

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145767.4 + phase: 0
(218 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
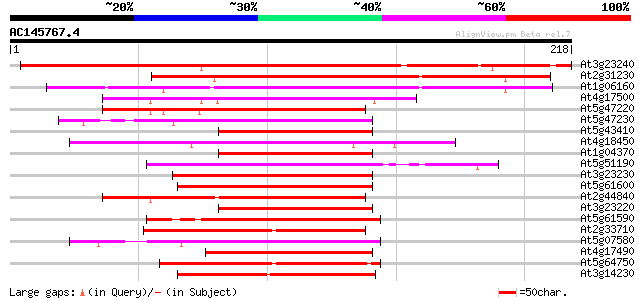
Score E
Sequences producing significant alignments: (bits) Value
At3g23240 ethylene response factor 1 234 3e-62
At2g31230 ethylene reponse factor-like AP2 domain transcription ... 158 2e-39
At1g06160 ethylene response factor, putative 157 3e-39
At4g17500 ethylene responsive element binding factor 1 (frameshi... 105 1e-23
At5g47220 ethylene responsive element binding factor 2 (ATERF2) ... 105 2e-23
At5g47230 ethylene responsive element binding factor 5 (ATERF5) ... 101 3e-22
At5g43410 Nicotiana EREBP-3 like 100 6e-22
At4g18450 EREBP - like protein 100 1e-21
At1g04370 hypothetical protein 99 2e-21
At5g51190 unknown protein 98 3e-21
At3g23230 ethylene responsive element binding protein, putative 98 3e-21
At5g61600 DNA binding protein - like 97 7e-21
At2g44840 putative ethylene response element binding protein (ER... 97 7e-21
At3g23220 ethylene responsive element binding protein, putative 97 9e-21
At5g61590 ethylene responsive element binding factor - like 96 1e-20
At2g33710 putative AP2 domain transcription factor 94 7e-20
At5g07580 transcription factor-like protein (emb|CAB87947.1) 92 2e-19
At4g17490 ethylene responsive element binding factor-like protein 92 2e-19
At5g64750 unknown protein 91 5e-19
At3g14230 transcription factor EREBP like protein 90 8e-19
>At3g23240 ethylene response factor 1
Length = 218
Score = 234 bits (597), Expect = 3e-62
Identities = 130/220 (59%), Positives = 158/220 (71%), Gaps = 11/220 (5%)
Query: 5 FIPFSSKSNFFPQSSFGSQDSISLNPKNHNEFLPFNENDPEEMLLYGMITSSPQEQTLSK 64
F+ S S F P+ S GS + ++N LPFNEND EEM LYG+I S Q+ +
Sbjct: 4 FLIQSPFSGFSPEYSIGSSPDSFSSSSSNNYSLPFNENDSEEMFLYGLIEQSTQQTYIDS 63
Query: 65 TSNEKTTKS-NKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYD 123
S + KS + K+ KSYRGVRRRPWGKFAAEIRDSTR+GIRVWLGTF+SAE AALAYD
Sbjct: 64 DSQDLPIKSVSSRKSEKSYRGVRRRPWGKFAAEIRDSTRNGIRVWLGTFESAEEAALAYD 123
Query: 124 QAAFSMRGSSATLNFSVEKVKESLRDMNYLLSNDNECSPVIALKRKHSMKRKMDEKKKKH 183
QAAFSMRGSSA LNFS E+V+ESL ++ Y + ++ CSPV+ALKRKHSM+R+M KK K
Sbjct: 124 QAAFSMRGSSAILNFSAERVQESLSEIKY--TYEDGCSPVVALKRKHSMRRRMTNKKTK- 180
Query: 184 DTD-----VRLDNLVVFEDLGADYLEQLLMSSSDDNQNIW 218
D+D V+LDN+VVFEDLG YLE+LL SS +N W
Sbjct: 181 DSDFDHRSVKLDNVVVFEDLGEQYLEELLGSS--ENSGTW 218
>At2g31230 ethylene reponse factor-like AP2 domain transcription
factor
Length = 243
Score = 158 bits (399), Expect = 2e-39
Identities = 93/189 (49%), Positives = 115/189 (60%), Gaps = 35/189 (18%)
Query: 56 SPQEQTLSKTSNEKTTKSNKDKN-----NKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLG 110
SP+ + N + S +++ +SYRGVR+RPWGKFAAEIRDSTR+GIRVWLG
Sbjct: 55 SPEMMIKEEIQNGDVSNSEEEEKVGIDEERSYRGVRKRPWGKFAAEIRDSTRNGIRVWLG 114
Query: 111 TFDSAEAAALAYDQAAFSMRGSSATLNFSVEKVKESLRDMNYLLSNDNECSPVIALKRKH 170
TFD AE AALAYDQAAF+ +GS ATLNF VE V+ESL+ M + +D SPV+ALKRKH
Sbjct: 115 TFDKAEEAALAYDQAAFATKGSLATLNFPVEVVRESLKKMENVNLHDGG-SPVMALKRKH 173
Query: 171 SMKRKMDEKKKKHDTDVRLDN-----------------------------LVVFEDLGAD 201
S++ + KK+ + N LVVFEDLGA+
Sbjct: 174 SLRNRPRGKKRSSSSSSSSSNSSSCSSSSSTSSTSRSSSKQSVVKQESGTLVVFEDLGAE 233
Query: 202 YLEQLLMSS 210
YLEQLLMSS
Sbjct: 234 YLEQLLMSS 242
>At1g06160 ethylene response factor, putative
Length = 244
Score = 157 bits (398), Expect = 3e-39
Identities = 102/241 (42%), Positives = 133/241 (54%), Gaps = 47/241 (19%)
Query: 15 FPQSSFGSQDSISLNPKNHNEFLPFNENDPEEMLLYGMITSSPQ----------EQTLSK 64
F F ++S S + + FL + E+ + + SSP E ++ K
Sbjct: 7 FLSGEFSPENSSSSSWSSQESFL-WEESFLHQSFDQSFLLSSPTDNYCDDFFAFESSIIK 65
Query: 65 TSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQ 124
++ T + +++ KSYRGVR+RPWGKFAAEIRDSTR GIRVWLGTFD+AEAAALAYDQ
Sbjct: 66 EEGKEATVAAEEEE-KSYRGVRKRPWGKFAAEIRDSTRKGIRVWLGTFDTAEAAALAYDQ 124
Query: 125 AAFSMRGSSATLNFSVEKVKESLRDMNYLLSNDNECSPVIALKRKHSMKRKMDEKKKKHD 184
AAF+++GS A LNF + V+ESLR M + ND E SPVIALKRKHSM+ + KKK
Sbjct: 125 AAFALKGSLAVLNFPADVVEESLRKMENVNLNDGE-SPVIALKRKHSMRNRPRGKKKSSS 183
Query: 185 TDVRLDN----------------------------------LVVFEDLGADYLEQLLMSS 210
+ + LVV EDLGA+YLE+L+ S
Sbjct: 184 SSTLTSSPSSSSSYSSSSSSSSLSSRSRKQSVVMTQESNTTLVVLEDLGAEYLEELMRSC 243
Query: 211 S 211
S
Sbjct: 244 S 244
>At4g17500 ethylene responsive element binding factor 1 (frameshift
!)
Length = 246
Score = 105 bits (263), Expect = 1e-23
Identities = 65/161 (40%), Positives = 92/161 (56%), Gaps = 39/161 (24%)
Query: 37 LPFNENDPEEMLLYGMI------------TSSPQEQTLSKTSNEKTTKS---------NK 75
LP END E+ML+YG++ +SS ++++ + +T +S K
Sbjct: 49 LPLKENDSEDMLVYGILNDAFHGGWEPSSSSSDEDRSSFPSVKIETPESFAAVDSVPVKK 108
Query: 76 DKNN-----------KSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQ 124
+K + K YRGVR+RPWGKFAAEIRD ++G RVWLGTF++AE AALAYD+
Sbjct: 109 EKTSPVSAAVTAAKGKHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDR 168
Query: 125 AAFSMRGSSATLNFSV-------EKVKESLRDMNYLLSNDN 158
AAF MRGS A LNF + + V+ + ++ SN+N
Sbjct: 169 AAFRMRGSRALLNFPLRVNSGEPDPVRIKSKRSSFSSSNEN 209
>At5g47220 ethylene responsive element binding factor 2 (ATERF2)
(sp|O80338)
Length = 243
Score = 105 bits (262), Expect = 2e-23
Identities = 63/126 (50%), Positives = 76/126 (60%), Gaps = 24/126 (19%)
Query: 37 LPFNENDPEEMLLYGMI-------TSSPQ------------EQTLSKTSNEKTTK----- 72
LP END E+ML+YG++ TSS E T + T+ E+ K
Sbjct: 48 LPLKENDSEDMLVYGLLKDAFHFDTSSSDLSCLFDFPAVKVEPTENFTAMEEKPKKAIPV 107
Query: 73 SNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGS 132
+ K YRGVR+RPWGKFAAEIRD ++G RVWLGTF++AE AALAYD AAF MRGS
Sbjct: 108 TETAVKAKHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDIAAFRMRGS 167
Query: 133 SATLNF 138
A LNF
Sbjct: 168 RALLNF 173
>At5g47230 ethylene responsive element binding factor 5 (ATERF5)
(sp|O80341)
Length = 300
Score = 101 bits (251), Expect = 3e-22
Identities = 60/135 (44%), Positives = 79/135 (58%), Gaps = 20/135 (14%)
Query: 20 FGSQDSIS---LNPKNHNEFLPFNENDPEEMLLYGMITSSPQEQTL----------SKTS 66
F S+ S+S P N N+ N+ +PE L I P + +L ++ +
Sbjct: 88 FDSEVSVSDFDFKPSNQNQ----NQFEPE---LKSQIRKPPLKISLPAKTEWIQFAAENT 140
Query: 67 NEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAA 126
+ TK ++ K YRGVR+RPWGKFAAEIRD + G RVWLGTFD+A AA AYD+AA
Sbjct: 141 KPEVTKPVSEEEKKHYRGVRQRPWGKFAAEIRDPNKRGSRVWLGTFDTAIEAARAYDEAA 200
Query: 127 FSMRGSSATLNFSVE 141
F +RGS A LNF +E
Sbjct: 201 FRLRGSKAILNFPLE 215
>At5g43410 Nicotiana EREBP-3 like
Length = 131
Score = 100 bits (249), Expect = 6e-22
Identities = 47/60 (78%), Positives = 52/60 (86%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNFSVE 141
YRGVRRRPWGK+AAEIRDS +HG RVWLGTFD+AE AA AYDQAA+SMRG +A LNF E
Sbjct: 15 YRGVRRRPWGKYAAEIRDSRKHGERVWLGTFDTAEEAARAYDQAAYSMRGQAAILNFPHE 74
>At4g18450 EREBP - like protein
Length = 303
Score = 99.8 bits (247), Expect = 1e-21
Identities = 60/158 (37%), Positives = 86/158 (53%), Gaps = 8/158 (5%)
Query: 24 DSISLNPKNHNEFLPFNENDPEEMLLYGMITSSPQEQTLSKTSNEK----TTKSNKDKNN 79
D I + + L + D E + I S P + +N + S
Sbjct: 48 DDIPEGSREMLQSLDMSTEDQEWTEILDAIASFPNKTNHDPLTNPTIDSCSLSSRVSCKT 107
Query: 80 KSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGS-SATLNF 138
+ YRGVR+RPWGKFAAEIRDSTR+G+RVWLGTF +AE AA+AYD+AA +RG+ A NF
Sbjct: 108 RKYRGVRKRPWGKFAAEIRDSTRNGVRVWLGTFQTAEEAAMAYDKAAVRIRGTQKAHTNF 167
Query: 139 SVEKVKESLR---DMNYLLSNDNECSPVIALKRKHSMK 173
+E V +++ + NY N++ S + RK ++
Sbjct: 168 QLETVIKAMEMDCNPNYYRMNNSNTSDPLRSSRKIGLR 205
>At1g04370 hypothetical protein
Length = 133
Score = 98.6 bits (244), Expect = 2e-21
Identities = 46/60 (76%), Positives = 52/60 (86%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNFSVE 141
YRGVRRRPWGK+AAEIRDS +HG RVWLGTFD+AE AA AYD+AA+SMRG +A LNF E
Sbjct: 20 YRGVRRRPWGKYAAEIRDSRKHGERVWLGTFDTAEDAARAYDRAAYSMRGKAAILNFPHE 79
>At5g51190 unknown protein
Length = 221
Score = 98.2 bits (243), Expect = 3e-21
Identities = 59/144 (40%), Positives = 82/144 (55%), Gaps = 16/144 (11%)
Query: 54 TSSPQEQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFD 113
T S ++ L+ + T ++ + + YRGVRRRPWGK+AAEIRD + G+RVWLGTFD
Sbjct: 44 TLSQRKPPLATIAVPTTAPVVQENDQRHYRGVRRRPWGKYAAEIRDPNKKGVRVWLGTFD 103
Query: 114 SAEAAALAYDQAAFSMRGSSATLNFSVEKVKESLRDMNYLLSNDNECSPVIALKRKHSMK 173
+A AA YD+AAF +RGS A LNF +E K D+ DN+ I+LK K +
Sbjct: 104 TAMEAARGYDKAAFKLRGSKAILNFPLEAGKH--EDL-----GDNK--KTISLKAKRKRQ 154
Query: 174 RKMDEKK-------KKHDTDVRLD 190
DE + K+ + V+ D
Sbjct: 155 VTEDESQLISRKAVKREEAQVQAD 178
>At3g23230 ethylene responsive element binding protein, putative
Length = 122
Score = 98.2 bits (243), Expect = 3e-21
Identities = 45/78 (57%), Positives = 59/78 (74%)
Query: 64 KTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYD 123
++SN + ++D +RGVRRRPWGKFAAEIRD +R+G R+WLGTF++AE AA AYD
Sbjct: 2 ESSNRSSNNQSQDDKQARFRGVRRRPWGKFAAEIRDPSRNGARLWLGTFETAEEAARAYD 61
Query: 124 QAAFSMRGSSATLNFSVE 141
+AAF++RG A LNF E
Sbjct: 62 RAAFNLRGHLAILNFPNE 79
>At5g61600 DNA binding protein - like
Length = 241
Score = 97.1 bits (240), Expect = 7e-21
Identities = 43/76 (56%), Positives = 56/76 (73%)
Query: 66 SNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQA 125
S +TK+ K++ + YRGVRRRPWGK+AAEIRD + G R+WLGT+D+A A AYDQA
Sbjct: 72 SRTVSTKTEKEEEERHYRGVRRRPWGKYAAEIRDPNKKGCRIWLGTYDTAVEAGRAYDQA 131
Query: 126 AFSMRGSSATLNFSVE 141
AF +RG A LNF ++
Sbjct: 132 AFQLRGRKAILNFPLD 147
>At2g44840 putative ethylene response element binding protein
(EREBP)
Length = 226
Score = 97.1 bits (240), Expect = 7e-21
Identities = 50/116 (43%), Positives = 71/116 (61%), Gaps = 15/116 (12%)
Query: 37 LPFNENDPEEMLLYGMI--------------TSSPQEQTLSKTSNEKTTKSNKDKNNKSY 82
LP + +D ++M +Y + +SP E+ + + + + K + Y
Sbjct: 34 LPLSVDDSQDMAIYNTLRDAVSSGWTPSVPPVTSPAEENKPPATKASGSHAPRQKGMQ-Y 92
Query: 83 RGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNF 138
RGVRRRPWGKFAAEIRD ++G RVWLGT+++ E AA+AYD+AAF +RGS A LNF
Sbjct: 93 RGVRRRPWGKFAAEIRDPKKNGARVWLGTYETPEDAAVAYDRAAFQLRGSKAKLNF 148
>At3g23220 ethylene responsive element binding protein, putative
Length = 128
Score = 96.7 bits (239), Expect = 9e-21
Identities = 45/60 (75%), Positives = 50/60 (83%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNFSVE 141
YRGVR+RPWGK+AAEIRDS RHG RVWLGTF++AE AA AYD+AAF MRG A LNF E
Sbjct: 3 YRGVRKRPWGKYAAEIRDSARHGARVWLGTFNTAEDAARAYDRAAFGMRGQRAILNFPHE 62
>At5g61590 ethylene responsive element binding factor - like
Length = 201
Score = 95.9 bits (237), Expect = 1e-20
Identities = 50/91 (54%), Positives = 63/91 (68%), Gaps = 5/91 (5%)
Query: 54 TSSPQEQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFD 113
T+SP+ +T+S N K TK + + + YRGVRRRPWGKFAAEIRD + G R+WLGTF+
Sbjct: 84 TTSPEVETVS---NRKKTK--RFEETRHYRGVRRRPWGKFAAEIRDPAKKGSRIWLGTFE 138
Query: 114 SAEAAALAYDQAAFSMRGSSATLNFSVEKVK 144
S AA AYD AAF +RG A LNF ++ K
Sbjct: 139 SDIDAARAYDYAAFKLRGRKAVLNFPLDAGK 169
>At2g33710 putative AP2 domain transcription factor
Length = 218
Score = 93.6 bits (231), Expect = 7e-20
Identities = 45/86 (52%), Positives = 58/86 (67%), Gaps = 1/86 (1%)
Query: 53 ITSSPQEQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTF 112
+T P ++ S+ T + + ++YRGVR+RPWGK+AAEIRD + RVWLGTF
Sbjct: 41 LTGQPIPSSIDDQSSSLTLQEKSNSRQRNYRGVRQRPWGKWAAEIRDPNK-AARVWLGTF 99
Query: 113 DSAEAAALAYDQAAFSMRGSSATLNF 138
D+AE AALAYD+AAF RG A LNF
Sbjct: 100 DTAEEAALAYDKAAFEFRGHKAKLNF 125
>At5g07580 transcription factor-like protein (emb|CAB87947.1)
Length = 207
Score = 92.4 bits (228), Expect = 2e-19
Identities = 56/127 (44%), Positives = 73/127 (57%), Gaps = 14/127 (11%)
Query: 24 DSISLNPKNH-NEFLPFNENDPEEMLLYGMITSSPQEQTLSKT-----SNEKTTKSNKDK 77
DS L+P + NEFL +SSP+ + S T S +K + ++
Sbjct: 54 DSPVLDPDSFVNEFLQVEGESSS--------SSSPELNSSSSTYETDQSVKKAERFEEEV 105
Query: 78 NNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLN 137
+ + YRGVRRRPWGKFAAEIRD + G R+WLGTF+S AA AYD AAF +RG A LN
Sbjct: 106 DARHYRGVRRRPWGKFAAEIRDPAKKGSRIWLGTFESDVDAARAYDCAAFKLRGRKAVLN 165
Query: 138 FSVEKVK 144
F ++ K
Sbjct: 166 FPLDAGK 172
>At4g17490 ethylene responsive element binding factor-like protein
Length = 282
Score = 92.0 bits (227), Expect = 2e-19
Identities = 43/65 (66%), Positives = 50/65 (76%)
Query: 77 KNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATL 136
+ + YRGVR RPWGKFAAEIRD TR G RVWLGTF++A AA AYD+ AF +RGS A L
Sbjct: 132 EEKRHYRGVRMRPWGKFAAEIRDPTRRGTRVWLGTFETAIEAARAYDKEAFRLRGSKAIL 191
Query: 137 NFSVE 141
NF +E
Sbjct: 192 NFPLE 196
>At5g64750 unknown protein
Length = 391
Score = 90.9 bits (224), Expect = 5e-19
Identities = 47/86 (54%), Positives = 62/86 (71%), Gaps = 2/86 (2%)
Query: 59 EQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAA 118
E T + T++ +T+ + D+ + YRGVR+RPWGK+AAEIRD + RVWLGTFD+AE+A
Sbjct: 162 EYTTTATASSETSSFSGDQPRRRYRGVRQRPWGKWAAEIRDPFK-AARVWLGTFDNAESA 220
Query: 119 ALAYDQAAFSMRGSSATLNFSVEKVK 144
A AYD+AA RG+ A LNF E VK
Sbjct: 221 ARAYDEAALRFRGNKAKLNFP-ENVK 245
>At3g14230 transcription factor EREBP like protein
Length = 375
Score = 90.1 bits (222), Expect = 8e-19
Identities = 46/77 (59%), Positives = 55/77 (70%), Gaps = 1/77 (1%)
Query: 66 SNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQA 125
S E+ KS+K K YRG+R+RPWGK+AAEIRD R G R WLGTFD+AE AA AYD A
Sbjct: 108 SAEQAEKSSKRKRKNQYRGIRQRPWGKWAAEIRDP-RKGSREWLGTFDTAEEAARAYDAA 166
Query: 126 AFSMRGSSATLNFSVEK 142
A +RG+ A +NF EK
Sbjct: 167 ARRIRGTKAKVNFPEEK 183
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.128 0.365
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,937,274
Number of Sequences: 26719
Number of extensions: 208506
Number of successful extensions: 901
Number of sequences better than 10.0: 167
Number of HSP's better than 10.0 without gapping: 150
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 620
Number of HSP's gapped (non-prelim): 188
length of query: 218
length of database: 11,318,596
effective HSP length: 95
effective length of query: 123
effective length of database: 8,780,291
effective search space: 1079975793
effective search space used: 1079975793
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC145767.4


