

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145513.12 + phase: 0
(304 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
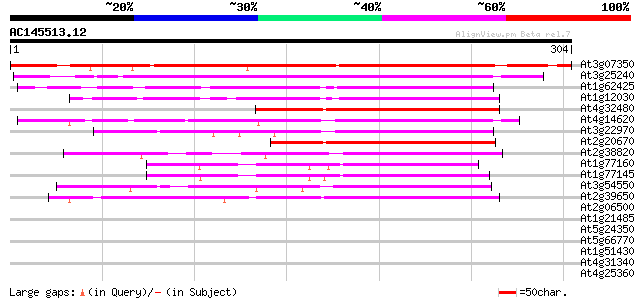
Sequences producing significant alignments: (bits) Value
At3g07350 unknown protein 260 6e-70
At3g25240 hypothetical protein 210 9e-55
At1g62425 unknown protein 156 1e-38
At1g12030 hypothetical protein 149 2e-36
At4g32480 putative protein 110 1e-24
At4g14620 unknown protein 109 2e-24
At3g22970 unknown protein 109 2e-24
At2g20670 unknown protein 105 3e-23
At2g38820 Unknown protein 105 4e-23
At1g77160 hypothetical protein 102 3e-22
At1g77145 unknown protein 102 3e-22
At3g54550 putative protein 98 6e-21
At2g39650 unknown protein 98 6e-21
At2g06500 Ac-like transposase 37 0.017
At1g21485 putative protein 32 0.33
At5g24350 putative protein 31 0.95
At5g66770 SCARECROW gene regulator 30 1.2
At1g51430 hypothetical protein 30 1.6
At4g31340 unknown protein 30 2.1
At4g25360 unknown protein (At4g25360) 30 2.1
>At3g07350 unknown protein
Length = 298
Score = 260 bits (665), Expect = 6e-70
Identities = 148/317 (46%), Positives = 196/317 (61%), Gaps = 32/317 (10%)
Query: 1 MNFARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTG----ECDSSPSLSELVHG 56
M F+R KRVTDP +E +ARLVG SSGSEH+G G E D SP LS+LV G
Sbjct: 1 MGFSRAKRVTDPLAEEVRARLVGCSF------SSGSEHTGDGIEDYEDDDSPCLSDLVQG 54
Query: 57 FLEEDDGNV------CDS-TGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLR 109
FLE++ V CD +G+D DS+ + + D +D+ +L + DSY +
Sbjct: 55 FLEDEVDTVDDESCWCDQDSGSDSDSD--SELGELPDFADDIAKLLRNSLREDSYGRTVL 112
Query: 110 LHVSEAAVKFDFLKKQSV--SVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFID 167
+HV+ A L Q +V+ R VMS LRE GHNAAICKT+W SSGGLTAGNHEFID
Sbjct: 113 VHVARAMEMLSSLGSQPEQRAVFQRKVMSLLRELGHNAAICKTKWKSSGGLTAGNHEFID 172
Query: 168 VVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALC 227
VV S SS R+ V+LDF+ +F+IARPTS+Y+ ++ +P +FVG ++LKR + +C
Sbjct: 173 VVYTPSASSQ-SVRFIVDLDFSSRFQIARPTSQYARVLQSLPAVFVGKGDDLKRILRLVC 231
Query: 228 GAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLVGF 287
A ++ LR+RGL++PPWRKNRYMQ +W GPY+RTTN + S V CR +GF
Sbjct: 232 DAARISLRNRGLTLPPWRKNRYMQTRWLGPYKRTTN------LTPSTSGVDTVMCRAIGF 285
Query: 288 DNAMSEIKHGVAFVRIK 304
DNA+ G FVR +
Sbjct: 286 DNAVG----GRLFVRTR 298
>At3g25240 hypothetical protein
Length = 281
Score = 210 bits (534), Expect = 9e-55
Identities = 117/287 (40%), Positives = 168/287 (57%), Gaps = 20/287 (6%)
Query: 3 FARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDD 62
F+R KRVTDP +D+AKAR++ S HS + DS P L ELVHGFLE D
Sbjct: 4 FSRTKRVTDPLDDDAKARIL-------------SSHSYILDQDS-PRLCELVHGFLE--D 47
Query: 63 GNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFL 122
G +D D S S ++ +++R++ +++D Y N+L HV A +
Sbjct: 48 GPEESFYDSDPDLSETSSAEHSGEASVEIVRMAVSFSDSDPYRNLLLAHVLRAVEVYSGF 107
Query: 123 KKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRY 182
+ ++ +V+ V SFLRE GH+AA+C ++W SS L AG++ FIDVV S + RY
Sbjct: 108 RSRNKTVFRDKVASFLRELGHDAAVCVSKWTSSSKLIAGSYHFIDVVHKPSDNDQKAVRY 167
Query: 183 FVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIP 242
V+LDFA +FEIARPT Y+ + +P +FVGN E L+ V C A K ++SRGLS+P
Sbjct: 168 LVDLDFASEFEIARPTREYTRGLQLLPNVFVGNEENLRTIVRESCDAAKRSMKSRGLSLP 227
Query: 243 PWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLVGFDN 289
PWR++ Y+Q+KWFGPY+R G+ + + V CR +GFD+
Sbjct: 228 PWRRSSYLQHKWFGPYKRKV----GSSLGVKPLNSDAVSCRSLGFDD 270
>At1g62425 unknown protein
Length = 283
Score = 156 bits (394), Expect = 1e-38
Identities = 95/258 (36%), Positives = 140/258 (53%), Gaps = 28/258 (10%)
Query: 5 RKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGN 64
R KR+ FN + A + D SSGS+HS P LS+LV F+E++
Sbjct: 7 RFKRIESAFN------VAAARVHPPCDNSSGSDHS--------PDLSDLVASFIEKEGQI 52
Query: 65 VCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKK 124
V + E S ++++ V + LR E + R+ + A ++
Sbjct: 53 VLR------EEEETSSDDNNLEDVNERLRKLLEGLSCGEE----RMRILSATMEVAGTFV 102
Query: 125 QSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFV 184
+S R++M+FLR KG +A +CK+ W+ G T G +E++DV R G + NRYFV
Sbjct: 103 GDISSSKRHLMAFLRNKGFDAGLCKSSWERFGKNTGGKYEYVDV---RCGGD-YNNRYFV 158
Query: 185 ELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPW 244
E + A +FEIARPT RY I+S VP +FVG SEELK V +C ++ ++ G+ +PPW
Sbjct: 159 ETNLAGEFEIARPTKRYLSILSQVPRVFVGTSEELKLLVRIMCHEMRRSMKHVGIHVPPW 218
Query: 245 RKNRYMQNKWFGPYRRTT 262
R+N YMQ KWFG Y+RT+
Sbjct: 219 RRNGYMQAKWFGFYKRTS 236
>At1g12030 hypothetical protein
Length = 295
Score = 149 bits (375), Expect = 2e-36
Identities = 84/233 (36%), Positives = 132/233 (56%), Gaps = 22/233 (9%)
Query: 33 SSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLL 92
SSGS+HS D + L +LV F++ + + + F E D + + V++ L
Sbjct: 29 SSGSDHSP----DDTEDLWDLVESFIDREVETLPEDA---FQEEEDDKSDEDYEDVKERL 81
Query: 93 RLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRW 152
R EN + R + + AV + + R+ M++LR KG +A +CK+RW
Sbjct: 82 REILENHGGEE-----RQRIMDEAVN----ASRVFAGEKRHFMAYLRNKGFDAGLCKSRW 132
Query: 153 DSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIF 212
+ G TAG +E++DV ++G +NRY VE + A +FEIARPT+RY +++ VP +F
Sbjct: 133 EKFGKNTAGKYEYVDV---KAGD---KNRYIVETNLAGEFEIARPTTRYLSVLAQVPRVF 186
Query: 213 VGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPV 265
VG EELK+ V +C ++ ++ + +PPWR+N YMQ KWFG Y+RT+N V
Sbjct: 187 VGTPEELKQLVRIMCFEIRRSMKRADIFVPPWRRNGYMQAKWFGHYKRTSNEV 239
>At4g32480 putative protein
Length = 287
Score = 110 bits (274), Expect = 1e-24
Identities = 53/133 (39%), Positives = 84/133 (62%), Gaps = 2/133 (1%)
Query: 134 VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFE 193
V S LRE G++ I K++W SS + AG HE+++VV +S S + R +EL F +FE
Sbjct: 120 VSSLLREAGYDCVISKSKWRSSHEIPAGEHEYLEVVD-KSVSKKGEIRVVIELCFRAEFE 178
Query: 194 IARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNK 253
+AR + Y ++ +P ++VG +E LK + LC A K C++ + + + PWRK++YMQ K
Sbjct: 179 MARGSEEYKRLIGMLPEVYVGKTERLKSLIKILCTAAKKCMKDKKMHMGPWRKHKYMQAK 238
Query: 254 WFGP-YRRTTNPV 265
WFG R++ +PV
Sbjct: 239 WFGTCERKSVSPV 251
>At4g14620 unknown protein
Length = 341
Score = 109 bits (273), Expect = 2e-24
Identities = 79/282 (28%), Positives = 138/282 (48%), Gaps = 28/282 (9%)
Query: 5 RKKRVTDPFNDEAKARLVGADLRRLSD-----VSSGSEHSGTGECDSSPSLSELVHGFLE 59
R + T P RL+ R+S+ +S +GT + PSL+++V ++E
Sbjct: 15 RVESSTKPVLKSRLKRLLDRPFTRISNSEKLLISGDGVVAGT---EFEPSLAKMVQNYME 71
Query: 60 EDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKF 119
E++ T N ++ R + + + D +D L + N S + V+
Sbjct: 72 ENNDK---QTKNGRNTHRCNCFNGNNDISDDELDFF-DYDNFKSLIQCGSFVEKSLLVEA 127
Query: 120 DFLKKQSVSVYNRN-----VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSG 174
+ +++ SV ++ V+ L G++++ICK++WD + + AG +E+IDV+
Sbjct: 128 TKIIEKNKSVKRKDELRKIVVDELSSLGYDSSICKSKWDKTRSIPAGEYEYIDVI----- 182
Query: 175 SSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCL 234
R +++DF +FEIAR TS Y E++ +P IFVG S+ +++ V + A K L
Sbjct: 183 --VNGERLIIDIDFRSEFEIARQTSGYKELLQSLPLIFVGKSDRIRQIVSIVSEASKQSL 240
Query: 235 RSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSS 276
+ +G+ PPWRK YM+ KW Y R + G P V S+
Sbjct: 241 KKKGMHFPPWRKADYMRAKWLSSYTRNS----GEKKPTVTSA 278
>At3g22970 unknown protein
Length = 370
Score = 109 bits (272), Expect = 2e-24
Identities = 70/226 (30%), Positives = 115/226 (49%), Gaps = 17/226 (7%)
Query: 46 SSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYL 105
SS L+++V F+EE++ N + ++ S D DL S + +A +L
Sbjct: 76 SSVCLAKMVQNFIEENNEKQAKCGRNRCNCFNGNN-DGSSDDESDLFGGSIDGCDASDHL 134
Query: 106 NMLR--LHVSEAAVKFDFLK--KQSVSVYNRNVMSFLREKG-----HNAAICKTRWDSSG 156
L V E + D K ++ SV ++ M + +G +N++ICK++WD S
Sbjct: 135 KSLIPCTTVGERNLLADAAKIVDKNKSVKRKDDMKKIVNEGLLSLNYNSSICKSKWDKSP 194
Query: 157 GLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNS 216
AG +E+IDV+ + R +++DF +F+IAR TS Y ++ +P IFVG S
Sbjct: 195 SFPAGEYEYIDVI-------IGEERLIIDVDFRSEFDIARQTSGYKVLLQSLPFIFVGKS 247
Query: 217 EELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTT 262
+ L + V + A K L+ +G+ PPWRK YM++KW Y R +
Sbjct: 248 DRLSQIVFLISEAAKQSLKKKGMPFPPWRKAEYMRSKWLSSYTRAS 293
>At2g20670 unknown protein
Length = 294
Score = 105 bits (262), Expect = 3e-23
Identities = 49/122 (40%), Positives = 77/122 (62%), Gaps = 1/122 (0%)
Query: 142 GHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRY 201
G++ I K++W S + AG HEFI++V RSGS + R +EL F +FEIA+ + Y
Sbjct: 130 GYDCVISKSKWRSCQDIPAGEHEFIEIVD-RSGSKKSEMRVVIELSFRAEFEIAKGSEEY 188
Query: 202 SEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRT 261
++S +P ++VG +E L+ + LC A K CLR + + + PWRK++YMQ KW G R+
Sbjct: 189 KRLISRLPEVYVGKTERLRSLIKILCIAGKKCLRDKKMHMAPWRKHKYMQAKWLGTCDRS 248
Query: 262 TN 263
++
Sbjct: 249 SS 250
>At2g38820 Unknown protein
Length = 288
Score = 105 bits (261), Expect = 4e-23
Identities = 67/246 (27%), Positives = 118/246 (47%), Gaps = 39/246 (15%)
Query: 30 SDVSSGSEHSGTGECD-SSPSLSELVHGFLEEDDGNVCDSTG----NDFDSERVDSVSDS 84
SDV + +G+ + SS L+++V F+E+++G G N F +S D
Sbjct: 53 SDVEAPLSRGNSGDFEPSSVCLAKMVLNFMEDNNGGEKQRCGRSRCNCFSGSGTESSDDE 112
Query: 85 MDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSF---LREK 141
+ + A L L L S+ RN+++ + E
Sbjct: 113 TE---------CSSGEACEILKSLVL---------------CKSIRVRNLLTDVTKIAET 148
Query: 142 GHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRY 201
++AA+CK+RW+ S AG +E++DV+ R +++DF +FEIAR T Y
Sbjct: 149 SYDAALCKSRWEKSPSCPAGEYEYVDVIMKGE-------RLLIDIDFKSKFEIARATKTY 201
Query: 202 SEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRT 261
++ +P IFVG ++ L++ ++ +C A K L+ +GL +PPWR+ Y+++KW + R
Sbjct: 202 KSMLQTLPYIFVGKADRLQKIIVLICKAAKQSLKKKGLHVPPWRRAEYVKSKWLSSHVRV 261
Query: 262 TNPVHG 267
+G
Sbjct: 262 DQNSNG 267
>At1g77160 hypothetical protein
Length = 258
Score = 102 bits (254), Expect = 3e-22
Identities = 63/197 (31%), Positives = 102/197 (50%), Gaps = 27/197 (13%)
Query: 75 SERVDSVSDSMDSVEDLLRLSAENANA-----DSYLNMLRLHVSEAAVKFDFLKKQSVSV 129
S+ +++ S ++ +++++LR SY+N RL K D + K
Sbjct: 24 SQHINTSSKALITLQEILRAKGVEEKEMEEKIRSYINRGRLSYEGDDEKRDVMNK----- 78
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAG--------NHEFIDVVRM----RSGSST 177
++S LR +G+NA++ KT WDSS G +E+ID + + R G S
Sbjct: 79 ----IVSKLRSEGYNASLSKTSWDSSFDHREGCRVFTCSRKYEYIDAMVIGDSDRDGVSK 134
Query: 178 WQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSR 237
+ R ++LDF QFE+AR T Y ++ +P +FV L+R V +CG +K ++
Sbjct: 135 LK-RVIIDLDFKTQFELARQTEAYKDMTEMLPTVFVATEGRLRRVVSLVCGEMKKSMKKE 193
Query: 238 GLSIPPWRKNRYMQNKW 254
G+S PPWR +RYMQ+KW
Sbjct: 194 GMSRPPWRTSRYMQSKW 210
>At1g77145 unknown protein
Length = 260
Score = 102 bits (254), Expect = 3e-22
Identities = 65/203 (32%), Positives = 104/203 (51%), Gaps = 27/203 (13%)
Query: 75 SERVDSVSDSMDSVEDLLRLSAENANAD-----SYLNMLRLHVSEAAVKFDFLKKQSVSV 129
S+ ++S + ++ +++++LR E S++N RL K D + K
Sbjct: 24 SQHIESSTTALITLKEILRAKGEKEKEMEEKIWSFINGGRLSYEGDDEKRDVMNK----- 78
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAG--------NHEFIDVV----RMRSGSST 177
++S LR +G++A++ KT WDSS G +E+IDV+ R G S
Sbjct: 79 ----IVSKLRSEGYDASLSKTSWDSSFDHREGCRVFRCSRKYEYIDVMVKGDSNRDGVSK 134
Query: 178 WQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSR 237
+ R ++LDF QFE+AR T Y ++ +P +FV L+R V +CG +K ++
Sbjct: 135 LK-RVIIDLDFKTQFELARQTEAYKDMTEMLPLVFVATEGRLRRVVSLVCGEMKKSMKKE 193
Query: 238 GLSIPPWRKNRYMQNKWFGPYRR 260
G+S PPWR RYMQ+KW RR
Sbjct: 194 GMSRPPWRTTRYMQSKWLPENRR 216
>At3g54550 putative protein
Length = 288
Score = 97.8 bits (242), Expect = 6e-21
Identities = 74/255 (29%), Positives = 118/255 (46%), Gaps = 36/255 (14%)
Query: 26 LRRLSDVSSGSEHSGTGECD-SSPSLSELVHGFLEEDDGN-----VCDSTGNDFDSERVD 79
L+ L+ V S + E + SS L +V ++E+ D + + N F D
Sbjct: 35 LKNLAGVDSSLSRENSEEIEPSSVCLRRMVQNYIEDPDSEKQSKCIVRNRCNCFSGSGTD 94
Query: 80 SVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNR------- 132
S SD D E++++ L L+ + A V L+ ++ + R
Sbjct: 95 S-SDEDDE---------ESSSSRRVLRSLKSLLLCANVSERDLETKTTEIVEREVEDKSR 144
Query: 133 --NVMSFLREKGHNAAICKTRWDSSGG----LTAGNHEFIDVVRMRSGSSTWQNRYFVEL 186
NV+ L G++AAICK+RW+ S + AG++E++DV + R ++
Sbjct: 145 LKNVVDELVALGYDAAICKSRWEKSKTKSYCVPAGDYEYLDV-------NIGGERVLIDF 197
Query: 187 DFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRK 246
DF +FEIAR + Y I +P +FVG + L + V+ L A K R +GL +PPWR+
Sbjct: 198 DFQSKFEIARSSETYESISKTLPYVFVGQVDRLTKVVVFLSKAAKTSFRKKGLFMPPWRR 257
Query: 247 NRYMQNKWFGPYRRT 261
Y+ KW Y RT
Sbjct: 258 AEYLLTKWVSQYDRT 272
>At2g39650 unknown protein
Length = 291
Score = 97.8 bits (242), Expect = 6e-21
Identities = 70/252 (27%), Positives = 116/252 (45%), Gaps = 16/252 (6%)
Query: 22 VGADLRRLSD---VSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSER- 77
+G DLR + ++ G + S GE D L +V FLE G+ + DS+
Sbjct: 1 MGMDLRVVPAEEWLNRGIDSSHDGEHD----LGLMVTDFLETGGGSGGAGSWCSSDSDSG 56
Query: 78 VDSVSDSMDSVEDLLRLSAEN-ANADSYLNMLRLHVSEA---AVKFDFLKKQSVSVYNRN 133
S D ++ L A++ S + L L + E +VK + Y
Sbjct: 57 FPDPSYLSDKIQYLKYSMAQHETEVLSVVRTLMLTIKEKDLHSVKSGTCNASCIRFY--- 113
Query: 134 VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFE 193
+ LR G++AA+C RW G + G++E+ID++ + +R V++DF FE
Sbjct: 114 LAKLLRLSGYDAAVCSARWQGGGKVPGGDNEYIDII-LSDTEVGQDDRLIVDIDFRSHFE 172
Query: 194 IARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNK 253
IAR Y IM +P ++VG L + + + A K L+ + +PPWR Y+++K
Sbjct: 173 IARAVDSYQRIMESLPVVYVGTVARLNQFLQVMVDAAKFSLKQNSMPLPPWRSLNYLRSK 232
Query: 254 WFGPYRRTTNPV 265
W P++R P+
Sbjct: 233 WHSPHKRHLGPI 244
>At2g06500 Ac-like transposase
Length = 582
Score = 36.6 bits (83), Expect = 0.017
Identities = 32/97 (32%), Positives = 44/97 (44%), Gaps = 22/97 (22%)
Query: 65 VCDSTGNDFDSERVDSVS-----DSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKF 119
V +S GN+ ER D V D++D V + EN + +A V F
Sbjct: 28 VINSDGNNNQQERADEVDPENKVDNLDEVVQVDENEFENKQQE-----------QAEVVF 76
Query: 120 DFLKKQSVSVYN--RNVMSFLREK--GHNAAICKTRW 152
+ KK++ SVYN RN+ L E H+ IC TRW
Sbjct: 77 E--KKKTTSVYNDWRNLSKRLEEHEGSHDHIICMTRW 111
>At1g21485 putative protein
Length = 181
Score = 32.3 bits (72), Expect = 0.33
Identities = 25/98 (25%), Positives = 43/98 (43%), Gaps = 10/98 (10%)
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFA 189
Y V++ R+K H A I KT W S G A ++ I+V+ + R ++D
Sbjct: 74 YLTKVINLFRQKHHVAEIIKTSWTSDDG-KAQTYDIIEVIHYPT-----NERLIADID-- 125
Query: 190 VQFEIARPTSRYSE-IMSYVPGIFVGNSEELKRTVLAL 226
++ E + VP FVG ++++R V +
Sbjct: 126 -MYDGLHQVMEIPENLEKDVPVAFVGTGDDIRRLVATI 162
>At5g24350 putative protein
Length = 2376
Score = 30.8 bits (68), Expect = 0.95
Identities = 33/119 (27%), Positives = 55/119 (45%), Gaps = 18/119 (15%)
Query: 31 DVSSGSEHSG--TGECDSSPSLSELVHGFLEEDDGNVCDSTG-----NDFDSERVDSVSD 83
D+SS + G G CD S+SEL+H + + D C++ G N + +++D VS
Sbjct: 1159 DISSRKQLLGFALGHCDDE-SISELLHAWKDFDLQGQCETLGMLSESNSPEFQKMDGVSC 1217
Query: 84 SMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKG 142
D + L + ++D L++ R S + V D SV ++ S L+E G
Sbjct: 1218 LTDFPQML-----DGLSSDQQLDLDRAKDSISCVAKDMPVDDSV-----DLESLLKENG 1266
>At5g66770 SCARECROW gene regulator
Length = 584
Score = 30.4 bits (67), Expect = 1.2
Identities = 30/106 (28%), Positives = 46/106 (43%), Gaps = 7/106 (6%)
Query: 11 DPFND-EAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDST 69
DPF+ ++ + +DL R+ D SS S S+ L H ++D DS
Sbjct: 148 DPFDTYPSRLSVQPSDLNRVIDTSSPLPPPTLWPPSSPLSIPPLTHESPTKEDPETNDSE 207
Query: 70 GNDFDSERVDSVSDSMDSVEDLLRLSAENAN-ADSYLNMLRLHVSE 114
+DFD E + ++ D R+S + N A L +R VSE
Sbjct: 208 DDDFDLE-----PPLLKAIYDCARISDSDPNEASKTLLQIRESVSE 248
>At1g51430 hypothetical protein
Length = 191
Score = 30.0 bits (66), Expect = 1.6
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 3/66 (4%)
Query: 102 DSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAG 161
++ +++L+ VS ++ D LK+ SVY N EK H D + G+TA
Sbjct: 71 EARISLLQDEVSTNGIEDDALKR---SVYKLNGRIDQGEKFHKTIDLSVGSDETTGITAN 127
Query: 162 NHEFID 167
N F+D
Sbjct: 128 NKAFVD 133
>At4g31340 unknown protein
Length = 437
Score = 29.6 bits (65), Expect = 2.1
Identities = 18/58 (31%), Positives = 24/58 (41%), Gaps = 3/58 (5%)
Query: 217 EELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVV 274
EE+ RT L K L + G +PPW + + F Y T HG P + V
Sbjct: 182 EEMLRTKLEATTKAKELLEAHGSWLPPWLAVHWFK---FQTYTETHWEAHGKPAVETV 236
>At4g25360 unknown protein (At4g25360)
Length = 533
Score = 29.6 bits (65), Expect = 2.1
Identities = 14/50 (28%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 2 NFARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLS 51
+F+ +++ P E K LV +D+ +DV SG + + + +PS S
Sbjct: 111 DFSSDRKLETPLTQE-KEDLVSSDITEKTDVQSGERETNVSKAEDTPSAS 159
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,095,280
Number of Sequences: 26719
Number of extensions: 308143
Number of successful extensions: 921
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 881
Number of HSP's gapped (non-prelim): 37
length of query: 304
length of database: 11,318,596
effective HSP length: 99
effective length of query: 205
effective length of database: 8,673,415
effective search space: 1778050075
effective search space used: 1778050075
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145513.12


