

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
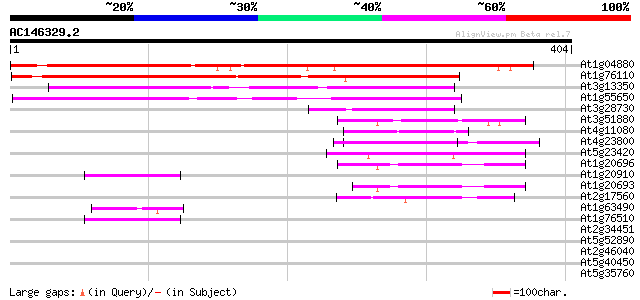
Query= AC146329.2 + phase: 0
(404 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g04880 unknown protein 410 e-115
At1g76110 hypothetical protein 268 5e-72
At3g13350 unknown protein 248 6e-66
At1g55650 unknown protein 197 9e-51
At3g28730 recombination signal sequence recognition protein, put... 59 5e-09
At3g51880 high mobility group protein 2-like 55 5e-08
At4g11080 98b like protein 55 9e-08
At4g23800 98b like protein 53 3e-07
At5g23420 unknown protein 50 2e-06
At1g20696 unknown protein 49 4e-06
At1g20910 unknown protein 49 6e-06
At1g20693 unknown protein 45 9e-05
At2g17560 putative HMG protein 43 3e-04
At1g63490 RB-binding protein -like 42 5e-04
At1g76510 putative DNA-binding protein 42 6e-04
At2g34451 putative HMG protein 40 0.002
At5g52890 putative protein 36 0.033
At2g46040 unknown protein 36 0.043
At5g40450 unknown protein 34 0.16
At5g35760 putative protein 34 0.16
>At1g04880 unknown protein
Length = 448
Score = 410 bits (1055), Expect = e-115
Identities = 228/402 (56%), Positives = 283/402 (69%), Gaps = 34/402 (8%)
Query: 1 MASTSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPV 60
MAS+SC + +PM ++ P ATYE VV +P+LF+ LE+LH+L+GTKFM+P+
Sbjct: 1 MASSSCLKQGSVPMNNVCVT------PEATYEAVVADPRLFMTSLERLHSLLGTKFMVPI 54
Query: 61 IGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYH 120
IGGR+LDLH+LFVEVTSRGG KI+ +R+WKEVT F FP TATNAS+VLRKYY SLL +
Sbjct: 55 IGGRDLDLHKLFVEVTSRGGINKILNERRWKEVTATFVFPPTATNASYVLRKYYFSLLNN 114
Query: 121 YEQIYYFKARDWTNTTSDVLQSQSSIPA-------PAPKMQFSH--PSPQVQPAVFPMGS 171
YEQIY+F++ D +QS S+ P P+ ++Q P P++ A F +G
Sbjct: 115 YEQIYFFRSNG--QIPPDSMQSPSARPCFIQGAIRPSQELQALTFTPQPKINTAEF-LGG 171
Query: 172 SSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQA-PQNPV---LPASHHSVPANNNN 227
S AGS VVGVIDGKF+SGYLVTVTIGSE+LKGVLYQ PQN V P H V N N
Sbjct: 172 SLAGSNVVGVIDGKFESGYLVTVTIGSEQLKGVLYQLLPQNTVSYQTPQQSHGVLPNTLN 231
Query: 228 VTASV-----GV-HRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDRE 281
++A+ GV RRRRRKKSE+K+RDP HPKPNRSGYNFFFAEQH RLKPLH GKDR+
Sbjct: 232 ISANPQGVAGGVTKRRRRRKKSEIKRRDPDHPKPNRSGYNFFFAEQHARLKPLHPGKDRD 291
Query: 282 ISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPE 341
ISR IGELWNKL E EK +YQ KA++DKERY TEME YREK KN ++IS+AVPL+QRLPE
Sbjct: 292 ISRMIGELWNKLNEDEKLIYQGKAMEDKERYRTEMEDYREKKKNGQLISNAVPLQQRLPE 351
Query: 342 PDTDMLNAE--ADSLQTPEQ----SSLDGSDDYEDDKAKEKD 377
+ DM A+ D ++ ++ S G + DD++ E D
Sbjct: 352 QNVDMAEADLPIDEVEEDDEEGDSSGSSGESEPHDDQSIETD 393
>At1g76110 hypothetical protein
Length = 338
Score = 268 bits (684), Expect = 5e-72
Identities = 148/327 (45%), Positives = 200/327 (60%), Gaps = 16/327 (4%)
Query: 2 ASTSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVI 61
A+ + SP +KE YP P+A +E VV + +F L + H++M TKFMIPVI
Sbjct: 12 ATVEMMATSPAKIKE-------YPEPLALHEVVVKDSSVFWDTLRRFHSIMSTKFMIPVI 64
Query: 62 GGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHY 121
GG+ELDLH L+VEVT RGG+EK++ ++KW+EV VF F +T T+ASFVLRK+Y +LL+HY
Sbjct: 65 GGKELDLHVLYVEVTRRGGYEKVVVEKKWREVGGVFRFSATTTSASFVLRKHYLNLLFHY 124
Query: 122 EQIYYFKARDWTNTTSDVLQSQSSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGV 181
EQ++ F AR + S ++++ PS + P S+ +G
Sbjct: 125 EQVHLFTARGPLLHPIATFHANPSTSKEMALVEYTPPSIRYH-NTHPPSQGSSSFTAIGT 183
Query: 182 IDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSVPA-NNNNVTASVGVHRRRRR 240
I+GKFD GYLV V +GSE L GVLY + Q P S A NN V V RRRRR
Sbjct: 184 IEGKFDCGYLVKVKLGSEILNGVLYHSAQ----PGPSSSPTAVLNNAVVPYVETGRRRRR 239
Query: 241 ---KKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESE 297
++ ++ DP +PKPNRSGYNFFFAE+H +LK L+ K+RE ++ IGE W+ L E
Sbjct: 240 LGKRRRSRRREDPNYPKPNRSGYNFFFAEKHCKLKSLYPNKEREFTKLIGESWSNLSTEE 299
Query: 298 KAVYQDKAVKDKERYITEMEYYREKLK 324
+ VYQD +KDKERY E+ YRE L+
Sbjct: 300 RMVYQDIGLKDKERYQRELNEYRETLR 326
>At3g13350 unknown protein
Length = 319
Score = 248 bits (632), Expect = 6e-66
Identities = 137/293 (46%), Positives = 177/293 (59%), Gaps = 22/293 (7%)
Query: 29 ATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDR 88
A Y+++V N LF L L +P +GG LDLHRLF+EVTSRGG E+++KDR
Sbjct: 34 AKYDDLVRNSALFWEKLRAFLGLTSKTLKVPTVGGNTLDLHRLFIEVTSRGGIERVVKDR 93
Query: 89 KWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYF-KARDWTNTTSDVLQSQSSIP 147
KWKEV F+FP+T T+ASFVLRKYY L+ E +YY K +T + L+S ++
Sbjct: 94 KWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLEHVYYLEKPVSSLQSTDEALKSLAN-E 152
Query: 148 APAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQ 207
+P P+ P G +V G IDGKFDSGYLVT+ +GS++LKGVLY
Sbjct: 153 SPNPEEGIDEP--------------QVGYEVQGFIDGKFDSGYLVTMKLGSQELKGVLYH 198
Query: 208 APQNPVLPASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQ 267
PQ P P +A V +RR RKKS++ D PK +RSGYNFFFAEQ
Sbjct: 199 IPQTPSQSQQTMETP------SAIVQSSQRRHRKKSKLAVVDTQKPKCHRSGYNFFFAEQ 252
Query: 268 HPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYR 320
+ RLKP + G++R I++ IG +W+ L ESEK VYQDK VKD ERY EM Y+
Sbjct: 253 YARLKPEYHGQERSITKKIGHMWSNLTESEKQVYQDKGVKDVERYRIEMLEYK 305
>At1g55650 unknown protein
Length = 337
Score = 197 bits (501), Expect = 9e-51
Identities = 116/325 (35%), Positives = 173/325 (52%), Gaps = 41/325 (12%)
Query: 3 STSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIG 62
ST +P +AL + Y+++V NP+LF L H KF IP++G
Sbjct: 2 STDSSQFEVVPANASALDNDVSSHMSMLYQDIVRNPELFWEMLRDFHESSDKKFKIPIVG 61
Query: 63 GRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYE 122
G+ LDLHRLF EVTSRGG EK+IKDR+ KEV FNF +T TN++FVLRK Y +L+ +E
Sbjct: 62 GKSLDLHRLFNEVTSRGGLEKVIKDRRCKEVIDAFNFKTTITNSAFVLRKSYLKMLFEFE 121
Query: 123 QIYYFKARDWTNTTSDVLQSQSSIPAPAPK-MQFSHPSPQVQPAVFPMGSSSAGSQVVGV 181
+YYF+A S + + ++ K S +++P G+ + G+
Sbjct: 122 HLYYFQA-----PLSTFWEKEKALKLLIEKSANRDKDSQELKP----------GTVITGI 166
Query: 182 IDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSVPANNNNVTASVGVHRRRRRK 241
IDGKF+SGYL++ +GSEKLKG+LY + +R +K
Sbjct: 167 IDGKFESGYLISTKVGSEKLKGMLYH------------------------ISPETKRGKK 202
Query: 242 KSEMKKRDP-AHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAV 300
K++ + D PK R+GYNFF AEQ R+K + G+ + G +W L ES++ V
Sbjct: 203 KAKSSQGDSHKPPKRQRTGYNFFVAEQSVRIKAENAGQKVSSPKNFGNMWTNLSESDRKV 262
Query: 301 YQDKAVKDKERYITEMEYYREKLKN 325
Y +K+ +D +RY E+ YR +++
Sbjct: 263 YYEKSREDGKRYKMEILQYRSLMES 287
>At3g28730 recombination signal sequence recognition protein,
putative
Length = 646
Score = 58.9 bits (141), Expect = 5e-09
Identities = 33/106 (31%), Positives = 53/106 (49%), Gaps = 5/106 (4%)
Query: 216 ASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLH 275
+S +P V A G +R++ KK K+DP PK SG+ FF + +K H
Sbjct: 529 SSSKGLPPKRKTVAADEGSSKRKKPKK----KKDPNAPKRAMSGFMFFSQMERDNIKKEH 584
Query: 276 RG-KDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYR 320
G E+ + +G+ W ++ +K Y+ KA DK+RY E+ Y+
Sbjct: 585 PGIAFGEVGKVLGDKWRQMSADDKEPYEAKAQVDKQRYKDEISDYK 630
>At3g51880 high mobility group protein 2-like
Length = 178
Score = 55.5 bits (132), Expect = 5e-08
Identities = 39/149 (26%), Positives = 70/149 (46%), Gaps = 21/149 (14%)
Query: 237 RRRRKKSEMKKRDPAHPKPNRSGYNFF-------FAEQHPRLKPLHRGKDREISRTIGEL 289
+R +K + K+DP PK S + F F +++P +K + + + G+
Sbjct: 37 KRETRKEKKAKKDPNKPKRAPSAFFVFLEDFRVTFKKENPNVKAVSA-----VGKAGGQK 91
Query: 290 WNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPD-----T 344
W + ++EKA Y++KA K K Y +M+ Y + L +E ++ R + + D
Sbjct: 92 WKSMSQAEKAPYEEKAAKRKAEYEKQMDAYNKNL--EEGSDESEKSRSEINDEDEASGEE 149
Query: 345 DMLNAEA--DSLQTPEQSSLDGSDDYEDD 371
++L EA D + E+ D DD E+D
Sbjct: 150 ELLEKEAAGDDEEEEEEEDDDDDDDEEED 178
>At4g11080 98b like protein
Length = 446
Score = 54.7 bits (130), Expect = 9e-08
Identities = 33/90 (36%), Positives = 48/90 (52%), Gaps = 2/90 (2%)
Query: 241 KKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAV 300
KK K +DP PK S Y + E+ LK ++ E+++ GE W L E +KA
Sbjct: 234 KKKAKKIKDPLKPKQPISAYLIYANERRAALKGENKSVI-EVAKMAGEEWKNLSEEKKAP 292
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVIS 330
Y A K+KE Y+ EME Y+ + K +E +S
Sbjct: 293 YDQMAKKNKEIYLQEMEGYK-RTKEEEAMS 321
Score = 40.0 bits (92), Expect = 0.002
Identities = 29/104 (27%), Positives = 46/104 (43%), Gaps = 4/104 (3%)
Query: 225 NNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REIS 283
N S + + KK + KK+D A K + Y + + +K + D +E S
Sbjct: 101 NMTFAFSQSLAQTEEEKKGKKKKKDCAETKRPSTPYILWCKDNWNEVKKQNPEADFKETS 160
Query: 284 RTIGELWNKLPESEKAVYQDKAVKDKERY---ITEMEYYREKLK 324
+G W + EK Y++K DKE Y IT+ + RE +K
Sbjct: 161 NILGAKWKGISAEEKKPYEEKYQADKEAYLQVITKEKREREAMK 204
Score = 40.0 bits (92), Expect = 0.002
Identities = 25/83 (30%), Positives = 37/83 (44%), Gaps = 1/83 (1%)
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRG-KDREISRTIGELWNKLPESEK 298
+ K + + DP PK S Y F + + H G + ++ I W +L E EK
Sbjct: 359 KNKKKNENVDPNKPKKPTSSYFLFCKDARKSVLEEHPGINNSTVTAHISLKWMELGEEEK 418
Query: 299 AVYQDKAVKDKERYITEMEYYRE 321
VY KA + E Y E+E Y +
Sbjct: 419 QVYNSKAAELMEAYKKEVEEYNK 441
Score = 28.5 bits (62), Expect = 6.9
Identities = 30/126 (23%), Positives = 52/126 (40%), Gaps = 15/126 (11%)
Query: 273 PLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLK-NDEVISD 331
P H K R + + + N++ ES + K K E+ + EM+ EK+K E D
Sbjct: 8 PAHAKKSRNSRKALKQK-NEIVESSPVSDKGKETKSFEKDLMEMQAMLEKMKIEKEKTED 66
Query: 332 AVPLRQ---RLPEPDTDMLNAEADSLQTPEQ----------SSLDGSDDYEDDKAKEKDF 378
+ + R E + + L E LQ ++ SL +++ + K K+KD
Sbjct: 67 LLKEKDEILRKKEVEQEKLKTELKKLQKMKEFKPNMTFAFSQSLAQTEEEKKGKKKKKDC 126
Query: 379 SVDSLP 384
+ P
Sbjct: 127 AETKRP 132
>At4g23800 98b like protein
Length = 456
Score = 52.8 bits (125), Expect = 3e-07
Identities = 38/141 (26%), Positives = 67/141 (46%), Gaps = 8/141 (5%)
Query: 241 KKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAV 300
KK K++DP PK S + + E+ L+ ++ E+++ GE W L + +KA
Sbjct: 243 KKKNKKEKDPLKPKHPVSAFLVYANERRAALREENKSVV-EVAKITGEEWKNLSDKKKAP 301
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y+ A K+KE Y+ ME Y+ + K +E +S Q+ E + L+ + ++
Sbjct: 302 YEKVAKKNKETYLQAMEEYK-RTKEEEALS------QKKEEEELLKLHKQEALQMLKKKE 354
Query: 361 SLDGSDDYEDDKAKEKDFSVD 381
D E K+K+ +VD
Sbjct: 355 KTDNLIKKEKATKKKKNENVD 375
Score = 48.1 bits (113), Expect = 8e-06
Identities = 28/90 (31%), Positives = 42/90 (46%), Gaps = 1/90 (1%)
Query: 234 VHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDRE-ISRTIGELWNK 292
+ + + KK + + DP PK S Y F ++ +L G + ++ I W +
Sbjct: 360 IKKEKATKKKKNENVDPNKPKKPASSYFLFSKDERKKLTEERPGTNNATVTALISLKWKE 419
Query: 293 LPESEKAVYQDKAVKDKERYITEMEYYREK 322
L E EK VY KA K E Y E+E Y +K
Sbjct: 420 LSEEEKQVYNGKAAKLMEAYKKEVEAYNKK 449
Score = 37.4 bits (85), Expect = 0.015
Identities = 34/155 (21%), Positives = 68/155 (42%), Gaps = 16/155 (10%)
Query: 227 NVTASVG---VHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REI 282
N+T + G + + + K ++ KK+D K S Y + +Q +K + D +E
Sbjct: 109 NMTFACGQSSLTQAEQEKANKKKKKDCPETKRPSSSYVLWCKDQWTEVKKENPEADFKET 168
Query: 283 SRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEP 342
S +G W L +K Y+++ +KE Y+ + + + + +++ D R +
Sbjct: 169 SNILGAKWKSLSAEDKKPYEERYQVEKEAYLQVIAKEKREKEAMKLLEDDQKQRTAM--- 225
Query: 343 DTDMLNAEADSLQTPEQSSLDGSDDYEDDKAKEKD 377
++L+ + +Q EQ D + KEKD
Sbjct: 226 --ELLDQYLNFVQEAEQ-------DNKKKNKKEKD 251
>At5g23420 unknown protein
Length = 241
Score = 50.1 bits (118), Expect = 2e-06
Identities = 38/148 (25%), Positives = 66/148 (43%), Gaps = 5/148 (3%)
Query: 229 TASVGVHRRRRRKKSEMKKRDPAHPKPNR--SGYNFFFAEQHPRLKPLHRGK-DREISRT 285
T S G +R +K ++ KK KP R + + F ++ K H G ++ ++
Sbjct: 89 TTSDGPKPKRLKKTNDEKKSSSTSNKPKRPLTAFFIFMSDFRKTFKSEHNGSLAKDAAKI 148
Query: 286 IGELWNKLPESEKAVYQDKAVKDKERYITEMEY--YREKLKNDEVISDAVPLRQRLPEPD 343
GE W L E EK VY DKA + K Y +E E+ +++E SD V + D
Sbjct: 149 GGEKWKSLTEEEKKVYLDKAAELKAEYNKSLESNDADEEEEDEEKQSDDVDDAEEKQVDD 208
Query: 344 TDMLNAEADSLQTPEQSSLDGSDDYEDD 371
D + + ++ +G ++ E++
Sbjct: 209 DDEVEEKEVENTDDDKKEAEGKEEEEEE 236
>At1g20696 unknown protein
Length = 141
Score = 49.3 bits (116), Expect = 4e-06
Identities = 39/142 (27%), Positives = 61/142 (42%), Gaps = 27/142 (19%)
Query: 237 RRRRKKSEMKKRDPAHPKPNRSGYNFF-------FAEQHPRLKPLHRGKDREISRTIGEL 289
++ K ++ +DP PK S + F + E+HP+ K + + + GE
Sbjct: 19 KKPAKGAKGAAKDPNKPKRPSSAFFVFMEDFRVTYKEEHPKNKSV-----AAVGKAGGEK 73
Query: 290 WNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNA 349
W L +SEKA Y KA K K Y M+ Y +KL+ P D +
Sbjct: 74 WKSLSDSEKAPYVAKADKRKVEYEKNMKAYNKKLEEG---------------PKEDEESD 118
Query: 350 EADSLQTPEQSSLDGSDDYEDD 371
++ S E + DGS++ EDD
Sbjct: 119 KSVSEVNDEDDAEDGSEEEEDD 140
>At1g20910 unknown protein
Length = 398
Score = 48.5 bits (114), Expect = 6e-06
Identities = 22/69 (31%), Positives = 38/69 (54%)
Query: 55 KFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYY 114
+F P G+ L++ +L+ V + GG+E + ++ W++V FN P T T S+ R +Y
Sbjct: 126 EFKPPKFYGQPLNILKLWRAVVNLGGYEVVTTNKLWRQVGESFNPPKTCTTVSYTFRNFY 185
Query: 115 TSLLYHYEQ 123
L YE+
Sbjct: 186 EKALLEYEK 194
>At1g20693 unknown protein
Length = 144
Score = 44.7 bits (104), Expect = 9e-05
Identities = 35/131 (26%), Positives = 56/131 (42%), Gaps = 27/131 (20%)
Query: 248 RDPAHPKPNRSGYNFF-------FAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAV 300
+DP PK S + F F +++P+ K + + + G+ W L +SEKA
Sbjct: 33 KDPNKPKRPASAFFVFMEDFRETFKKENPKNKSV-----ATVGKAAGDKWKSLSDSEKAP 87
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y KA K K Y ++ Y +KL+ P D + ++ S E
Sbjct: 88 YVAKAEKRKVEYEKNIKAYNKKLEEG---------------PKEDEESDKSVSEVNDEDD 132
Query: 361 SLDGSDDYEDD 371
+ DGS++ EDD
Sbjct: 133 AEDGSEEEEDD 143
>At2g17560 putative HMG protein
Length = 138
Score = 43.1 bits (100), Expect = 3e-04
Identities = 33/131 (25%), Positives = 54/131 (41%), Gaps = 14/131 (10%)
Query: 236 RRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREIS---RTIGELWNK 292
+ R RK + K+DP PK S + F F E + L ++ ++ + G W
Sbjct: 18 KTRGRKAGKKTKKDPNQPKRPPSAF-FVFLEDFRKEFNLANPNNKSVATVGKAAGARWKA 76
Query: 293 LPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEAD 352
+ + +KA Y KA K YI ++ Y KL + E D+D +E D
Sbjct: 77 MTDEDKAPYVAKAESRKTEYIKNVQQYNLKLASG----------TNREEDDSDKSKSEVD 126
Query: 353 SLQTPEQSSLD 363
+ E++ D
Sbjct: 127 EAVSEEEAEDD 137
>At1g63490 RB-binding protein -like
Length = 1458
Score = 42.4 bits (98), Expect = 5e-04
Identities = 26/70 (37%), Positives = 37/70 (52%), Gaps = 7/70 (10%)
Query: 60 VIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATN----ASFVLRKYYT 115
V G ELDL +LF V GG+EK++K +KW E V+ F S+ A VL + Y
Sbjct: 121 VFEGEELDLCKLFNAVKRFGGYEKVVKGKKWGE---VYQFMSSGEKISKCAKHVLCQLYK 177
Query: 116 SLLYHYEQIY 125
L+ +E +
Sbjct: 178 EHLHDFENYH 187
>At1g76510 putative DNA-binding protein
Length = 434
Score = 42.0 bits (97), Expect = 6e-04
Identities = 20/69 (28%), Positives = 35/69 (49%)
Query: 55 KFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYY 114
+F P G+ L+ +L+ V GG++ + + W++V F+ P T T S+ R +Y
Sbjct: 162 EFKAPKFYGQPLNCLKLWRAVIKLGGYDVVTTSKLWRQVGESFHPPKTCTTVSWTFRIFY 221
Query: 115 TSLLYHYEQ 123
L YE+
Sbjct: 222 EKALLEYEK 230
>At2g34451 putative HMG protein
Length = 151
Score = 40.4 bits (93), Expect = 0.002
Identities = 31/116 (26%), Positives = 50/116 (42%), Gaps = 23/116 (19%)
Query: 228 VTASVGVHRRRR------RKKSEMKKRDPAH-----PKPNRSGYNFF-------FAEQHP 269
V +S G R R RK K+ P PK + + FF + E++P
Sbjct: 27 VKSSEGARRSTRLRLQPLRKPKTSPKKKPVKLQTKMPKKPATAFFFFLDDFRKQYQEENP 86
Query: 270 RLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKN 325
+K + REI +T GE W + EK Y D A + +E + M Y +++++
Sbjct: 87 DVKSM-----REIGKTCGEKWKTMTYEEKVKYYDIATEKREEFHRAMTEYTKRMES 137
>At5g52890 putative protein
Length = 385
Score = 36.2 bits (82), Expect = 0.033
Identities = 21/59 (35%), Positives = 35/59 (58%), Gaps = 5/59 (8%)
Query: 163 QPAVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIG--SEKLKGVLYQAPQ--NPVLPAS 217
QP P+ + G V GV++G F++GY + V + ++LKGV++ PQ P+ PA+
Sbjct: 38 QPVNPPVDENLIGRVVSGVVEGSFEAGYFLNVKVADTEKQLKGVVF-LPQKVTPLTPAT 95
>At2g46040 unknown protein
Length = 562
Score = 35.8 bits (81), Expect = 0.043
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 63 GRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPS-TATNASFVLRKY 113
GR +DL LF+ VT +GGF+ + ++ W EV S + +A + KY
Sbjct: 76 GRTVDLFNLFLNVTHKGGFDAVSENGSWDEVVQESGLESYDSASAKLIYVKY 127
>At5g40450 unknown protein
Length = 2910
Score = 33.9 bits (76), Expect = 0.16
Identities = 28/130 (21%), Positives = 60/130 (45%), Gaps = 11/130 (8%)
Query: 281 EISRTIGELWNKLPESEKAVYQDKAV--------KDKERYITEMEYYREKLKNDEVISDA 332
E S+T+ E + PE E +YQ+ V K++ + E E+ + + + D
Sbjct: 875 ETSKTVDEKIEEKPEEEVTLYQEGQVDGSYGLETKEETVSVPESIELEEQPQEERSVIDP 934
Query: 333 VPLRQRLPEPDTDMLNAEADSLQTPEQSSLDGSDDYEDDKAKEKDFSVDSLPVIGLGAES 392
PL++ E +++L +S +T ++ + +D E + +++ SV L + +
Sbjct: 935 TPLQKPTLESPSEVLE---ESSKTVDEKIEEKTDSIELGEIAQEERSVTDLTPLQEESSQ 991
Query: 393 MDSVERSSKL 402
+ E+ +KL
Sbjct: 992 PNEQEKETKL 1001
Score = 28.5 bits (62), Expect = 6.9
Identities = 25/111 (22%), Positives = 50/111 (44%), Gaps = 5/111 (4%)
Query: 291 NKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAE 350
+K E+EK + ++ + K I E+ E + +V + + Q LP + +
Sbjct: 2276 DKSGEAEK-IKEESGLAGKSLPIEEINLQEEHKEEVKVQEETREIAQVLPREE---ILIS 2331
Query: 351 ADSLQTPEQSSLDGSDDYEDDKAKEKDFSVDSLPVIGLGAESMDSVERSSK 401
+ L EQ + SD+ ++++ ++DF+ + I L E + E S K
Sbjct: 2332 SSPLSAEEQEHVI-SDEKQEEREPQQDFNGSTSEKISLQVEHLKDFETSKK 2381
>At5g35760 putative protein
Length = 448
Score = 33.9 bits (76), Expect = 0.16
Identities = 29/97 (29%), Positives = 40/97 (40%), Gaps = 7/97 (7%)
Query: 226 NNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFF--AEQHPRLKPLHRGKD-REI 282
N V A G+H R K+ +KK P P GY FF +P + L +GK R
Sbjct: 295 NFVLAGRGLHANRMSKR--VKKNAIMIPSPRSGGYRSFFNSLPPYPPITKLLQGKAIRLK 352
Query: 283 SRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYY 319
SR G KLP ++ + R + + YY
Sbjct: 353 SRRCG--GEKLPPDTNGIFDPAKRSQRRRSLFSIPYY 387
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,743,284
Number of Sequences: 26719
Number of extensions: 459871
Number of successful extensions: 1486
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 1425
Number of HSP's gapped (non-prelim): 99
length of query: 404
length of database: 11,318,596
effective HSP length: 102
effective length of query: 302
effective length of database: 8,593,258
effective search space: 2595163916
effective search space used: 2595163916
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146329.2


