

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.6 + phase: 2 /partial
(753 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
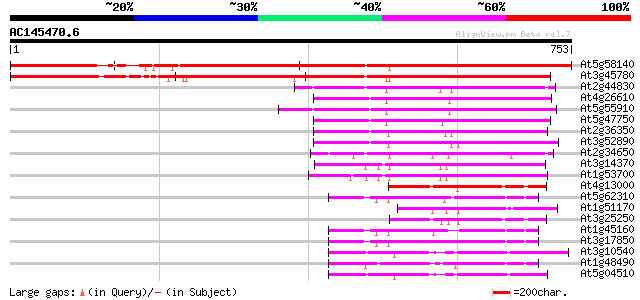
Score E
Sequences producing significant alignments: (bits) Value
At5g58140 non phototropic hypocotyl 1-like (NPL1) 1037 0.0
At3g45780 nonphototropic hypocotyl 1 (NPH1) 835 0.0
At2g44830 protein kinase like protein 275 7e-74
At4g26610 putative protein kinase 271 1e-72
At5g55910 serine/threonine-specific protein kinase ATPK64 (pir||... 269 4e-72
At5g47750 protein kinase (EC 2.7.1.37) 5 (pir||JN0505) 269 5e-72
At2g36350 putative protein kinase 266 3e-71
At3g52890 protein kinase KIPK 264 2e-70
At2g34650 protein kinase PINOID 238 7e-63
At3g14370 putative protein kinase 231 1e-60
At1g53700 auxin-induced protein kinase like protein 227 2e-59
At4g13000 unknown protein 188 8e-48
At5g62310 IRE (root hair elongation) 183 4e-46
At1g51170 putative serine/threonine protein kinase 181 1e-45
At3g25250 protein kinase, putative 180 2e-45
At1g45160 similar to IRE homolog 1 dbj|BAA89784.1 176 4e-44
At3g17850 protein kinase like protein 172 8e-43
At3g10540 3-phosphoinositide-dependent protein kinase-1 like 170 3e-42
At1g48490 IRE - like protein kinase 170 3e-42
At5g04510 3-phosphoinositide-dependent protein kinase-1 (PDK1) 164 1e-40
>At5g58140 non phototropic hypocotyl 1-like (NPL1)
Length = 915
Score = 1037 bits (2682), Expect = 0.0
Identities = 546/779 (70%), Positives = 611/779 (78%), Gaps = 63/779 (8%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGP+TD+NEVAKIRD KNGKSYCGRLLNYKK+GTPFWNLLTVTPIKDD+GNTIKFI
Sbjct: 171 RFLQGPDTDKNEVAKIRDCVKNGKSYCGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFI 230
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEGVN+KALRPNGL KSLIRYDARQKE+A+ SITEVVQT+R+ KS ++
Sbjct: 231 GMQVEVSKYTEGVNDKALRPNGLSKSLIRYDARQKEKALDSITEVVQTIRHRKSQVQ--- 287
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTT-TPGRQTPLKFHGDNNNMSRFSSYEERNN 179
E++++D + V+P + TT TPGRQT + +
Sbjct: 288 -----------ESVSNDTM----VKPDSSTTPTPGRQT---------------RQSDEAS 317
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMT------KEIEWSKYELRERDIR 233
KS R G S K SS R +D +EPE LM + W + RERDIR
Sbjct: 318 KSFRTPGRVSTP-TGSKLKSSNNRHEDLLRMEPEELMLSTEVIGQRDSWDLSD-RERDIR 375
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
QGIDLATTLERIEKNFVISDPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGPETD
Sbjct: 376 QGIDLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETD 435
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
QATV +IRDAI+DQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH
Sbjct: 436 QATVQKIRDAIRDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 495
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
+EPL+NRLSE +E+QS+KLVKATA NVD AVRELPDAN RPEDLWA HS+ V P PH ++
Sbjct: 496 VEPLQNRLSERTEMQSSKLVKATATNVDEAVRELPDANTRPEDLWAAHSKPVYPLPHNKE 555
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+ SW AI+KI A GE +GLHHF PI+PLG GDTGSVHLVEL+GTGELYAMKAMEK++MLN
Sbjct: 556 STSWKAIKKIQASGETVGLHHFKPIKPLGSGDTGSVHLVELKGTGELYAMKAMEKTMMLN 615
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHY--TPHF-----------------KQPMKI 514
RNK+ + IER + L H ++ + + + H +QPMKI
Sbjct: 616 RNKA--HRACIEREIISLLDHPFLPTLYASFQTSTHVCLITDFCPGGELFALLDRQPMKI 673
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
L EDSARFYAAEVVIGLEYLHCLGI+YRDLKPEN+LL+KDGHIVL DFDLSF+T+C PQ+
Sbjct: 674 LTEDSARFYAAEVVIGLEYLHCLGIVYRDLKPENILLKKDGHIVLADFDLSFMTTCTPQL 733
Query: 575 VKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGIL 634
+ + P RRRS+SQP P FV+EP TQSNSFVGTEEYIAPEIITGA HTSAIDWW LGIL
Sbjct: 734 IIPAAPSKRRRSKSQPLPTFVAEPSTQSNSFVGTEEYIAPEIITGAGHTSAIDWWALGIL 793
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYEMLYGRTPFRGKNRQKTF+NILHKDLTFPSSIP SL RQLIN LL RDP+SRLGS
Sbjct: 794 LYEMLYGRTPFRGKNRQKTFANILHKDLTFPSSIPVSLVGRQLINTLLNRDPSSRLGSKG 853
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSIDMDI 753
G+NEIKQH FFRGINWPLIR MSPPPLD PL I KDP AKD KWEDDGVL S D+DI
Sbjct: 854 GANEIKQHAFFRGINWPLIRGMSPPPLDAPLSIIEKDPNAKDIKWEDDGVLVNSTDLDI 912
Score = 124 bits (310), Expect = 3e-28
Identities = 78/260 (30%), Positives = 134/260 (51%), Gaps = 24/260 (9%)
Query: 141 PKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNKS-----SRKSGITSLK---- 191
P S + + +T+ + PL D NN S + E + + + + G++++K
Sbjct: 28 PSSGKETHGSTSSSSKPPL----DGNNKGSSSKWMEFQDSAKITERTAEWGLSAVKPDSG 83
Query: 192 --GVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRERDIRQGIDLATTLERIEKNF 249
G+ K S V R K+ + + E S R +L T L +++ F
Sbjct: 84 DDGISFKLSSEVERSKN--------MSRRSSEESTSSESGAFPRVSQELKTALSTLQQTF 135
Query: 250 VISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVNRIRDAIKDQRE 309
V+SD P CPI++AS F +T Y+ +EI+GRNCRFLQGP+TD+ V +IRD +K+ +
Sbjct: 136 VVSDATQPHCPIVYASSGFFTMTGYSSKEIVGRNCRFLQGPDTDKNEVAKIRDCVKNGKS 195
Query: 310 ITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLRNRLSEGSEIQS 369
+L+NY K G FWNL + P++D +G FIG+Q++ S + E + ++ + + S
Sbjct: 196 YCGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFIGMQVEVSKYTEGVNDKALRPNGL-S 254
Query: 370 AKLVKATAENVDGAVRELPD 389
L++ A + A+ + +
Sbjct: 255 KSLIRYDARQKEKALDSITE 274
>At3g45780 nonphototropic hypocotyl 1 (NPH1)
Length = 996
Score = 835 bits (2158), Expect = 0.0
Identities = 438/753 (58%), Positives = 547/753 (72%), Gaps = 43/753 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +E+AKIR+ G +YCGR+LNYKK+GT FWNLLT+ PIKD+ G +KFI
Sbjct: 235 RFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFI 294
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSK+TEG EKALRPNGLP+SLIRYDARQK+ A S+TE+V+ V+ P+++ S N
Sbjct: 295 GMQVEVSKHTEGAKEKALRPNGLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTN 354
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTP-GRQTPLKFHGDNNNMSRFSSYEERNN 179
+H D + K +++ P GR+ G N+M R + E
Sbjct: 355 ------LHPFMTKSESDELPKKPARRMSENVVPSGRRN--SGGGRRNSMQRINEIPE--- 403
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVE-----PEVLMTKEIEWSKYE-LRERDIR 233
K SRKS + S G+K KS S+ D +E E+ E S + +R++++R
Sbjct: 404 KKSRKSSL-SFMGIKKKS-ESLDESIDDGFIEYGEEDDEISDRDERPESVDDKVRQKEMR 461
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
+GIDLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGPETD
Sbjct: 462 KGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETD 521
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
TV +IR+AI +Q E+TVQLINYTKSGKKFWN+FHLQPMRDQKGE+QYFIGVQLDGS H
Sbjct: 522 LTTVKKIRNAIDNQTEVTVQLINYTKSGKKFWNIFHLQPMRDQKGEVQYFIGVQLDGSKH 581
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
+EP+RN + E + + LVK TA N+D AVRELPDAN+ PEDLWA HS+ V +PH++D
Sbjct: 582 VEPVRNVIEETAVKEGEDLVKKTAVNIDEAVRELPDANMTPEDLWANHSKVVHCKPHRKD 641
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+P W+AIQK+ GE IGL HF P++PLG GDTGSVHLVEL GT +L+AMKAM+K+VMLN
Sbjct: 642 SPPWIAIQKVLESGEPIGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLN 701
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKI 514
RNK + ER + + L H ++ + + T ++ +QP K+
Sbjct: 702 RNK--VHRARAEREILDLLDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKV 759
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
LKED+ RFYAA+VV+ LEYLHC GIIYRDLKPEN+L+Q +G I L+DFDLS +TSCKPQ+
Sbjct: 760 LKEDAVRFYAAQVVVALEYLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQL 819
Query: 575 VKQSLPGNRRR--SRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLG 632
+ S+ +++ +SQ PIF++EP+ SNSFVGTEEYIAPEII+GA HTSA+DWW LG
Sbjct: 820 LIPSIDEKKKKKQQKSQQTPIFMAEPMRASNSFVGTEEYIAPEIISGAGHTSAVDWWALG 879
Query: 633 ILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGS 692
IL+YEMLYG TPFRGK RQKTF+N+L KDL FP+SIPASL +QLI LLQRDP RLG
Sbjct: 880 ILMYEMLYGYTPFRGKTRQKTFTNVLQKDLKFPASIPASLQVKQLIFRLLQRDPKKRLGC 939
Query: 693 ATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
G+NE+KQH FF+GINW LIR +PP L+ P+
Sbjct: 940 FEGANEVKQHSFFKGINWALIRCTNPPELETPI 972
Score = 112 bits (281), Expect = 6e-25
Identities = 65/186 (34%), Positives = 102/186 (53%), Gaps = 12/186 (6%)
Query: 223 SKYELRERDI---RQGI-----DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEY 274
S E+ + D+ R GI DL L ++ FV+SD PD PI++AS F +T Y
Sbjct: 165 SSGEMSDGDVPGGRSGIPRVSEDLKDALSTFQQTFVVSDATKPDYPIMYASAGFFNMTGY 224
Query: 275 TREEILGRNCRFLQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMR 334
T +E++GRNCRFLQG TD + +IR+ + +++NY K G FWNL + P++
Sbjct: 225 TSKEVVGRNCRFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIK 284
Query: 335 DQKGELQYFIGVQLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVD---GAVRELPDAN 391
D+ G++ FIG+Q++ S H E + + + + + L++ A D +V EL +A
Sbjct: 285 DESGKVLKFIGMQVEVSKHTEGAKEKALRPNGLPES-LIRYDARQKDMATNSVTELVEAV 343
Query: 392 LRPEDL 397
RP L
Sbjct: 344 KRPRAL 349
>At2g44830 protein kinase like protein
Length = 765
Score = 275 bits (703), Expect = 7e-74
Identities = 160/416 (38%), Positives = 226/416 (53%), Gaps = 71/416 (17%)
Query: 383 AVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLG 442
A R + L E W+ + +++ +PHK ++P W AI I R +G+ HF ++ LG
Sbjct: 312 ASRASDSSGLSEESSWSNFTGSLN-KPHKGNDPWWNAILAIRTRDGILGMSHFKLLKRLG 370
Query: 443 CGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQ 502
CGD GSV+L EL GT +A+K M+K+ + +R K ER + + L H ++ +
Sbjct: 371 CGDIGSVYLAELSGTRCHFAVKVMDKASLEDRKK--LNRAQTERDILQLLDHPFLPTLYT 428
Query: 503 HY-------------------TPHFKQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRD 543
H+ T +QP K E +ARFYAAEV++ LEYLH LG++YRD
Sbjct: 429 HFETDRFSCLVMEYCPGGDLHTLRQRQPGKHFSEYAARFYAAEVLLALEYLHMLGVVYRD 488
Query: 544 LKPENLLLQKDGHIVLTDFDLSFITSCKPQVVK--------------------------- 576
LKPEN+L++ DGHI+L+DFDLS + P ++K
Sbjct: 489 LKPENVLVRDDGHIMLSDFDLSLRCAVSPTLIKTFDSDPSRRGAFCVQPACMEPTSACII 548
Query: 577 --------QSLPGNRRRSRSQPP------------PIFVSEPVTQSNSFVGTEEYIAPEI 616
P ++++S+ P V+EP T+S SFVGT EY+APEI
Sbjct: 549 QPSCFLPRSIFPNKNKKNKSRKTQADFFKSHSGSLPELVAEPNTRSMSFVGTHEYLAPEI 608
Query: 617 ITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQ 676
I G H SA+DWWT GI ++E+LYG+TPF+G + T N++ + L FP S S A R
Sbjct: 609 IKGEGHGSAVDWWTFGIFVHELLYGKTPFKGSGNRATLFNVVGEQLKFPESPATSYAGRD 668
Query: 677 LINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDP 732
LI ALL +DP +RLG+ G+ EIKQHPFF G+NW LIR +PP +VP Q + P
Sbjct: 669 LIQALLVKDPKNRLGTKRGATEIKQHPFFEGVNWALIRCSTPP--EVPRQMETEPP 722
>At4g26610 putative protein kinase
Length = 506
Score = 271 bits (692), Expect = 1e-72
Identities = 148/385 (38%), Positives = 210/385 (54%), Gaps = 67/385 (17%)
Query: 408 RPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAME 467
+PHK ++ W AIQ + + +GL+HF ++ LGCGD G+VHL EL GT +AMK M+
Sbjct: 96 KPHKANDVRWEAIQAVRTKHGVLGLNHFRLLKRLGCGDIGTVHLAELHGTRCFFAMKVMD 155
Query: 468 KSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQHY-------------------TPHF 508
K + +R K + ER + + L H ++ + H+ T
Sbjct: 156 KGALASRKKLLRAQT--EREILQCLDHPFLPTLYSHFETEKFSCLVMEFCPGGDLHTLRQ 213
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
+QP K E +A+FY AEV++ +EYLH LGIIYRDLKPEN+L++ DGH++L+DFDLS
Sbjct: 214 RQPGKRFSEQAAKFYVAEVLLAMEYLHMLGIIYRDLKPENVLVRDDGHVMLSDFDLSLRC 273
Query: 569 SCKPQVVKQSLPGNRRRSRS---------------------------------------- 588
+ P VV+ ++ + + S
Sbjct: 274 TVSPTVVRSTVLASEGQKNSGYCAQPACIQQPSCISAPTTCFSPRYFSSKSKKDKKMKNE 333
Query: 589 -----QPPPIFVSEPVT-QSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGR 642
P P V+EP + +S SFVGT EY+APEII G H SA+DWWT GI LYE+L+G+
Sbjct: 334 TGNQVSPLPELVAEPTSARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLFGK 393
Query: 643 TPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQH 702
TPF+G + T N++ + L FP S S AAR LI +LL ++P RL G+ E+KQH
Sbjct: 394 TPFKGSGNRATLFNVVGQPLRFPESPVVSFAARDLIRSLLVKEPQHRLAYKRGATEMKQH 453
Query: 703 PFFRGINWPLIRNMSPPPLDVPLQF 727
PFF G+NW L+R SPP + P+ +
Sbjct: 454 PFFEGVNWALVRCASPPEIPKPVDY 478
>At5g55910 serine/threonine-specific protein kinase ATPK64
(pir||S20918)
Length = 498
Score = 269 bits (688), Expect = 4e-72
Identities = 155/441 (35%), Positives = 229/441 (51%), Gaps = 72/441 (16%)
Query: 362 SEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWVAIQ 421
S + ++K+ + R +++ E + S +++ +PHK ++ W AIQ
Sbjct: 37 SSDGKTGNSKINEQGESGKSSTCRPSTSSDISDESTCSSFSSSIN-KPHKANDVRWEAIQ 95
Query: 422 KITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYY 481
+ + +GL+HF ++ LGCGD G+VHL EL GT +AMK M+K+ + +R K +
Sbjct: 96 AVRTKHGGLGLNHFRLLKRLGCGDIGTVHLAELNGTRCYFAMKVMDKTALASRKKLLRAQ 155
Query: 482 RFIERALKERLYHYWIILFFQHY-------------------TPHFKQPMKILKEDSARF 522
ER + + L H ++ + H+ T +QP K E +A+F
Sbjct: 156 T--EREILQCLDHPFLPTLYSHFETEKFSCLVMEFCPGGDLHTLRQRQPGKRFTEQAAKF 213
Query: 523 YAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGN 582
Y AEV++ +EYLH LGIIYRDLKPEN+L++ DGH++L+DFDLS + +V+ + G+
Sbjct: 214 YVAEVLLAMEYLHMLGIIYRDLKPENVLVRDDGHVMLSDFDLSLRCTVSLSIVRSANVGS 273
Query: 583 RRRSRSQ-------------------------------------------------PPPI 593
S++ P P
Sbjct: 274 EGLSKNSVSCSQQPACIQQPSCISMAPTSCFGPRFFSSKSKKDKKPKTENGNHQVTPLPE 333
Query: 594 FVSEPV-TQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQK 652
V+EP +S SFVGT EY+APEII G H SA+DWWT GI LYE+L+G+TPF+G +
Sbjct: 334 LVAEPTGARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLFGKTPFKGSGNRA 393
Query: 653 TFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPL 712
T N++ + L FP S S AAR LI +LL ++P RL G+ EIKQHPFF G+NW L
Sbjct: 394 TLFNVVGQPLRFPESPVVSFAARDLIRSLLVKEPQHRLAYKRGATEIKQHPFFEGVNWAL 453
Query: 713 IRNMSPPPLDVPLQFIGKDPT 733
+R SPP + P+ +PT
Sbjct: 454 VRCASPPEIPKPVDLEALNPT 474
>At5g47750 protein kinase (EC 2.7.1.37) 5 (pir||JN0505)
Length = 586
Score = 269 bits (687), Expect = 5e-72
Identities = 150/387 (38%), Positives = 209/387 (53%), Gaps = 70/387 (18%)
Query: 408 RPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAME 467
+PHK ++ W AIQ + R +GL HF ++ LGCGD GSV+L EL GT +AMK M+
Sbjct: 164 KPHKANDLRWEAIQAVRVRDGLLGLSHFRLLKRLGCGDIGSVYLSELSGTKCYFAMKVMD 223
Query: 468 KSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQHY-------------------TPHF 508
K+ + +R K + ER + + L H ++ + H+ T
Sbjct: 224 KTSLASRKKLLRAQT--EREILQSLDHPFLPTLYTHFETEKFSCLVMEFCPGGDLHTLRQ 281
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
+QP K E + +FY AE ++ LEYLH LGI+YRDLKPEN+L+++DGHI+L+DFDLS
Sbjct: 282 RQPGKHFSEQAVKFYIAESLLALEYLHMLGIVYRDLKPENVLVREDGHIMLSDFDLSLRC 341
Query: 569 SCKPQVVKQSL------------------------------------------------P 580
P +VK + P
Sbjct: 342 LVSPTLVKSAAIESDPLRKNVYCVQPACIEPSCIQPSCTVPTTCFSPRLFSSKSKKDRKP 401
Query: 581 GNRRRSRSQPPPIFVSEPV-TQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEML 639
N ++ +P P V+EP +S SFVGT EY+APEII G H SA+DWWT GI LYE+L
Sbjct: 402 KNDTANQVRPLPELVAEPTDARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELL 461
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEI 699
+GRTPF+G ++T N++ + L FP + S AAR LI LL ++P RLG G+ E+
Sbjct: 462 FGRTPFKGSGNRQTLFNVVGQPLRFPETPVVSFAARDLIRGLLMKEPQQRLGFKRGATEV 521
Query: 700 KQHPFFRGINWPLIRNMSPPPLDVPLQ 726
KQHPFF G+NW LIR +PP + P++
Sbjct: 522 KQHPFFEGVNWALIRCATPPEIPKPVE 548
>At2g36350 putative protein kinase
Length = 949
Score = 266 bits (680), Expect = 3e-71
Identities = 147/385 (38%), Positives = 213/385 (55%), Gaps = 73/385 (18%)
Query: 408 RPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAME 467
+PH + W A++ + + +GL HF+ ++ LGCGD G+V+L EL GT L+A+K M+
Sbjct: 532 KPHMSMDVRWEAVKHVKLQYGSLGLRHFNLLKKLGCGDIGTVYLAELVGTNCLFAIKVMD 591
Query: 468 KSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQHYTP-------------------HF 508
+ R K+ ERA+ + L H ++ + +T
Sbjct: 592 NEFLARRKKTPRAQA--ERAILKMLDHPFLPTLYAQFTSDNLSCLVMEYCPGGDLHVLRQ 649
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
KQ + E + RFY AE+++ LEYLH LG+IYRDLKPEN+L+++DGHI+LTDFDLS
Sbjct: 650 KQLSRCFSEPATRFYVAEILLALEYLHMLGVIYRDLKPENILVREDGHIMLTDFDLSLRC 709
Query: 569 SCKPQVVKQSLPGNR-----------------------------------RRSRSQPP-- 591
+ P +++ + P + ++R+Q P
Sbjct: 710 AVNPTLLRSTSPPEKDPARMSGPYSTSNCIQPLCIEPSCRVPCFSPRLLSTQARNQKPRK 769
Query: 592 --------------PIFVSEPV-TQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLY 636
P V+EP +SNSFVGT EY+APEII G H +A+DWWT G+LLY
Sbjct: 770 PKRPDLLTQQFRSLPQLVAEPTEARSNSFVGTHEYLAPEIIKGEGHGAAVDWWTFGVLLY 829
Query: 637 EMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGS 696
E+LYG+TPF+G + ++T SN+++++L FP S S A++LI LL +DP SRLGS G+
Sbjct: 830 ELLYGKTPFKGYDNEETLSNVVYQNLKFPDSPLVSFQAKELIRRLLVKDPESRLGSEKGA 889
Query: 697 NEIKQHPFFRGINWPLIRNMSPPPL 721
EIK+HPFF G+NW LIR PP L
Sbjct: 890 AEIKRHPFFEGLNWALIRCAIPPEL 914
>At3g52890 protein kinase KIPK
Length = 744
Score = 264 bits (674), Expect = 2e-70
Identities = 153/402 (38%), Positives = 212/402 (52%), Gaps = 75/402 (18%)
Query: 408 RPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAME 467
+PH + W AI+ I + +GL HF+ ++ LGCGD G+V+L EL GT L+A+K M+
Sbjct: 321 KPHMSMDVRWEAIKHIKVQYGSLGLRHFNLLKKLGCGDIGTVYLAELIGTNCLFAIKVMD 380
Query: 468 KSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQHYTP-------------------HF 508
+ R KS ER + + L H ++ + +T
Sbjct: 381 NEFLARRKKSPRAQA--EREILKMLDHPFLPTLYAQFTSDNLSCLVMEYCPGGDLHVLRQ 438
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
KQ + E +ARFY AE+++ LEYLH LGIIYRDLKPEN+L+++DGHI+LTDFDLS
Sbjct: 439 KQLGRCFPEPAARFYVAEILLALEYLHMLGIIYRDLKPENILVREDGHIMLTDFDLSLRC 498
Query: 569 SCKPQVVKQSLPGNRRRSRSQPP--------PIFVSEPVTQ------------------- 601
+ P +V+ + P + +R P P ++EP Q
Sbjct: 499 AVNPTLVRSNSPPGKDPARISGPYNTSNCIQPFCITEPSCQVSCFSPRLSSNQQQGRKPK 558
Query: 602 ---------------------------SNSFVGTEEYIAPEIITGARHTSAIDWWTLGIL 634
SNSFVGT EY+APEII G H +A+DWWT G+L
Sbjct: 559 RGDHLSKTQQHLSRSLPQLVAEPTEARSNSFVGTHEYLAPEIIKGEGHGAAVDWWTFGVL 618
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYE+LYG+TPF+G N +T +N++ ++L FP S S A+ LI LL ++P +RLGS
Sbjct: 619 LYELLYGKTPFKGYNNDETLANVVLQNLKFPDSPLVSFQAKDLIRGLLVKEPENRLGSEK 678
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKD 736
GS EIK+HPFF G+NW LIR PP L ++ G A D
Sbjct: 679 GSVEIKRHPFFEGLNWALIRCAIPPELPDFYEYGGGPEAAAD 720
>At2g34650 protein kinase PINOID
Length = 438
Score = 238 bits (608), Expect = 7e-63
Identities = 142/377 (37%), Positives = 210/377 (55%), Gaps = 57/377 (15%)
Query: 405 VSPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGE----- 459
+S +PH+ + ++ I++ +G + F +R +G GD G+V+L L G E
Sbjct: 47 LSLKPHRSSDFAYAEIRRRKKQG--LTFRDFRLMRRIGAGDIGTVYLCRLAGDEEESRSS 104
Query: 460 LYAMKAMEKSVMLNRNKSVFYYRFIERALKERLYHYWI-ILFFQHYTPHF---------- 508
+AMK ++K + + K + +E+ + + L H ++ L+ + HF
Sbjct: 105 YFAMKVVDKEALALKKK--MHRAEMEKTILKMLDHPFLPTLYAEFEASHFSCIVMEYCSG 162
Query: 509 --------KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLT 560
+QP + SARFYAAEV++ LEYLH LGIIYRDLKPEN+L++ DGHI+L+
Sbjct: 163 GDLHSLRHRQPHRRFSLSSARFYAAEVLVALEYLHMLGIIYRDLKPENILVRSDGHIMLS 222
Query: 561 DFDLSF-------ITSCKPQVVKQSLPGNRRRSR----------------SQPPPIFVSE 597
DFDLS + S Q L RR +R +P +FV+E
Sbjct: 223 DFDLSLCSDSIAAVESSSSSPENQQLRSPRRFTRLARLFQRVLRSKKVQTLEPTRLFVAE 282
Query: 598 PVT-QSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSN 656
PVT +S SFVGT EY+APE+ +G H +A+DWW G+ LYEM+YG+TPF N
Sbjct: 283 PVTARSGSFVGTHEYVAPEVASGGSHGNAVDWWAFGVFLYEMIYGKTPFVAPTNDVILRN 342
Query: 657 ILHKDLTFPSSIPAS---LAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLI 713
I+ + L+FP+ PA+ L AR LI+ LL +DP RLGS G+ E+K HPFF+G+N+ LI
Sbjct: 343 IVKRQLSFPTDSPATMFELHARNLISGLLNKDPTKRLGSRRGAAEVKVHPFFKGLNFALI 402
Query: 714 RNMSPPPLDVPLQFIGK 730
R ++PP ++P + K
Sbjct: 403 RTLTPP--EIPSSVVKK 417
>At3g14370 putative protein kinase
Length = 480
Score = 231 bits (589), Expect = 1e-60
Identities = 138/352 (39%), Positives = 191/352 (54%), Gaps = 44/352 (12%)
Query: 410 HKRDNPSWVAIQ--KITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAME 467
H+R +P W AI+ K+ + I L H IR LG G+ G V L L+ + +A+K ++
Sbjct: 61 HRRHDPHWSAIKSAKLLSSDGNIHLRHLKLIRHLGTGNLGRVFLCNLRDSSARFALKVID 120
Query: 468 KSVMLNRNK------SVFYYRFIERALKERLY------HYWIILFFQHYTPHF------- 508
++ + K ++ LY HY +L Y P+
Sbjct: 121 RNCLTTEKKLSQVETEAEILSLLDHPFLPTLYARIDESHYTCLLI--DYAPNGDLHSLLR 178
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
KQP L RF+AAEV++ LEYLH +GI+YRDLKPEN+LL++DGH++L+DFDL F +
Sbjct: 179 KQPGNRLPIQPVRFFAAEVLVALEYLHAMGIVYRDLKPENVLLREDGHVMLSDFDLCFKS 238
Query: 569 SCKPQVVKQ------SLPGNRRR-------------SRSQPPPIFVSEPVTQ-SNSFVGT 608
P + S P RRR R + F +EPVT S S VGT
Sbjct: 239 DVVPTFKSRRYRRSSSSPSLRRRRSGCFSVAAEKKYEREEIVSEFAAEPVTAFSRSCVGT 298
Query: 609 EEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSI 668
EY+APE+++G H S +DWW GI LYE+LYG TPF+G+++++T NI+ T +
Sbjct: 299 HEYLAPELVSGNGHGSGVDWWAFGIFLYELLYGTTPFKGESKEQTLRNIVSTTKTASFHM 358
Query: 669 PASL-AARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPP 719
L AR LI LL +DP RLG A G+ +IK+HPFF GI WPLIR+ PP
Sbjct: 359 DGDLDEARDLIEKLLVKDPRKRLGCARGAQDIKRHPFFDGIKWPLIRHYKPP 410
>At1g53700 auxin-induced protein kinase like protein
Length = 476
Score = 227 bits (579), Expect = 2e-59
Identities = 141/363 (38%), Positives = 193/363 (52%), Gaps = 47/363 (12%)
Query: 402 SQAVSPRPHKRDNPSWVAIQKITARGE--KIGLHHFSPIRPLGCGDTGSVHLVELQG--- 456
S A + H+R +P W +I+ T ++ L HF +R LG G+ G V L L+
Sbjct: 58 SSAATTLHHRRYDPHWTSIRAATTLSSDGRLHLRHFKLVRHLGTGNLGRVFLCHLRDCPN 117
Query: 457 -TGELYAMKAMEKSVMLNRNKS-----VFYYRFIERALKERLY------HYWIILFFQHY 504
TG +A+K +++ V+ + S ++ LY HY +L Y
Sbjct: 118 PTG--FALKVIDRDVLTAKKISHVETEAEILSLLDHPFLPTLYARIDASHYTCLLI--DY 173
Query: 505 TPHF-------KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHI 557
P+ KQP L RF+AAEV++ LEYLH LGI+YRDLKPEN+L+++DGHI
Sbjct: 174 CPNGDLHSLLRKQPNNRLPISPVRFFAAEVLVALEYLHALGIVYRDLKPENILIREDGHI 233
Query: 558 VLTDFDLSFITSCKPQVVKQ------SLPGNRRR-----------SRSQPPPIFVSEPVT 600
+L+DFDL F P + S P RR R + F +EPVT
Sbjct: 234 MLSDFDLCFKADVVPTFRSRRFRRTSSSPRKTRRGGGCFSTEVEYEREEIVAEFAAEPVT 293
Query: 601 Q-SNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNIL- 658
S S VGT EY+APE++ G H S +DWW GI LYEMLYG TPF+G +++T NI+
Sbjct: 294 AFSKSCVGTHEYLAPELVAGNGHGSGVDWWAFGIFLYEMLYGTTPFKGGTKEQTLRNIVS 353
Query: 659 HKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSP 718
+ D+ F + A+ LI LL +DP RLG A G+ +IK+H FF GI WPLIRN P
Sbjct: 354 NDDVAFTLEEEGMVEAKDLIEKLLVKDPRKRLGCARGAQDIKRHEFFEGIKWPLIRNYKP 413
Query: 719 PPL 721
P +
Sbjct: 414 PEI 416
>At4g13000 unknown protein
Length = 372
Score = 188 bits (478), Expect = 8e-48
Identities = 101/235 (42%), Positives = 146/235 (61%), Gaps = 32/235 (13%)
Query: 509 KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFIT 568
KQ ++ ++ RFYAAE+VI LEYLH GI+YRDLKP+N+++Q++GH++L DFDLS T
Sbjct: 112 KQSEEMFSDEIIRFYAAELVIALEYLHNQGIVYRDLKPDNVMIQENGHLMLVDFDLS--T 169
Query: 569 SCKPQVVKQSLPGNRRRSRS--QPPPIFVSEPV---------------------TQSNSF 605
+ P+ + S + R S + + IF + +SNSF
Sbjct: 170 NLPPRTPQSSFSSSPRLSTATKKERSIFAFSGLCNSGISPDDSVSRSSESEFSGEKSNSF 229
Query: 606 VGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFP 665
VGTEEY+APE+ITG+ H A+DWW+LG++LYEMLYG TPFRG NR++TF IL + P
Sbjct: 230 VGTEEYVAPEVITGSGHDFAVDWWSLGVVLYEMLYGATPFRGSNRKETFLKILTEP---P 286
Query: 666 SSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPP 720
S + + + R L+ LL++DP+ R+ IK H FF+G++W L+ +S PP
Sbjct: 287 SLVGETTSLRDLVRKLLEKDPSRRI----NVEGIKGHDFFKGLDWDLVLKVSRPP 337
>At5g62310 IRE (root hair elongation)
Length = 1168
Score = 183 bits (464), Expect = 4e-46
Identities = 111/314 (35%), Positives = 175/314 (55%), Gaps = 43/314 (13%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERA 487
++ + F I+P+ G G V L + + TG+L+A+K ++K+ M+ +N +E
Sbjct: 747 DRTSIEDFEIIKPISRGAFGRVFLAKKRATGDLFAIKVLKKADMIRKNA-------VESI 799
Query: 488 LKER-----LYHYWIILFFQHYTPH-----------------FKQPMKILKEDSARFYAA 525
L ER + + +++ FF +T + + L ED AR Y A
Sbjct: 800 LAERNILISVRNPFVVRFFYSFTCRENLYLVMEYLNGGDLFSLLRNLGCLDEDMARIYIA 859
Query: 526 EVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFI--TSCKPQVVKQSLPGN- 582
EVV+ LEYLH + II+RDLKP+NLL+ +DGHI LTDF LS + + + +S GN
Sbjct: 860 EVVLALEYLHSVNIIHRDLKPDNLLINQDGHIKLTDFGLSKVGLINSTDDLSGESSLGNS 919
Query: 583 ----RRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEM 638
S++Q + + ++ VGT +Y+APEI+ G H DWW++G++L+E+
Sbjct: 920 GFFAEDGSKAQHSQ---GKDSRKKHAVVGTPDYLAPEILLGMGHGKTADWWSVGVILFEV 976
Query: 639 LYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLA--ARQLINALLQRDPASRLGSATGS 696
L G PF + Q+ F NI+++D+ +P ++P ++ A LIN LL +P RLG ATG+
Sbjct: 977 LVGIPPFNAETPQQIFENIINRDIPWP-NVPEEISYEAHDLINKLLTENPVQRLG-ATGA 1034
Query: 697 NEIKQHPFFRGINW 710
E+KQH FF+ INW
Sbjct: 1035 GEVKQHHFFKDINW 1048
>At1g51170 putative serine/threonine protein kinase
Length = 404
Score = 181 bits (459), Expect = 1e-45
Identities = 103/243 (42%), Positives = 144/243 (58%), Gaps = 33/243 (13%)
Query: 521 RFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLS------------FIT 568
+FY AE+V L++LH +GI YRDLKPEN+LLQ+ GH+ LTDFDLS ++
Sbjct: 131 KFYLAEIVCALDHLHTMGIAYRDLKPENILLQESGHVTLTDFDLSCSLNKPTRPEFYHLS 190
Query: 569 SCKPQVVKQS-LPGNRR-----RSRSQPPPIFVSEPVTQ----------SNSFVGTEEYI 612
+P +S L N++ R + + P+T+ SNSFVGT+EYI
Sbjct: 191 DPEPDPNPESNLSHNKKSLRIFRQKKKKTKSARVNPITRRRLSFSGGERSNSFVGTDEYI 250
Query: 613 APEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASL 672
+PE+I G H A+DWW LG+L YEM+YG TPF+G+N+++TF N+L K+ F P+ L
Sbjct: 251 SPEVIRGDGHDFAVDWWALGVLTYEMMYGETPFKGRNKKETFRNVLVKEPEFAGK-PSDL 309
Query: 673 AARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDP 732
LI LL +DP R G G+ EIK+H FF+G+ W L+ + PP +PL+ G D
Sbjct: 310 T--DLIRRLLVKDPTKRFGFWRGAAEIKEHAFFKGVRWELLTEVLRPPF-IPLRDDG-DL 365
Query: 733 TAK 735
T K
Sbjct: 366 TGK 368
>At3g25250 protein kinase, putative
Length = 421
Score = 180 bits (457), Expect = 2e-45
Identities = 102/238 (42%), Positives = 143/238 (59%), Gaps = 36/238 (15%)
Query: 510 QPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITS 569
Q + ++ RFYAAE+V+ L+YLH GI+YRDLKP+N+++Q++GH++L DFDLS T+
Sbjct: 116 QSESMFSDEIIRFYAAELVLALDYLHNQGIVYRDLKPDNVMIQENGHLMLIDFDLS--TN 173
Query: 570 CKPQVVKQS------LPGNRRRSR------------SQPPPIFVSEPVT---------QS 602
P+ + S P +R+ R S I V T +S
Sbjct: 174 LAPRTPQPSPSLSKPSPTMKRKKRLFRFTSFCNSGISPQESISVHSSSTLAVSDSSGEKS 233
Query: 603 NSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDL 662
NSFVGTEEY+APE+I+G H A+DWW+LG++LYEMLYG TPFRG NR++TF IL K
Sbjct: 234 NSFVGTEEYVAPEVISGDGHDFAVDWWSLGVVLYEMLYGATPFRGSNRKETFYRILSKP- 292
Query: 663 TFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPP 720
P+ + + R LI LL++DP+ R+ EIK H FFRG++W + +S PP
Sbjct: 293 --PNLTGETTSLRDLIRRLLEKDPSRRI----NVEEIKGHDFFRGVDWEKVILVSRPP 344
>At1g45160 similar to IRE homolog 1 dbj|BAA89784.1
Length = 1067
Score = 176 bits (446), Expect = 4e-44
Identities = 111/320 (34%), Positives = 166/320 (51%), Gaps = 59/320 (18%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERA 487
++I + F I+P+ G G V L + TG+ +A+K ++K M+ +N IER
Sbjct: 663 DRISIDDFEIIKPISRGAFGKVFLARKRTTGDFFAIKVLKKLDMIRKND-------IERI 715
Query: 488 LKER-----LYHYWIILFFQHYTPH-----------------FKQPMKILKEDSARFYAA 525
L+ER + + +++ FF +T Q + L E+ AR Y A
Sbjct: 716 LQERNILITVRYPFLVRFFYSFTCRDNLYLVMEYLNGGDLYSLLQKVGCLDEEIARIYIA 775
Query: 526 EVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFI-------------TSCKP 572
E+V+ LEYLH L I++RDLKP+NLL+ +GHI LTDF LS I + P
Sbjct: 776 ELVLALEYLHSLKIVHRDLKPDNLLIAYNGHIKLTDFGLSKIGLINNTIDLSGHESDVSP 835
Query: 573 QVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLG 632
+ N+ R + +S VGT +Y+APEI+ G H A DWW+ G
Sbjct: 836 RTNSHHFQKNQEEERIR-------------HSAVGTPDYLAPEILLGTEHGYAADWWSAG 882
Query: 633 ILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLA--ARQLINALLQRDPASRL 690
I+L+E+L G PF +K F NIL+ + +P +P ++ A+ LIN LL +P RL
Sbjct: 883 IVLFELLTGIPPFTASRPEKIFDNILNGKMPWP-DVPGEMSYEAQDLINRLLVHEPEKRL 941
Query: 691 GSATGSNEIKQHPFFRGINW 710
G A G+ E+K HPFF+G++W
Sbjct: 942 G-ANGAAEVKSHPFFQGVDW 960
>At3g17850 protein kinase like protein
Length = 1296
Score = 172 bits (435), Expect = 8e-43
Identities = 104/311 (33%), Positives = 168/311 (53%), Gaps = 37/311 (11%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERA 487
++ + F I+P+ G G V L + + TG+L+A+K ++K+ M+ +N +E
Sbjct: 875 DRTSIDDFEIIKPISRGAFGRVFLAKKRTTGDLFAIKVLKKADMIRKNA-------VESI 927
Query: 488 LKER-----LYHYWIILFFQHYTPH-----------------FKQPMKILKEDSARFYAA 525
L ER + + +++ FF +T + + L+ED R Y A
Sbjct: 928 LAERDILINVRNPFVVRFFYSFTCRDNLYLVMEYLNGGDLYSLLRNLGCLEEDIVRVYIA 987
Query: 526 EVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFI--TSCKPQVVKQSLPGNR 583
EVV+ LEYLH G+++RDLKP+NLL+ DGHI LTDF LS + + + ++ G
Sbjct: 988 EVVLALEYLHSEGVVHRDLKPDNLLIAHDGHIKLTDFGLSKVGLINSTDDLAGPAVSGTS 1047
Query: 584 RRSRSQPPPIFVSEPV--TQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYG 641
+ E + + S VGT +Y+APEI+ G H + DWW++GI+L+E++ G
Sbjct: 1048 LLDEEESRLAASEEQLERRKKRSAVGTPDYLAPEILLGTGHGATADWWSVGIILFELIVG 1107
Query: 642 RTPFRGKNRQKTFSNILHKDLTFPSSIPASLA--ARQLINALLQRDPASRLGSATGSNEI 699
PF ++ Q+ F NIL++ + +P +P ++ A +I+ L DP RLG A G+ E+
Sbjct: 1108 IPPFNAEHPQQIFDNILNRKIPWP-HVPEEMSAEAHDIIDRFLTEDPHQRLG-ARGAAEV 1165
Query: 700 KQHPFFRGINW 710
KQH FF+ INW
Sbjct: 1166 KQHIFFKDINW 1176
>At3g10540 3-phosphoinositide-dependent protein kinase-1 like
Length = 486
Score = 170 bits (430), Expect = 3e-42
Identities = 113/340 (33%), Positives = 168/340 (49%), Gaps = 39/340 (11%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERA 487
E H F + G G V + + G +YA+K M+K + NK+ Y +ER
Sbjct: 38 ENFTYHDFELGKIYGVGSYSKVVRAKKKDNGTVYALKIMDKKFITKENKTA--YVKLERI 95
Query: 488 LKERLYHYWIILFFQHYTPHFKQPMKI-----------------LKEDSARFYAAEVVIG 530
+ ++L H I+ F + M + L ED ARFY+AEVV
Sbjct: 96 VLDQLEHPGIVKLFFTFQDTQSLYMALESCEGGELFDQITRKGRLSEDEARFYSAEVVDA 155
Query: 531 LEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQP 590
LEY+H +G+I+RD+KPENLLL DGHI + DF S KP + S+
Sbjct: 156 LEYIHNMGLIHRDIKPENLLLTLDGHIKIADFG-----SVKPM----------QDSQITV 200
Query: 591 PPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNR 650
P S+ ++ +FVGT Y+ PE++ + T D W LG LY+ML G +PF+ +
Sbjct: 201 LPNAASD--DKACTFVGTAAYVPPEVLNSSPATFGNDLWALGCTLYQMLSGTSPFKDASE 258
Query: 651 QKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT-GSNEIKQHPFFRGIN 709
F I+ +D+ FP+ S AAR LI+ LL DP+ R G+ + G + +K+HPFF+G++
Sbjct: 259 WLIFQRIIARDIKFPNHF--SEAARDLIDRLLDTDPSRRPGAGSEGYDSLKRHPFFKGVD 316
Query: 710 WPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSI 749
W +R+ +PP L P W V +TS+
Sbjct: 317 WKNLRSQTPPKLAPDPASQSASPERDGSPWNPTHVGDTSV 356
>At1g48490 IRE - like protein kinase
Length = 1235
Score = 170 bits (430), Expect = 3e-42
Identities = 109/308 (35%), Positives = 166/308 (53%), Gaps = 37/308 (12%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN------------ 475
++I + F ++ + G G V L TG+L+A+K + K+ M+ +N
Sbjct: 821 DRISIDDFEVMKSISRGAFGHVILARKNTTGDLFAIKVLRKADMIRKNAVESILAERDIL 880
Query: 476 ---KSVFYYRFIER-ALKERLYHYWIILFFQHYTPHFKQPMKI--LKEDSARFYAAEVVI 529
++ F RF E LY +++ + + + KI L E +AR Y AEVV+
Sbjct: 881 INARNPFVVRFFYSFTCSENLY---LVMEYLNGGDFYSMLRKIGCLDEANARVYIAEVVL 937
Query: 530 GLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQ 589
LEYLH G+++RDLKP+NLL+ DGH+ LTDF LS K ++ + + S
Sbjct: 938 ALEYLHSEGVVHRDLKPDNLLIAHDGHVKLTDFGLS-----KVGLINNT--DDLSGPVSS 990
Query: 590 PPPIFVSE-----PVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTP 644
+ V E + S VGT +Y+APEI+ G H + DWW++GI+LYE L G P
Sbjct: 991 ATSLLVEEKPKLPTLDHKRSAVGTPDYLAPEILLGTGHGATADWWSVGIILYEFLVGIPP 1050
Query: 645 FRGKNRQKTFSNILHKDLTFPSSIPASLA--ARQLINALLQRDPASRLGSATGSNEIKQH 702
F + Q+ F NIL++++ +P +P ++ AR LI+ LL DP RLG A G+ E+KQH
Sbjct: 1051 FNADHPQQIFDNILNRNIQWP-PVPEDMSHEARDLIDRLLTEDPHQRLG-ARGAAEVKQH 1108
Query: 703 PFFRGINW 710
FF+ I+W
Sbjct: 1109 SFFKDIDW 1116
>At5g04510 3-phosphoinositide-dependent protein kinase-1 (PDK1)
Length = 491
Score = 164 bits (416), Expect = 1e-40
Identities = 108/312 (34%), Positives = 159/312 (50%), Gaps = 39/312 (12%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERA 487
E H F + G G V + + TG +YA+K M+K + NK+ Y +ER
Sbjct: 37 ENFTSHDFEFGKIYGVGSYSKVVRAKKKETGTVYALKIMDKKFITKENKTA--YVKLERI 94
Query: 488 LKERLYHYWIILFFQHYTPHFKQPMKI-----------------LKEDSARFYAAEVVIG 530
+ ++L H II + + M + L ED ARFY AEVV
Sbjct: 95 VLDQLEHPGIIKLYFTFQDTSSLYMALESCEGGELFDQITRKGRLSEDEARFYTAEVVDA 154
Query: 531 LEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQP 590
LEY+H +G+I+RD+KPENLLL DGHI + DF S KP + S+
Sbjct: 155 LEYIHSMGLIHRDIKPENLLLTSDGHIKIADFG-----SVKPM----------QDSQITV 199
Query: 591 PPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNR 650
P S+ ++ +FVGT Y+ PE++ + T D W LG LY+ML G +PF+ +
Sbjct: 200 LPNAASD--DKACTFVGTAAYVPPEVLNSSPATFGNDLWALGCTLYQMLSGTSPFKDASE 257
Query: 651 QKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT-GSNEIKQHPFFRGIN 709
F I+ +D+ FP+ S AAR LI+ LL +P+ R G+ + G +K+HPFF G++
Sbjct: 258 WLIFQRIIARDIKFPNHF--SEAARDLIDRLLDTEPSRRPGAGSEGYVALKRHPFFNGVD 315
Query: 710 WPLIRNMSPPPL 721
W +R+ +PP L
Sbjct: 316 WKNLRSQTPPKL 327
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.136 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,640,616
Number of Sequences: 26719
Number of extensions: 788313
Number of successful extensions: 3986
Number of sequences better than 10.0: 906
Number of HSP's better than 10.0 without gapping: 637
Number of HSP's successfully gapped in prelim test: 269
Number of HSP's that attempted gapping in prelim test: 2264
Number of HSP's gapped (non-prelim): 1448
length of query: 753
length of database: 11,318,596
effective HSP length: 107
effective length of query: 646
effective length of database: 8,459,663
effective search space: 5464942298
effective search space used: 5464942298
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC145470.6


