

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.5 + phase: 0
(507 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
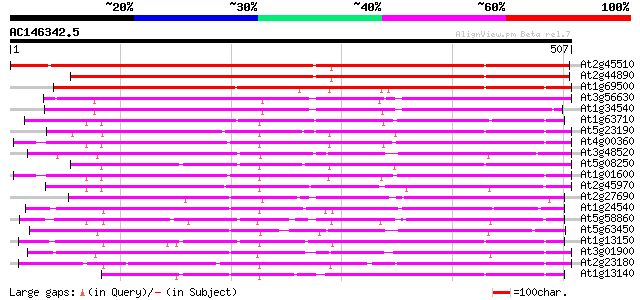
Score E
Sequences producing significant alignments: (bits) Value
At2g45510 putative cytochrome P450 515 e-146
At2g44890 putative cytochrome P450 485 e-137
At1g69500 unknown protein 386 e-107
At3g56630 cytochrome P450-like protein 321 5e-88
At1g34540 cytochrome p450, putative 309 2e-84
At1g63710 cytochrome P450 like protein 298 5e-81
At5g23190 cytochrome P450-like protein 293 2e-79
At4g00360 probable cytochrome P450 291 4e-79
At3g48520 cytochrome P450-like protein 290 1e-78
At5g08250 cytochrome P450-like protein 290 1e-78
At1g01600 unknown protein 284 7e-77
At2g45970 putative cytochrome P450 280 1e-75
At2g27690 putative cytochrome P450 276 1e-74
At1g24540 putative cytochrome P450 275 4e-74
At5g58860 cytochrome P450 CYP86A1 272 3e-73
At5g63450 cytochrome P450-like protein 272 4e-73
At1g13150 putative cytochrome P450 monooxygenase 267 9e-72
At3g01900 putative cytochrome P450 258 7e-69
At2g23180 putative cytochrome P450 247 1e-65
At1g13140 cytochrome P450 monooxygenase like protein 244 6e-65
>At2g45510 putative cytochrome P450
Length = 511
Score = 515 bits (1326), Expect = e-146
Identities = 263/513 (51%), Positives = 355/513 (68%), Gaps = 12/513 (2%)
Query: 1 MDFLSNPLLFAALSASLTLLL-VQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLH 59
M+ L++ + A + + L + L++R KS + K+Y PV TVF+ + + + L+
Sbjct: 1 MEILTSIAITVATTIFIVLCFTIYLMIRIFTGKSRN--DKRYAPVHATVFDLLFHSDELY 58
Query: 60 HYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGI 119
Y T++AR+ TYR L+P +SE+ T++P NVE+ILKT F+NY KG + +N+ DLLG GI
Sbjct: 59 DYETEIAREKPTYRFLSPGQSEILTADPRNVEHILKTRFDNYSKGHSSRENMADLLGHGI 118
Query: 120 FTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFM 179
F VDGEKWR+QRK+SS EFSTR+LRDFS S+FR+NA+K+ VSE A S + QD+ M
Sbjct: 119 FAVDGEKWRQQRKLSSFEFSTRVLRDFSCSVFRRNASKLVGFVSEFALSGKAFDAQDLLM 178
Query: 180 KSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSE 239
+ TLDSIF V FG E+ + G S+EG+ F +FD + T R++D WK+K F NIGS+
Sbjct: 179 RCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGSQ 238
Query: 240 AALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYD-----TKYLR 294
+ L+ + +++FV LI T+ +++ + VR+ DILSRFL E D KYLR
Sbjct: 239 SKLKKSIATIDKFVYSLITTKRKELAKEQNTVVRE--DILSRFLVESEKDPENMNDKYLR 296
Query: 295 DIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREAT-NTKTVSSCTEFVSSVT 353
DIILNF+IAGKDTT LSWF+YMLCK P VQEK QE+R+ T + + + FV S+
Sbjct: 297 DIILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESIN 356
Query: 354 DEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMK 413
+EA+++M+Y+HA L+ETLRLYP +P D + DD LPDG+ V KGD + Y YAMGRM
Sbjct: 357 EEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMT 416
Query: 414 FIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGC 473
+IWG DAEEF+PERWL ++G+FQPE PFKF +F AGPRICLGK+FAYRQMKI S LL
Sbjct: 417 YIWGQDAEEFKPERWL-KDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLHF 475
Query: 474 FRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
FRFK+ DE V YK M+TLH+DGGL + A+ R
Sbjct: 476 FRFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>At2g44890 putative cytochrome P450
Length = 490
Score = 485 bits (1248), Expect = e-137
Identities = 242/457 (52%), Positives = 323/457 (69%), Gaps = 9/457 (1%)
Query: 56 NRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLL 115
++L+ Y T++AR T+R L+P +SE++T++P NVE+ILKT F NY KG NL DLL
Sbjct: 34 HKLYDYETEIARTKPTFRFLSPGQSEIFTADPRNVEHILKTRFHNYSKGPVGTVNLADLL 93
Query: 116 GDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQ 175
G GIF VDGEKW++QRK+ S EFSTR+LR+FS S+FR +A+K+ ++E A S + Q
Sbjct: 94 GHGIFAVDGEKWKQQRKLVSFEFSTRVLRNFSYSVFRTSASKLVGFIAEFALSGKSFDFQ 153
Query: 176 DIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLN 235
D+ MK TLDSIF V FG E+ + G S+EG+ F +FD + T R D FWK+K FLN
Sbjct: 154 DMLMKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLN 213
Query: 236 IGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYD-----T 290
IGSE+ L+ + I+++FV LI T+ +++ + SVR+ DILS+FL E D
Sbjct: 214 IGSESRLKKSIAIIDKFVYSLITTKRKELSKEQNTSVRE--DILSKFLLESEKDPENMND 271
Query: 291 KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNT-KTVSSCTEFV 349
KYLRDIILN ++AGKDTT +LSWF+YMLCK P VQEK QE+R+ T++ + + F+
Sbjct: 272 KYLRDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFI 331
Query: 350 SSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAM 409
SVT+EA+ +M Y+HA L+ET+RLYP +P + DD LPDG+ V KGD + Y YAM
Sbjct: 332 ESVTEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAM 391
Query: 410 GRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 469
GRM +IWG DAEEF+PERWL ++G+FQPE FKF +F AGPRIC+GK+FAYRQMKI S
Sbjct: 392 GRMTYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMA 450
Query: 470 LLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
LL FRFK+ DE V+YK M+TLH+DGGL + A+ R
Sbjct: 451 LLHFFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>At1g69500 unknown protein
Length = 524
Score = 386 bits (992), Expect = e-107
Identities = 212/499 (42%), Positives = 306/499 (60%), Gaps = 34/499 (6%)
Query: 40 KYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFE 99
K P+ G Q+ NF+R+H ++ + +T + PF + Y ++P NVEY+LKTNF
Sbjct: 29 KTWPLVGAAIEQLTNFDRMHDWLVEYLYNSRTVVVPMPFTTYTYIADPINVEYVLKTNFS 88
Query: 100 NYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVA 159
NY KG + ++ LLGDGIF DGE WR+QRK +S EF+++ LRDFST +F++ + K+
Sbjct: 89 NYPKGETYHSYMEVLLGDGIFNSDGELWRKQRKTASFEFASKNLRDFSTVVFKEYSLKLF 148
Query: 160 NIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALT 219
I+S+A+ ++++Q++ M+ TLDSI V FG EI ++ E +FA +FD A+ +
Sbjct: 149 TILSQASFKEQQVDMQELLMRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIV 207
Query: 220 LYRYVDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTR--------IQQMKNSKGDS 271
R++D WK+KKFLNIGSEA L + +++N+F +I R I N+ ++
Sbjct: 208 TLRFIDPLWKMKKFLNIGSEALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNN 267
Query: 272 VRKGGDILSRFLQVKE-----YDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQ 326
+ DILSRF+++ + K LRDI+LNFVIAG+DTT TL+W +YM+ V
Sbjct: 268 NKVKHDILSRFIEISDDPDSKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVA 327
Query: 327 EKAAQEVR-------EATNTKT-----------VSSCTEFVSSVTDEAIEKMNYVHAVLT 368
EK E++ EATNT TEF + +++ K++Y+HAV+T
Sbjct: 328 EKLYSELQELEKESAEATNTSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVIT 387
Query: 369 ETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERW 428
ETLRLYPA+P D K DD LP+G VK G MV+Y PY+MGRM++ WG DA F+PERW
Sbjct: 388 ETLRLYPAVPQDPKGVLEDDMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERW 447
Query: 429 LDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYK 488
L ++G+FQ PFKFTAFQAGPRICLGK+ AY QMK+ A+L ++F L V Y+
Sbjct: 448 L-KDGVFQNASPFKFTAFQAGPRICLGKDSAYLQMKMAMAILCRFYKFHLVPNHP-VKYR 505
Query: 489 TMITLHIDGGLEIKALYRN 507
M L + GL++ R+
Sbjct: 506 MMTILSMAHGLKVTVSRRS 524
>At3g56630 cytochrome P450-like protein
Length = 499
Score = 321 bits (823), Expect = 5e-88
Identities = 190/489 (38%), Positives = 284/489 (57%), Gaps = 25/489 (5%)
Query: 31 NKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLL--NPFRSE-VYTSEP 87
N S+ K Y P+ G++ + N +R + + + T + P + + V T+ P
Sbjct: 24 NSSSEFGFKSY-PIVGSLPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKLQFVMTANP 82
Query: 88 SNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFS 147
+NVEY+LKT FE++ KG L+D LG GIF DGE W +QRK +S+EFST+ LRDF
Sbjct: 83 ANVEYMLKTKFESFPKGERFISILEDFLGRGIFNSDGEMWWKQRKTASYEFSTKSLRDFV 142
Query: 148 TS-IFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGK 206
S + + ++ +++EAA + +++QDI + D+I + F + + G
Sbjct: 143 MSNVTVEINTRLVPVLAEAATNGKLIDLQDILERFAFDNICKLAFNVDSACLGDDGAAGV 202
Query: 207 NFANSFDNASALTLYRYVDVF---WKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQ 263
NF +F+ A+ + R+ V WKIKK LNIGSE LR + I+++F +++ RI+Q
Sbjct: 203 NFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSERVLRESIMIVHKFADEIVRNRIEQ 262
Query: 264 MKNSKGDSVRKGGDILSRFLQVKEYDT-KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKY 322
K S D+LSRF+ +E ++ + LRDI+++F++AG+DTT LSWF ++L +
Sbjct: 263 GKVSDHKE-----DLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMH 317
Query: 323 PAVQEKAAQE---VREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPF 379
P V++K QE +RE T K + F E ++ MNY+HA +TE+LRLYP +P
Sbjct: 318 PEVKDKILQELNSIRERTG-KRIGEVYGF------EDLKLMNYLHAAITESLRLYPPVPV 370
Query: 380 DAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDE-NGIFQPE 438
D C D+ LPDG + K +SY YAMGRM+ IWG D + F PERW+DE NG F+ E
Sbjct: 371 DTMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESIWGKDCDRFDPERWIDETNGGFRGE 430
Query: 439 CPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGG 498
P+KF AF AGPR+CLGKE AY QMK A +L F ++ +K+ +TL I GG
Sbjct: 431 NPYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLERFVVEVPGKKERPEILMSVTLRIRGG 490
Query: 499 LEIKALYRN 507
L ++ R+
Sbjct: 491 LNVRVQERS 499
>At1g34540 cytochrome p450, putative
Length = 498
Score = 309 bits (792), Expect = 2e-84
Identities = 182/479 (37%), Positives = 278/479 (57%), Gaps = 23/479 (4%)
Query: 32 KSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLL--NPFRSE-VYTSEPS 88
KS+S K +P+ G+ + N +R + + + T + P + + + T+ PS
Sbjct: 24 KSSSEFGFKSYPIVGSFPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKQQLIMTANPS 83
Query: 89 NVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFST 148
NVEY+LKT FE++ KG L+D LG GIF DG+ W +QRK +S+EFST+ LRDF
Sbjct: 84 NVEYMLKTKFESFPKGQQFTSVLEDFLGHGIFNSDGDMWWKQRKTASYEFSTKSLRDFVM 143
Query: 149 S-IFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKN 207
S + + ++ ++ EAA + +++QDI + D+I + F + + G N
Sbjct: 144 SNVTVEINTRLVPVLVEAATTGKLIDLQDILERFAFDNICKLAFNVDCACLGHDGAVGVN 203
Query: 208 FANSFDNASALTLYRYVDVF---WKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQM 264
F +F+ A+ + R+ V W+IKK LNIGSE LR + +++F +++ RI Q
Sbjct: 204 FMRAFETAATIISQRFRSVASCAWRIKKKLNIGSERVLRESIATVHKFADEIVRNRIDQG 263
Query: 265 KNSKGDSVRKGGDILSRFLQVKEYDT-KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYP 323
++S D+LSRF+ +E ++ + LRDI+++F++AG+DTT LSWF ++L +P
Sbjct: 264 RSSDHKE-----DLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMHP 318
Query: 324 AVQEKAAQEVRE--ATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDA 381
V++K QE+ A K + F E ++ MNY+HA +TE+LRLYP +P D
Sbjct: 319 EVEDKILQELNSIRARTGKRIGEVYGF------EHLKMMNYLHAAITESLRLYPPVPVDI 372
Query: 382 KICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDE-NGIFQPECP 440
K C D+ LPDG V KG ++Y +AMGRM+ IWG D + F PERW+DE NG F+ E P
Sbjct: 373 KSCAEDNVLPDGTFVGKGWAITYNIFAMGRMESIWGKDCDRFDPERWIDETNGCFRGEDP 432
Query: 441 FKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGL 499
KF AF AGPR+C+GK+ AY QMK A +L F ++ +++ +M TL I GGL
Sbjct: 433 SKFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERFVVEVPGKERPEILLSM-TLRIKGGL 490
>At1g63710 cytochrome P450 like protein
Length = 523
Score = 298 bits (763), Expect = 5e-81
Identities = 183/505 (36%), Positives = 290/505 (57%), Gaps = 24/505 (4%)
Query: 14 SASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARK----Y 69
+A++ L L+ + + S K + P+ G++ + N +R+H ++ D R Y
Sbjct: 5 TAAIILTLIVTYIIWFVSLRRSYKGPRVWPLVGSLPALITNAHRMHDFIADNLRMCGGTY 64
Query: 70 KTYRLLNPFRSE-----VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDG 124
+T PF ++ T +P N+E+ILKT F+NY KG DLLGDGIF DG
Sbjct: 65 QTCIFPIPFLAKKQGHVTVTCDPKNLEHILKTRFDNYPKGPSWQSVFHDLLGDGIFNSDG 124
Query: 125 EKWREQRKISSHEFSTRMLRD-FSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTL 183
+ WR QRK ++ EF+TR LR + + R ++ I+ A + +++QD+ ++ T
Sbjct: 125 DTWRFQRKTAALEFTTRTLRQAMARWVDRAIKNRLVPILESARSRAEPIDLQDVLLRLTF 184
Query: 184 DSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGSEAA 241
D+I + FG + ++ E FA +FD A+ TL R++ + WKI+K+L +G E
Sbjct: 185 DNICGLTFGKDPRTLSPEFPEN-GFAVAFDGATEATLQRFIMPEFIWKIRKWLRLGLEDD 243
Query: 242 LRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKE-YDTKYLRDIILNF 300
+ + ++ ++ ++INTR ++ + D R D+LSRF++ KE Y KYL+ + LNF
Sbjct: 244 MSRSISHVDNYLSEIINTRKLELLGQQQDESRHD-DLLSRFMKKKESYSDKYLKYVALNF 302
Query: 301 VIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREA---TNTKTVSSCTEFVSSVTDEAI 357
++AG+DT+ +SWF +++ P V+EK E+ T VS T+ +T + I
Sbjct: 303 ILAGRDTSSVAMSWFFWLVSLNPRVEEKIINEICTILIKTRDTNVSKWTD--EPLTFDEI 360
Query: 358 EKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWG 417
+++ Y+ A L+ETLRLYP++P D+K A+D LPDG V G V+Y Y++GRMKFIWG
Sbjct: 361 DQLVYLKAALSETLRLYPSVPEDSKFVVANDVLPDGTFVPSGSNVTYSIYSVGRMKFIWG 420
Query: 418 DDAEEFRPERWLDENGIFQPEC-PFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRF 476
+D EF+PERWL+E+ +C +KF AF AGPRICLGK+ AY QMK +A +L R
Sbjct: 421 EDCLEFKPERWLEESR--DEKCNQYKFVAFNAGPRICLGKDLAYLQMKSITASILLRHRL 478
Query: 477 KLNDEKKNVTYKTMITLHIDGGLEI 501
+ + V K +TL + GL++
Sbjct: 479 TVAPGHR-VEQKMSLTLFMKFGLKM 502
>At5g23190 cytochrome P450-like protein
Length = 559
Score = 293 bits (749), Expect = 2e-79
Identities = 173/496 (34%), Positives = 291/496 (57%), Gaps = 28/496 (5%)
Query: 34 NSMKKKKYHPVAGTVFNQMMNF------NRLHHYMTD-LARKYKTYRLLNPFRSEV---Y 83
+++++KKY + F M+ ++ +++D L + T++ P+ S +
Sbjct: 52 HALRQKKYQGLPVWPFLGMLPSLAFGLRGNIYEWLSDVLCLQNGTFQFRGPWFSSLNSTI 111
Query: 84 TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRML 143
T +P NVE++LK F + KG Y NL+DLLGDGIF D E W+ QRK +S EF +
Sbjct: 112 TCDPRNVEHLLKNRFSVFPKGSYFRDNLRDLLGDGIFNADDETWQRQRKTASIEFHSAKF 171
Query: 144 RDFST-SIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMC-GT 201
R +T S+F ++ ++ + S++ +++QD+ ++ T D++ + FG +D C G
Sbjct: 172 RQLTTQSLFELVHKRLLPVLETSVKSSSPIDLQDVLLRLTFDNVCMIAFG--VDPGCLGP 229
Query: 202 SEEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINT 259
+ FA +F++A+ + R+V WK ++L+IG+E L+ + + +++F ++I T
Sbjct: 230 DQPVIPFAKAFEDATEAAVVRFVMPTCVWKFMRYLDIGTEKKLKESIKGVDDFADEVIRT 289
Query: 260 RIQQMKNSKGDSVRKGGDILSRFLQVKE-----YDTKYLRDIILNFVIAGKDTTGGTLSW 314
R +++ + +G++ ++ D+L+ F+ +++ + K+LRDI +NF++AG+DT+ LSW
Sbjct: 290 RKKEL-SLEGETTKRS-DLLTVFMGLRDEKGESFSDKFLRDICVNFILAGRDTSSVALSW 347
Query: 315 FMYMLCKYPAVQEKAAQEVREATNTKTVSSCTE---FVSSVTDEAIEKMNYVHAVLTETL 371
F ++L K P V+EK E+ + + E + E I+KM+Y+ A L+E L
Sbjct: 348 FFWLLEKNPEVEEKIMVEMCKILRQRDDHGNAEKSDYEPVFGPEEIKKMDYLQAALSEAL 407
Query: 372 RLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDE 431
RLYP++P D K DD PDG +KKGD V Y YAMGRM+ IWG D EFRPERWL
Sbjct: 408 RLYPSVPVDHKEVQEDDVFPDGTMLKKGDKVIYAIYAMGRMEAIWGKDCLEFRPERWL-R 466
Query: 432 NGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMI 491
+G F E +KFTAF GPR+CLGK+FAY QMK +A ++ ++ K+ + K V K +
Sbjct: 467 DGRFMSESAYKFTAFNGGPRLCLGKDFAYYQMKSTAAAIVYRYKVKVVNGHK-VEPKLAL 525
Query: 492 TLHIDGGLEIKALYRN 507
T+++ GL + + R+
Sbjct: 526 TMYMKHGLMVNLINRS 541
>At4g00360 probable cytochrome P450
Length = 553
Score = 291 bits (746), Expect = 4e-79
Identities = 178/522 (34%), Positives = 289/522 (55%), Gaps = 30/522 (5%)
Query: 4 LSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
+SN +L A+ A+ L ++ S +K + PV G++ + +R+H ++T
Sbjct: 3 VSNTMLLVAVVAAYWLWFQRI--------SRWLKGPRVWPVLGSLPGLIEQRDRMHDWIT 54
Query: 64 DLARK----YKTYRLLNPFRSE-----VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 114
+ R Y+T PF ++ T +P N+E++LKT F+NY KG D
Sbjct: 55 ENLRACGGTYQTCICAVPFLAKKQGLVTVTCDPKNIEHMLKTRFDNYPKGPTWQAVFHDF 114
Query: 115 LGDGIFTVDGEKWREQRKISSHEFSTRMLRD-FSTSIFRKNAAKVANIVSEAANSNTKLE 173
LG GIF DG+ W QRK ++ EF+TR LR + R + I+ A N+ ++
Sbjct: 115 LGQGIFNSDGDTWLFQRKTAALEFTTRTLRQAMGRWVNRGIKLRFCPILETAQNNYEPVD 174
Query: 174 IQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWKIK 231
+QD+ ++ T D+I + FG + + C FA++FD A+ +L R++ + W++K
Sbjct: 175 LQDLILRLTFDNICGLAFGKDTRT-CAPGLPENGFASAFDRATEASLQRFILPEFLWRLK 233
Query: 232 KFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKE--YD 289
K+L +G E +L + ++ ++ +INTR Q++ + + V++ D+LSRF++ K+ Y
Sbjct: 234 KWLGLGLEVSLSRSLGEIDGYLDAVINTRKQELLSQRESGVQRHDDLLSRFMKKKDQSYS 293
Query: 290 TKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREA---TNTKTVSSCT 346
+LR + LNF++AG+DT+ LSWF +++ +P V++K +E+ T VSS T
Sbjct: 294 ETFLRHVALNFILAGRDTSSVALSWFFWLITTHPTVEDKIVREICSVLIETRGTDVSSWT 353
Query: 347 EFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQP 406
+ + ++++ Y+ A L+ETLRLYP++P D+K DD LPDG V G V+Y
Sbjct: 354 --AEPLEFDEVDRLVYLKAALSETLRLYPSVPEDSKHVVNDDILPDGTFVPAGSSVTYSI 411
Query: 407 YAMGRMKFIWGDDAEEFRPERWLD-ENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKI 465
YA GRMK WG+D EF+PERW+ ++G F ++F AF AGPRICLGK+ AY QMK
Sbjct: 412 YAAGRMKSTWGEDCLEFKPERWISPDDGKFVNHDQYRFVAFNAGPRICLGKDLAYLQMKT 471
Query: 466 FSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYRN 507
+A +L R + K V K +TL + GL + R+
Sbjct: 472 IAAAVLLRHRLTVAPGHK-VEQKMSLTLFMKNGLLVNVHKRD 512
>At3g48520 cytochrome P450-like protein
Length = 506
Score = 290 bits (743), Expect = 1e-78
Identities = 173/512 (33%), Positives = 276/512 (53%), Gaps = 37/512 (7%)
Query: 17 LTLLLVQLLLRKLNNKSNSMKKKKY---------HPVAGTVFNQMMNFNRLHHYMTDLAR 67
L+ L++ L+ + S+S KK +P+ G++ + N +RL + T+L R
Sbjct: 5 LSFLILAFLITIIFFLSSSSTKKVQENTTYGPPSYPLIGSILSFNKNRHRLLQWYTELLR 64
Query: 68 KYKTYRLLNPF---RSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDG 124
+ +L P R + T+ P NVEYILKTNF N+ KG L DLLG GIF VDG
Sbjct: 65 LSPSQTILVPLLGNRRTIITTNPLNVEYILKTNFFNFPKGKPFTDLLGDLLGGGIFNVDG 124
Query: 125 EKWREQRKISSHEFSTRMLRDFSTSIFRKNAA-KVANIVSEAANSNTKLEIQDIFMKSTL 183
W QRK++SHEFSTR LR F+ + + ++ ++S AA+ T +++QD+ +
Sbjct: 125 HSWSSQRKLASHEFSTRSLRSFAFEVLKDEVENRLVPVLSTAADVGTTVDLQDVLKRFAF 184
Query: 184 DSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVD---VFWKIKKFLNIGSEA 240
D + V G + D + T +FD A+ ++ R + WK K+ LN+GSE
Sbjct: 185 DVVCKVSLGWDPDCLDLTRPVNP-LVEAFDTAAEISARRATEPIYAVWKTKRVLNVGSER 243
Query: 241 ALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKYLRDIILNF 300
LR ++ V +++ + + ++ G ++ D+LSRFL ++ + +RD++++F
Sbjct: 244 KLREAIRTVHVLVSEIVRAKKKSLEIGTGAEAKQ--DLLSRFLAAG-HNGEAVRDMVISF 300
Query: 301 VIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAIEKM 360
++AG+DTT ++W ++L + V+ K +EV + + E +++M
Sbjct: 301 IMAGRDTTSAAMTWLFWLLTENDDVERKILEEVDPLVSL-----------GLGFEDLKEM 349
Query: 361 NYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDA 420
Y A L E +RLYP + +D+K DD LPDG VK+GD V+Y PY MGRM+ +WG D+
Sbjct: 350 AYTKACLCEAMRLYPPVSWDSKHAANDDVLPDGTRVKRGDKVTYFPYGMGRMETLWGTDS 409
Query: 421 EEFRPERWLDE-----NGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFR 475
EEF P RW D + +P P+KF FQAGPR+C+GKE A+ QMK +L F
Sbjct: 410 EEFNPNRWFDSEPGSTRPVLKPISPYKFPVFQAGPRVCVGKEMAFMQMKYVVGSVLSRFE 469
Query: 476 FKLNDEKKNVTYKTMITLHIDGGLEIKALYRN 507
+ K + ++T H+ GGL++K R+
Sbjct: 470 I-VPVNKDRPVFVPLLTAHMAGGLKVKIKRRS 500
>At5g08250 cytochrome P450-like protein
Length = 550
Score = 290 bits (742), Expect = 1e-78
Identities = 165/468 (35%), Positives = 273/468 (58%), Gaps = 23/468 (4%)
Query: 56 NRLHHYMTD-LARKYKTYRLLNPFRSE---VYTSEPSNVEYILKTNFENYGKGLYNYQNL 111
+ ++ +++D L + T+R P+ S V T +P NVE++LKT F Y KG Y + +
Sbjct: 81 SNIYEWLSDVLISQNGTFRFRGPWFSTLNCVVTCDPRNVEHLLKTRFSIYPKGSYFRETM 140
Query: 112 KDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTK 171
+DLLGDGIF D W+ QRK +S EF + R ++ + V N + ++ K
Sbjct: 141 QDLLGDGIFNTDDGTWQRQRKAASVEFHSAKFRQLTSQSLHE---LVHNRLLPVLETSGK 197
Query: 172 LEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWK 229
+++QDI ++ T D++ + FG + + E FA +F++A+ T+ R+V WK
Sbjct: 198 IDLQDILLRLTFDNVCMIAFGVDPGCLSPKLPEIP-FAKAFEDATEATVVRFVMPKFVWK 256
Query: 230 IKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKE-- 287
+ + LN+G+E L+ + +++F ++I TR ++M S + K D+L+ F+ +++
Sbjct: 257 LMRSLNLGTEKKLKESINGVDDFAEEVIRTRKKEM--SLETEIAKRPDLLTIFMGLRDEN 314
Query: 288 ---YDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSS 344
+ K+LRDI +NF++AG+DT+ LSWF +++ K P V+EK + + +
Sbjct: 315 GQKFSDKFLRDICVNFILAGRDTSSVALSWFFWLIEKNPEVEEKIMMGICKILEQRVDHG 374
Query: 345 CT----EFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGD 400
T E+ E I+KM+Y+ A L+ETLRLYP++P D K DD PDG +KKG+
Sbjct: 375 DTKKNMEYEPVFRPEEIKKMDYLQAALSETLRLYPSVPVDHKEVLEDDVFPDGTKLKKGE 434
Query: 401 MVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAY 460
V Y YAMGRM+ IWG D EF+PERWL +G + E +KFTAF GPR+CLGK+FAY
Sbjct: 435 KVIYAIYAMGRMETIWGKDCREFKPERWL-RDGRYMSESAYKFTAFNGGPRLCLGKDFAY 493
Query: 461 RQMKIFSAVLLGCFRFKLNDE-KKNVTYKTMITLHIDGGLEIKALYRN 507
QM+ +A ++ ++ +++D+ V K +T+++ GL++ + R+
Sbjct: 494 YQMRYVAAAIIYRYKVRVDDKGGHKVEPKMALTMYMKHGLKVNMVKRS 541
>At1g01600 unknown protein
Length = 554
Score = 284 bits (727), Expect = 7e-77
Identities = 178/524 (33%), Positives = 286/524 (53%), Gaps = 32/524 (6%)
Query: 4 LSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
+SN +L A+ L ++ S +K + P+ G++ + +R+H ++T
Sbjct: 3 ISNAMLLVAIVTGYWLWFKRI--------SRWLKGPRVWPLLGSLPGLIEQRDRMHEWIT 54
Query: 64 DLARK----YKTYRLLNPFRSE-----VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 114
+ R Y+T PF ++ T +P N+E++LKT F+NY KG DL
Sbjct: 55 ENLRACGGTYQTCIFAVPFLAKKQGLVTVTCDPKNLEHMLKTRFDNYPKGPTWQSVFHDL 114
Query: 115 LGDGIFTVDGEKWREQRKISSHEFSTRMLRD-FSTSIFRKNAAKVANIVSEAANSNTKLE 173
LG GIF DG+ W QRK ++ EF+TR LR + R + I++ A ++ ++
Sbjct: 115 LGQGIFNSDGDTWLFQRKTAALEFTTRTLRQAMGRWVNRGIKLRFCPILATAQDNAEPVD 174
Query: 174 IQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWKIK 231
+QD+ ++ T D+I + FG + + C FA++FD A+ +L R++ WK+K
Sbjct: 175 LQDLILRLTFDNICGLAFGKDTRT-CAPGLPENGFASAFDRATEASLQRFIIPKFMWKLK 233
Query: 232 KFLNIGSEAALRNNTEILNEFVIKLINTRIQQ-MKNSKGDSVRKGGDILSRFLQVK--EY 288
K+L +G E +L + ++E++ +INTR Q+ M + + ++ D+LSRF+ K Y
Sbjct: 234 KWLGLGLEVSLSRSLGEIDEYLAAVINTRKQELMSQQESGTHQRHDDLLSRFMMKKTESY 293
Query: 289 DTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVR----EATNTKTVSS 344
+L+ + LNF++AG+DT+ LSWF +++ +P V++K +E+ E T V+S
Sbjct: 294 SDTFLQHVALNFILAGRDTSSVALSWFFWLITMHPTVEDKIVREICSVLIETRGTDDVAS 353
Query: 345 CTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSY 404
TE + + I+++ Y+ A ++ETLRLYP++P D+K DD LPDG V G V+Y
Sbjct: 354 WTE--EPLGFDEIDRLVYLKAAISETLRLYPSVPEDSKHVENDDVLPDGTFVPAGSSVTY 411
Query: 405 QPYAMGRMKFIWGDDAEEFRPERWLDE-NGIFQPECPFKFTAFQAGPRICLGKEFAYRQM 463
YA GRMK WG+D EF PERW+ +G F ++F AF AGPRICLGK+ AY QM
Sbjct: 412 SIYAAGRMKSTWGEDCLEFNPERWISPIDGKFINHDQYRFVAFNAGPRICLGKDLAYLQM 471
Query: 464 KIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYRN 507
K +A +L R + K V K +TL + GL + R+
Sbjct: 472 KTIAAAVLLRHRLTVVPGHK-VEQKMSLTLFMKNGLLVNLYKRD 514
>At2g45970 putative cytochrome P450
Length = 537
Score = 280 bits (717), Expect = 1e-75
Identities = 174/496 (35%), Positives = 278/496 (55%), Gaps = 29/496 (5%)
Query: 33 SNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARK----YKTYRLLNPFRSE-----VY 83
S +K + P+ G++ + N R+H +++D R Y+T PF ++
Sbjct: 24 SRCLKGPRVWPILGSLPGLIENCERMHDWISDNLRACSGTYQTCICAIPFLAKKQGLVTV 83
Query: 84 TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRML 143
T +P N+E+ILK F+NY KG DLLG GIF DG+ W QRK ++ EF+TR L
Sbjct: 84 TCDPRNLEHILKNRFDNYPKGPTWQAVFHDLLGQGIFNSDGDTWLFQRKTAALEFTTRTL 143
Query: 144 RD-FSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTS 202
R + + R + I+ A + +++QD+ ++ T D+I + FG + C
Sbjct: 144 RQAMARWVNRAIKLRFLPILENARLGSEPIDLQDLLLRLTFDNICGLTFGKD-PRTCAPG 202
Query: 203 EEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTR 260
FA +FD A+ +L R++ ++ WK K++L +G E +L + ++ ++ ++I TR
Sbjct: 203 LPVNTFAVAFDRATEASLQRFILPEILWKFKRWLRLGLEVSLTRSLVQVDNYLSEIITTR 262
Query: 261 IQQMKNSKGDSVRKGGDILSRFLQVKE-YDTKYLRDIILNFVIAGKDTTGGTLSWFMYML 319
++M + + D+LSRF++ KE Y + L+ + LNF++AG+DT+ LSWF +++
Sbjct: 263 KEEMMTQHNNG-KHHDDLLSRFIKKKESYSDETLQRVALNFILAGRDTSSVALSWFFWLI 321
Query: 320 CKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAI-----EKMNYVHAVLTETLRLY 374
++PA+++K +E+ T V + + V+ TDE + +++ ++ A L+ETLRLY
Sbjct: 322 TQHPAIEDKILREIC----TVLVETRGDDVALWTDEPLSCEELDRLVFLKAALSETLRLY 377
Query: 375 PALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDEN-- 432
P++P D+K DD LPDG V G ++Y Y+ GRMK WG+D EF+PERW+ ++
Sbjct: 378 PSVPEDSKRAVKDDVLPDGTFVPAGSSITYSIYSAGRMKSTWGEDCLEFKPERWISQSDG 437
Query: 433 GIFQPECPFKFTAFQAGPRICLGKEFAYRQMK-IFSAVLLGCFRFKLNDEKKNVTYKTMI 491
G F PFKF AF AGPRICLGK+ AY QMK I SAVLL + K V K +
Sbjct: 438 GRFINHDPFKFVAFNAGPRICLGKDLAYLQMKSIASAVLLRHRLTVVTGHK--VEQKMSL 495
Query: 492 TLHIDGGLEIKALYRN 507
TL + GL + R+
Sbjct: 496 TLFMKYGLLVNVHERD 511
>At2g27690 putative cytochrome P450
Length = 495
Score = 276 bits (707), Expect = 1e-74
Identities = 169/456 (37%), Positives = 255/456 (55%), Gaps = 22/456 (4%)
Query: 54 NFNRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKD 113
+F L + T L R+ T + + V T+ PSNVE+ILKTNF NY KG L D
Sbjct: 48 DFINLSDWYTHLLRRSPTSTIKVHVLNSVITANPSNVEHILKTNFHNYPKGKQFSVILGD 107
Query: 114 LLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAK--VANIVSEAANSNTK 171
LLG GIF DG+ WR QRK++S E + +R F+ I + + + S + N +
Sbjct: 108 LLGRGIFNSDGDTWRFQRKLASLELGSVSVRVFAHEIVKTEIETRLLPILTSFSDNPGSV 167
Query: 172 LEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVF---W 228
L++QD+F + + D+I + FG + D + FA +FD AS L+ R + F W
Sbjct: 168 LDLQDVFRRFSFDTISKLSFGFDPDCL-RLPFPISEFAVAFDTASLLSAKRALAPFPLLW 226
Query: 229 KIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQV-KE 287
K K+ L IGSE L+ + ++N LI R ++ G + D++SRF+ V E
Sbjct: 227 KTKRLLRIGSEKKLQESINVINRLAGDLIKQR--RLTGLMGKN-----DLISRFMAVVAE 279
Query: 288 YDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTE 347
D +YLRDI+++F++AG+DT L+ F ++L ++P V+ + +E+ T S
Sbjct: 280 DDDEYLRDIVVSFLLAGRDTVAAGLTGFFWLLTRHPEVENRIREELDRVMGTGFDS---- 335
Query: 348 FVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPY 407
V++ DE + +M+Y+HA L E++RL+P + FD+K DD L DG V G V+Y Y
Sbjct: 336 -VTARCDE-MREMDYLHASLYESMRLFPPVQFDSKFALNDDVLSDGTFVNSGTRVTYHAY 393
Query: 408 AMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFS 467
AMGRM IWG D EEF+PERWLD G F+PE P K+ FQAG R+C+GKE A +MK +
Sbjct: 394 AMGRMDRIWGPDYEEFKPERWLDNEGKFRPENPVKYPVFQAGARVCIGKEMAIMEMKSIA 453
Query: 468 AVLLGCFRFKLNDEKKNVT--YKTMITLHIDGGLEI 501
++ F ++ + T + +T ++GGL +
Sbjct: 454 VAIIRRFETRVASPETTETLRFAPGLTATVNGGLPV 489
>At1g24540 putative cytochrome P450
Length = 522
Score = 275 bits (703), Expect = 4e-74
Identities = 175/505 (34%), Positives = 272/505 (53%), Gaps = 30/505 (5%)
Query: 15 ASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTD-LARKYKTYR 73
A + L L+ L KL +K + PV G + +N + L + T + R T+
Sbjct: 23 ALVGLFLLSYLREKLVSKGGPVM----WPVLGIIPMLALNKHDLFTWCTRCVVRSGGTFH 78
Query: 74 LLNPFRSEVY---TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQ 130
+ Y T++P+NVE+ILKTNF+NY KG + + +DLL DGIF D E W+E+
Sbjct: 79 YRGIWFGGAYGIMTADPANVEHILKTNFKNYPKGAFYRERFRDLLEDGIFNADDELWKEE 138
Query: 131 RKISSHEF-STRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNV 189
R+++ E S+R L T++ K+ ++ + S ++QD+ ++ T D+I
Sbjct: 139 RRVAKTEMHSSRFLEHTFTTMRDLVDQKLVPLMENLSTSKRVFDLQDLLLRFTFDNICIS 198
Query: 190 VFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGSEAALRNNTE 247
FG S+ T FA +F++A+ TL R++ WK +FL IG E L N
Sbjct: 199 AFGVYPGSL-ETGLPEIPFAKAFEDATEYTLARFLIPPFVWKPMRFLGIGYERKLNNAVR 257
Query: 248 ILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQ---VKEYDT--------KYLRDI 296
I++ F K + R +M+ K ++ D+LSR +Q KE DT KY R+
Sbjct: 258 IVHAFANKTVRERRNKMR--KLGNLNDYADLLSRLMQREYEKEEDTTRGNYFSDKYFREF 315
Query: 297 ILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEA 356
+F+IAG+DTT L WF +++ K+P V+++ +E+RE T + + E
Sbjct: 316 CTSFIIAGRDTTSVALVWFFWLVQKHPEVEKRILREIREIKRKLTTQETEDQFEA---ED 372
Query: 357 IEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIW 416
+M Y+ A LTE+LRLYP++P + K DD LPDG VKKG + Y Y+MGR++ IW
Sbjct: 373 FREMVYLQAALTESLRLYPSVPMEMKQALEDDVLPDGTRVKKGARIHYSVYSMGRIESIW 432
Query: 417 GDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRF 476
G D EEF+PERW+ E I E FK+ F GPR+C+GK+FAY QMK+ +A +L +
Sbjct: 433 GKDWEEFKPERWIKEGRIVS-EDQFKYVVFNGGPRLCVGKKFAYTQMKMVAAAILMRYSV 491
Query: 477 KLNDEKKNVTYKTMITLHIDGGLEI 501
K+ + + + K TL++ G+ +
Sbjct: 492 KV-VQGQEIVPKLTTTLYMKNGMNV 515
>At5g58860 cytochrome P450 CYP86A1
Length = 513
Score = 272 bits (696), Expect = 3e-73
Identities = 189/522 (36%), Positives = 276/522 (52%), Gaps = 59/522 (11%)
Query: 10 FAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARK- 68
+A + S+ L L R+L K P G++ + N +R+H ++ D R
Sbjct: 11 YAVAALSVYALWFYFLSRRLTGP-------KVLPFVGSLPYLIANRSRIHDWIADNLRAT 63
Query: 69 ---YKTYRLLNPFRSEVY-----TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIF 120
Y+T ++ PF ++ T P NVE+ILKT F+NY KG DLLG GIF
Sbjct: 64 GGTYQTCTMVIPFVAKAQGFYTVTCHPKNVEHILKTRFDNYPKGPMWRAAFHDLLGQGIF 123
Query: 121 TVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVAN----IVSEAANSNTKLEIQD 176
DG+ W QRK ++ EF+TR LR ++ R + N I+ A +N +++QD
Sbjct: 124 NSDGDTWLMQRKTAALEFTTRTLRQ---AMARWVNGTIKNRLWLILDRAVQNNKPVDLQD 180
Query: 177 IFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYR--YVDVFWKIKKFL 234
+F++ T D+I + FG + +++ + F+ +FD A+ TL R Y W+I+K +
Sbjct: 181 LFLRLTFDNICGLTFGKDPETLSLDLPDNP-FSVAFDTATEATLKRLLYTGFLWRIQKAM 239
Query: 235 NIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYD----- 289
IGSE L+ + E++ ++ N I KNS D D+LSRFL+ ++ +
Sbjct: 240 GIGSEDKLKKSLEVVETYM----NDAIDARKNSPSD------DLLSRFLKKRDVNGNVLP 289
Query: 290 TKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEV-------REATNTKTV 342
T L+ I LNFV+AG+DT+ LSWF +++ V+ K E+ R K
Sbjct: 290 TDVLQRIALNFVLAGRDTSSVALSWFFWLVMNNREVETKIVNELSMVLKETRGNDQEKWT 349
Query: 343 SSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMV 402
EF DEA +++ Y+ A L ETLRLYP++P D K DD LPDG V +G V
Sbjct: 350 EEPLEF-----DEA-DRLVYLKAALAETLRLYPSVPQDFKYVVDDDVLPDGTFVPRGSTV 403
Query: 403 SYQPYAMGRMKFIWGDDAEEFRPERWLDENG--IFQPECPFKFTAFQAGPRICLGKEFAY 460
+Y Y++GRMK IWG+D EFRPERWL +G P+ +KF AF AGPR CLGK+ AY
Sbjct: 404 TYSIYSIGRMKTIWGEDCLEFRPERWLTADGERFETPKDGYKFVAFNAGPRTCLGKDLAY 463
Query: 461 RQMK-IFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEI 501
QMK + SAVLL F + + V K +TL + GL +
Sbjct: 464 NQMKSVASAVLLRYRVFPVPGHR--VEQKMSLTLFMKNGLRV 503
>At5g63450 cytochrome P450-like protein
Length = 510
Score = 272 bits (695), Expect = 4e-73
Identities = 167/504 (33%), Positives = 265/504 (52%), Gaps = 43/504 (8%)
Query: 19 LLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLLNPF 78
+L+ + L K+ S + + G++ + N +RL + TDL R + +
Sbjct: 18 VLIFSFPTKTLKAKTASPSNPTSYQLIGSILSFNKNRHRLLQWYTDLLRLSPSQTITVDL 77
Query: 79 ---RSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISS 135
R + T+ P NVE+ILKTNF N+ KG L DLLG GIF DGE W QRK++S
Sbjct: 78 LFGRRTIITANPENVEHILKTNFYNFPKGKPFTDLLGDLLGGGIFNSDGELWSSQRKLAS 137
Query: 136 HEFSTRMLRDFSTSIFRKNAA-KVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTE 194
HEF+ R LR+F+ I R+ ++ ++S A + ++ Q++ + D + V G +
Sbjct: 138 HEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLKRFAFDVVCKVSLGWD 197
Query: 195 IDSMCGTSEEGKNFANSFDNASALTLYRYVD---VFWKIKKFLNIGSEAALRNNTEILNE 251
D + + +FD A+ ++ R + WK+K+FLN+GSE LR
Sbjct: 198 PDCL-DLTRPVPELVKAFDVAAEISARRATEPVYAVWKVKRFLNVGSEKRLRE------- 249
Query: 252 FVIKLINTRIQQMKNSKGDSVRKGGDI------LSRFLQVKEYDTKYLRDIILNFVIAGK 305
IK ++ + ++ +K S+ GGD+ LSRFL + + +RD +++F++AG+
Sbjct: 250 -AIKTVHLSVSEIIRAKKKSLDIGGDVSDKQDLLSRFLAAG-HGEEAVRDSVISFIMAGR 307
Query: 306 DTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVHA 365
DTT ++W ++L + V+ K E+R + + E + +M+Y A
Sbjct: 308 DTTSAAMTWLFWLLSQNDDVETKILDELRNKGSL-----------GLGFEDLREMSYTKA 356
Query: 366 VLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRP 425
L E +RLYP + +D+K DD LPDG +KKGD V+Y PY MGRM+ +WG D +EF+P
Sbjct: 357 CLCEAMRLYPPVAWDSKHAANDDILPDGTPLKKGDKVTYFPYGMGRMEKVWGKDWDEFKP 416
Query: 426 ERWLDE------NGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLN 479
RW +E + + FKF FQAGPR+C+GKE A+ QMK +L RFK+
Sbjct: 417 NRWFEEEPSYGTKPVLKSVSSFKFPVFQAGPRVCIGKEMAFTQMKYVVGSVLS--RFKII 474
Query: 480 DEKKN-VTYKTMITLHIDGGLEIK 502
N + ++T H+ GGL++K
Sbjct: 475 PVCNNRPVFVPLLTAHMAGGLKVK 498
>At1g13150 putative cytochrome P450 monooxygenase
Length = 529
Score = 267 bits (683), Expect = 9e-72
Identities = 170/512 (33%), Positives = 273/512 (53%), Gaps = 31/512 (6%)
Query: 9 LFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARK 68
LF L + L L + L KL NK M PV G + N ++ ++T +K
Sbjct: 13 LFDLLLSLLGLFVFCCLREKLTNKRGPM----LWPVFGITLEFFFHINDVYGWVTRSLKK 68
Query: 69 YK-TYRLLNPFRSEVY---TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDG 124
+ T+ + Y T P+NVEY+LKTNF+N+ KG + DLL +GIF D
Sbjct: 69 CRGTFLYRGVWLDGSYGAVTCVPANVEYMLKTNFKNFPKGTFFKSRFNDLLEEGIFNADD 128
Query: 125 EKWREQRKISSHEFSTR--MLRDFSTS--IFRKNAAKVANIVSEAANSNTKLEIQDIFMK 180
+ W+EQR+I E + + F T+ + RK K+ ++ A S ++QD+F++
Sbjct: 129 DSWKEQRRIIITEMHSTGFVEHSFQTTQHLVRK---KLLKVMESFAKSQEAFDLQDVFLR 185
Query: 181 STLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGS 238
T D I G + +++ + FA +F+ A+ TL+R++ WK +FL+ G
Sbjct: 186 LTFDIICLAGLGADPETLAVDLPQVP-FAKAFEEATESTLFRFMIPPFIWKPMRFLDTGY 244
Query: 239 EAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDT-------- 290
E LR +++ FV K+I RI ++K +++ D+LSR +Q++ +
Sbjct: 245 EKGLRIAVGVVHGFVDKMIVDRICELKEE--ETLDNRSDVLSRIIQIESHKRENEIDPST 302
Query: 291 -KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFV 349
++ R +F++AG+DT+ LSWF +++ K+P V+ K E+RE + S ++
Sbjct: 303 IRFFRQFCTSFILAGRDTSSVALSWFCWVIQKHPEVENKIICEIREILRQRGDSPTSKNE 362
Query: 350 SSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAM 409
S T + + M Y+ A L+ETLRL+P +P + K DD LPDG V+KG V + YAM
Sbjct: 363 SLFTVKELNNMVYLQAALSETLRLFPPIPMEMKQAIEDDVLPDGTFVRKGSRVYFSIYAM 422
Query: 410 GRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 469
GRM+ IWG D E FRPERW+ + G F + FK+ F AGPR+C+GK FAY QMK+ +A
Sbjct: 423 GRMESIWGKDCEIFRPERWI-QAGKFVSDDQFKYVVFNAGPRLCIGKTFAYLQMKMIAAS 481
Query: 470 LLGCFRFKLNDEKKNVTYKTMITLHIDGGLEI 501
+L + K+ + + + L++ GL++
Sbjct: 482 VLLRYSIKVVQDHV-IAPRVTTNLYMKYGLKV 512
>At3g01900 putative cytochrome P450
Length = 496
Score = 258 bits (658), Expect = 7e-69
Identities = 160/502 (31%), Positives = 264/502 (51%), Gaps = 33/502 (6%)
Query: 17 LTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLLN 76
L LLLV + + NS K +PV G + + N NRL + T+L + + ++
Sbjct: 12 LVLLLVSAGKHVIYSCRNSTPKT--YPVIGCLISFYTNRNRLLDWYTELLTESPSRTVVI 69
Query: 77 ---PFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKI 133
R V T+ PSNVEYILKTNF+NY KG + L D LG+GIF VDG W +QR++
Sbjct: 70 RRLAARRTVVTANPSNVEYILKTNFDNYPKGKPFTEILGDFLGNGIFNVDGNLWLKQRRL 129
Query: 134 SSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGT 193
++H+F+ + LR++ T + + ++ ++ AA + ++Q++ + T + + V G
Sbjct: 130 ATHDFTPKSLREYVTVLRNEVEKELLAFLNAAAEDSQPFDLQELLRRFTFNIVCIVFLGI 189
Query: 194 EIDSMCGTSEEGKNFANSFDNASALTLYR---YVDVFWKIKKFLNIGSEAALRNNTEILN 250
+ ++ S F +F ASA++ R + WK K+ + GSE LR ++
Sbjct: 190 DRCTL-NPSSPVSEFDRAFQTASAVSAGRGSAPLSFVWKFKRLVGFGSEKELRKAVGEVH 248
Query: 251 EFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKYLRDIILNFVIAGKDTTGG 310
V ++I + ++ N D LSR + E D + +RD++++ ++AG+DTT
Sbjct: 249 NCVDEIIRDKKRKPANQ---------DFLSRLIVAGESD-ETVRDMVISIIMAGRDTTSA 298
Query: 311 TLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTET 370
+ +++ + + E+R E E+++K++ + A L E
Sbjct: 299 VATRLFWLITGHEETEHDLVSEIRSVKE--------EITGGFDYESLKKLSLLKACLCEV 350
Query: 371 LRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLD 430
+RLYP +P+D+K DD LPDG V+ GD V+Y PY MGRM+ +WG+D +EF+P RW +
Sbjct: 351 MRLYPPVPWDSKHALTDDRLPDGTLVRAGDRVTYFPYGMGRMEELWGEDWDEFKPNRWAE 410
Query: 431 ENG-----IFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNV 485
+ + PFKF FQAGPR+CLG+E AY QMK A +L F + K
Sbjct: 411 SYDKTCCRVLKKVNPFKFPVFQAGPRVCLGEEMAYVQMKYIVASILDRFEIEPIPTDK-P 469
Query: 486 TYKTMITLHIDGGLEIKALYRN 507
+ M+T H+ GG++++ R+
Sbjct: 470 DFVPMLTAHMAGGMQVRVHRRD 491
>At2g23180 putative cytochrome P450
Length = 516
Score = 247 bits (630), Expect = 1e-65
Identities = 157/523 (30%), Positives = 262/523 (50%), Gaps = 36/523 (6%)
Query: 9 LFAALSASLTLLLVQLLLRK-LNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLAR 67
L L S++LL L L +K P G + ++ R++ ++T+L
Sbjct: 3 LITLLEVSISLLFFSFLYGYFLISKKPHRSFLTNWPFLGMLPGLLVEIPRVYDFVTELL- 61
Query: 68 KYKTYRLLNPFRSEVY-------TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIF 120
+ L PF+ + T +P+N+ +I+ +NF NY KG ++ + D+LGDGIF
Sbjct: 62 --EASNLTYPFKGPCFGGLDMLITVDPANIHHIMSSNFANYPKGT-EFKKIFDVLGDGIF 118
Query: 121 TVDGEKWREQRKISSHEFSTRMLRDFST-SIFRKNAAKVANIVSEAANSNTKLEIQDIFM 179
D E W++ RK + + + + F+ +I K + ++ A +++QD+F
Sbjct: 119 NADSELWKDLRKSAQSMMTHQDFQRFTLRTIMSKLEKGLVPLLDYVAEKKQVVDLQDVFQ 178
Query: 180 KSTLDSIFNVVFGTEIDSMCGTSEEGK-NFANSFDNASALTLYRYV--DVFWKIKKFLNI 236
+ T D+ F V T +D C ++E + FA + D A +R+V ++ WK+++F+
Sbjct: 179 RFTFDTSF--VLATGVDPGCLSTEMPQIEFARALDEAEEAIFFRHVKPEIVWKMQRFIGF 236
Query: 237 GSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEY-------- 288
G E ++ + K I ++ ++ N + D+L ++ V
Sbjct: 237 GDELKMKKAHSTFDRVCSKCIASKRDEITNGVINIDSSSKDLLMCYMNVDTICHTTKYKL 296
Query: 289 ----DTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSS 344
D K+LRD+IL+F++AG+DTT L+WF ++L K P KA ++R+ NT+
Sbjct: 297 LNPSDDKFLRDMILSFMLAGRDTTSSALTWFFWLLSKNP----KAITKIRQEINTQLSPR 352
Query: 345 CTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSY 404
+F S + + K+ YVH L E LRLYP +PF K D LP G+ V + +
Sbjct: 353 TNDF-DSFNAQELNKLVYVHGALCEALRLYPPVPFQHKSPTKSDVLPSGHRVDASSKIVF 411
Query: 405 QPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMK 464
Y++GRMK +WG+DA EF+PERW+ E+G FKF +F AGPR CLGKE A QMK
Sbjct: 412 CLYSLGRMKSVWGEDASEFKPERWISESGRLIHVPSFKFLSFNAGPRTCLGKEVAMTQMK 471
Query: 465 IFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYRN 507
+ ++ + K+ + K + I LH+ GL++ R+
Sbjct: 472 TVAVKIIQNYEIKVVEGHK-IEPVPSIILHMKHGLKVTVTKRS 513
>At1g13140 cytochrome P450 monooxygenase like protein
Length = 519
Score = 244 bits (624), Expect = 6e-65
Identities = 148/425 (34%), Positives = 232/425 (53%), Gaps = 22/425 (5%)
Query: 84 TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEF-STRM 142
T P+NVEY+LKTNF+N+ KG + + DLL DGIF D E W+EQR+I E STR
Sbjct: 95 TCVPANVEYMLKTNFKNFPKGAFFKERFNDLLEDGIFNADAESWKEQRRIIITEMHSTRF 154
Query: 143 LR-DFSTS--IFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMC 199
+ F T+ + RK K+ ++ A S ++QD+ ++ T D+I G + ++
Sbjct: 155 VEHSFQTTQDLVRK---KLLKVMESFARSQEAFDLQDVLLRLTFDNICIAGLGDDPGTL- 210
Query: 200 GTSEEGKNFANSFDNASALTLYRYV--DVFWKIKKFLNIGSEAALRNNTEILNEFVIKLI 257
+ FA +F+ A+ T++R++ WK KF +IG E LR ++ + +
Sbjct: 211 DSDLPLVPFAQAFEEATESTMFRFMIPPFIWKPLKFFDIGYEKGLRKAVDVSMSLSTRWL 270
Query: 258 NTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDT-KYLRDIILNFVIAGKDTTGGTLSWFM 316
I + K + K D K+ T K+ R +F++AG+DT+ L+WF
Sbjct: 271 --WIVSASSKKKEQSHKTTD-------EKDPSTIKFFRQFCTSFILAGRDTSSVALTWFF 321
Query: 317 YMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPA 376
+++ K+P V+ K +E+ E + S ++ S T + + M Y+ A L+ET+RLYP
Sbjct: 322 WVIQKHPEVENKIIREISEILRQRGDSPTSKNESLFTVKELNDMVYLQAALSETMRLYPP 381
Query: 377 LPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQ 436
+P + K DD PDG ++KG V + YAMGRM+ IWG D E F+PERW+ ++G F
Sbjct: 382 IPMEMKQAIEDDVFPDGTFIRKGSRVYFATYAMGRMESIWGKDCESFKPERWI-QSGNFV 440
Query: 437 PECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHID 496
+ FK+ F AGPR+CLGK FAY QMK +A +L + K+ + V + TL++
Sbjct: 441 NDDQFKYVVFNAGPRLCLGKTFAYLQMKTIAASVLSRYSIKVAKDHV-VVPRVTTTLYMR 499
Query: 497 GGLEI 501
GL++
Sbjct: 500 HGLKV 504
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,028,364
Number of Sequences: 26719
Number of extensions: 474417
Number of successful extensions: 1909
Number of sequences better than 10.0: 255
Number of HSP's better than 10.0 without gapping: 216
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 1158
Number of HSP's gapped (non-prelim): 276
length of query: 507
length of database: 11,318,596
effective HSP length: 104
effective length of query: 403
effective length of database: 8,539,820
effective search space: 3441547460
effective search space used: 3441547460
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146342.5


