

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.11 + phase: 0
(327 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
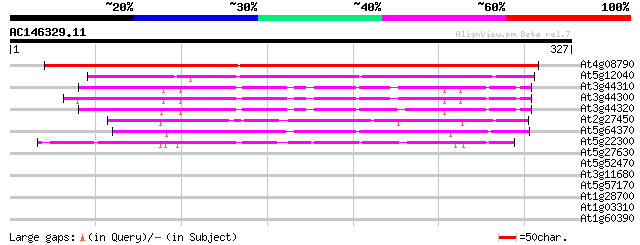
Score E
Sequences producing significant alignments: (bits) Value
At4g08790 nitrilase 1 like protein 430 e-121
At5g12040 unknown protein 148 3e-36
At3g44310 nitrilase 1 80 1e-15
At3g44300 nitrilase 2 79 4e-15
At3g44320 nitrilase 3 78 7e-15
At2g27450 nitrilase like protein 77 1e-14
At5g64370 beta-ureidopropionase 67 1e-11
At5g22300 Nitrilase 4 (sp|P46011) 67 1e-11
At5g27630 unknown protein 32 0.61
At5g52470 fibrillarin homolog (Fbr1) 30 2.3
At3g11680 unknown protein 29 4.0
At5g57170 light-inducible protein ATLS1-like 28 5.2
At1g28700 unknown protein 28 5.2
At1g03310 putative isoamylase 28 6.8
At1g60390 28 8.9
>At4g08790 nitrilase 1 like protein
Length = 307
Score = 430 bits (1105), Expect = e-121
Identities = 211/288 (73%), Positives = 246/288 (85%), Gaps = 1/288 (0%)
Query: 21 SIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLC 80
S+ R+ S + +T +VRVAAAQMTS+ DL +NF+TCSRLV+EAA AGAKL+C
Sbjct: 14 SLFTRITLSSQIPLTMATTVNKTVRVAAAQMTSVNDLMTNFATCSRLVQEAALAGAKLIC 73
Query: 81 FPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTH 140
FPE FSFVG K+G+SV IA+PLDGP+M++YCSLAR+S+IWLSLGGFQE+ D HL NTH
Sbjct: 74 FPENFSFVGDKEGESVKIAEPLDGPVMERYCSLARDSNIWLSLGGFQERFDDT-HLCNTH 132
Query: 141 VVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLR 200
VV+DD G I+ TY+K+HLFDVDVPGG YKES+FT G IV+VDSP+GRLGL+VCYDLR
Sbjct: 133 VVIDDAGMIRDTYQKMHLFDVDVPGGSSYKESSFTVPGTKIVSVDSPVGRLGLTVCYDLR 192
Query: 201 FPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKR 260
FP++YQ LRF+ AQ+LLVP+AFTKVTGEAHWEILLRARAIE QCYVIAAAQAG HN+KR
Sbjct: 193 FPKIYQQLRFEQKAQVLLVPSAFTKVTGEAHWEILLRARAIETQCYVIAAAQAGKHNEKR 252
Query: 261 ESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
ESYGDTLIIDPWGTVVGRLPDR+STGIVVADID SL+DSVR KMPI K
Sbjct: 253 ESYGDTLIIDPWGTVVGRLPDRVSTGIVVADIDFSLIDSVRTKMPIDK 300
>At5g12040 unknown protein
Length = 369
Score = 148 bits (374), Expect = 3e-36
Identities = 85/266 (31%), Positives = 140/266 (51%), Gaps = 11/266 (4%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDG- 104
+ Q++ +D N S + ++EAAS GAKL+ PE ++ + D V A+ +D
Sbjct: 90 IGLCQLSVTSDKKRNISHAKKAIEEAASKGAKLVLLPEIWNSPYSNDSFPV-YAEEIDAG 148
Query: 105 ----PIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFD 160
P +++ I + G E+ D L+NT V G+++ +RKIHLFD
Sbjct: 149 GDASPSTAMLSEVSKRLKITIIGGSIPERVGD--RLYNTCCVFGSDGELKAKHRKIHLFD 206
Query: 161 VDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVP 220
+D+PG + ES +G+ VD+ +GR+G+ +CYD+RF EL ++ GA +L P
Sbjct: 207 IDIPGKITFMESKTLTAGETPTIVDTDVGRIGIGICYDIRFQEL-AMIYAARGAHLLCYP 265
Query: 221 AAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLP 280
AF TG HWE+L RARA +NQ YV + A ++G + ++ P+G V+
Sbjct: 266 GAFNMTTGPLHWELLQRARATDNQLYVATCSPARDSGAGYTAWGHSTLVGPFGEVLATTE 325
Query: 281 DRLSTGIVVADIDLSLVDSVREKMPI 306
I++A+ID S+++ R +P+
Sbjct: 326 H--EEAIIIAEIDYSILEQRRTSLPL 349
>At3g44310 nitrilase 1
Length = 346
Score = 80.5 bits (197), Expect = 1e-15
Identities = 78/302 (25%), Positives = 137/302 (44%), Gaps = 56/302 (18%)
Query: 41 TNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFV---GAKDGDSV 96
+ +VRV Q +++ D + + + EAAS GA+L+ FPE F G + G +V
Sbjct: 22 STTVRVTIVQSSTVYNDTPATIDKAEKYIVEAASKGAELVLFPEGFIGGYPRGFRFGLAV 81
Query: 97 SI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHV 141
+ A + GP + + +AR++ ++L +G +++G L+ T +
Sbjct: 82 GVHNEEGRDEFRKYHASAIHVPGPEVARLADVARKNHVYLVMGAIEKEGYT---LYCTVL 138
Query: 142 VVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRF 201
G+ +RK+ ++ ++ + + G I D+PIG+LG ++C++ R
Sbjct: 139 FFSPQGQFLGKHRKLMPTSLE---RCIWGQGD----GSTIPVYDTPIGKLGAAICWENRM 191
Query: 202 PELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ--------- 252
P LY+ + G ++ P A G W+ + AIE C+V++A Q
Sbjct: 192 P-LYRTALYAKGIELYCAPTA----DGSKEWQSSMLHIAIEGGCFVLSACQFCQRKHFPD 246
Query: 253 ------AGTHNDKRE----SYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVRE 302
++DK S G ++II P G V+ P+ S G+V ADIDL D R
Sbjct: 247 HPDYLFTDWYDDKEHDSIVSQGGSVIISPLGQVLAG-PNFESEGLVTADIDLG--DIARA 303
Query: 303 KM 304
K+
Sbjct: 304 KL 305
>At3g44300 nitrilase 2
Length = 339
Score = 78.6 bits (192), Expect = 4e-15
Identities = 78/316 (24%), Positives = 143/316 (44%), Gaps = 61/316 (19%)
Query: 32 VTTGEST-----MATNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAF 85
++T E+T ++ VR Q +++ D + ++ + EAAS G++L+ FPEAF
Sbjct: 1 MSTSENTPFNGVASSTIVRATIVQASTVYNDTPATLEKANKFIVEAASKGSELVVFPEAF 60
Query: 86 SFV---GAKDGDSVSI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQ 127
G + G V + A + GP +++ LA +++++L +G +
Sbjct: 61 IGGYPRGFRFGLGVGVHNEEGRDEFRKYHASAIKVPGPEVEKLAELAGKNNVYLVMGAIE 120
Query: 128 EKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSP 187
+ G L+ T + G+ +RK+ ++ ++ + + G I D+P
Sbjct: 121 KDGYT---LYCTALFFSPQGQFLGKHRKLMPTSLE---RCIWGQGD----GSTIPVYDTP 170
Query: 188 IGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYV 247
IG+LG ++C++ R P LY+ + G ++ P A G W+ + AIE C+V
Sbjct: 171 IGKLGAAICWENRMP-LYRTALYAKGIELYCAPTA----DGSKEWQSSMLHIAIEGGCFV 225
Query: 248 IAAAQ---------------AGTHNDKRE----SYGDTLIIDPWGTVVGRLPDRLSTGIV 288
++A Q ++DK S G ++II P G V+ P+ S G++
Sbjct: 226 LSACQFCLRKDFPDHPDYLFTDWYDDKEPDSIVSQGGSVIISPLGQVLAG-PNFESEGLI 284
Query: 289 VADIDLSLVDSVREKM 304
AD+DL D R K+
Sbjct: 285 TADLDLG--DVARAKL 298
>At3g44320 nitrilase 3
Length = 346
Score = 77.8 bits (190), Expect = 7e-15
Identities = 76/302 (25%), Positives = 136/302 (44%), Gaps = 56/302 (18%)
Query: 41 TNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSF---VGAKDGDSV 96
+++VRV Q +++ D + + + EAAS GAKL+ FPEAF G + G +V
Sbjct: 22 SSTVRVTIVQSSTVYNDTPATLDKAEKFIVEAASKGAKLVLFPEAFIGGYPRGFRFGLAV 81
Query: 97 SI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHV 141
+ A + GP +++ LA ++++ L +G ++ G L+ T +
Sbjct: 82 GVHNEEGRDEFRNYHASAIKVPGPEVERLAELAGKNNVHLVMGAIEKDGYT---LYCTAL 138
Query: 142 VVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRF 201
G+ +RK+ ++ ++ + + G I D+PIG++G ++C++ R
Sbjct: 139 FFSPQGQFLGKHRKVMPTSLE---RCIWGQGD----GSTIPVYDTPIGKIGAAICWENRM 191
Query: 202 PELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ--------- 252
P LY+ + G +I P A + W+ + A+E C+V++A Q
Sbjct: 192 P-LYRTALYAKGIEIYCAPTADYSL----EWQASMIHIAVEGGCFVLSAHQFCKRREFPE 246
Query: 253 ----------AGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVRE 302
+D S G ++II P G V+ P+ S G+V AD+DL D R
Sbjct: 247 HPDYLFNDIVDTKEHDPTVSGGGSVIISPLGKVLAG-PNYESEGLVTADLDLG--DIARA 303
Query: 303 KM 304
K+
Sbjct: 304 KL 305
>At2g27450 nitrilase like protein
Length = 326
Score = 77.4 bits (189), Expect = 1e-14
Identities = 69/263 (26%), Positives = 120/263 (45%), Gaps = 28/263 (10%)
Query: 58 ASNFSTCSRLVKEAASAGAKLLCFPEAFS---FVGAKDGDSVSIAQPLDG-PIMDQYCSL 113
+S+F LV+EA + GA ++ E F F A+ D A+P P + + L
Sbjct: 51 SSSFKFPYALVREAHAKGANIILIQELFEGYYFCQAQREDFFKRAKPYKNHPTIARMQKL 110
Query: 114 ARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESN 173
A+E + + + F+E ++ H +N+ ++D G YRK H +P G Y+E
Sbjct: 111 AKELGVVIPVSFFEE--ANTAH-YNSIAIIDADGTDLGIYRKSH-----IPDGPGYQEKF 162
Query: 174 FTESGKDIVAV-DSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTK------V 226
+ G V + ++G+++C+D FPE + + Q GA+IL P A +
Sbjct: 163 YFNPGDTGFKVFQTKFAKIGVAICWDQWFPEAARAMVLQ-GAEILFYPTAIGSEPQDQGL 221
Query: 227 TGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRES-------YGDTLIIDPWGTVVGRL 279
HW +++ A N ++A+ + G + E YG + I P G +V
Sbjct: 222 DSRDHWRRVMQGHAGANVVPLVASNRIGKEIIETEHGPSQITFYGTSFIAGPTGEIVAEA 281
Query: 280 PDRLSTGIVVADIDLSLVDSVRE 302
D+ S ++VA DL ++ S R+
Sbjct: 282 DDK-SEAVLVAQFDLDMIKSKRQ 303
>At5g64370 beta-ureidopropionase
Length = 408
Score = 67.0 bits (162), Expect = 1e-11
Identities = 62/261 (23%), Positives = 110/261 (41%), Gaps = 24/261 (9%)
Query: 61 FSTCSRLVKEAASAGAKLLCFPEAFSFVGA---KDGDSVSIAQPLDGPIMDQYCSLARES 117
F ++ A AG +LC EA++ A ++ A+P+DG LA++
Sbjct: 115 FDKLKPIIDAAGVAGVNILCLQEAWTMPFAFCTRERRWCEFAEPVDGESTKFLQELAKKY 174
Query: 118 SIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTES 177
++ + + L+NT V++ + G I +RK H+ V G + + + E
Sbjct: 175 NMVIVSPILERDIDHGEVLWNTAVIIGNNGNIIGKHRKNHIPRV----GDFNESTYYMEG 230
Query: 178 GKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLR 237
++ G++ +++CY P L L +GA+I+ P+A E W I R
Sbjct: 231 DTGHPVFETVFGKIAVNICYGRHHP-LNWLAFGLNGAEIVFNPSATVGELSEPMWPIEAR 289
Query: 238 ARAIENQCYVIAAAQAGT---------------HNDKRESYGDTLIIDPWGTVVGRLPDR 282
AI N +V + + GT HND YG + P + L R
Sbjct: 290 NAAIANSYFVGSINRVGTEVFPNPFTSGDGKPQHNDFGHFYGSSHFSAPDASCTPSL-SR 348
Query: 283 LSTGIVVADIDLSLVDSVREK 303
G++++D+DL+L ++K
Sbjct: 349 YKDGLLISDMDLNLCRQYKDK 369
>At5g22300 Nitrilase 4 (sp|P46011)
Length = 355
Score = 67.0 bits (162), Expect = 1e-11
Identities = 77/315 (24%), Positives = 134/315 (42%), Gaps = 59/315 (18%)
Query: 17 TNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGA 76
TNG ++ + ++ G+S+ + V A+ T D + RL+ EAA G+
Sbjct: 16 TNG----HQIFPEIDMSAGDSSSIVRATVVQAS--TVFYDTPATLDKAERLLSEAAENGS 69
Query: 77 KLLCFPEAFS-----------FVG---AKDGDSV----SIAQPLDGPIMDQYCSLARESS 118
+L+ FPEAF +G AK D + A + GP +++ +A++
Sbjct: 70 QLVVFPEAFIGGYPRGSTFELAIGSRTAKGRDDFRKYHASAIDVPGPEVERLALMAKKYK 129
Query: 119 IWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESG 178
++L +G + +G L+ T + D G +RK+ +P F + G
Sbjct: 130 VYLVMGVIEREGYT---LYCTVLFFDSQGLFLGKHRKL------MPTALERCIWGFGD-G 179
Query: 179 KDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRA 238
I D+PIG++G ++C++ R P L + + G +I P A ++ T W +
Sbjct: 180 STIPVFDTPIGKIGAAICWENRMPSL-RTAMYAKGIEIYCAPTADSRET----WLASMTH 234
Query: 239 RAIENQCYVIAAAQAGTHND-----------KRESY--------GDTLIIDPWGTVVGRL 279
A+E C+V++A Q D ES G + II P G V+
Sbjct: 235 IALEGGCFVLSANQFCRRKDYPSPPEYMFSGSEESLTPDSVVCAGGSSIISPLGIVLAG- 293
Query: 280 PDRLSTGIVVADIDL 294
P+ ++ AD+DL
Sbjct: 294 PNYRGEALITADLDL 308
>At5g27630 unknown protein
Length = 648
Score = 31.6 bits (70), Expect = 0.61
Identities = 38/137 (27%), Positives = 57/137 (40%), Gaps = 38/137 (27%)
Query: 148 KIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCY--DLRFPELY 205
KIQT K L ++D +YKE +S ++ +A + S C+ ++ EL
Sbjct: 547 KIQTLQLKEELAEIDTRNTELYKE---LQSVRNQLAAEQ-------SRCFKLEVEVAELR 596
Query: 206 QLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGD 265
Q L+ T T + E+L R RA +A+ QA T N KR+S G
Sbjct: 597 QKLQ--------------TMETLQKELELLQRQRA-------VASEQAATMNAKRQSSGG 635
Query: 266 TLIIDPWGTVVGRLPDR 282
WG + G P +
Sbjct: 636 V-----WGWLAGTPPPK 647
>At5g52470 fibrillarin homolog (Fbr1)
Length = 308
Score = 29.6 bits (65), Expect = 2.3
Identities = 36/148 (24%), Positives = 65/148 (43%), Gaps = 21/148 (14%)
Query: 157 HLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLL-------- 208
H+ D+ P G VY SG+D+V + + + + D R P Y++L
Sbjct: 160 HVSDLVGPEGCVYAVEFSHRSGRDLVNMAKKRTNV-IPIIEDARHPAKYRMLVGMVDVIF 218
Query: 209 ---RFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESY-- 263
A+IL + A+F TG H+ I ++A I++ A Q+ ++E +
Sbjct: 219 SDVAQPDQARILALNASFFLKTG-GHFVISIKANCIDSTVAAEAVFQSEVKKLQQEQFKP 277
Query: 264 GDTLIIDPW----GTVVG--RLPDRLST 285
+ + ++P+ VVG R+P + T
Sbjct: 278 AEQVTLEPFERDHACVVGGYRMPKKQKT 305
>At3g11680 unknown protein
Length = 331
Score = 28.9 bits (63), Expect = 4.0
Identities = 21/73 (28%), Positives = 36/73 (48%), Gaps = 3/73 (4%)
Query: 166 GRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTK 225
G V+ F G+ IV + + LG + + FP + Q R+ +GA I ++ +F
Sbjct: 115 GAVHLARFFGHQGEPIV-LGILVFSLGAAATFSRFFPRIKQ--RYDYGALIFILTFSFVA 171
Query: 226 VTGEAHWEILLRA 238
++G EIL+ A
Sbjct: 172 ISGYRTDEILIMA 184
>At5g57170 light-inducible protein ATLS1-like
Length = 115
Score = 28.5 bits (62), Expect = 5.2
Identities = 13/45 (28%), Positives = 26/45 (56%)
Query: 55 TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIA 99
T++ + TCS ++K+A A AK++ PE++ + G ++ A
Sbjct: 8 TNIPVDAVTCSDILKDATKAVAKIIGKPESYVMILLNSGVPIAFA 52
>At1g28700 unknown protein
Length = 307
Score = 28.5 bits (62), Expect = 5.2
Identities = 19/69 (27%), Positives = 35/69 (50%), Gaps = 7/69 (10%)
Query: 129 KGSDPRHLFNTHVVV--DDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDS 186
+G R L N +VV D T + +R++H + +D G + E + D++ + S
Sbjct: 99 EGEGTRPLLNHLMVVAADQTAYDRCLFRRLHCYKMDTEGVDLEGEKD-----TDVMWLRS 153
Query: 187 PIGRLGLSV 195
P+ RL +S+
Sbjct: 154 PLSRLNVSL 162
>At1g03310 putative isoamylase
Length = 882
Score = 28.1 bits (61), Expect = 6.8
Identities = 12/48 (25%), Positives = 25/48 (52%)
Query: 264 GDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPIAKVIV 311
G+ +++DP+ TVVG+ + G + + V +P+ K++V
Sbjct: 319 GEPIVLDPYATVVGKSVSQKYLGSLSKSPSFDWGEDVSPNIPLEKLLV 366
>At1g60390
Length = 624
Score = 27.7 bits (60), Expect = 8.9
Identities = 18/54 (33%), Positives = 24/54 (44%), Gaps = 2/54 (3%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIA 99
+ A T LASN + + + AKL CFPE + K GD V+ A
Sbjct: 71 LTAVDSTRFASLASNHALNTH--HSDFCSAAKLFCFPELAAHSLEKHGDDVNFA 122
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,056,076
Number of Sequences: 26719
Number of extensions: 287841
Number of successful extensions: 600
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 576
Number of HSP's gapped (non-prelim): 16
length of query: 327
length of database: 11,318,596
effective HSP length: 100
effective length of query: 227
effective length of database: 8,646,696
effective search space: 1962799992
effective search space used: 1962799992
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146329.11


