

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146333.2 - phase: 0
(615 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
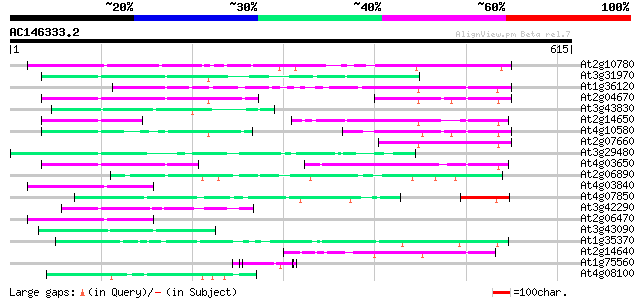
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 167 2e-41
At3g31970 hypothetical protein 98 1e-20
At1g36120 putative reverse transcriptase gb|AAD22339.1 96 6e-20
At2g04670 putative retroelement pol polyprotein 87 3e-17
At3g43830 putative protein 84 2e-16
At2g14650 putative retroelement pol polyprotein 84 2e-16
At4g10580 putative reverse-transcriptase -like protein 82 7e-16
At2g07660 putative retroelement pol polyprotein 82 9e-16
At3g29480 hypothetical protein 82 1e-15
At4g03650 putative reverse transcriptase 79 7e-15
At2g06890 putative retroelement integrase 70 3e-12
At4g03840 putative transposon protein 66 5e-11
At4g07850 putative polyprotein 65 8e-11
At3g42290 putative protein 65 8e-11
At2g06470 putative retroelement pol polyprotein 65 8e-11
At3g43090 putative protein 63 6e-10
At1g35370 hypothetical protein 63 6e-10
At2g14640 putative retroelement pol polyprotein 59 8e-09
At1g75560 DNA-binding protein 56 7e-08
At4g08100 putative polyprotein 55 9e-08
>At2g10780 pseudogene
Length = 1611
Score = 167 bits (422), Expect = 2e-41
Identities = 153/578 (26%), Positives = 240/578 (41%), Gaps = 91/578 (15%)
Query: 20 VGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKV 79
V QPN +LE + F G DP A W ++R F+ + E+ +
Sbjct: 154 VNPQPNHGRKRGDYLSLLEHVSRLGTRHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQR 213
Query: 80 RFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQ 139
H L +A +WW+ + G E V ++A F EF ++YF + + E +L+L Q
Sbjct: 214 DIAVHFLEGDAHNWWLTVEKRRGDE--VRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQ 271
Query: 140 GNMSVTEYAAKFVELSKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVN 199
GN +V EY +F L ++ E E ++ +F GLR++I+ + +LV
Sbjct: 272 GNRTVREYDEEFNRLRRYVGRELEE--EQAQLRRFIRGLRIEIRNHCLVRTFNSVSELVE 329
Query: 200 SCRIYEEDTKAHYKVVNERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFN 259
+ EE + + +N K R + PAD K++ D K DA C
Sbjct: 330 RAAMIEEGIEEE-RYLNREKA---PIRNNQSTKPAD--KKRKFDKVDNTKSDAKTGECVT 383
Query: 260 CGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTD-----------IVCFNCNGEGHISS 308
CG K H S C + I C RCG K H + C + + CF C GH+
Sbjct: 384 CG-KNH-SGTCWKAIGACGRCGSKDHAIQSCPKMEPGQSKVLGEETRTCFYCGKTGHLKR 441
Query: 309 QCTQ---------------------PKRAPTTGRVFALT--GTQTENEDRLIRGTCYINN 345
+C + PKR RV+ L+ N + G N
Sbjct: 442 ECPKLTAEKQAGQRDNRGGNGLPPPPKRQAVASRVYELSEEANDAGNFRAITGGFRKEPN 501
Query: 346 TPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLSMFG 405
T + V A G G+ + T ++ + + G
Sbjct: 502 TD---------------YGMVRAAG-------GQAMYPTGLVRGIS---------VVVNG 530
Query: 406 RDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFSKSVH---------FSSVEEESGAEF 456
+ DL+ +PL DVILGM+WL I+C + F + SG+
Sbjct: 531 VNMPADLIIVPLKKHDVILGMDWLGKYKAHIDCHRGRIQFERDEGMLKFQGIRTTSGSLV 590
Query: 457 LSTKQLKQLERDGILMF-SLMATLSIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEF 515
+S Q +++ G + + + T + A + +P+V +F +VF + VPP+R F
Sbjct: 591 ISAIQAERMLGKGCEAYLATITTKEVGASAELKDIPIVNEFSDVFA-AVSGVPPDRSDPF 649
Query: 516 SIDLVPGTKPVSMAPYRISASE---LKKQLEDLLEKKF 550
+I+L PGT P+S APYR++ +E LKKQLE+LL+K F
Sbjct: 650 TIELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGF 687
Score = 29.6 bits (65), Expect = 5.2
Identities = 10/34 (29%), Positives = 23/34 (67%)
Query: 434 VLINCFSKSVHFSSVEEESGAEFLSTKQLKQLER 467
V+++ +KS HF ++ ++ GAE ++ K + ++ R
Sbjct: 1256 VVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIVR 1289
>At3g31970 hypothetical protein
Length = 1329
Score = 98.2 bits (243), Expect = 1e-20
Identities = 105/419 (25%), Positives = 156/419 (37%), Gaps = 54/419 (12%)
Query: 35 RMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWW 94
RM+E + F G P+ A +W +E F + E +V H L +A WW
Sbjct: 145 RMMEQLQRIGTRYFFGGTSPEEADSWKSRVEHNFGSSRCPAEYRVDLAVHFLEGDAHLWW 204
Query: 95 VALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVEL 154
++ T + A M+WA F EF +YF ++ + E FLEL QG SV EY +F L
Sbjct: 205 RSV--TARRRQAHMSWADFVAEFNAKYFPQEALDRMEARFLELTQGERSVREYEREFNRL 262
Query: 155 SKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKV 214
+ ++ R +F GLR D++ Q LV + EED +
Sbjct: 263 LVYAGRGMQDDQAQMR--RFLRGLRPDLRVRCRVLQYATKAALVETAAEVEEDLQRQVVG 320
Query: 215 VN---ERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACP 271
V+ + K QQ P PA K+K E+GH CP
Sbjct: 321 VSPAVKPKKTQQQVAPSKGGKPAQGQKRK---------------------ERGHLRPNCP 359
Query: 272 EEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPTTGRVFALTGTQTE 331
+ + V + + +R + A T
Sbjct: 360 KLQRMAVAVVQPA---------------------VQHGAQVQQRVQQLAHIAAAPQGYTT 398
Query: 332 NEDRLIRGTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVETPAKGSVT 391
E I T T + D+GA+HCFI + S G D ++ A G
Sbjct: 399 RE---IGSTSKRAITEAHVLFDSGASHCFITPESASR-GNIRGDPGEQLGAVKVAGGQFL 454
Query: 392 TSLVCLK-CPLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFSKSVHFSSVE 449
L K + + G +DL+ P+ DVILGM+WL++ V ++C V F E
Sbjct: 455 AVLGRAKGVDIQIAGESMPVDLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPE 513
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 95.9 bits (237), Expect = 6e-20
Identities = 117/454 (25%), Positives = 186/454 (40%), Gaps = 63/454 (13%)
Query: 113 FRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSRCI 172
F EF +YF ++ + E FLEL +G +SV EY +F L + ++ R
Sbjct: 157 FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKRLLVYAGRGMEDDQAQMR-- 214
Query: 173 KFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKVVNERKGKGQQSRPKPYSA 232
+F GLR D++ Q L + EED + V+ Q + + A
Sbjct: 215 RFLRGLRPDLRVRCRVSQYATKAAL--TAAEVEEDLQRQVVAVSPSV---QPKKIQQQVA 269
Query: 233 PADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNR 292
P+ GK V R+ F G+ G +A + + A R
Sbjct: 270 PSKGGKPAQVQKRKWDHP-------FRAGQGGRAGSAADDSSTSTAGGAAQS---AGAAR 319
Query: 293 TDIVCFNCNGEGHISSQCTQPKRAPTTGRVFALTGTQTENEDRLIRGTCYINNTPLVAII 352
+ C +S+ C+ TT +A LI G C + +
Sbjct: 320 SAAEC------AAVSTYCSG-----TTWLYYA----------DLISGRCRGH-----VLF 353
Query: 353 DTGATHCFIAFDCVSALGL--DLSDMNGEMVVETPAKGSVTTSLVCLK-CPLSMFGRDFE 409
D+GA+HCFI + S + + + G + + A G T L K + +
Sbjct: 354 DSGASHCFITPESASRDNIRGEPGEQFGAVKI---AGGQFLTVLGRAKGVNIQIEEESMP 410
Query: 410 MDLVCLPLSGMDVILGMNWLEYNHVLINCFSKSVHFS---------SVEEESGAEFLSTK 460
DL+ P+ D ILGM+WL++ V ++C V F V S + +S
Sbjct: 411 ADLIINPVVLYDAILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPTSRSLVISEV 470
Query: 461 QLKQLERDGILMFSLMATLSIE-NQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDL 519
Q++++ G + + ++S Q + + VV +F +VF + +PP R F+I+L
Sbjct: 471 QVEKMIEKGCEAYLVTISMSESVGQVAVSDIWVVQEFEDVF-QSLQGLPPSRSDPFTIEL 529
Query: 520 VPGTKPVSMAPYRI---SASELKKQLEDLLEKKF 550
GT P+S PYR+ +ELKKQLEDLL K F
Sbjct: 530 ELGTAPLSKTPYRMVPAEIAELKKQLEDLLGKGF 563
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 87.0 bits (214), Expect = 3e-17
Identities = 57/165 (34%), Positives = 87/165 (52%), Gaps = 18/165 (10%)
Query: 401 LSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFSKSVHFS---------SVEEE 451
+ + G DL+ P+ DVILGM+WL++ V ++C V F V
Sbjct: 384 IQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPT 443
Query: 452 SGAEFLSTKQLKQLERDGILMFSLMATLSIEN---QAVIDRLPVVCDFPEVFPDEIPDVP 508
SG+ +S Q K++ G + + T+S+ Q + + V+ +F +VF + +P
Sbjct: 444 SGSLVISAVQAKKMIEKGCEAY--LVTISMPESLGQVAVSDIRVIQEFEDVF-QSLQGLP 500
Query: 509 PEREVEFSIDLVPGTKPVSMAPYRIS---ASELKKQLEDLLEKKF 550
P R F+I+L PGT P+S APYR++ +ELKKQLEDLL K F
Sbjct: 501 PSRSDPFTIELEPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGF 545
Score = 81.6 bits (200), Expect = 1e-15
Identities = 68/241 (28%), Positives = 99/241 (40%), Gaps = 9/241 (3%)
Query: 35 RMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWW 94
RM+E + F P+ A +W ++R F + E +V H L +A WW
Sbjct: 125 RMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDAHLWW 184
Query: 95 VALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVEL 154
++ T + A M+WA F EF +YF ++ + E FLEL QG SV EY +F L
Sbjct: 185 RSV--TARRRQADMSWADFVAEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRL 242
Query: 155 SKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKV 214
+ ++ R +F GLR D++ Q LV + EED +
Sbjct: 243 LAYAGRGMEDDQAQMR--RFLRGLRPDLRVQCRVSQYATKAALVETAAEVEEDLQRQVVG 300
Query: 215 VN---ERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACP 271
V+ + K QQ P PA K+K R + A F+CG H C
Sbjct: 301 VSPAVQTKNTQQQVTPSKGGKPAQGQKRKWDHPSRAGQGGRAGY--FSCGSLDHTVADCT 358
Query: 272 E 272
E
Sbjct: 359 E 359
>At3g43830 putative protein
Length = 329
Score = 84.3 bits (207), Expect = 2e-16
Identities = 67/247 (27%), Positives = 97/247 (39%), Gaps = 45/247 (18%)
Query: 47 TFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGA 106
TF G DP A W +E F+ M+ E + L EA +WW + Q G
Sbjct: 115 TFNGGTDPVKAYNWRNMLECKFKSMRCPVEFWRDLASCYLRGEAQEWWERVKQR-EQVGC 173
Query: 107 VMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENA 166
V W+ F+ EF RRY E+ E++FL L+QG +V +Y +F L +F +
Sbjct: 174 VDQWSFFKEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFHSLERF---ERRKRG 230
Query: 167 EFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVN---SCRIYEEDTKAHYKVVNERKGKGQ 223
E KF +GLR+DI+ + DLV S I E+ + ++ E++ G
Sbjct: 231 EHELIHKFISGLRVDIRLCCHVRDFDNMIDLVEKAASLEIGLEEEARYKRIAQEKEAMG- 289
Query: 224 QSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKK 283
+ G+ GH C E C RCG
Sbjct: 290 -----------------------------------SYGQTGHSKRRCQE--VTCYRCGVA 312
Query: 284 GHVVADC 290
GH+ DC
Sbjct: 313 GHIARDC 319
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 84.3 bits (207), Expect = 2e-16
Identities = 73/250 (29%), Positives = 109/250 (43%), Gaps = 40/250 (16%)
Query: 310 CTQPKRAPTTGRVFALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATHCFIAFDCVSAL 369
CT + T+ R A+TG + GT + + D+GA+HCFI + S
Sbjct: 280 CTTREIGGTSNR--AITGFLAHE---VCVGTLLVGGVEAHVLFDSGASHCFITPESASR- 333
Query: 370 GLDLSDMNGEMVVETPAKGSVTTSLVCLK-CPLSMFGRDFEMDLVCLPLSGMDVILGMNW 428
G D ++ A G L K + + G DL+ P+ DVILGM+W
Sbjct: 334 GNIRGDPGEQLGAVKVAGGQFVAVLGRTKGVDIQIAGESMPADLIISPVELYDVILGMDW 393
Query: 429 LEYNHVLINCFSKSVHFS---------SVEEESGAEFLSTKQLKQLERDGILMFSLMATL 479
L++ V ++C V F V SG+ +S Q +++ G +
Sbjct: 394 LDHYRVHLDCHRGRVSFERPEGSLVYQGVRPTSGSLVISAVQAEKMIEKGCEAY------ 447
Query: 480 SIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMAPYRIS---AS 536
+F +VF + +PP R F+I+L PGT P+S APYR++ +
Sbjct: 448 --------------LEFEDVF-QSLQGLPPSRSDPFTIELEPGTAPLSKAPYRMAPAEMA 492
Query: 537 ELKKQLEDLL 546
ELKKQLEDLL
Sbjct: 493 ELKKQLEDLL 502
Score = 58.9 bits (141), Expect = 8e-09
Identities = 33/111 (29%), Positives = 53/111 (47%), Gaps = 2/111 (1%)
Query: 35 RMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWW 94
RM+E + F G P+ A +W +ER F + E ++ H L +A WW
Sbjct: 143 RMMEQLQRIGTGYFSGGTSPEVADSWRSRVERNFGSSRCPAEYRIDLAVHFLEGDAHLWW 202
Query: 95 VALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVT 145
++ T + A ++WA F EF +YF ++ + E FLEL QG ++
Sbjct: 203 RSV--TARRRQADISWADFVAEFNAKYFPQEALDRMEAHFLELTQGRGDIS 251
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 82.4 bits (202), Expect = 7e-16
Identities = 65/202 (32%), Positives = 98/202 (48%), Gaps = 29/202 (14%)
Query: 365 CVSALGLDLSDMNGE-MVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVI 423
C S LD D G+ + V AKG + + G DL+ P+ DVI
Sbjct: 331 CFSCGSLDHKDAGGQFLAVLGRAKG----------VDIQIAGESMPADLIISPVELYDVI 380
Query: 424 LGMNWLEYNHVLINCFSKSVHFSSVEEE---------SGAEFLSTKQLKQLERDGILMFS 474
LGM+WL+Y V ++ V F E SG+ +S Q +++ G +
Sbjct: 381 LGMDWLDYYRVHLDWHRGRVFFERPEGRLVYQGVRPISGSLVISAVQAEKMIEKGCEAY- 439
Query: 475 LMATLSIEN---QAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMAPY 531
+ T+S+ Q + + VV +F +VF + +PP + F+I+L PGT P+S APY
Sbjct: 440 -LVTISMPESVGQVAVSDIRVVQEFQDVF-QSLQGLPPSQSDPFTIELEPGTAPLSKAPY 497
Query: 532 RIS---ASELKKQLEDLLEKKF 550
R++ +ELKKQL+DLL K F
Sbjct: 498 RMAPAEMAELKKQLKDLLGKGF 519
Score = 57.0 bits (136), Expect = 3e-08
Identities = 59/235 (25%), Positives = 90/235 (38%), Gaps = 36/235 (15%)
Query: 35 RMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWW 94
RM+E + + F G P+ A +W + R F + E +V H L +A WW
Sbjct: 139 RMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLEGDAHLWW 198
Query: 95 VALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVEL 154
++ T + M+WA F EF +YF ++ ++ YA + +E
Sbjct: 199 RSV--TARRRQTDMSWADFVAEFKAKYFPQE-----------------ALDPYAGQGME- 238
Query: 155 SKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKV 214
+ A+ R F GLR D++ Q LV + EED +
Sbjct: 239 --------DDQAQMRR---FLRGLRPDLRVRCRVSQYATKAALVETAAEVEEDFQRQVVG 287
Query: 215 VN---ERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHK 266
V+ + K QQ P PA K+K R + A CF+CG HK
Sbjct: 288 VSPVVQPKKTQQQVTPSKGGKPAQGQKRKWDHPSRAGQGGRAR--CFSCGSLDHK 340
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 82.0 bits (201), Expect = 9e-16
Identities = 54/159 (33%), Positives = 85/159 (52%), Gaps = 14/159 (8%)
Query: 405 GRDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFSKSVHFS---------SVEEESGAE 455
G + DL+ +PL DVILGM+WL I+C + + F + SG+
Sbjct: 25 GVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHRERIQFERDEGMLKFQGIRTTSGSL 84
Query: 456 FLSTKQLKQLERDGILMF-SLMATLSIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVE 514
+S Q +++ G + + + T + A + + +V +F +VF + VPP+R
Sbjct: 85 VISAIQAERMLGKGCEAYLATITTKEVGASAELKDILIVNEFSDVFA-AVSGVPPDRSDP 143
Query: 515 FSIDLVPGTKPVSMAPYRIS---ASELKKQLEDLLEKKF 550
F+I+L PGT P+S APYR++ +ELKKQLE+LL K F
Sbjct: 144 FTIELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGF 182
Score = 29.6 bits (65), Expect = 5.2
Identities = 10/34 (29%), Positives = 23/34 (67%)
Query: 434 VLINCFSKSVHFSSVEEESGAEFLSTKQLKQLER 467
V+++ +KS HF ++ ++ GAE ++ K + ++ R
Sbjct: 648 VVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIMR 681
>At3g29480 hypothetical protein
Length = 718
Score = 81.6 bits (200), Expect = 1e-15
Identities = 104/447 (23%), Positives = 158/447 (35%), Gaps = 94/447 (21%)
Query: 2 AGRNDAALAAALQAVAQAVGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWL 61
AG + A + V Q+ V + RM+E + F G P+ A +W
Sbjct: 5 AGGAQVPVQAPVAPRVAEVQQRAAVAEEVPSYLRMMEQLQRIGIGYFSGGTSPEEADSWR 64
Query: 62 KEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRY 121
+ER F + E +V H L +A WW ++ T + A M+WA F EF +Y
Sbjct: 65 SRVERNFGSSRCPVEYRVDLAVHFLEGDAHLWWKSV--TTRRRQANMSWADFVAEFNAKY 122
Query: 122 FLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSRCIKFENGLRLD 181
F ++ + E FLEL QG SV EY +F
Sbjct: 123 FPQEALDRMEAHFLELTQGERSVREYDREF------------------------------ 152
Query: 182 IKRAIGYQQLRVFQDLVNSCRI-YEEDTKAHYKVVNERKGKGQQSRPKPYSAPADKGKQK 240
R + Y + R S RI + K H +V + GK Q + + + P+ G+
Sbjct: 153 -NRLLVYAECR-------STRIPVVQPKKTHQQVAPSKGGKPAQGQKRSWDHPSRAGQ-- 202
Query: 241 MVDVRRPKKKDAAEIVCFNCGEKGHKSNACPE--EIKKCVRCGKKGHVVADCNRTDIVCF 298
CF CG HK C + E ++ C K+ H+ +C + +
Sbjct: 203 -----------GGRAGCFFCGSLDHKVADCTQRAETREYYHCRKRRHLRPNCPKLQRMAV 251
Query: 299 NCNGEGHISSQCTQPKRAPTTGRVFALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATH 358
H + Q + A V L + R NN + A+ G
Sbjct: 252 ---AVVHAAVQ----QGAQVQQNVQQLAHIAPAPQGYTTREIGVTNNRAITAVKVVGGQ- 303
Query: 359 CFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLS 418
F+A A G+D+ M P ++F E+
Sbjct: 304 -FLAV-LGRAKGVDIQIARESM-------------------PANLFLSPEEL-------- 334
Query: 419 GMDVILGMNWLEYNHVLINCFSKSVHF 445
DVILGM+WL+Y V ++C V F
Sbjct: 335 -YDVILGMDWLDYYRVHLDCHRGRVSF 360
>At4g03650 putative reverse transcriptase
Length = 839
Score = 79.0 bits (193), Expect = 7e-15
Identities = 69/228 (30%), Positives = 103/228 (44%), Gaps = 16/228 (7%)
Query: 324 ALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVE 383
AL T E E+ L R + +P V T HC + SA ++ GE +
Sbjct: 300 ALVETAAEVEEDLQRQV--VGVSPAVQPKKTQQQHCIASLPPESASRGNIRGDPGEQLGA 357
Query: 384 TPAKGSVTTSLV--CLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFSK 441
G +++ + + G DL+ + DVILGM+WL++ V ++C
Sbjct: 358 VKVAGGQFLAVLGGAKGVDIQIAGESLPADLIISHVELYDVILGMDWLDHYRVHLDCHRG 417
Query: 442 SVHFSSVEEESGAEFLSTKQLKQLERDGILMFSLMATLSIENQAVIDRLPVVCDFPEVFP 501
V F E +G E + L+ L D I + Q + + VV +F VF
Sbjct: 418 RVSFERPEGSAGRE-NDREGLRGLPGDDIY-------AGVCGQVAVSDIRVVQEFQYVF- 468
Query: 502 DEIPDVPPEREVEFSIDLVPGTKPVSMAPYRIS---ASELKKQLEDLL 546
+ +PP R F+I+L P T P+S APYR++ +ELKKQLEDLL
Sbjct: 469 QSLQGLPPSRSDPFTIELEPKTAPLSKAPYRMAPAKMAELKKQLEDLL 516
Score = 72.8 bits (177), Expect = 5e-13
Identities = 52/173 (30%), Positives = 77/173 (44%), Gaps = 4/173 (2%)
Query: 35 RMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWW 94
RM++ + F G P+ A +W ++ER F + E +V H L +A WW
Sbjct: 143 RMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWW 202
Query: 95 VALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVEL 154
++ T + A M+WA F EF +YF + + E FLEL QG SV EY KF L
Sbjct: 203 RSV--TARRRQADMSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRL 260
Query: 155 SKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEED 207
+ ++ R +F GLR +++ Q LV + EED
Sbjct: 261 LVYAGRGMEDDQAQMR--RFLRGLRPNLRVRCRVSQYATKAALVETAAEVEED 311
>At2g06890 putative retroelement integrase
Length = 1215
Score = 70.1 bits (170), Expect = 3e-12
Identities = 106/497 (21%), Positives = 185/497 (36%), Gaps = 106/497 (21%)
Query: 111 AVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSR 170
A+ R+ F+ ++ ++ ++ L QGN SV EY + L A SR
Sbjct: 3 AIMRKRFVPSHYHRELH----LKLRNLTQGNRSVEEYYKEMETLMLRADISEDREATLSR 58
Query: 171 CIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTK--------------AHYKVVN 216
F L DI+ + Q +++++ ++E+ K A
Sbjct: 59 ---FLGDLNRDIQDRLETQYYVQIEEMLHKAILFEQQVKRKSSSRSSYGSGTIAKPTYQR 115
Query: 217 ERKGKGQQSRP--------KPYSAPAD-KGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKS 267
E + ++P KPY+A D KGK ++ R ++ C+ C KGH +
Sbjct: 116 EERTSSYHNKPIVSPRAESKPYAAVQDHKGKAEISTSR------VRDVRCYKCQGKGHYA 169
Query: 268 NACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPTTGRVFALTG 327
N CP K+ + G + + D S + + P G +
Sbjct: 170 NECPN--KRVMILLDNGEIEPEEEIPDS-----------PSSLKENEELPAQGELLVARR 216
Query: 328 T--------QTENEDRLIRGTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGE 379
T + E L C+++ IID G+ + V LGL + +G+
Sbjct: 217 TLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVASETMVKKLGLKWLNDSGK 276
Query: 380 MVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVC--LPLSGMDVILGMNWLEYNHVLIN 437
M V K V +V K +E +++C LP+ ++LG W V+ +
Sbjct: 277 MRV----KNQVVVPIVIGK---------YEDEILCDVLPMEAGHILLGRPWQSDRKVMHD 323
Query: 438 CFS----------KSVHFSSVEEESGAEFLSTKQLKQL-------------------ERD 468
F+ K++ S E + + KQ K+ +
Sbjct: 324 GFTNRHSFEFKGGKTILVSMTPHEVYQDQIHLKQKKEQVVKQPNFFAKSGEVKSAYSSKQ 383
Query: 469 GILMFSLMATL-SIENQAVI---DRLPVVCDFPEVFPDEIPD-VPPEREVEFSIDLVPGT 523
+L+F L S+ N A + + ++ D+ +VFP++ P +PP R +E ID VPG
Sbjct: 384 PMLLFVFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGA 443
Query: 524 KPVSMAPYRISASELKK 540
+ YR + E K+
Sbjct: 444 SLPNRPAYRTNPVETKE 460
>At4g03840 putative transposon protein
Length = 973
Score = 66.2 bits (160), Expect = 5e-11
Identities = 40/138 (28%), Positives = 64/138 (45%), Gaps = 2/138 (1%)
Query: 20 VGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKV 79
V QPN +LE + F G DP A W ++R F+ + E+ +
Sbjct: 119 VNPQPNHGRKRGDYLSLLEHVSRLGTRHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQR 178
Query: 80 RFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQ 139
H L +A +WW+ + G E V ++A F EF ++YF + + E +L+L Q
Sbjct: 179 DIAVHFLERDAHNWWLTVEKRRGDE--VRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQ 236
Query: 140 GNMSVTEYAAKFVELSKF 157
GN +V EY +F L ++
Sbjct: 237 GNRTVREYDEEFNRLRRY 254
>At4g07850 putative polyprotein
Length = 1138
Score = 65.5 bits (158), Expect = 8e-11
Identities = 88/385 (22%), Positives = 141/385 (35%), Gaps = 60/385 (15%)
Query: 72 QYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEG--AVMTWAVFRREFLRRYFLEDVRGK 129
+YTE KV+ + + A WW ++ T + G ++ TW + RR+ +
Sbjct: 89 EYTEANKVKVAATEFYDYALSWWDQVVTTKRRLGDDSIETWNQLKNIMKRRFVPSHYHRE 148
Query: 130 KEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQ 189
L QGN +V EY F E+ + + +F GL DI
Sbjct: 149 LHQRLRNLVQGNRTVEEY---FKEMETLMLRADVQEECEATMSRFMGGLNRDILDRFEVI 205
Query: 190 QLRVFQDLVNSCRIYEEDTKAHYKVVNERKGKGQQSRPKP-YSAPADKGKQKMVDVRRPK 248
++L + ++E K + R K + KP Y G QK +P
Sbjct: 206 HYENLEELFHKAVMFE-------KQIKRRSAKPSYNSSKPSYQREEKSGFQKEY---KPF 255
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISS 308
K E + KG + KC +C GH ++C+ I+ +GE + S
Sbjct: 256 VKPKVEEI----SSKGKEKEVTRTRDLKCFKCHGLGHYASECSNKRIMIIRDSGE--VES 309
Query: 309 QCTQPKRAP----------TTGRVFALTGTQTENEDR--LIRGTCYINNTPLVAIIDTGA 356
+ +P+ + T RV ++ E R L C I IID G+
Sbjct: 310 EDEKPEESDVEEAPKGELLVTMRVLSVLNKAEEQAQRENLFHTRCLIKGKVCSLIIDGGS 369
Query: 357 THCFIAFDCVSALGLD---------LSDMN--GEMVVETPAKGSVTTSLVCLKCPLSMFG 405
+ V LGL+ L +N GEM V ++ PL++
Sbjct: 370 CTNVASETMVQKLGLEEFPHPKPYKLQWLNESGEMAVTRQ-----------VQVPLAI-- 416
Query: 406 RDFEMDLVC--LPLSGMDVILGMNW 428
+E +++C LPL V+LG W
Sbjct: 417 GKYEDEILCDILPLEASHVLLGRPW 441
Score = 45.8 bits (107), Expect = 7e-05
Identities = 23/58 (39%), Positives = 36/58 (61%), Gaps = 4/58 (6%)
Query: 495 DFPEVFPDEIPD-VPPEREVEFSIDLVPGTKPVSMAPYR---ISASELKKQLEDLLEK 548
D+ +VFP+E P+ +PP R +E ID VPG + YR + EL+KQ+ +L+E+
Sbjct: 447 DYQDVFPEENPEGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELEKQVNELMER 504
>At3g42290 putative protein
Length = 462
Score = 65.5 bits (158), Expect = 8e-11
Identities = 54/211 (25%), Positives = 90/211 (42%), Gaps = 27/211 (12%)
Query: 57 AQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRRE 116
A +W ++R F + E +V H L E+A WW ++ T + A M+WA F E
Sbjct: 163 ADSWRSRVKRNFSSSRCLMEYRVDLAVHFLEEDAHLWWKSV--TARRRQADMSWADFVVE 220
Query: 117 FLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSRCIKFEN 176
F +YFL+D + E+ FLEL SV EY +F + Y + ++ + + +F
Sbjct: 221 FNAKYFLQDELDRMEVRFLELTLVERSVWEYDREFNRV-LVYAGWGMKDGQ-AELRRFMR 278
Query: 177 GLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKVVNERKGKGQQSRPKPYSAPADK 236
GLR D++ Q + LV + +V + GK Q + + ++ +
Sbjct: 279 GLRPDLRVRCRMSQYALKAALVETAA----------EVAPRKGGKPMQGKKRSWNHLSRA 328
Query: 237 GKQKMVDVRRPKKKDAAEIVCFNCGEKGHKS 267
G+ CF+CG HK+
Sbjct: 329 GQ-------------GGRAGCFSCGNLAHKN 346
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 65.5 bits (158), Expect = 8e-11
Identities = 40/138 (28%), Positives = 64/138 (45%), Gaps = 2/138 (1%)
Query: 20 VGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKV 79
V QPN +LE + F G DP A W ++R F+ + E+ +
Sbjct: 119 VNPQPNHGRKRGDYLSLLEHVSRLGTRHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQR 178
Query: 80 RFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQ 139
H L +A +WW+ + G E V ++A F EF ++YF + + E +L+L Q
Sbjct: 179 DIAVHFLEGDAHNWWLTVEKRRGDE--VRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQ 236
Query: 140 GNMSVTEYAAKFVELSKF 157
GN +V EY +F L ++
Sbjct: 237 GNRTVREYDEEFNRLRRY 254
>At3g43090 putative protein
Length = 357
Score = 62.8 bits (151), Expect = 6e-10
Identities = 50/194 (25%), Positives = 75/194 (37%), Gaps = 3/194 (1%)
Query: 32 AEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEAD 91
A+ +++ F G DP A W K +E F + E + H L EEA
Sbjct: 115 AQMEIMKVMRTMGTEYFTGGIDPVKADDWRKLLENNFENARCPVEFQKDIIVHFLREEAS 174
Query: 92 DWWVALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKF 151
WW +L + + W FR EF R++F ++ E +F EL+Q V E +
Sbjct: 175 HWWDGVLGNTPVQHVIF-WEDFREEFNRKFFPQEAMDSLEDDFEELRQDTKKVRENEREL 233
Query: 152 VELSKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAH 211
LS+F A E S + GLR DI ++ DLV E
Sbjct: 234 SHLSRF--SVRAGRGEQSMIRRLMRGLRPDILTRCASREFFSMIDLVEKAARIESGLVLE 291
Query: 212 YKVVNERKGKGQQS 225
K + + + Q+
Sbjct: 292 AKYLKNTQARTTQA 305
>At1g35370 hypothetical protein
Length = 1447
Score = 62.8 bits (151), Expect = 6e-10
Identities = 117/528 (22%), Positives = 206/528 (38%), Gaps = 74/528 (14%)
Query: 51 RYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTW 110
R+D WL ++E F V EE KV+ A W + + + W
Sbjct: 121 RFDGSRINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATWHHSFIQSGIGLDVFFNW 180
Query: 111 AVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSR 170
+ + R+ ED E +L++ + + EY +F EL K + + E
Sbjct: 181 PEYVKLLKDRF--EDACDDPMAELKKLQETD-GIVEYHQQF-ELIKVRLNLSEEYL---- 232
Query: 171 CIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKVVNERKGKGQQSRPKPY 230
+ GLR D + + + + +D + + YE +AH K + +R
Sbjct: 233 VSVYLAGLRTDTQMHVRMFEPKTVRDCLRLGKYYE---RAHPKKTVSSTWSQKGTRSGGS 289
Query: 231 SAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADC 290
P + +QK + +C+ C EK PE H
Sbjct: 290 YRPVKEVEQKSDHLG----------LCYFCDEK-----FTPEHYLV--------HKKTQL 326
Query: 291 NRTDIVCFNCNGEGHISSQCTQPKRAP--TTGRVFALTGTQTENEDRLIRGTCYINNTPL 348
R D+ + +S + K P + V ++G +T ++GT ++ L
Sbjct: 327 FRMDVDEEFEDAVEVLSDDDHEQKPMPQISVNAVSGISGYKTMG----VKGT--VDKRDL 380
Query: 349 VAIIDTGATHCFIAFDCVSALGLDLSDMN-GEMVVETPAKGSVTTSLVCLKCPLSMFGRD 407
+ID+G+TH FI + LG + ++ V K +V + L
Sbjct: 381 FILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIKGFTWKLQ--STT 438
Query: 408 FEMDLVCLPLSGMDVILGMNWL--------EYNHVLINCFSKS--VHFSSVEEESGAEFL 457
F+ D++ +PL G+D++LG+ WL E+ + + F K+ V + S +
Sbjct: 439 FQSDILLIPLQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWLHGIITGSVRDIK 498
Query: 458 STK-QLKQLERDGILMFSLMATLSIENQAV--IDRLP-----------VVCDFPEVFPDE 503
+ K Q Q ++ + M + +S E Q + I L +V +FP+VF E
Sbjct: 499 AHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFA-E 557
Query: 504 IPDVPPEREV-EFSIDLVPGTKPVSMAPYRI---SASELKKQLEDLLE 547
D+PP RE + I L+ G PV+ PYR E+ K ++D+++
Sbjct: 558 PTDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIK 605
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 58.9 bits (141), Expect = 8e-09
Identities = 69/250 (27%), Positives = 112/250 (44%), Gaps = 31/250 (12%)
Query: 301 NGEGHISSQCTQPKRAPTTGRVFALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATHCF 360
N + + + T+ + P ++AL G T + R +R T + N L+A+I++G+TH F
Sbjct: 286 NDDEAVEEESTEAEGKPEI-TLYALEGVDTTSTIR-VRATIHRNR--LIALINSGSTHNF 341
Query: 361 IAFDCVSALGLDLSDMNGEMVVETPAKGSVTTS--LVCLK----CPLSMFGRDFEMDLVC 414
I V MN + P V LVC P+ M G F + L
Sbjct: 342 IGEKAVRG-------MNLKATTTKPFTVRVVNGMPLVCRSRYEAIPVVMGGVVFPVTLYA 394
Query: 415 LPLSGMDVILGMNWLE-YNHVLINCFSKSVHFSSVEEE--------SGAEFLSTKQLKQL 465
LPL G+D+ +G+ WL L N +++ F +E +G + K + +
Sbjct: 395 LPLMGLDLAMGVQWLSTLGPTLCNWKEQTLQFHWAGDEVRLMGIKPTGLRGVEHKTITKK 454
Query: 466 ERDGILMFSL-MATLSIE-NQAVIDRLPVVCDFPEVFPDEIP-DVPPEREVEFSIDLVPG 522
R G +F++ MA S + N + ++ +F +F E P +P ER + I L G
Sbjct: 455 ARMGHTIFAITMAHNSSDPNTPDGEIKKLIEEFAGLF--ETPTQLPSERPIVHRIALKKG 512
Query: 523 TKPVSMAPYR 532
T PV++ PYR
Sbjct: 513 TDPVNVRPYR 522
Score = 35.4 bits (80), Expect = 0.094
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Query: 60 WLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALL---PTLGQEGAV 107
WL + ++ F + EQ VRF T L A+ WW A L T+GQ G++
Sbjct: 119 WLSKAKQYFEYHEGPVEQAVRFTTFHLDGVANGWWQATLRRSATIGQTGSL 169
>At1g75560 DNA-binding protein
Length = 257
Score = 55.8 bits (133), Expect = 7e-08
Identities = 27/68 (39%), Positives = 37/68 (53%), Gaps = 7/68 (10%)
Query: 253 AEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDI------VCFNCNGEGHI 306
AE C+NC E GH ++ C E C CGK GH DC+ +D +C NC +GH+
Sbjct: 91 AESRCWNCREPGHVASNCSNE-GICHSCGKSGHRARDCSNSDSRAGDLRLCNNCFKQGHL 149
Query: 307 SSQCTQPK 314
++ CT K
Sbjct: 150 AADCTNDK 157
Score = 52.0 bits (123), Expect = 1e-06
Identities = 22/67 (32%), Positives = 35/67 (51%), Gaps = 2/67 (2%)
Query: 245 RRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEG 304
R P++ + +C NC GH + C + C CG GH+ A+C + C+NC G
Sbjct: 45 REPRRAFSQGNLCNNCKRPGHFARDC-SNVSVCNNCGLPGHIAAECT-AESRCWNCREPG 102
Query: 305 HISSQCT 311
H++S C+
Sbjct: 103 HVASNCS 109
Score = 47.4 bits (111), Expect = 2e-05
Identities = 21/55 (38%), Positives = 30/55 (54%), Gaps = 2/55 (3%)
Query: 256 VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQC 310
+C NC ++GH + C + K C C GH+ DC R D VC C+ GH++ C
Sbjct: 139 LCNNCFKQGHLAADCTND-KACKNCRTSGHIARDC-RNDPVCNICSISGHVARHC 191
Score = 38.9 bits (89), Expect = 0.009
Identities = 16/39 (41%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Query: 277 CVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKR 315
C C + GH DC+ VC NC GHI+++CT R
Sbjct: 57 CNNCKRPGHFARDCSNVS-VCNNCGLPGHIAAECTAESR 94
Score = 33.5 bits (75), Expect = 0.36
Identities = 17/53 (32%), Positives = 28/53 (52%), Gaps = 2/53 (3%)
Query: 221 KGQQSRPKPYSAPADKGKQK--MVDVRRPKKKDAAEIVCFNCGEKGHKSNACP 271
KG + S D G Q+ + + R ++ +A I+C NCG +GH++ CP
Sbjct: 193 KGDSNYSDRGSRVRDGGMQRGGLSRMSRDREGVSAMIICHNCGGRGHRAYECP 245
>At4g08100 putative polyprotein
Length = 1054
Score = 55.5 bits (132), Expect = 9e-08
Identities = 57/255 (22%), Positives = 102/255 (39%), Gaps = 38/255 (14%)
Query: 41 MKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPT 100
+K P F G +PD W ++IE +F + + KV+ + A WW L+ +
Sbjct: 92 LKLKIPAFHGTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYALSWWDQLVTS 151
Query: 101 --LGQEGAVMTW----AVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVEL 154
++ + TW V R+ F+ Y+ ++ + L QG+ +V EY F+E+
Sbjct: 152 RRRTRDYPIKTWNQLKFVMRKRFVPSYYHRELHQR----LRNLVQGSKTVEEY---FLEM 204
Query: 155 SKFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTK----- 209
+ + +F GL +I+ + Q +++++ ++E+ K
Sbjct: 205 ETLMLRADLQEDGEAVMSRFMGGLNREIQDRLETQHYVELEEMLHKAVMFEQQIKRKNAR 264
Query: 210 AHYKVVNERKGK---------GQQSRPKPYSAPA-----DKGKQKMVDVRRPKKKDAAEI 255
+ + N GK G Q+ KP+ P KGK K V R A +I
Sbjct: 265 SSHTKTNYSSGKPSYQKEEKFGYQNDSKPFVKPKPVDLDPKGKGKEVITR------ARDI 318
Query: 256 VCFNCGEKGHKSNAC 270
CF H ++ C
Sbjct: 319 RCFKSQGLRHYASKC 333
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,568,447
Number of Sequences: 26719
Number of extensions: 590400
Number of successful extensions: 1992
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 53
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 1638
Number of HSP's gapped (non-prelim): 237
length of query: 615
length of database: 11,318,596
effective HSP length: 105
effective length of query: 510
effective length of database: 8,513,101
effective search space: 4341681510
effective search space used: 4341681510
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146333.2


