

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.5 + phase: 0
(639 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
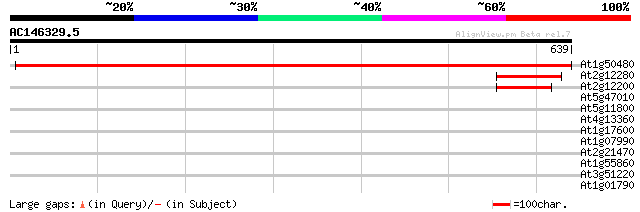
Score E
Sequences producing significant alignments: (bits) Value
At1g50480 10-formyltetrahydrofolate synthetase 1123 0.0
At2g12280 10-formyltetrahydrofolate synthetase 136 4e-32
At2g12200 10-formyltetrahydrofolate synthetase 88 1e-17
At5g47010 UPF1 31 1.9
At5g11800 potassium transport protein like 30 3.2
At4g13360 3-hydroxyisobutyryl-coenzyme A hydrolase - like protein 30 5.4
At1g17600 disease resistance protein, putative 30 5.4
At1g07990 unknown protein 30 5.4
At2g21470 ubiquitin activating enzyme like protein 29 7.0
At1g55860 ubiquitin-protein ligase 1, putative 29 7.0
At3g51220 unknown protein 29 9.2
At1g01790 K Efflux antiporter KEA1 29 9.2
>At1g50480 10-formyltetrahydrofolate synthetase
Length = 634
Score = 1123 bits (2904), Expect = 0.0
Identities = 545/633 (86%), Positives = 602/633 (95%)
Query: 7 SSVMRKLDVVSPVPSDIDIANSVQPIHISEIANHLNLTPNHYDLYGKYKAKVLLSALDEL 66
SS RKL+VVSPVP+DIDIANSV+P+HISEIA LN+ P HYDLYGKYKAKVLLSA DEL
Sbjct: 2 SSSTRKLEVVSPVPADIDIANSVEPLHISEIAKDLNINPLHYDLYGKYKAKVLLSAFDEL 61
Query: 67 QESKDGYYVVVGGITPTPLGEGKSTTTVGLCQALGAFLDKKVVTCIRQPSQGPTFGIKGG 126
Q +DGYYVVVGGITPTPLGEGKSTTTVGLCQALGA+LDKKVVTC+RQPSQGPTFGIKGG
Sbjct: 62 QGQEDGYYVVVGGITPTPLGEGKSTTTVGLCQALGAYLDKKVVTCLRQPSQGPTFGIKGG 121
Query: 127 AAGGGYSQVIPMDEFNLHLTGDIHAITAANNLLAAAIDTRIFHESTQSDKALFNRLCPPN 186
AAGGGYSQVIPMDEFNLHLTGDIHAITA+NNLLAAAIDTRIFHE++QSDKALFNRLCPPN
Sbjct: 122 AAGGGYSQVIPMDEFNLHLTGDIHAITASNNLLAAAIDTRIFHETSQSDKALFNRLCPPN 181
Query: 187 KEGKRSFSDVMFRRLKKFGISKTNPDDLTPEEVNKFARLDIDPDSITWRRVMDINDRFLR 246
KEGKRSFSD+MFRRL K GISKT+P++LTPEE+ KFARLDIDP SITWRRVMD+NDRFLR
Sbjct: 182 KEGKRSFSDIMFRRLTKLGISKTSPEELTPEEIKKFARLDIDPASITWRRVMDVNDRFLR 241
Query: 247 KITVGQGPDEKGMVRETAFDISVASEIMAVLALTTSLTDMRERLGKMVIGNSKSGDPVTA 306
KIT+GQGP+EKGM RET FDISVASEIMAVLALTTSL DMRERLGKMVIGNSK+GDP+TA
Sbjct: 242 KITIGQGPEEKGMTRETGFDISVASEIMAVLALTTSLGDMRERLGKMVIGNSKAGDPITA 301
Query: 307 DDLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG 366
DDLG+GGALTVLMKDAI+PTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG
Sbjct: 302 DDLGVGGALTVLMKDAINPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG 361
Query: 367 GFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMHGGGPAVVAGKPLDHA 426
GFVVTEAGFG+DIGTEKFMNIKCRYSGLTPQCAI+VAT+RALKMHGGGP VVAG+PLD A
Sbjct: 362 GFVVTEAGFGSDIGTEKFMNIKCRYSGLTPQCAIVVATVRALKMHGGGPDVVAGRPLDRA 421
Query: 427 YLTENVALVEAGCANLARHILNSKAYGANVIVAINKFSTDTDAELNAVKKAALDAGAYDA 486
Y++ENV+LVEAGC NLA+HI N+KAYG NVIVA+N F+TDT+AELNAV+K ++DAGA+DA
Sbjct: 422 YVSENVSLVEAGCVNLAKHISNTKAYGVNVIVAVNMFATDTEAELNAVRKFSMDAGAFDA 481
Query: 487 VICSHHAHGGRGAVDLGIAVQKACENATQPLKFLYPLDIGIKEKIEAIAKSYGASGVEYS 546
V+CSHHAH G+GAVDLGIAV+KAC+N TQPL+FLYPLDIGIK+KIEAIAKSYGASGVEYS
Sbjct: 482 VVCSHHAHSGKGAVDLGIAVEKACQNITQPLRFLYPLDIGIKDKIEAIAKSYGASGVEYS 541
Query: 547 EQAEKKIELYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYP 606
+QAEK+IE+YT+QGFS LPICM+KTQYSFS +A+ KGAP+GF+LPIRDVR SIGAGFIYP
Sbjct: 542 DQAEKQIEMYTQQGFSNLPICMSKTQYSFSHDASKKGAPSGFVLPIRDVRGSIGAGFIYP 601
Query: 607 LVGTMSTMPGLPTRPCFYDIDLDTTTGQVIGLS 639
LVGTMSTMPGLPTRPCFY+ID+DT TG+V GLS
Sbjct: 602 LVGTMSTMPGLPTRPCFYEIDIDTETGKVRGLS 634
>At2g12280 10-formyltetrahydrofolate synthetase
Length = 90
Score = 136 bits (342), Expect = 4e-32
Identities = 61/74 (82%), Positives = 70/74 (94%)
Query: 555 LYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYPLVGTMSTM 614
+YT+QGFS LPICM+KTQYSFS +A+ KGAP+GF+LPIRDVR SIGAGFIYPLVGTMSTM
Sbjct: 1 MYTQQGFSNLPICMSKTQYSFSHDASKKGAPSGFVLPIRDVRGSIGAGFIYPLVGTMSTM 60
Query: 615 PGLPTRPCFYDIDL 628
PGLPTRPCFY+ID+
Sbjct: 61 PGLPTRPCFYEIDI 74
>At2g12200 10-formyltetrahydrofolate synthetase
Length = 63
Score = 88.2 bits (217), Expect = 1e-17
Identities = 40/63 (63%), Positives = 51/63 (80%)
Query: 555 LYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYPLVGTMSTM 614
+YT+QGFS LPICM+KTQYSFS + + K AP+GF+LPIRDVR SIGAGFIY T++ +
Sbjct: 1 MYTQQGFSNLPICMSKTQYSFSHDTSKKRAPSGFVLPIRDVRGSIGAGFIYTTKKTVNPL 60
Query: 615 PGL 617
G+
Sbjct: 61 AGM 63
>At5g47010 UPF1
Length = 1254
Score = 31.2 bits (69), Expect = 1.9
Identities = 27/94 (28%), Positives = 36/94 (37%), Gaps = 2/94 (2%)
Query: 308 DLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPGG 367
D G GGA +A P+ + G V G +SS D +A VG
Sbjct: 60 DRGEGGAAADHHSEASSPSSLSAGAGNGAKVGRGGVGGSGGVSSSSQVDALAAG-VGNLN 118
Query: 368 FVVTEAGFGADIGTEKFMNIKCRYSGLT-PQCAI 400
F T G D G F C+Y G++ P C +
Sbjct: 119 FEETGDDDGFDYGKNDFTEHACKYCGISNPACVV 152
>At5g11800 potassium transport protein like
Length = 597
Score = 30.4 bits (67), Expect = 3.2
Identities = 28/112 (25%), Positives = 49/112 (43%), Gaps = 7/112 (6%)
Query: 280 TTSLTDMRERLGKMVIGNSKSGDPVTADDLGIGGALTVLMKDAIHPTLMQTLEGTPVL-- 337
T +L D ++ + +I NSKS PV DL + L V++ A + G PV+
Sbjct: 148 TPTLIDRKDNV--FIISNSKSKYPVLQLDLRLISDLVVVIVSATCGGIAFACAGQPVITG 205
Query: 338 -VHAGPFANIAHGNSSIVADKIALKLVGPGGFVVTEAGFGADIGTEKFMNIK 388
+ AG I G + +++ + ++ V G V G + T K ++
Sbjct: 206 YLLAGSI--IGPGGLNFISEMVQVETVAQFGVVFLLFALGLEFSTAKLKVVR 255
>At4g13360 3-hydroxyisobutyryl-coenzyme A hydrolase - like protein
Length = 381
Score = 29.6 bits (65), Expect = 5.4
Identities = 24/72 (33%), Positives = 34/72 (46%), Gaps = 10/72 (13%)
Query: 527 IKEKIEAIAK---SYGASGVEYSEQAEKKIELYTKQGFSGLPICMAKTQYSFSDNAAAKG 583
IKE IE + K S +S VE++ +A K +E G P + TQ FS+ A AK
Sbjct: 254 IKETIEELKKYQQSTESSVVEWANEALKGLE-------KGAPFSLYLTQKYFSNVACAKS 306
Query: 584 APTGFILPIRDV 595
P + + V
Sbjct: 307 KPENELATLNGV 318
>At1g17600 disease resistance protein, putative
Length = 960
Score = 29.6 bits (65), Expect = 5.4
Identities = 20/77 (25%), Positives = 35/77 (44%), Gaps = 4/77 (5%)
Query: 380 GTEKFMNIKCRYSGLTPQ--CAIIVATIRALKMHGGGPAVVAGKPLDHAYLTENVAL--V 435
G K +KC Y L+P+ + I+++ G K L + L +++ L V
Sbjct: 215 GIGKTSIVKCLYDQLSPKFPAHCFIENIKSVSKDNGHDLKHLQKELLSSILCDDIRLWSV 274
Query: 436 EAGCANLARHILNSKAY 452
EAGC + + + N K +
Sbjct: 275 EAGCQEIKKRLGNQKVF 291
>At1g07990 unknown protein
Length = 802
Score = 29.6 bits (65), Expect = 5.4
Identities = 28/95 (29%), Positives = 41/95 (42%), Gaps = 17/95 (17%)
Query: 5 SSSSVMRKLDVVSPVPSDIDIANSVQPIHISEIANHLNLTPNHY----DLYGKYKAKVLL 60
SS ++ R LD+ P + + + V+ I +S + N +L NH DL GK+ LL
Sbjct: 360 SSGTIKRTLDLFFEYPYNNALHHQVESIILSCLENKSDLMVNHILRDCDLIGKF----LL 415
Query: 61 SALDELQESKDGYYVVVGGITPTPLGEGKSTTTVG 95
S D ++G PT GK VG
Sbjct: 416 SDRDS---------NLLGDSQPTVAASGKKKPRVG 441
>At2g21470 ubiquitin activating enzyme like protein
Length = 625
Score = 29.3 bits (64), Expect = 7.0
Identities = 14/45 (31%), Positives = 24/45 (53%)
Query: 49 DLYGKYKAKVLLSALDELQESKDGYYVVVGGITPTPLGEGKSTTT 93
DL + K+ + +E E K+ +V+ G TP+P G+S +T
Sbjct: 517 DLQQELSCKINVKHREEFDEEKEPEGMVLSGWTPSPATNGESAST 561
>At1g55860 ubiquitin-protein ligase 1, putative
Length = 3891
Score = 29.3 bits (64), Expect = 7.0
Identities = 11/20 (55%), Positives = 15/20 (75%)
Query: 374 GFGADIGTEKFMNIKCRYSG 393
G+GA G EK +++KCRY G
Sbjct: 1307 GYGAAAGHEKSLSVKCRYLG 1326
>At3g51220 unknown protein
Length = 186
Score = 28.9 bits (63), Expect = 9.2
Identities = 17/53 (32%), Positives = 30/53 (56%), Gaps = 1/53 (1%)
Query: 211 PDDLTPEEVNKFARLDIDPDSITWRR-VMDINDRFLRKITVGQGPDEKGMVRE 262
PD++T ++N+F R ++ D + RR V N L K+ VG+ + MV++
Sbjct: 112 PDNITEIKMNRFDRNEVYGDRLEKRRSVKFANPPLLTKVIVGKEEKNQVMVKK 164
>At1g01790 K Efflux antiporter KEA1
Length = 618
Score = 28.9 bits (63), Expect = 9.2
Identities = 28/126 (22%), Positives = 49/126 (38%), Gaps = 6/126 (4%)
Query: 304 VTADDLGIGGALTVLMKDAIHPTLMQTLE-GTPVLVHAGPFANIAHGNSSIVADKIALKL 362
V ++ + L +L+ I L Q + G+PVL + I SI+ + +
Sbjct: 7 VNEEEASLFDFLWLLLASVIFVPLFQKIPGGSPVLGYLAAGILIGPYGLSIIRNVHGTRA 66
Query: 363 VGPGGFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMH-----GGGPAV 417
+ G V G ++ E+ ++K GL ++ A + L H G A+
Sbjct: 67 IAEFGVVFLLFNIGLELSVERLSSMKKYVFGLGSAQVLVTAAVVGLLAHYVAGQAGPAAI 126
Query: 418 VAGKPL 423
V G L
Sbjct: 127 VIGNGL 132
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,081,832
Number of Sequences: 26719
Number of extensions: 618253
Number of successful extensions: 1464
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1458
Number of HSP's gapped (non-prelim): 13
length of query: 639
length of database: 11,318,596
effective HSP length: 106
effective length of query: 533
effective length of database: 8,486,382
effective search space: 4523241606
effective search space used: 4523241606
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146329.5


