

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.4 + phase: 0
(457 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
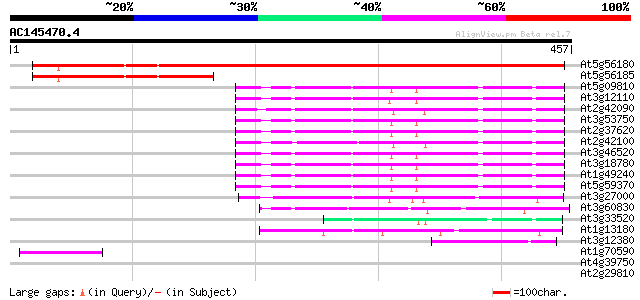
Score E
Sequences producing significant alignments: (bits) Value
At5g56180 putative protein 572 e-163
At5g56185 unknown protein 136 2e-32
At5g09810 ACTIN 2/7 (sp|P53492) 126 2e-29
At3g12110 unknown protein 125 6e-29
At2g42090 putative actin 124 8e-29
At3g53750 actin (ACT3) 122 3e-28
At2g37620 actin 3 122 3e-28
At2g42100 putative actin 120 2e-27
At3g46520 actin 12 117 1e-26
At3g18780 actin 2 116 2e-26
At1g49240 actin 11 like protein 116 2e-26
At5g59370 actin 4 115 4e-26
At3g27000 actin related protein, putative 88 1e-17
At3g60830 actin - like protein 80 2e-15
At3g33520 actin like protein 63 3e-10
At1g13180 actin-like protein 59 4e-09
At3g12380 actin like protein 43 4e-04
At1g70590 unknown protein 42 7e-04
At4g39750 putative protein 38 0.010
At2g29810 hypothetical protein 38 0.013
>At5g56180 putative protein
Length = 471
Score = 572 bits (1473), Expect = e-163
Identities = 290/441 (65%), Positives = 346/441 (77%), Gaps = 9/441 (2%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFLEFGNVETPIYTRLRHFFATVYGRMKV 192
Q+P +IIIDGGSGYCKFGWSK P GR ATFLEFGN+E+PIY RL+ FFAT++ RM+V
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFLEFGNIESPIYARLQQFFATIFTRMQV 189
Query: 193 KPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQATLALYAANQ 252
KPS Q +VVSLPLCH+DDTESAKASR+QLK AI LFDMNVPAVCA+NQA LALYAA +
Sbjct: 190 KPSMQPIVVSLPLCHFDDTESAKASRRQLKTAIFNVLFDMNVPAVCAVNQAVLALYAARR 249
Query: 253 TSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYFESLYTV 312
TSGI VNIGFQV +++PIL+GKVMR+VGVEV+G GALKLTGFL+EKMQ+NN+ F+SLYTV
Sbjct: 250 TSGIVVNIGFQVITILPILHGKVMRQVGVEVIGFGALKLTGFLKEKMQENNISFQSLYTV 309
Query: 313 RTLKEKLCYVAYDYEAELSKDTHASFE-AAEGKFTLSKERFQTGEILFQPRLAGVRAMSL 371
RTLKEKLCYVA DY+AELSKDT AS E + EG FTLSKERFQTGEILFQPRLAG+RAMSL
Sbjct: 310 RTLKEKLCYVALDYKAELSKDTQASVEVSGEGWFTLSKERFQTGEILFQPRLAGMRAMSL 369
Query: 372 HHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRV 431
H A++LCMDHC +A L G WFKTVVL+GGSACLPGL+ERLE+EL LP +SNGIRV
Sbjct: 370 HQAVSLCMDHCDAAGLTGDDSWFKTVVLTGGSACLPGLSERLERELQDHLPSSISNGIRV 429
Query: 432 IPPPHGADTAWFGAKLIGSVS 452
IPPP+G DT+W GAKLI ++S
Sbjct: 430 IPPPYGVDTSWHGAKLISNLS 450
>At5g56185 unknown protein
Length = 165
Score = 136 bits (343), Expect = 2e-32
Identities = 74/154 (48%), Positives = 95/154 (61%), Gaps = 8/154 (5%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFL 166
Q+P +IIIDGGSGYCKFGWSK P GR ATFL
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFL 163
>At5g09810 ACTIN 2/7 (sp|P53492)
Length = 377
Score = 126 bits (317), Expect = 2e-29
Identities = 89/278 (32%), Positives = 141/278 (50%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + NVPA+ QA
Sbjct: 91 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NVPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDSLMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL YVA DYE EL S ++E +G+ T+ ERF+
Sbjct: 200 YMFTTTAEREIVRDIKEKLAYVALDYEQELETAKSSSSVEKNYELPDGQVITIGAERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP L G+ A +H + C ++ D + +VLSGGS PG+A+R+
Sbjct: 260 PEVLFQPSLIGMEAPGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGSTMFPGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At3g12110 unknown protein
Length = 377
Score = 125 bits (313), Expect = 6e-29
Identities = 86/278 (30%), Positives = 141/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NTPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT +L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDYLMKILTERG 199
Query: 304 LYFES---LYTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL Y+A DYE E+ S S+E +G+ T+ ERF+
Sbjct: 200 YSFTTSAEREIVRDVKEKLAYIALDYEQEMETANTSSSVEKSYELPDGQVITIGGERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP L G+ A +H + C ++ D + +VLSGG+ PG+A+R+
Sbjct: 260 PEVLFQPSLVGMEAAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFPGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At2g42090 putative actin
Length = 366
Score = 124 bits (312), Expect = 8e-29
Identities = 80/277 (28%), Positives = 146/277 (51%), Gaps = 24/277 (8%)
Query: 185 TVYGRMKVKPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQAT 244
T Y ++V P V+++ + + KA+R+++ + + + +VPA+ Q+
Sbjct: 80 TFYNELRVDPEEHPVLLT------EAPYNPKANREKMTQIMFESF---DVPAMYVSMQSV 130
Query: 245 LALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNL 304
L LY++ +T+G+ +++G +V+ VP+ G + G+ + LG LT +L E M +
Sbjct: 131 LYLYSSGRTTGVVLDLGERVSHTVPVYEGYALPH-GILRLDLGGRDLTDYLIEIMTERGY 189
Query: 305 YFESLYT---VRTLKEKLCYVAYDYEAELSKDTHA-----SFEAAEG-KFTLSKERFQTG 355
+ + VR +KEKLCYVA DYE E+ K T ++ +G + T+ ERF
Sbjct: 190 TYTTSAEREIVRDIKEKLCYVALDYEQEMEKTTKGWTIDKTYVLPDGQEITIEAERFMCP 249
Query: 356 EILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEK 415
E+LFQP + G + +H A + C + D + ++++GG+ L G+ ER+ K
Sbjct: 250 EVLFQPSVIGKESSGIHEATRNSILKC---PVDTRRDMYGNILMTGGTTMLHGIKERMTK 306
Query: 416 ELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
EL++L+P M ++V+ PP + W G ++ S+S
Sbjct: 307 ELNALVPSSMK--VKVVVPPESECSVWIGGSILASLS 341
>At3g53750 actin (ACT3)
Length = 377
Score = 122 bits (307), Expect = 3e-28
Identities = 83/278 (29%), Positives = 140/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P +++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRVAPEEHPILLTEAPL-------NPKANREKMTQIMFETF---NAPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDALMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKLCY+A DYE EL S ++E +G+ T+ ERF+
Sbjct: 200 YSFTTTAEREIVRDIKEKLCYIALDYEQELETAKTSSSVEKNYELPDGQVITIGSERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+L+QP + G+ +H + C ++ D + +VLSGG+ PG+A+R+
Sbjct: 260 PEVLYQPSMIGMENAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFPGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At2g37620 actin 3
Length = 377
Score = 122 bits (307), Expect = 3e-28
Identities = 83/278 (29%), Positives = 140/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P +++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRVAPEEHPILLTEAPL-------NPKANREKMTQIMFETF---NAPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDALMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKLCY+A DYE EL S ++E +G+ T+ ERF+
Sbjct: 200 YSFTTTAEREIVRDIKEKLCYIALDYEQELETAKTSSSVEKNYELPDGQVITIGSERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+L+QP + G+ +H + C ++ D + +VLSGG+ PG+A+R+
Sbjct: 260 PEVLYQPSMIGMENAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFPGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At2g42100 putative actin
Length = 378
Score = 120 bits (301), Expect = 2e-27
Identities = 81/278 (29%), Positives = 144/278 (51%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++++P +++ PL + K +R+++ + + + P++ QA
Sbjct: 92 TFYNELRLEPEEHPILLTEAPL-------NPKVNREKMTQIMFESFA---FPSMYIGIQA 141
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LY++ +T+GI ++ G V+ VPI G + + + L LT +L + M +
Sbjct: 142 VLSLYSSGRTTGIVLDSGDGVSHTVPIYEGYALPHAILRL-DLAGRDLTEYLTKIMMERG 200
Query: 304 LYFESLYT---VRTLKEKLCYVAYDYEAELSKDTHAS-----FEAAEGK-FTLSKERFQT 354
+ + VR +KEKLCY+A DYE E+ K T +S +E +G+ T+ ERF+
Sbjct: 201 YTYTTSAEREIVRDIKEKLCYIAVDYEQEMEKATTSSAIDRTYELPDGQVITIGAERFRC 260
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQ L G+ +H + C ++ D + +VLSGG+ PG+A+R+
Sbjct: 261 PEVLFQTSLIGMETSGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFPGIADRMN 317
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+++L PP M I+V+ PP + W G ++ S+S
Sbjct: 318 KEINALAPPSMK--IKVVAPPERKYSVWVGGSILASLS 353
>At3g46520 actin 12
Length = 377
Score = 117 bits (293), Expect = 1e-26
Identities = 84/278 (30%), Positives = 137/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NTPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDHLMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL Y+A DYE EL S SFE +G+ T+ ERF+
Sbjct: 200 YSFTTTAEREIVRDMKEKLSYIALDYEQELETSKTSSSVEKSFELPDGQVITIGAERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP + G+ +H + C ++ D + +VLSGG+ G+ +R+
Sbjct: 260 PEVLFQPSMIGMENPGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFGGIGDRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At3g18780 actin 2
Length = 377
Score = 116 bits (291), Expect = 2e-26
Identities = 82/278 (29%), Positives = 139/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y +++ P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRIAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NSPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT +L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGFSLPH-AILRLDLAGRDLTDYLMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL +VA DYE E+ S ++E +G+ T+ ERF+
Sbjct: 200 YMFTTTAEREIVRDIKEKLSFVAVDYEQEMETSKTSSSIEKNYELPDGQVITIGAERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP G+ A +H + C ++ D + +VLSGG+ G+A+R+
Sbjct: 260 PEVLFQPSFVGMEAAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFSGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At1g49240 actin 11 like protein
Length = 377
Score = 116 bits (291), Expect = 2e-26
Identities = 82/278 (29%), Positives = 139/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y +++ P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRIAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NSPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT +L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGFSLPH-AILRLDLAGRDLTDYLMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL +VA DYE E+ S ++E +G+ T+ ERF+
Sbjct: 200 YMFTTTAEREIVRDIKEKLSFVAVDYEQEMETSKTSSSIEKNYELPDGQVITIGAERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP G+ A +H + C ++ D + +VLSGG+ G+A+R+
Sbjct: 260 PEVLFQPSFVGMEAAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFSGIADRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At5g59370 actin 4
Length = 377
Score = 115 bits (289), Expect = 4e-26
Identities = 83/278 (29%), Positives = 137/278 (48%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 91 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NTPAMYVAIQA 140
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 141 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDHLMKILTERG 199
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL Y+A D+E EL S SFE +G+ T+ ERF+
Sbjct: 200 YSFTTTAEREIVRDMKEKLSYIALDFEQELETSKTSSSVEKSFELPDGQVITIGAERFRC 259
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP + G+ +H + C ++ D + +VLSGG+ G+ +R+
Sbjct: 260 PEVLFQPSMIGMENPGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFGGIGDRMS 316
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 317 KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 352
>At3g27000 actin related protein, putative
Length = 389
Score = 87.8 bits (216), Expect = 1e-17
Identities = 71/284 (25%), Positives = 126/284 (44%), Gaps = 33/284 (11%)
Query: 187 YGRMKVKPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDM-NVPAVCALNQATL 245
Y +K+ PS ++++ P + + +E + +F+ N V QA L
Sbjct: 92 YNELKINPSDCKILLTDP----------PLNPSKNREKMIETMFEKYNFAGVFIQIQAVL 141
Query: 246 ALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLY 305
LYA +G+ ++ G VT VVP+++G + + + + +T +L + + +
Sbjct: 142 TLYAQGLLTGLVIDSGDGVTHVVPVVDGYSFPHL-TKRMNVAGRHITAYLVDLLSRRGYA 200
Query: 306 FES---LYTVRTLKEKLCYVAYDY--EAELSKDTH---ASFEAAEGK-FTLSKERFQTGE 356
TVR +KEKLCY++YDY E++L +T ++ +G+ + ERFQ E
Sbjct: 201 MNKTADFETVREIKEKLCYISYDYKRESQLGLETTILVKNYTLPDGRVIKVGTERFQAPE 260
Query: 357 ILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKE 416
LF P L V + + C+ ++ ++ +VLSGGS PGL RLEKE
Sbjct: 261 ALFTPELIDVEGDGMADMVFRCI---QEMDIDNRMMLYQHIVLSGGSTMYPGLPSRLEKE 317
Query: 417 LHSLLPPYMSNG---------IRVIPPPHGADTAWFGAKLIGSV 451
+ + G +R+ PP + G ++ +
Sbjct: 318 IQDRYLDTVLKGNKDGLKKLRLRIEDPPRRKHMVYLGGAVLAGI 361
>At3g60830 actin - like protein
Length = 363
Score = 80.1 bits (196), Expect = 2e-15
Identities = 70/261 (26%), Positives = 113/261 (42%), Gaps = 22/261 (8%)
Query: 204 PLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQ 263
PLC + KA R+QL + + NV A QA L+LYA + SG V+IG
Sbjct: 96 PLC------TPKAIREQLVQLMFETF---NVSGFYASEQAVLSLYAVGRISGCTVDIGHG 146
Query: 264 VTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYFE-SLYTVRTLKEKLCYV 322
+ P+L G V + + + LG +LT +++ + N S+ V LKE+
Sbjct: 147 KIDIAPVLEGAV-QHIASKRFELGGTELTKLFAQELGKTNPSMNLSMSDVEKLKEQYANC 205
Query: 323 AYDYEAELSKDTHASFE---AAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALC 378
A D E K + E +G+ ++ +ER+ GE LFQP + G+ H +
Sbjct: 206 AED-EIAYKKTQNCEIEQHTLPDGQVISIGRERYSVGEALFQPSILGLEE---HGIVEQL 261
Query: 379 MDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKELH---SLLPPYMSNGIRVIPPP 435
+ + + VL GG+ + G R +KE + S + P + +P
Sbjct: 262 VRIISTVSSENHRQLLENTVLCGGTTSMTGFESRFQKEANLCSSAIRPTLVKPPEYMPEN 321
Query: 436 HGADTAWFGAKLIGSVSYPLN 456
G +AW G ++ V +P N
Sbjct: 322 LGMYSAWVGGAILAKVVFPQN 342
>At3g33520 actin like protein
Length = 421
Score = 63.2 bits (152), Expect = 3e-10
Identities = 57/245 (23%), Positives = 95/245 (38%), Gaps = 56/245 (22%)
Query: 256 IAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNL-YFESLYTVRT 314
+ V+ GF T VP+L+ + ++ + LG T +L+E + ++ + + V
Sbjct: 155 LVVDCGFSFTHAVPVLHNFTLNHA-IKRIDLGGKAFTNYLKELVSYRSINVMDETFLVDD 213
Query: 315 LKEKLCYVAYDYEAELS--------------------------KDTHAS----------- 337
KEKLC+V+ D +L KD A+
Sbjct: 214 AKEKLCFVSLDLLRDLRLARNGNTLIKSTYVLPDGVTHTKGYVKDPQAAKRFLSLSEKES 273
Query: 338 ------------FEAAEGKFTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSA 385
+ + + L+ ERF E LFQP G+ L I ++ CHS
Sbjct: 274 VVVMDKVGERKKADMNKNEIDLTNERFLVPETLFQPADLGMNQAGLAECIVRAINSCHSY 333
Query: 386 ELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGA 445
+++++L+GGS P L ERLE EL L+P + +++ W G
Sbjct: 334 LQPV---LYQSIILTGGSTLFPQLKERLEGELRPLVPDHFD--VKITTQEDPILGVWRGG 388
Query: 446 KLIGS 450
L+ S
Sbjct: 389 SLLAS 393
>At1g13180 actin-like protein
Length = 427
Score = 59.3 bits (142), Expect = 4e-09
Identities = 69/283 (24%), Positives = 115/283 (40%), Gaps = 40/283 (14%)
Query: 204 PLCHYDDTESAKASRQQLKEAICAALFD-MNVPAVCALNQATLALYAANQTS-----GIA 257
P HY + + + +E LF+ NVP + + LAL A TS G+
Sbjct: 117 PEDHYFLLTESPLTPPESREYTGEILFETFNVPGLYIAVNSVLALAAGYTTSKCEMTGVV 176
Query: 258 VNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQN--NLYFESLYTV-RT 314
V++G T VVP+ G V+ ++ + + +T F+++ M++ N+ E + V R
Sbjct: 177 VDVGDGATHVVPVAEGYVIGSC-IKSIPIAGKDVTLFIQQLMRERGENIPPEDSFDVARK 235
Query: 315 LKEKLCYVAYDYEAELSK-DTHASFEAAEGKFTLSK-----------ERFQTGEILFQPR 362
+KE CY D E +K D + + K K ERF E+ F P
Sbjct: 236 VKEMYCYTCSDIVKEFNKHDKEPAKYIKQWKGVKPKTGAPYTCDVGYERFLGPEVFFNPE 295
Query: 363 LAGVRAMSLHHAIALCMDHC-HSAELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLL 421
+ + + +D C SA + +K +VLSGGS RL+++L ++
Sbjct: 296 ---IYSNDFTTTLPAVIDKCIQSAPIDTRRALYKNIVLSGGSTMFKDFGRRLQRDLKKIV 352
Query: 422 PP-YMSNGIR-------------VIPPPHGADTAWFGAKLIGS 450
++N R V+ P WFG ++ S
Sbjct: 353 DARVLANNARTGGEITSQPVEVNVVSHPVQRFAVWFGGSVLSS 395
>At3g12380 actin like protein
Length = 590
Score = 42.7 bits (99), Expect = 4e-04
Identities = 29/102 (28%), Positives = 45/102 (43%), Gaps = 2/102 (1%)
Query: 344 KFTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGS 403
K + ERF+ EILF P L G+ + L + E +++++GG
Sbjct: 454 KIVIGIERFRCPEILFHPNLIGIDQVGLDEMAGTSIRRLPHDEKELEERLTSSILMTGGC 513
Query: 404 ACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGA 445
+ LPG+ ERLE + + P + I V+ AW GA
Sbjct: 514 SLLPGMNERLECGIRMIRP--CGSPINVVRAMDPVLDAWRGA 553
Score = 38.5 bits (88), Expect = 0.008
Identities = 26/99 (26%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 247 LYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYF 306
L+ + GI + GF T +P ++G+ + K G +G +T +L++ + +
Sbjct: 29 LHGICKKDGIVLCPGFTTTHSIPFVDGEPIYK-GSSRTNIGGYHVTDYLKQLLSLKYPFH 87
Query: 307 ESLYT---VRTLKEKLCYVAYDYEAELSKDTHASFEAAE 342
S +T LK + CY+A DY +E+ EA E
Sbjct: 88 SSRFTWEKAEDLKLEHCYIAPDYASEIRLFQEGRKEAEE 126
>At1g70590 unknown protein
Length = 351
Score = 42.0 bits (97), Expect = 7e-04
Identities = 25/67 (37%), Positives = 32/67 (47%)
Query: 9 SASSSIKRSTNTTNSSSSSRSHQHHHHHAPSPLGLFDSLPPDILLKITRLLGPKHAAKLC 68
S SSI ++T + +SS S S + G F LP DIL+KI +
Sbjct: 30 SRISSINKATAKSTTSSRSSSSSSSSRPPSNEFGDFSMLPYDILMKIAAPFSHPNLQAAS 89
Query: 69 LVCKSWR 75
LVCKSWR
Sbjct: 90 LVCKSWR 96
>At4g39750 putative protein
Length = 912
Score = 38.1 bits (87), Expect = 0.010
Identities = 30/116 (25%), Positives = 50/116 (42%), Gaps = 7/116 (6%)
Query: 4 RKLCDS-----ASSSIKRSTNTTNSSSSSRSHQHHHHHAPSPLGLFDSLPPDILLKITRL 58
R++C+ A S + N + S + P+ F SLP +IL+
Sbjct: 513 RRVCEDKYICCALISFNKRKNGQVWAMDSEAEPPQEKKKPNSCPSFLSLPEEILVNCLAR 572
Query: 59 LGPKHAAKLCLVCKSWRSLVSDNELWAH--FLQTHQPIHFHSILFSETNLTSGYPL 112
+ + KL LVCKS+ SL+ EL+ +L TH+ + + + L S + L
Sbjct: 573 IPKSYYPKLSLVCKSFCSLILSMELYVERLYLGTHEDVLHVCLQLPDRRLPSWFSL 628
>At2g29810 hypothetical protein
Length = 383
Score = 37.7 bits (86), Expect = 0.013
Identities = 32/127 (25%), Positives = 57/127 (44%), Gaps = 11/127 (8%)
Query: 17 STNTTNSSSSSRSHQHHHHHAPSPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCKSWRS 76
S +N ++++ Q + P + LP ++++ I L+ H KL L+ K++R
Sbjct: 8 SDEGSNGGAANKKPQEEEENIPP---IPKELPEELIVIIVALVRRYHYPKLSLISKAYRD 64
Query: 77 LVSDNELW--AHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHREQ 134
L+S EL+ L +P+ + SI F +L S Y L Q FK +
Sbjct: 65 LISSPELFQTRSRLGFTEPVLYTSIGFPPFDLPSWYILHRISLQ------FKQITSLPSM 118
Query: 135 LPPAIII 141
LP + ++
Sbjct: 119 LPGSAVV 125
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,562,570
Number of Sequences: 26719
Number of extensions: 457391
Number of successful extensions: 2386
Number of sequences better than 10.0: 132
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 2098
Number of HSP's gapped (non-prelim): 194
length of query: 457
length of database: 11,318,596
effective HSP length: 103
effective length of query: 354
effective length of database: 8,566,539
effective search space: 3032554806
effective search space used: 3032554806
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC145470.4


