

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145753.3 - phase: 0
(383 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
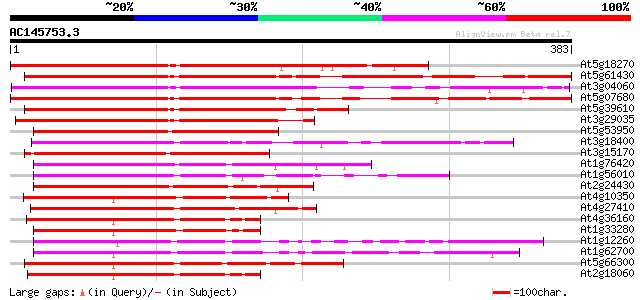
Score E
Sequences producing significant alignments: (bits) Value
At5g18270 NAM (no apical meristem)-like protein 313 1e-85
At5g61430 NAM, no apical meristem, - like protein 307 7e-84
At3g04060 NAM-like protein (no apical meristem) 304 5e-83
At5g07680 NAM (no apical meristem)-like protein 296 1e-80
At5g39610 NAM / CUC2 - like protein 266 2e-71
At3g29035 NAM-like protein (No Apical Meristem) 260 8e-70
At5g53950 CUC2 (dbj|BAA19529.1) 242 3e-64
At3g18400 organ separation protein, putative 238 5e-63
At3g15170 NAM-like protein 232 3e-61
At1g76420 unknown protein 223 1e-58
At1g56010 NAC1 216 2e-56
At2g24430 NAM (no apical meristem)-like protein 213 1e-55
At4g10350 NAM/NAP like protein 189 2e-48
At4g27410 unknown protein 187 1e-47
At4g36160 NAM like protein 186 2e-47
At1g33280 hypothetical protein 182 2e-46
At1g12260 unknown protein 182 3e-46
At1g62700 unknown protein 181 5e-46
At5g66300 NAM (no apical meristem)-like protein 181 6e-46
At2g18060 putative NAM (no apical meristem)-like protein 181 8e-46
>At5g18270 NAM (no apical meristem)-like protein
Length = 335
Score = 313 bits (801), Expect = 1e-85
Identities = 170/313 (54%), Positives = 212/313 (67%), Gaps = 33/313 (10%)
Query: 1 MEEPIVVNKGEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPW 60
+EE +V+N G ++ +DLPPGFRFHPTD EII CYL EKV NS+F+A A+GEADLNKCEPW
Sbjct: 5 VEEGVVLNHGGEELVDLPPGFRFHPTDEEIITCYLKEKVLNSRFTAVAMGEADLNKCEPW 64
Query: 61 DLPKKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGM 120
DLPK+AKMGEKE+YFFCQ+DRKYPTGMRTNRATESGYWKATGKDKEI+ KGKG LVGM
Sbjct: 65 DLPKRAKMGEKEFYFFCQRDRKYPTGMRTNRATESGYWKATGKDKEIF-KGKGC--LVGM 121
Query: 121 KKTLVFYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI 180
KKTLVFY+GRAPKGEKTNWVMHE+RLEGK++ +NLP +DEWVV RVFHKN Q
Sbjct: 122 KKTLVFYRGRAPKGEKTNWVMHEYRLEGKYSYYNLPKSARDEWVVCRVFHKNNPSTTTQP 181
Query: 181 SSGL-----LRINSIGH-DDLLDYSSLPPLMDPSYTND----DFKGITT----------- 219
+ + R++S+ + D LLD+SSLPPL+DPS+ + +FK I
Sbjct: 182 MTRIPVEDFTRMDSLENIDHLLDFSSLPPLIDPSFMSQTEQPNFKPINPPTYDISSPIQP 241
Query: 220 ---NQQISSTKSQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYD---QHEINPTMMNYT 273
N S Q G S NNN+ +IK E + ++ + ++N M T
Sbjct: 242 HHFNSYQSIFNHQVFGSASGSTYNNNNE---MIKMEQSLVSVSQETCLSSDVNANMTTTT 298
Query: 274 SISNQSNLNNPIG 286
+S+ + +G
Sbjct: 299 EVSSGPVMKQEMG 311
>At5g61430 NAM, no apical meristem, - like protein
Length = 336
Score = 307 bits (786), Expect = 7e-84
Identities = 179/373 (47%), Positives = 231/373 (60%), Gaps = 46/373 (12%)
Query: 11 EDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGE 70
E++ +DLPPGFRFHPTD E+I YL +KV ++ FSA AIGE DLNK EPW+LP AKMGE
Sbjct: 10 EEEQMDLPPGFRFHPTDEELITHYLHKKVLDTSFSAKAIGEVDLNKSEPWELPWMAKMGE 69
Query: 71 KEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGR 130
KEWYFFC +DRKYPTG+RTNRATE+GYWKATGKDKEIY +GK +LVGMKKTLVFY+GR
Sbjct: 70 KEWYFFCVRDRKYPTGLRTNRATEAGYWKATGKDKEIY-RGK---SLVGMKKTLVFYRGR 125
Query: 131 APKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSI 190
APKG+KTNWVMHE+RLEGKF+ HNLP K+EWV+ RVF K+ KK ISS L+RI S+
Sbjct: 126 APKGQKTNWVMHEYRLEGKFSAHNLPKTAKNEWVICRVFQKSAGGKKIPISS-LIRIGSL 184
Query: 191 GHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIK 250
G D + S LP L D S ND K++++ Y+P FS +Q+Q
Sbjct: 185 GTD--FNPSLLPSLTDSSPYND--------------KTKTEPVYVPCFSNQTDQNQGTTL 228
Query: 251 PEDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRM 310
+ +N Q +I + Y + S Q ++N L P + ++ L+ + NR
Sbjct: 229 NCFSSPVLNSIQADIFHRIPLYQTQSLQVSMN------LQSPVLTQEHSVLHAMIENNRR 282
Query: 311 QSSMPTNVYGSGKNNECKVEQFSSNQSQDTGLSNDTSSAVSKLDMERNRALYDDLEGPSS 370
QS +V SQ+TG+S D ++ +S D E + +D E PSS
Sbjct: 283 QSLKTMSV------------------SQETGVSTDMNTDISS-DFEFGKRRFDSQEDPSS 323
Query: 371 VAPLSDLDSFWDY 383
DL+ FW+Y
Sbjct: 324 STGPVDLEPFWNY 336
>At3g04060 NAM-like protein (no apical meristem)
Length = 338
Score = 304 bits (779), Expect = 5e-83
Identities = 189/389 (48%), Positives = 230/389 (58%), Gaps = 67/389 (17%)
Query: 2 EEPIVVNKGEDQ-TLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPW 60
E +VVN+G DQ +DLPPGFRFHPTD EII YL EKV N +F+A AIG+ADLNK EPW
Sbjct: 4 EGGVVVNQGGDQEVVDLPPGFRFHPTDEEIITHYLKEKVFNIRFTAAAIGQADLNKNEPW 63
Query: 61 DLPKKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGM 120
DLPK AKMGEKE+YFFCQ+DRKYPTGMRTNRAT SGYWKATGKDKEI+ +GKG LVGM
Sbjct: 64 DLPKIAKMGEKEFYFFCQRDRKYPTGMRTNRATVSGYWKATGKDKEIF-RGKGC--LVGM 120
Query: 121 KKTLVFYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI 180
KKTLVFY GRAPKGEKTNWVMHE+RL+GK++ HNLP +DEWVV RVFHKN
Sbjct: 121 KKTLVFYTGRAPKGEKTNWVMHEYRLDGKYSYHNLPKTARDEWVVCRVFHKNAPSTTITT 180
Query: 181 SSGLLRINSIGH-DDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFS 239
+ L RI+S+ + D LLD+SSLPPL+DP + G PSFS
Sbjct: 181 TKQLSRIDSLDNIDHLLDFSSLPPLIDPGFL---------------------GQPGPSFS 219
Query: 240 INNNQHQFLIKPEDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNP 299
QH++ P + + T+ P+ NT Q
Sbjct: 220 GAR------------------QQHDLKPVLHHPTTA--------PVDNTYLPTQA----- 248
Query: 300 NLNYFMYQNRMQSSMPTNVYGSGKNNE--CKVEQFSSNQSQDTGLSNDTSSA----VSKL 353
LN+ + S GSG NN+ K+E + SQ+TGLS+D ++ +S
Sbjct: 249 -LNFPYHSVHNSGSDFGYGAGSGNNNKGMIKLEHSLVSVSQETGLSSDVNTTATPEISSY 307
Query: 354 DMERNRALYDDLEGPSSVAPLSDLDSFWD 382
M N A+ D S+ L DL FW+
Sbjct: 308 PMMMNPAMMDG--SKSACDGLDDL-IFWE 333
>At5g07680 NAM (no apical meristem)-like protein
Length = 329
Score = 296 bits (759), Expect = 1e-80
Identities = 179/386 (46%), Positives = 234/386 (60%), Gaps = 60/386 (15%)
Query: 1 MEEPIVVNKGEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPW 60
ME V +K +D+ +DLPPGFRFHPTD E+I YL +KV + FSA AIGE DLNK EPW
Sbjct: 1 METFGVFHKEDDEQMDLPPGFRFHPTDEELITHYLHKKVLDLGFSAKAIGEVDLNKAEPW 60
Query: 61 DLPKKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGM 120
+LP KAK+GEKEWYFFC +DRKYPTG+RTNRAT++GYWKATGKDKEI+ +GK +LVGM
Sbjct: 61 ELPYKAKIGEKEWYFFCVRDRKYPTGLRTNRATQAGYWKATGKDKEIF-RGK---SLVGM 116
Query: 121 KKTLVFYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI 180
KKTLVFY+GRAPKG+KTNWVMHE+RL+GK + HNLP K+EWV+ RVFHK KK I
Sbjct: 117 KKTLVFYRGRAPKGQKTNWVMHEYRLDGKLSAHNLPKTAKNEWVICRVFHKTAGGKKIPI 176
Query: 181 SSGLLRINSIGHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSI 240
S+ L+RI S G SSLPPL D S ND K++++ Y+P FS
Sbjct: 177 ST-LIRIGSYGTG-----SSLPPLTDSSPYND--------------KTKTEPVYVPCFS- 215
Query: 241 NNNQHQFLIKPEDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLP--QPQIRIQN 298
+Q E T++N S + S++ +P QPQ
Sbjct: 216 --------------------NQAETRGTILNCFSNPSLSSIQPDFLQMIPLYQPQ----- 250
Query: 299 PNLNYFMYQNRMQSSMPTNVYGSGKNNECKVEQFSS-NQSQDTGLSNDTSSAVSKLDMER 357
+LN N + + + + +NN + + F + + SQ+TG+SN +S+V E
Sbjct: 251 -SLNISESSNPVLTQEQSVLQAMMENN--RRQNFKTLSISQETGVSNTDNSSV----FEF 303
Query: 358 NRALYDDLEGPSSVAPLSDLDSFWDY 383
R +D E PS + DL+ FW+Y
Sbjct: 304 GRKRFDHQEVPSPSSGPVDLEPFWNY 329
>At5g39610 NAM / CUC2 - like protein
Length = 285
Score = 266 bits (679), Expect = 2e-71
Identities = 137/221 (61%), Positives = 160/221 (71%), Gaps = 11/221 (4%)
Query: 11 EDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGE 70
+++ +DLPPGFRFHPTD E+I YL KV N+ FSATAIGE DLNK EPWDLP KAKMGE
Sbjct: 14 DEEHIDLPPGFRFHPTDEELITHYLKPKVFNTFFSATAIGEVDLNKIEPWDLPWKAKMGE 73
Query: 71 KEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGR 130
KEWYFFC +DRKYPTG+RTNRATE+GYWKATGKDKEI+ KGK +LVGMKKTLVFYKGR
Sbjct: 74 KEWYFFCVRDRKYPTGLRTNRATEAGYWKATGKDKEIF-KGK---SLVGMKKTLVFYKGR 129
Query: 131 APKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSI 190
APKG KTNWVMHE+RLEGK+ NLP K+EWV+ RVF K D K +S IN
Sbjct: 130 APKGVKTNWVMHEYRLEGKYCIENLPQTAKNEWVICRVFQKRADGTKVPMSMLDPHINR- 188
Query: 191 GHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSD 231
++ + LP LMD S D F G +++ S + D
Sbjct: 189 -----MEPAGLPSLMDCS-QRDSFTGSSSHVTCFSDQETED 223
>At3g29035 NAM-like protein (No Apical Meristem)
Length = 318
Score = 260 bits (665), Expect = 8e-70
Identities = 136/205 (66%), Positives = 151/205 (73%), Gaps = 22/205 (10%)
Query: 5 IVVNKGED-QTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLP 63
IV + ED + +DLPPGFRFHPTD E+I YL KV NS FSA AIGE DLNK EPWDLP
Sbjct: 11 IVEGEVEDSEKIDLPPGFRFHPTDEELITHYLRPKVVNSFFSAIAIGEVDLNKVEPWDLP 70
Query: 64 KKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKT 123
KAK+GEKEWYFFC +DRKYPTG+RTNRAT++GYWKATGKDKEI+ KGK +LVGMKKT
Sbjct: 71 WKAKLGEKEWYFFCVRDRKYPTGLRTNRATKAGYWKATGKDKEIF-KGK---SLVGMKKT 126
Query: 124 LVFYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSG 183
LVFYKGRAPKG KTNWVMHE+RLEGKFA NL K+E V+SRVFH TD K +S G
Sbjct: 127 LVFYKGRAPKGVKTNWVMHEYRLEGKFAIDNLSKTAKNECVISRVFHTRTDGTKEHMSVG 186
Query: 184 LLRINSIGHDDLLDYSSLPPLMDPS 208
LPPLMD S
Sbjct: 187 -----------------LPPLMDSS 194
>At5g53950 CUC2 (dbj|BAA19529.1)
Length = 375
Score = 242 bits (617), Expect = 3e-64
Identities = 114/167 (68%), Positives = 129/167 (76%), Gaps = 2/167 (1%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYFF 76
LPPGFRFHPTD E+I YL KV + FS+ AI E DLNKCEPW LP +AKMGEKEWYFF
Sbjct: 17 LPPGFRFHPTDEELITHYLLRKVLDGCFSSRAIAEVDLNKCEPWQLPGRAKMGEKEWYFF 76
Query: 77 CQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGEK 136
+DRKYPTG+RTNRATE+GYWKATGKD+EI+ LVGMKKTLVFYKGRAPKGEK
Sbjct: 77 SLRDRKYPTGLRTNRATEAGYWKATGKDREIF--SSKTCALVGMKKTLVFYKGRAPKGEK 134
Query: 137 TNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSG 183
+NWVMHE+RLEGKF+ H + KDEWV+SRVF K T +S G
Sbjct: 135 SNWVMHEYRLEGKFSYHFISRSSKDEWVISRVFQKTTLASTGAVSEG 181
>At3g18400 organ separation protein, putative
Length = 314
Score = 238 bits (606), Expect = 5e-63
Identities = 147/333 (44%), Positives = 183/333 (54%), Gaps = 43/333 (12%)
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYF 75
+LPPGFRFHPTD E+I YL KV + F+ A+ + DLNKCEPWDLP KA MGEKEWYF
Sbjct: 4 NLPPGFRFHPTDEELITHYLCRKVSDIGFTGKAVVDVDLNKCEPWDLPAKASMGEKEWYF 63
Query: 76 FCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGE 135
F Q+DRKYPTG+RTNRATE+GYWK TGKDKEIY G LVGMKKTLVFYKGRAPKGE
Sbjct: 64 FSQRDRKYPTGLRTNRATEAGYWKTTGKDKEIYRSGV----LVGMKKTLVFYKGRAPKGE 119
Query: 136 KTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIGHDDL 195
K+NWVMHE+RLE K N NKE EWVV RVF K+T KK Q +
Sbjct: 120 KSNWVMHEYRLESK-QPFNPTNKE--EWVVCRVFEKSTAAKKAQ--------------EQ 162
Query: 196 LDYSSLPPLMDPSYTN----DDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKP 251
SS P P N ++F+ I ++S S D NN+ HQ+
Sbjct: 163 QPQSSQPSFGSPCDANSSMANEFEDIDELPNLNSNSSTID--------YNNHIHQY---- 210
Query: 252 EDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQ 311
+ + + + + +N +NL + + L P I + L F +N
Sbjct: 211 --SQRNVYSEDNTTSTAGLNMNMNMASTNLQSWTTSLLGPPLSPINSLLLKAFQIRN--S 266
Query: 312 SSMPTNVYGSGKNNECKVEQFSSNQSQDTGLSN 344
S P + S N ++Q SN Q+ S+
Sbjct: 267 YSFPKEMIPS--FNHSSLQQGVSNMIQNASSSS 297
>At3g15170 NAM-like protein
Length = 310
Score = 232 bits (591), Expect = 3e-61
Identities = 109/167 (65%), Positives = 135/167 (80%), Gaps = 3/167 (1%)
Query: 11 EDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGE 70
ED++L +PPGFRFHPTD E+I YL +KV +S FS AI + DLNK EPW+LP+KAKMGE
Sbjct: 15 EDESL-MPPGFRFHPTDEELITYYLLKKVLDSNFSCAAISQVDLNKSEPWELPEKAKMGE 73
Query: 71 KEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGR 130
KEWYFF +DRKYPTG+RTNRATE+GYWKATGKD+EI K ++L+GMKKTLVFYKGR
Sbjct: 74 KEWYFFTLRDRKYPTGLRTNRATEAGYWKATGKDREI--KSSKTKSLLGMKKTLVFYKGR 131
Query: 131 APKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKK 177
APKGEK+ WVMHE+RL+GKF+ H + + KDEWV+ +V K+ V +
Sbjct: 132 APKGEKSCWVMHEYRLDGKFSYHYISSSAKDEWVLCKVCLKSGVVSR 178
>At1g76420 unknown protein
Length = 334
Score = 223 bits (568), Expect = 1e-58
Identities = 121/259 (46%), Positives = 156/259 (59%), Gaps = 33/259 (12%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYFF 76
LPPGFRFHPTD E+I YL K+ + S I E DLN+CEPW+LP+ AKMGE+EWYF+
Sbjct: 22 LPPGFRFHPTDEELITFYLASKIFHGGLSGIHISEVDLNRCEPWELPEMAKMGEREWYFY 81
Query: 77 CQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGEK 136
+DRKYPTG+RTNRAT +GYWKATGKDKE++ G G LVGMKKTLVFYKGRAP+G K
Sbjct: 82 SLRDRKYPTGLRTNRATTAGYWKATGKDKEVFSGGGG--QLVGMKKTLVFYKGRAPRGLK 139
Query: 137 TNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI---SSGLLRINSIGHD 193
T WVMHE+RLE H+ + K+EWV+ RVF+K D K + L +S+
Sbjct: 140 TKWVMHEYRLEN---DHSHRHTCKEEWVICRVFNKTGDRKNVGLIHNQISYLHNHSLSTT 196
Query: 194 DLLDYSSLPPLMDPS------------------YTNDDFKGITTNQQISSTK-------S 228
+ +LP L++PS Y N++F ++ I K S
Sbjct: 197 HHHHHEALPLLIEPSNKTLTNFPSLLYDDPHQNYNNNNFLHGSSGHNIDELKALINPVVS 256
Query: 229 QSDGYYLPSFSINNNQHQF 247
Q +G PS + NN++ F
Sbjct: 257 QLNGIIFPSGNNNNDEDDF 275
>At1g56010 NAC1
Length = 324
Score = 216 bits (549), Expect = 2e-56
Identities = 126/289 (43%), Positives = 168/289 (57%), Gaps = 57/289 (19%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEK-VKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYF 75
LPPGFRFHP D E++ YL + + N+ + + DLNKCEPWD+PK A +G K+WYF
Sbjct: 19 LPPGFRFHPKDDELVCDYLMRRSLHNNHRPPLVLIQVDLNKCEPWDIPKMACVGGKDWYF 78
Query: 76 FCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGE 135
+ Q+DRKY TG+RTNRAT +GYWKATGKD+ I KGK LVGM+KTLVFY+GRAP+G
Sbjct: 79 YSQRDRKYATGLRTNRATATGYWKATGKDRTILRKGK----LVGMRKTLVFYQGRAPRGR 134
Query: 136 KTNWVMHEFRLEGKFATHNLPN----KEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIG 191
KT+WVMHEFRL+G +H+ PN K++WV+ RVFHKNT+ + R N
Sbjct: 135 KTDWVMHEFRLQG---SHHPPNHSLSSPKEDWVLCRVFHKNTE-------GVICRDNMGS 184
Query: 192 HDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKP 251
D +SLPPLMDP Y N D Q YL
Sbjct: 185 CFDETASASLPPLMDP-YINFD---------------QEPSSYL---------------- 212
Query: 252 EDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPN 300
D++H I IN + ++++S LN+ + N++ + +I +NPN
Sbjct: 213 SDDHHYI------INEHVPCFSNLSQNQTLNSNLTNSVSELKIPCKNPN 255
>At2g24430 NAM (no apical meristem)-like protein
Length = 316
Score = 213 bits (543), Expect = 1e-55
Identities = 110/206 (53%), Positives = 134/206 (64%), Gaps = 21/206 (10%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYFF 76
LPPGFRFHPTD E+I YL K+ + F+ AI + DLNK EPW+LP+KAKMG KEWYFF
Sbjct: 16 LPPGFRFHPTDEELISYYLVNKIADQNFTGKAIADVDLNKSEPWELPEKAKMGGKEWYFF 75
Query: 77 CQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGEK 136
+DRKYPTG+RTNRAT +GYWK TGKDKEI++ LVGMKKTLVFY+GRAP+GEK
Sbjct: 76 SLRDRKYPTGVRTNRATNTGYWKTTGKDKEIFN--STTSELVGMKKTLVFYRGRAPRGEK 133
Query: 137 TNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQIS--------------- 181
T WVMHE+RL K + ++DEWVV RVF K T+ K IS
Sbjct: 134 TCWVMHEYRLHSK---SSYRTSKQDEWVVCRVF-KKTEATKKYISTSSSSTSHHHNNHTR 189
Query: 182 SGLLRINSIGHDDLLDYSSLPPLMDP 207
+ +L N+ + D LPP + P
Sbjct: 190 ASILSTNNNNPNYSSDLLQLPPHLQP 215
>At4g10350 NAM/NAP like protein
Length = 341
Score = 189 bits (480), Expect = 2e-48
Identities = 98/184 (53%), Positives = 120/184 (64%), Gaps = 11/184 (5%)
Query: 10 GEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG 69
G +PPGFRFHPTD E++ YL +K+ KF I E DLNK EPWDL ++ K+G
Sbjct: 2 GSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIG 61
Query: 70 ---EKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVF 126
+ EWYFF KDRKYPTG RTNRAT +G+WKATG+DK I + K I GM+KTLVF
Sbjct: 62 STPQNEWYFFSHKDRKYPTGSRTNRATHAGFWKATGRDKCIRNSYKKI----GMRKTLVF 117
Query: 127 YKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLR 186
YKGRAP G+KT+W+MHE+RLE A N +D WVV RVF K K ++ G
Sbjct: 118 YKGRAPHGQKTDWIMHEYRLED--ADDPQANPSEDGWVVCRVFMKKNLFK--VVNEGSSS 173
Query: 187 INSI 190
INS+
Sbjct: 174 INSL 177
>At4g27410 unknown protein
Length = 297
Score = 187 bits (474), Expect = 1e-47
Identities = 98/206 (47%), Positives = 126/206 (60%), Gaps = 19/206 (9%)
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L LPPGFRF+PTD E++V YL KV FS IG+ DL K +PWDLP KA GEKEWY
Sbjct: 12 LSLPPGFRFYPTDEELLVQYLCRKVAGYHFSLQVIGDIDLYKFDPWDLPSKALFGEKEWY 71
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NR SGYWKATG DK I G+ VG+KK LVFY G+APKG
Sbjct: 72 FFSPRDRKYPNGSRPNRVAGSGYWKATGTDKIITADGR----RVGIKKALVFYAGKAPKG 127
Query: 135 EKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI-----------SSG 183
KTNW+MHE+RL +H + + D+WV+ R++ K + ++ + ++G
Sbjct: 128 TKTNWIMHEYRLIEHSRSHG--SSKLDDWVLCRIYKKTSGSQRQAVTPVQACREEHSTNG 185
Query: 184 LLRINSIGHDDLLDYSSLPPLMDPSY 209
+S DD+LD S P + D S+
Sbjct: 186 SSSSSSSQLDDVLD--SFPEIKDQSF 209
>At4g36160 NAM like protein
Length = 377
Score = 186 bits (472), Expect = 2e-47
Identities = 89/163 (54%), Positives = 114/163 (69%), Gaps = 11/163 (6%)
Query: 12 DQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG-- 69
DQ+ +PPGFRFHPTD E++ YL +KV + K I + DL + EPWDL + ++G
Sbjct: 5 DQSCSVPPGFRFHPTDEELVGYYLRKKVASQKIDLDVIRDIDLYRIEPWDLQESCRIGYE 64
Query: 70 -EKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYK 128
EWYFF KD+KYPTG RTNRAT +G+WKATG+DK +Y K K L+GM+KTLVFYK
Sbjct: 65 ERNEWYFFSHKDKKYPTGTRTNRATMAGFWKATGRDKAVYDKSK----LIGMRKTLVFYK 120
Query: 129 GRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHK 171
GRAP G+KT+W+MHE+RLE + N P +E + WVV R F K
Sbjct: 121 GRAPNGQKTDWIMHEYRLE---SDENAPPQE-EGWVVCRAFKK 159
>At1g33280 hypothetical protein
Length = 305
Score = 182 bits (463), Expect = 2e-46
Identities = 92/158 (58%), Positives = 111/158 (70%), Gaps = 12/158 (7%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG---EKEW 73
+PPGFRFHPTD E++ YL +K+ KF I E DLNK EPWDL + K+G + EW
Sbjct: 8 VPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEW 67
Query: 74 YFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPK 133
YFF KDRKYPTG RTNRAT SG+WKATG+DK I + K I GM+KTLVFYKGRAP
Sbjct: 68 YFFSHKDRKYPTGSRTNRATHSGFWKATGRDKCIRNSYKKI----GMRKTLVFYKGRAPH 123
Query: 134 GEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHK 171
G+KT+W+MHE+R+E T + P +D WVV RVF K
Sbjct: 124 GQKTDWIMHEYRIED---TEDDPC--EDGWVVCRVFKK 156
>At1g12260 unknown protein
Length = 395
Score = 182 bits (461), Expect = 3e-46
Identities = 120/352 (34%), Positives = 186/352 (52%), Gaps = 62/352 (17%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKE---W 73
+PPGFRFHPTD E++ YL +KV + + I + DL K EPWDL + K+G +E W
Sbjct: 7 VPPGFRFHPTDEELVDYYLRKKVASKRIEIDFIKDIDLYKIEPWDLQELCKIGHEEQSDW 66
Query: 74 YFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPK 133
YFF KD+KYPTG RTNRAT++G+WKATG+DK IY + +L+GM+KTLVFYKGRAP
Sbjct: 67 YFFSHKDKKYPTGTRTNRATKAGFWKATGRDKAIYLR----HSLIGMRKTLVFYKGRAPN 122
Query: 134 GEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIGHD 193
G+K++W+MHE+RLE T +++ WVV RVF K L + +G
Sbjct: 123 GQKSDWIMHEYRLE----TDENGTPQEEGWVVCRVFKKR-----------LAAVRRMG-- 165
Query: 194 DLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKPED 253
DY S PS+ DD Q+S S+ + N Q + L
Sbjct: 166 ---DYDS-----SPSHWYDD--------QLSFMASELE---------TNGQRRIL----P 196
Query: 254 NYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSS 313
N+H+ +H+ + M Y ++ LNNP N + ++ + + N+ + +N +
Sbjct: 197 NHHQQQQHEHQQH---MPYGLNASAYALNNP--NLQCKQELEL---HYNHLVQRNHLLDE 248
Query: 314 MPTNVYGSGKNNECKVEQFSSN-QSQDTGLSNDTSSAVSKLDMERNRALYDD 364
+ + K++Q +SN S G SN +++ +++++ +++
Sbjct: 249 SHLSFLQLPQLESPKIQQDNSNCNSLPYGTSNIDNNSSHNANLQQSNIAHEE 300
>At1g62700 unknown protein
Length = 394
Score = 181 bits (460), Expect = 5e-46
Identities = 123/338 (36%), Positives = 168/338 (49%), Gaps = 52/338 (15%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG---EKEW 73
+PPGFRFHPTD E++ YL +KV + + I + DL K EP DL + K+G + EW
Sbjct: 7 VPPGFRFHPTDEELVDYYLRKKVASKRIEIDIIKDVDLYKIEPCDLQELCKIGNEEQSEW 66
Query: 74 YFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPK 133
YFF KD+KYPTG RTNRAT++G+WKATG+DK IY + +L+GM+KTLVFYKGRAP
Sbjct: 67 YFFSHKDKKYPTGTRTNRATKAGFWKATGRDKAIYIR----HSLIGMRKTLVFYKGRAPN 122
Query: 134 GEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIGHD 193
G+K++W+MHE+RLE T +++ WVV RVF K + +G
Sbjct: 123 GQKSDWIMHEYRLE----TSENGTPQEEGWVVCRVFKKKL----------AATVRKMG-- 166
Query: 194 DLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKPED 253
DY S P + DD ++ ISS+ Q +LP+ N + HQ +
Sbjct: 167 ---DYHSSP----SQHWYDDQLSFMASEIISSSPRQ----FLPNHHYNRHHHQQTLPCGL 215
Query: 254 NYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSS 313
N N NP + + N Q + QN +L M+ Q
Sbjct: 216 NAFNNN------NPNLQCKQELELHYN---------QMVQHQQQNHHLRESMFLQLPQLE 260
Query: 314 MPTNVYGSGKNNECK---VEQFSSNQSQDTGLSNDTSS 348
PT+ S NN + Q SSN S + L S
Sbjct: 261 SPTSNCNSDNNNNTRNISNLQKSSNISHEEQLQQGNQS 298
>At5g66300 NAM (no apical meristem)-like protein
Length = 292
Score = 181 bits (459), Expect = 6e-46
Identities = 93/221 (42%), Positives = 133/221 (60%), Gaps = 15/221 (6%)
Query: 11 EDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG- 69
+D + +PPGFRFHPTD E++ YL +K+ + + I E DL K EPWDL ++ ++G
Sbjct: 6 QDYSCSIPPGFRFHPTDEELVGYYLKKKIASQRIDLDVIREIDLYKIEPWDLQERCRIGY 65
Query: 70 --EKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFY 127
+ EWYFF +D+KYPTG RTNRAT +G+WKATG+DK +Y K L+GM+KTLVFY
Sbjct: 66 EEQTEWYFFSHRDKKYPTGTRTNRATVAGFWKATGRDKAVYLNSK----LIGMRKTLVFY 121
Query: 128 KGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRI 187
+GRAP G+K++W++HE+ +H +++ WVV R F K T + P L
Sbjct: 122 RGRAPNGQKSDWIIHEYY---SLESHQNSPPQEEGWVVCRAFKKRTTI--PTKRRQLWDP 176
Query: 188 NSIGHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKS 228
N + +DD + L PL + N DF Q++ S S
Sbjct: 177 NCLFYDDA---TLLEPLDKRARHNPDFTATPFKQELLSEAS 214
>At2g18060 putative NAM (no apical meristem)-like protein
Length = 365
Score = 181 bits (458), Expect = 8e-46
Identities = 85/162 (52%), Positives = 115/162 (70%), Gaps = 11/162 (6%)
Query: 13 QTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG--- 69
++ +PPGFRFHPTD E++ YL +K+ + K I + DL + EPWDL ++ ++G
Sbjct: 5 ESCSVPPGFRFHPTDEELVGYYLRKKIASQKIDLDVIRDIDLYRIEPWDLQEQCRIGYEE 64
Query: 70 EKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKG 129
+ EWYFF KD+KYPTG RTNRAT +G+WKATG+DK +Y K K L+GM+KTLVFYKG
Sbjct: 65 QNEWYFFSHKDKKYPTGTRTNRATMAGFWKATGRDKAVYDKTK----LIGMRKTLVFYKG 120
Query: 130 RAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHK 171
RAP G+K++W+MHE+RLE + N P +E + WVV R F K
Sbjct: 121 RAPNGKKSDWIMHEYRLE---SDENAPPQE-EGWVVCRAFKK 158
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.132 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,616,668
Number of Sequences: 26719
Number of extensions: 447337
Number of successful extensions: 1182
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 873
Number of HSP's gapped (non-prelim): 145
length of query: 383
length of database: 11,318,596
effective HSP length: 101
effective length of query: 282
effective length of database: 8,619,977
effective search space: 2430833514
effective search space used: 2430833514
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145753.3


