGKDCEIFRPERWIQAGKFVSDDQFKYVVFNAGPRLCIGKTFAYLQMKMIAA 480
Query: 295 ALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
+++ + K+ + + + R L++ +GL + TPR
Sbjct: 481 SVLLRYSIKVVQD-HVIAPRVTTNLYMKYGLKVTITPR 517
>At2g45970 putative cytochrome P450
Length = 537
Score = 207 bits (528), Expect = 5e-54
Identities = 122/334 (36%), Positives = 198/334 (58%), Gaps = 19/334 (5%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D+I + FG + N F AF+ + +R++ P LWK KR L
Sbjct: 180 LRLTFDNICGLTFGKDPRTCAPGLP-VNTFAVAFDRATEASLQRFILPEILWKFKRWLRL 238
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRE-LLSMQKDFSDKEDILSRFLLESKKDSSNMTDKY 117
G E +L +++ +D+++ +I TR+E +++ + +D+LSRF+ K + +D+
Sbjct: 239 GLEVSLTRSLVQVDNYLSEIITTRKEEMMTQHNNGKHHDDLLSRFI----KKKESYSDET 294
Query: 118 LRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNL--DEF 175
L+ + LNF++AG+DTS+ LSWFF+++ ++P I++K+ +E+ V ++ ++ L DE
Sbjct: 295 LQRVALNFILAGRDTSSVALSWFFWLITQHPAIEDKILREICTVLVETRGDDVALWTDE- 353
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
+S LD++ +L AAL+ETLRLYP+VP + A + D+LPDG V G ++ Y Y+
Sbjct: 354 --PLSCEELDRLVFLKAALSETLRLYPSVPEDSKRAVKDDVLPDGTFVPAGSSITYSIYS 411
Query: 236 MGRMPYIWGDDAQEFLPERWLKD---GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIV 292
GRM WG+D EF PERW+ G F FKF++F+AGPRICLGKD AY QMK +
Sbjct: 412 AGRMKSTWGEDCLEFKPERWISQSDGGRFINHDPFKFVAFNAGPRICLGKDLAYLQMKSI 471
Query: 293 SMALVRFFRFKLENET-NDVTYRTMFTLHIDHGL 325
+ A++ R +L T + V + TL + +GL
Sbjct: 472 ASAVL--LRHRLTVVTGHKVEQKMSLTLFMKYGL 503
>At5g58860 cytochrome P450 CYP86A1
Length = 513
Score = 207 bits (527), Expect = 7e-54
Identities = 126/349 (36%), Positives = 194/349 (55%), Gaps = 34/349 (9%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKR--YLDPLWKLKRVLNF 58
++ T D+I + FG + L + N F AF+ + KR Y LW++++ +
Sbjct: 183 LRLTFDNICGLTFGKDPETLSLDLPD-NPFSVAFDTATEATLKRLLYTGFLWRIQKAMGI 241
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
GSE LKK++++++ +++ I R+ S +D+LSRFL + + + + L
Sbjct: 242 GSEDKLKKSLEVVETYMNDAIDARKN--------SPSDDLLSRFLKKRDVNGNVLPTDVL 293
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQ--------ESNL 170
+ I LNF++AG+DTS+ LSWFF+++ N ++ K+ E+ V ++ E L
Sbjct: 294 QRIALNFVLAGRDTSSVALSWFFWLVMNNREVETKIVNELSMVLKETRGNDQEKWTEEPL 353
Query: 171 NLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVY 230
DE D++ YL AAL ETLRLYP+VP + + D+LPDG V +G TV
Sbjct: 354 EFDE---------ADRLVYLKAALAETLRLYPSVPQDFKYVVDDDVLPDGTFVPRGSTVT 404
Query: 231 YLSYAMGRMPYIWGDDAQEFLPERWL-KDG--IFQPESSFKFISFHAGPRICLGKDFAYR 287
Y Y++GRM IWG+D EF PERWL DG P+ +KF++F+AGPR CLGKD AY
Sbjct: 405 YSIYSIGRMKTIWGEDCLEFRPERWLTADGERFETPKDGYKFVAFNAGPRTCLGKDLAYN 464
Query: 288 QMK-IVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRCDV 335
QMK + S L+R+ F + + V + TL + +GL +Y PR +V
Sbjct: 465 QMKSVASAVLLRYRVFPVPG--HRVEQKMSLTLFMKNGLRVYLQPRGEV 511
>At4g39480 cytochrome P450 - like protein
Length = 516
Score = 207 bits (526), Expect = 9e-54
Identities = 114/337 (33%), Positives = 192/337 (56%), Gaps = 9/337 (2%)
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNFGSE 61
T D+ F + G + CL E F +A +D+ +F R+ P +WK++R++ G+E
Sbjct: 180 TFDTTFVLATGYDPGCLSVEMPEIE-FARALDDAEEAIFYRHFKPEVVWKMQRLIGVGAE 238
Query: 62 AALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDIL-SRFLLESKKDS--SNMTDKYL 118
LK+ I D I ++R+ +S D S +D+L S +++ K + D++L
Sbjct: 239 LKLKRAHAIFDRVCSKCIASKRDEISQGIDSSSSKDLLMSSINVDTTKYKLLNPSDDRFL 298
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEV-INVTSTSQESNLNLDEFVS 177
RD IL+FM+AG+DT+ + L+WFF++LC N K+ QE+ N+ ++ + ++
Sbjct: 299 RDTILSFMLAGRDTTGSALTWFFWLLCNNQEAMTKIRQEINTNLFPRNKTDDGSVSYDSD 358
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
+ + + K+ YLH A+ E LRLYP VP + ++ + D+LP G+ V + + Y++G
Sbjct: 359 SFNPQEVKKLVYLHGAVCEALRLYPPVPFNHKSPAKPDVLPSGHKVKANSRILFCLYSLG 418
Query: 238 RMPYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMAL 296
RM +WG+DA EF PERW+ + G E S+KF+SF+AGPR CLGK+ A QMK V++ +
Sbjct: 419 RMKSVWGEDAMEFKPERWISESGRSVHEPSYKFLSFNAGPRTCLGKEVAMTQMKTVAVKI 478
Query: 297 VRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRC 333
++ + + E + + LH+ HGL + + RC
Sbjct: 479 IQNYDINVV-EGHKIKPAPSVILHMKHGLKVTVSKRC 514
>At4g39490 cytochrome P450 - like protein
Length = 479
Score = 205 bits (522), Expect = 2e-53
Identities = 120/342 (35%), Positives = 195/342 (56%), Gaps = 16/342 (4%)
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNFGSE 61
T D+ F + G + CL E F +A +D+ +F R++ P W+L+ +L G E
Sbjct: 140 TFDTTFVLATGYDPGCLSVEMPEVE-FARALDDAEEAIFFRHVKPEIFWRLQGLLGLGDE 198
Query: 62 AALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDK--EDILSRFL-LESKKDS--SNMTDK 116
+ K +D I +R+ +S + D +D+L+ ++ L++ K + ++
Sbjct: 199 KKMTKARSTLDRVCSKYIAIKRDEVSRGTNNVDSHSKDLLTSYMNLDTTKYKLLNPSDER 258
Query: 117 YLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFV 176
+LRD IL FM+AG+DT+ + L+WFF++L KNP + K+ QE+ T+ Q S ++ D
Sbjct: 259 FLRDTILTFMLAGRDTTGSGLTWFFWLLIKNPEVIAKIRQEIN--TALFQRSKVDDDASN 316
Query: 177 SNISDA----TLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYL 232
+N SD+ L K+ YLH A+ E+LRLYP VP ++ + D+LP G+ V+ + +
Sbjct: 317 NNDSDSFSPQELKKLVYLHGAICESLRLYPPVPFQHKSPTKPDVLPSGHKVDANSKILFC 376
Query: 233 SYAMGRMPYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKI 291
Y++GRM +WG+DA EF PERW+ + G E S+KF+SF+AGPR CLGK+ A QMK
Sbjct: 377 LYSLGRMKSVWGEDALEFKPERWISESGNSVHEPSYKFLSFNAGPRTCLGKEVAMMQMKS 436
Query: 292 VSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRC 333
V++ +++ + K+ E + LH+ HGL + T RC
Sbjct: 437 VAVKIIQNYEMKIV-EGQQIEPAPSVILHMKHGLKVTVTKRC 477
>At2g27690 putative cytochrome P450
Length = 495
Score = 204 bits (519), Expect = 6e-53
Identities = 123/336 (36%), Positives = 193/336 (56%), Gaps = 26/336 (7%)
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP---LWKLKRVLNFGS 60
+ D+I K+ FG + +CL + F AF+ ++ KR L P LWK KR+L GS
Sbjct: 178 SFDTISKLSFGFDPDCLRLPFPISE-FAVAFDTASLLSAKRALAPFPLLWKTKRLLRIGS 236
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E L+++I +I+ LIK RR M K+ D++SRF+ +D D+YLRD
Sbjct: 237 EKKLQESINVINRLAGDLIKQRRLTGLMGKN-----DLISRFMAVVAEDD----DEYLRD 287
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQES-NLNLDEFVSNI 179
I+++F++AG+DT A L+ FF++L ++P ++ ++ +E+ V T +S DE
Sbjct: 288 IVVSFLLAGRDTVAAGLTGFFWLLTRHPEVENRIREELDRVMGTGFDSVTARCDE----- 342
Query: 180 SDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRM 239
+ +M YLHA+L E++RL+P V + A D+L DG VN G V Y +YAMGRM
Sbjct: 343 ----MREMDYLHASLYESMRLFPPVQFDSKFALNDDVLSDGTFVNSGTRVTYHAYAMGRM 398
Query: 240 PYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVR 298
IWG D +EF PERWL +G F+PE+ K+ F AG R+C+GK+ A +MK +++A++R
Sbjct: 399 DRIWGPDYEEFKPERWLDNEGKFRPENPVKYPVFQAGARVCIGKEMAIMEMKSIAVAIIR 458
Query: 299 FFRFKLEN--ETNDVTYRTMFTLHIDHGLPLYATPR 332
F ++ + T + + T ++ GLP+ R
Sbjct: 459 RFETRVASPETTETLRFAPGLTATVNGGLPVMIQER 494
>At2g23180 putative cytochrome P450
Length = 516
Score = 202 bits (514), Expect = 2e-52
Identities = 118/344 (34%), Positives = 196/344 (56%), Gaps = 21/344 (6%)
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNFGSE 61
T D+ F + GV+ CL + F +A +++ +F R++ P +WK++R + FG E
Sbjct: 181 TFDTSFVLATGVDPGCLSTEMPQIE-FARALDEAEEAIFFRHVKPEIVWKMQRFIGFGDE 239
Query: 62 AALKKNIKIIDDFVHSLIKTRRELLS---MQKDFSDKEDILSRFLLES------KKDSSN 112
+KK D I ++R+ ++ + D S K+ ++ +++ K +
Sbjct: 240 LKMKKAHSTFDRVCSKCIASKRDEITNGVINIDSSSKDLLMCYMNVDTICHTTKYKLLNP 299
Query: 113 MTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNL 172
DK+LRD+IL+FM+AG+DT+++ L+WFF++L KNP K+ QE+ T S +N +
Sbjct: 300 SDDKFLRDMILSFMLAGRDTTSSALTWFFWLLSKNPKAITKIRQEIN--TQLSPRTN-DF 356
Query: 173 DEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYL 232
D F + L+K+ Y+H AL E LRLYP VP ++ + D+LP G+ V+ + +
Sbjct: 357 DSFNAQ----ELNKLVYVHGALCEALRLYPPVPFQHKSPTKSDVLPSGHRVDASSKIVFC 412
Query: 233 SYAMGRMPYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKI 291
Y++GRM +WG+DA EF PERW+ + G SFKF+SF+AGPR CLGK+ A QMK
Sbjct: 413 LYSLGRMKSVWGEDASEFKPERWISESGRLIHVPSFKFLSFNAGPRTCLGKEVAMTQMKT 472
Query: 292 VSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRCDV 335
V++ +++ + K+ E + + LH+ HGL + T R ++
Sbjct: 473 VAVKIIQNYEIKVV-EGHKIEPVPSIILHMKHGLKVTVTKRSNL 515
>At4g32170 cytochrome p450 - like protein
Length = 506
Score = 201 bits (510), Expect = 6e-52
Identities = 123/338 (36%), Positives = 186/338 (54%), Gaps = 20/338 (5%)
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNFGSE 61
T D+IF + G + L E F KA +D + R+ P LWKL+ + FG E
Sbjct: 179 TFDTIFILVTGSDPRSLSIEMPEDE-FAKALDDVGEGILYRHFKPRFLWKLQNWIGFGQE 237
Query: 62 AALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD--KEDILSRFL-LESKKDS--SNMTDK 116
L + D I +RE + + S+ +D+L+ F+ L++ K + DK
Sbjct: 238 KKLTEANATFDRVCAKYISAKREEIKRSQGTSNGGSQDLLTSFIKLDTTKYKLLNPSDDK 297
Query: 117 YLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFV 176
+LRD IL F++AG+DT+A LSWFF++L +NP + K+ QE+ N+N D
Sbjct: 298 FLRDNILAFILAGRDTTATALSWFFWLLSENPHVVAKIHQEI----------NINTDLSR 347
Query: 177 SNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAM 236
+ S +DK+ YLH AL E +RLYP V ++ + D+LP G+ V+ + YA+
Sbjct: 348 TGNSQENVDKLVYLHGALCEAMRLYPPVSFGRKSPIKSDVLPSGHKVDANSKIIICLYAL 407
Query: 237 GRMPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
GRM +WG+DA +F PERW+ ++G + E SFKF+SF+AGPR CLGK A QMKIV++
Sbjct: 408 GRMRAVWGEDASQFKPERWISENGGIKHEPSFKFLSFNAGPRTCLGKHLAMTQMKIVAVE 467
Query: 296 LVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRC 333
++R + K+ + + F L + HGL + T RC
Sbjct: 468 ILRNYDIKV-LQGQKIVPALGFILSMKHGLQITVTKRC 504
>At1g47620 hypothetical protein
Length = 520
Score = 196 bits (498), Expect = 2e-50
Identities = 112/322 (34%), Positives = 180/322 (55%), Gaps = 28/322 (8%)
Query: 30 FMKAFNDSNAFVFKRYLDP--LWKLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLS 87
F KA +D + R++ P +WKL++ + G+E + K D +I +RE L
Sbjct: 213 FSKALDDVGDAIVHRHITPRFVWKLQKWIGIGTEKKMLKAHATFDRVCEKIIAAKREELG 272
Query: 88 MQ----KDFSDKEDILSRFLLESKKDSSNMT------DKYLRDIILNFMIAGKDTSANTL 137
Q ++ED+L+ F+ K D++ DK+LRD + FM AG+D++A+TL
Sbjct: 273 SQGITYNSNGEREDLLTSFI---KLDATKYEVLKPSHDKFLRDFTIGFMAAGRDSTASTL 329
Query: 138 SWFFYMLCKNPLIQEKVAQEV-INVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTE 196
+WFF+ L KNP + K+ QE+ N+ T + +++ + L+K+ YLH AL+E
Sbjct: 330 TWFFWNLSKNPNVLTKILQEINTNLPRTGSDQDMS----------SYLNKLVYLHGALSE 379
Query: 197 TLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWL 256
++RLYP +P ++ + D+LP G+ V + YAMGRM IWG+DA EF PERW+
Sbjct: 380 SMRLYPPIPFQRKSPIKEDVLPSGHKVKSNINIMIFIYAMGRMKTIWGEDAMEFKPERWI 439
Query: 257 KD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRT 315
+ G + E S+KF+SF+AGPR CLGK+ A MK V + +++ + K+ + + +
Sbjct: 440 SETGGVRHEPSYKFLSFNAGPRTCLGKNLAMNLMKTVIVEILQNYEIKIVS-GQKIEPKP 498
Query: 316 MFTLHIDHGLPLYATPRCDVAE 337
LH+ HGL + T +C E
Sbjct: 499 GLILHMKHGLKVTMTKKCSSLE 520
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,314,064
Number of Sequences: 26719
Number of extensions: 304365
Number of successful extensions: 1430
Number of sequences better than 10.0: 255
Number of HSP's better than 10.0 without gapping: 230
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 760
Number of HSP's gapped (non-prelim): 263
length of query: 337
length of database: 11,318,596
effective HSP length: 100
effective length of query: 237
effective length of database: 8,646,696
effective search space: 2049266952
effective search space used: 2049266952
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146342.8


