

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
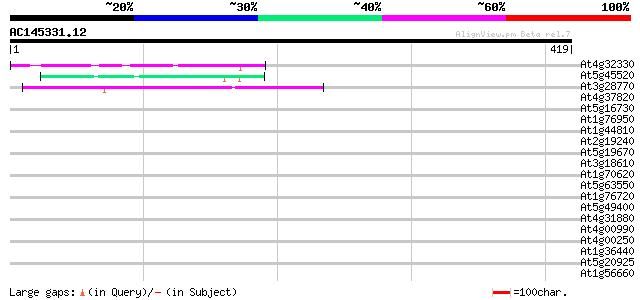
Query= AC145331.12 - phase: 0
(419 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g32330 unknown protein 56 3e-08
At5g45520 putative protein 43 3e-04
At3g28770 hypothetical protein 43 4e-04
At4g37820 unknown protein 41 0.001
At5g16730 putative protein 39 0.004
At1g76950 zinc finger protein (PRAF1) 39 0.004
At1g44810 unknown protein 39 0.004
At2g19240 hypothetical protein 39 0.005
At5g19670 putative protein 39 0.007
At3g18610 unknown protein 38 0.009
At1g70620 unknown protein 38 0.012
At5g63550 unknown protein 37 0.016
At1g76720 putative translation initiation factor IF-2 37 0.020
At5g49400 putative protein 36 0.035
At4g31880 unknown protein 36 0.035
At4g00990 unknown protein 36 0.035
At4g00250 unknown protein (At4g00250) 36 0.045
At1g36440 hypothetical protein 36 0.045
At5g20925 protein kinase (TOUSLED) 35 0.059
At1g56660 hypothetical protein 35 0.077
>At4g32330 unknown protein
Length = 436
Score = 56.2 bits (134), Expect = 3e-08
Identities = 54/207 (26%), Positives = 91/207 (43%), Gaps = 37/207 (17%)
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 60
MDP + + ADG D P+N G + + + V+ ET S+N N
Sbjct: 1 MDPESIMAADGTDSA--------PANGGLAMENVCVKENGAVSVETVDTTSESQNENSAN 52
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 120
S S + IE + +G+ + + V + +AQ+ P K + ++S
Sbjct: 53 S-----STLDTIEHVKEAAEGTQV-----EHVDDSKCMKGEKAQRKPRHEKLSGGKNNSS 102
Query: 121 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKK--------- 171
V+ +K SK GK A A SNG+ A + + P+K++S N R++Q++K+
Sbjct: 103 VH---IKKSKEGKSADAKVAASNGSVAPNVQTTNPLKSKSFNGREAQVTKQGKHDSAPAE 159
Query: 172 -------KPKSLKKGPLDKVQGEGESS 191
KPKS KK + + + +SS
Sbjct: 160 SADGEKVKPKSQKKQAHETSEDDTQSS 186
>At5g45520 putative protein
Length = 1167
Score = 43.1 bits (100), Expect = 3e-04
Identities = 46/174 (26%), Positives = 70/174 (39%), Gaps = 12/174 (6%)
Query: 24 PSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSN 83
PS DA+ N D H + + DG E +N E + +KE DNV G +
Sbjct: 818 PSRESTDAIENKPDDHQRGDKQEEKGDGEKEKVNLEEWKK--HDEIKEESSKQDNVTGGD 875
Query: 84 LTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSN 143
+ S KE K T +S+ +K + G A+ + K D +A + S+
Sbjct: 876 VKKSPPKESK---DTMESKRDDQKENSKVQEKGDVDKGKAADLDEGKKENDVKAESSKSD 932
Query: 144 GTSALDSRPRQPIKNR----SSNDRQSQLSK--KKPKSLKKG-PLDKVQGEGES 190
D P K++ S D ++SK +K K G L K+ G GE+
Sbjct: 933 KVIEGDEEKNPPQKSKDIIQSKPDDHREISKVQEKVDGEKNGDDLKKLDGGGEA 986
>At3g28770 hypothetical protein
Length = 2081
Score = 42.7 bits (99), Expect = 4e-04
Identities = 39/233 (16%), Positives = 99/233 (41%), Gaps = 10/233 (4%)
Query: 10 DGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAM 69
+ ++ ++ V + + G+ A+ +L+ V +E + ++ + A GN+ +
Sbjct: 267 ENVESNNEKEVEGQGESIGDSAIEKNLESKEDVKSEVEAAKNDGSSMTENLGEAQGNNGV 326
Query: 70 ------KEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNA 123
KE+EG ++++ S++ + E + +K E ++ + K + ++GV+
Sbjct: 327 STIDNEKEVEGQGESIEDSDIEKNLESKEDVKSEVEAAKNAGSSMTGKLEEAQRNNGVST 386
Query: 124 SLVKNSK-IGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLD 182
+ NS+ G + + + N T+ + ++ + N+ +S K + K G +
Sbjct: 387 NETMNSENKGSGESTNDKMVNATTNDEDHKKENKEETHENNGES--VKGENLENKAGNEE 444
Query: 183 KVQGEG-ESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLK 234
++GE E+ + L+ S + + +R + + N + K K
Sbjct: 445 SMKGENLENKVGNEELKGNASVEAKTNNESSKEEKREESQRSNEVYMNKETTK 497
Score = 37.0 bits (84), Expect = 0.020
Identities = 38/177 (21%), Positives = 74/177 (41%), Gaps = 7/177 (3%)
Query: 57 NQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVG 116
N+ E+T + NS +KE N S + SK +E K + ++S+ ++ K K
Sbjct: 977 NKKETTKSENSKLKEENKDNKEKKESEDSASKNREKK-EYEEKKSKTKEEAKKEKKKSQD 1035
Query: 117 SSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSL 176
S + SK K+K+ S + +++ ++ +N S ++ + + KS+
Sbjct: 1036 KKREEKDSEERKSK--KEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEHEDNKSM 1093
Query: 177 KKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRL 233
KK K + + E S RK E + K Q ++ K ++ + +L
Sbjct: 1094 KKEEDKKEKKKHEES----KSRKKEEDKKDMEKLEDQNSNKKKEDKNEKKKSQHVKL 1146
Score = 34.3 bits (77), Expect = 0.13
Identities = 29/145 (20%), Positives = 65/145 (44%), Gaps = 10/145 (6%)
Query: 4 VNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDL-----DPHVTVNTETFVPDGNSENINQ 58
+N + D L D +NG +E +N ED+V +++ + + N+ + P+G+ + N+
Sbjct: 1754 INGVKDDSLKDDSKNGDINEINNGKEDSVKDNVTEIQGNDNSLTNSTSSEPNGDKLDTNK 1813
Query: 59 LESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
+ N+ M+ GSN D +N + KE + ++ + + + N + +
Sbjct: 1814 ---DSMKNNTMEAQGGSNG--DSTNGETEETKESNVSMNNQNMQDVGSNENSMNNQTTGT 1868
Query: 119 SGVNASLVKNSKIGKDKQASPAVSN 143
S +++ K+ + +SN
Sbjct: 1869 GDDIISTTTDTESNTSKEVTSFISN 1893
Score = 33.9 bits (76), Expect = 0.17
Identities = 45/230 (19%), Positives = 82/230 (35%), Gaps = 28/230 (12%)
Query: 26 NSGEDAVSNDL----DPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
N+GE + +L D + E+ N+E + + T I + G
Sbjct: 592 NNGESVKNENLENKEDKKELKDDESVGAKTNNETSLEEKREQTQKGHDNSINSKIVDNKG 651
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
N +KEKEV + ST + + +V + G + + + KD +
Sbjct: 652 GNADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSEKGEEGKENNKDSMEDKKL 711
Query: 142 SNGTSALDSRPRQPIKNR---------SSNDRQSQLSKKKPKSLKKGPLDK--------- 183
N S DS+ + + ++ S D +S +K K K K+ K
Sbjct: 712 ENKESQTDSKDDKSVDDKQEEAQIYGGESKDDKSVEAKGKKKESKENKKTKTNENRVRNK 771
Query: 184 ---VQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKK 230
VQG + S + KGE + K + ++++ +N K+
Sbjct: 772 EENVQGNKKES---EKVEKGEKKESKDAKSVETKDNKKLSSTENRDEAKE 818
Score = 32.3 bits (72), Expect = 0.50
Identities = 33/179 (18%), Positives = 69/179 (38%), Gaps = 16/179 (8%)
Query: 60 ESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQK-------------- 105
E + N K+ E ++ D N ++ KE++ K K E+S+++K
Sbjct: 1069 EKKESENHKSKKKEDKKEHED--NKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQ 1126
Query: 106 GPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQ 165
K K K + LVK K+K+ + S S+ ++ ++
Sbjct: 1127 NSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKKSS 1186
Query: 166 SQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQN 224
KKK K +K+ K++ E S+ + + +++ + + ++ KQ+
Sbjct: 1187 KDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDKKNTTKQS 1245
Score = 31.2 bits (69), Expect = 1.1
Identities = 37/193 (19%), Positives = 75/193 (38%), Gaps = 19/193 (9%)
Query: 17 QNGVHDEPSNSGEDAVSN----DLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEI 72
+N +E S GE+ + +L + +V +T E + + + +
Sbjct: 437 ENKAGNEESMKGENLENKVGNEELKGNASVEAKTNNESSKEEKREESQRSNEVYMNKETT 496
Query: 73 EGSNDNVDGSNL-------TVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASL 125
+G N N+ G ++ ++ +++VK KV +S +++ A+V ++GV+
Sbjct: 497 KGENVNIQGESIGDSTKDNSLENKEDVKPKVDANESDGNSTKERHQEAQV--NNGVSTED 554
Query: 126 VKNSKIG------KDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKG 179
IG DK ++G + R N S ++ +K+ K LK
Sbjct: 555 KNLDNIGADEQKKNDKSVEVTTNDGDHTKEKREETQGNNGESVKNENLENKEDKKELKDD 614
Query: 180 PLDKVQGEGESSL 192
+ E+SL
Sbjct: 615 ESVGAKTNNETSL 627
>At4g37820 unknown protein
Length = 532
Score = 41.2 bits (95), Expect = 0.001
Identities = 45/216 (20%), Positives = 85/216 (38%), Gaps = 24/216 (11%)
Query: 20 VHDEPSNSGEDAVSNDLDPHVTVNT-ETFVPDGNSENINQLESTATGNSAMKEIEGSNDN 78
VH++ S + + + V++NT E DG + + TG + SN
Sbjct: 202 VHEDESGPKNEVLEGSVIKEVSLNTTENGSDDGEQQETKSELDSKTGEKGFSD---SNGE 258
Query: 79 VDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
+ +NL+ S E +TE S + +++ G S+G + KN + K+K S
Sbjct: 259 LPETNLSTSNATE-----TTESSGS------DESGSSGKSTGYQQT--KNEEDEKEKVQS 305
Query: 139 PAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLR 198
S +S+ ++ KN SK++ KK QGEG+ +
Sbjct: 306 -------SEEESKVKESGKNEKDASSSQDESKEEKPERKKKEESSSQGEGKEEEPEKREK 358
Query: 199 KGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLK 234
+ SS ++ + + + + Q +K+ +K
Sbjct: 359 EDSSSQEESKEEEPENKEKEASSSQEENEIKETEIK 394
Score = 32.3 bits (72), Expect = 0.50
Identities = 28/140 (20%), Positives = 59/140 (42%), Gaps = 3/140 (2%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTA--TGNSAMKEIEGSNDNV 79
+EP N ++A S+ + + + +S+ N+ + T + S KE S +
Sbjct: 370 EEPENKEKEASSSQEENEIKETEIKEKEESSSQEGNENKETEKKSSESQRKENTNSEKKI 429
Query: 80 DGSNLTVSKEKEVKIKVSTEQSRAQKG-PVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
+ T S + + T++S+ + G NK + SS + +N++ G+ ++
Sbjct: 430 EQVESTDSSNTQKGDEQKTDESKRESGNDTSNKETEDDSSKTESEKKEENNRNGETEETQ 489
Query: 139 PAVSNGTSALDSRPRQPIKN 158
SAL+ Q +K+
Sbjct: 490 NEQEQTKSALEISHTQDVKD 509
Score = 30.0 bits (66), Expect = 2.5
Identities = 36/188 (19%), Positives = 71/188 (37%), Gaps = 3/188 (1%)
Query: 25 SNSGEDAVSNDLDPHVTVNTET-FVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSN 83
SN+ E S+ D + T + N E+ + ++ S +KE G N+ S+
Sbjct: 267 SNATETTESSGSDESGSSGKSTGYQQTKNEEDEKEKVQSSEEESKVKE-SGKNEKDASSS 325
Query: 84 LTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPA-VS 142
SKE++ + K E S +G + + S + K+K+AS +
Sbjct: 326 QDESKEEKPERKKKEESSSQGEGKEEEPEKREKEDSSSQEESKEEEPENKEKEASSSQEE 385
Query: 143 NGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGES 202
N + + ++ ++ N+ + K K+ + + E S + +KG+
Sbjct: 386 NEIKETEIKEKEESSSQEGNENKETEKKSSESQRKENTNSEKKIEQVESTDSSNTQKGDE 445
Query: 203 SSLRLRKR 210
KR
Sbjct: 446 QKTDESKR 453
>At5g16730 putative protein
Length = 853
Score = 39.3 bits (90), Expect = 0.004
Identities = 37/140 (26%), Positives = 68/140 (48%), Gaps = 13/140 (9%)
Query: 54 ENINQLESTATGN--SAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNK 111
E N+LE +A+ + S MK++EGSND + + ++ KE + + T ++ QK ++
Sbjct: 334 EEANKLERSASVSLESVMKQLEGSNDKLHDTETEITDLKERIVTLETTVAK-QKEDLEVS 392
Query: 112 NAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKK 171
++GS V + KN +K+ S + + + R K + + R +LS++
Sbjct: 393 EQRLGS---VEEEVSKN-----EKEVEKLKSELETVKEEKNRALKKEQDATSRVQRLSEE 444
Query: 172 KPKSLKKGPLDKVQGEGESS 191
K K L L+ + E E S
Sbjct: 445 KSKLL--SDLESSKEEEEKS 462
>At1g76950 zinc finger protein (PRAF1)
Length = 1103
Score = 39.3 bits (90), Expect = 0.004
Identities = 30/117 (25%), Positives = 55/117 (46%), Gaps = 6/117 (5%)
Query: 56 INQLESTATGNSAMKEIEGSN-DNVDGSNLTVSKEKEVKIKVSTE-QSRAQKGPVKNKNA 113
++++ T NSA+ + G N D +D S + ++K + + + S+A K K
Sbjct: 696 VSEINDTNRRNSAVPRLSGENRDRLDKSEIRLAKFGTSNMDLIKQLDSKAAKQGKKTDTF 755
Query: 114 KVGSSSGVNASL----VKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQS 166
+G +S + + L S IG ++A+P ++ S + SR P RSS R +
Sbjct: 756 SLGRNSQLPSLLQLKDAVQSNIGDMRRATPKLAQAPSGISSRSVSPFSRRSSPPRSA 812
>At1g44810 unknown protein
Length = 296
Score = 39.3 bits (90), Expect = 0.004
Identities = 37/148 (25%), Positives = 65/148 (43%), Gaps = 8/148 (5%)
Query: 52 NSENINQLESTATGNSAM---KEIEGSNDNVDGSNLTVSKEKEVKIK---VSTEQSRAQK 105
N + +N LE T +S+ +EI D + + + S E+E ++K T S + +
Sbjct: 2 NKKLLNPLEDPPTASSSEDVDEEISSGEDEKEHISNSSSSEEENELKDLSTQTLNSPSTE 61
Query: 106 GPVKNKNAKVGSSSGVNASLV-KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDR 164
P + ++ S S L + K G D + S GTS+ D + + + N +
Sbjct: 62 APTLDSGSETNSDSDKPIVLTSQKKKEGTDSSGTKRASEGTSSKDIKRAKKVSGDDDNKK 121
Query: 165 -QSQLSKKKPKSLKKGPLDKVQGEGESS 191
QS +K+ SL +G +D G S+
Sbjct: 122 FQSLWTKEDEISLLQGMIDFKAETGTSA 149
>At2g19240 hypothetical protein
Length = 840
Score = 38.9 bits (89), Expect = 0.005
Identities = 40/153 (26%), Positives = 62/153 (40%), Gaps = 32/153 (20%)
Query: 16 HQNGVHDEPSNSG-------EDAVSN-DLDPHVTVNTETFVPDGNSENINQL-----EST 62
H+ G+ + +NS ED+ ++D HVTVN E P ENI + ES
Sbjct: 524 HKLGIGERKANSSPVTQSLLEDSSEQLNVDCHVTVNKENIHPQETEENIMEFHSADEESI 583
Query: 63 ATGNSAMKE-----------------IEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQK 105
+G+S +E IE + + GSNL ++ + + +S E S
Sbjct: 584 VSGSSPSEESSFVSLDPTSPVRCSTKIENDSVSSAGSNLLPDEDDKSIVSISEENSSVSS 643
Query: 106 GPVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
P+ A + SG S+ KD+Q S
Sbjct: 644 DPI--SPAIDSNYSGKYLDCCTGSENDKDQQTS 674
>At5g19670 putative protein
Length = 600
Score = 38.5 bits (88), Expect = 0.007
Identities = 42/199 (21%), Positives = 89/199 (44%), Gaps = 22/199 (11%)
Query: 5 NSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTAT 64
N+L + +D +G+H N VS + + + E FV + + E+ ++ +
Sbjct: 66 NTLAVNVSEDSAVSGIHVLEKNG---YVSGFGLRNESEDDEGFVGNVDFESFEDVKDSII 122
Query: 65 GNSAMKEIEGSNDNVDGSNLTVSKEKEVKI--------KVSTEQSRAQKGPVKNKNAKVG 116
+KE+ GS+DN+ S TV +++ V V+ + + K + + + +
Sbjct: 123 ----IKEVAGSSDNLFPSETTVMQKESVSTSNNGYQVQNVTVQSQKNVKSSILSGGSSIA 178
Query: 117 SSSGVNASLVKNSKIGKDKQ-ASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKS 175
S + N+SL+ + K+ K K+ + +D R ++R ++ L+ + K
Sbjct: 179 SPASGNSSLLVSKKVSKKKKMRCDLPPKSVTTIDEMNRILARHRRTSRAMEILTAR--KE 236
Query: 176 LKKGPLDKVQGEGESSLYP 194
++ P+ K++ E LYP
Sbjct: 237 IENAPVAKLERE----LYP 251
>At3g18610 unknown protein
Length = 636
Score = 38.1 bits (87), Expect = 0.009
Identities = 46/183 (25%), Positives = 71/183 (38%), Gaps = 28/183 (15%)
Query: 23 EPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGS 82
+P G+ DLD VT + E I+ ++ + K++E S+ + S
Sbjct: 20 KPLKKGKRDAEEDLDMQVTKKQK-------KELIDVVQKEKAEKTVPKKVESSSSDASDS 72
Query: 83 NLTVSKEKEVKIKVSTEQSR--------------AQKGPVKNKNAKVGSSSGVNASLVKN 128
+ K KE K+ E S A+K P K AKV SSS + S
Sbjct: 73 D-EEEKTKETPSKLKDESSSEEEDDSSSDEEIAPAKKRPEPIKKAKVESSSSDDDSTSDE 131
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEG 188
KQ PAV S + SS+D ++ KK+P L+K ++ +
Sbjct: 132 ETAPVKKQ--PAVLEKAKVESSSS----DDDSSSDEETVPVKKQPAVLEKAKIESSSSDD 185
Query: 189 ESS 191
+SS
Sbjct: 186 DSS 188
Score = 32.3 bits (72), Expect = 0.50
Identities = 29/140 (20%), Positives = 66/140 (46%), Gaps = 16/140 (11%)
Query: 52 NSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNK 111
+ +++ ++E+ A+ +K+ G D + ++ V+K+++ ++ ++ +A+K K
Sbjct: 5 SKKSVTEVETPASMTKPLKK--GKRDAEEDLDMQVTKKQKKELIDVVQKEKAEKTVPK-- 60
Query: 112 NAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKK 171
KV SSS +AS D S L ++ SS+D + +KK
Sbjct: 61 --KVESSSS-DAS---------DSDEEEKTKETPSKLKDESSSEEEDDSSSDEEIAPAKK 108
Query: 172 KPKSLKKGPLDKVQGEGESS 191
+P+ +KK ++ + +S+
Sbjct: 109 RPEPIKKAKVESSSSDDDST 128
Score = 29.6 bits (65), Expect = 3.2
Identities = 48/224 (21%), Positives = 88/224 (38%), Gaps = 40/224 (17%)
Query: 22 DEPSNSGEDAVSNDLD---------PHVTVNTETFVPDGNSENINQLESTATGNSAMKE- 71
DE S+ ED S+D + P E+ D +S + + + +++
Sbjct: 87 DESSSEEEDDSSSDEEIAPAKKRPEPIKKAKVESSSSDDDSTSDEETAPVKKQPAVLEKA 146
Query: 72 -IEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSK 130
+E S+ + D S S E+ V +K K P + AK+ SSS + S
Sbjct: 147 KVESSSSDDDSS----SDEETVPVK---------KQPAVLEKAKIESSSSDDDSSSDEET 193
Query: 131 IGKDKQAS---PAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGE 187
+ KQ + A + +S+ D SS+D + +KK+P +KK D+ +
Sbjct: 194 VPMKKQTAVLEKAKAESSSSDDG---------SSSDEEPTPAKKEPIVVKKDSSDESSSD 244
Query: 188 GESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKP 231
E+ + ++K ++ ++ K + P KKP
Sbjct: 245 EETPV----VKKKPTTVVKDAKAESSSSEEESSSDDEPTPAKKP 284
>At1g70620 unknown protein
Length = 897
Score = 37.7 bits (86), Expect = 0.012
Identities = 52/232 (22%), Positives = 93/232 (39%), Gaps = 21/232 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVT---VNTETFV-PDGNSENINQL-ESTA 63
+ GLDD + P + D + +PHV V ++ + D N L + +
Sbjct: 631 SSGLDDDTSGSRKEHPDRTDSDKDAILDEPHVKNSGVKSDCNLRQDSNKPYGKDLSDEVS 690
Query: 64 TGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVS-----TEQSRAQKGP-VKNKNAKVGS 117
T S + E +G + D N + + KE +K + E ++ P VK + V
Sbjct: 691 TDRSRIVETKGGKEKGDSQNDSKDRMKENDLKSAEKVKGVESNKKSTDPHVKKDSRDVER 750
Query: 118 SSGVNASLVKNSKIGKDKQASPAVSNGT--SALDSRPRQPIKNRSSNDRQSQLSKKKPKS 175
N+ + + K+K+ + S+ D R R P N SS+D + + S+ + +S
Sbjct: 751 PHRTNSKEDRGKRKEKEKEEERSRHRRAENSSKDKRRRSPTSNESSDDSKRK-SRSRRRS 809
Query: 176 LKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRR 227
+ P+ + S ESS RK +R+ R + ++ RR
Sbjct: 810 VSPSPVRSRRKRSSPS-------SDESSDDSKRKSSSKRKNRSPSPGKSRRR 854
Score = 28.9 bits (63), Expect = 5.5
Identities = 40/168 (23%), Positives = 62/168 (36%), Gaps = 13/168 (7%)
Query: 46 TFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGS---NLTVSKEKEVKIKVSTEQSR 102
T G+ ++ L S A+ + S+ N D + +L V V + STE+
Sbjct: 517 TKASSGSPADVLGLASYASDDDDADTDAASDANADENGVESLGVGSRHNVSQQPSTEKL- 575
Query: 103 AQKGPVKNKNAKVGSSSGVNASLVKNSKIG-KDKQASPAVSNGTSALDSRPRQPIKNRSS 161
P +AK+ + GVNA+ KNSK G +D P + S + S
Sbjct: 576 --PDPEAMASAKLDPAVGVNANSGKNSKSGLEDYSQMPGSTRKDDEAGSTKISDVSASSG 633
Query: 162 NDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRK 209
D + S+K+ D + E P G S LR+
Sbjct: 634 LDDDTSGSRKEHPDRTDSDKDAILDE------PHVKNSGVKSDCNLRQ 675
>At5g63550 unknown protein
Length = 530
Score = 37.4 bits (85), Expect = 0.016
Identities = 34/135 (25%), Positives = 58/135 (42%), Gaps = 14/135 (10%)
Query: 50 DGNSENIN---QLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKG 106
D +SE N + + A + E ++D D + +K+ K + E+S KG
Sbjct: 296 DADSEGTNDPHEEDDAAPEEESDHEKTDTDDEKDEVEVEKPSKKKSSSKKTVEESSGSKG 355
Query: 107 PVKNKNAKVGSSSGVNAS--LVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDR 164
K +AK + SG +S + K++ KQ V + +P +P
Sbjct: 356 KDKQPSAKGSARSGEKSSKQIAKSTSSPAKKQKVDHVESSKEKSKKQPSKP--------- 406
Query: 165 QSQLSKKKPKSLKKG 179
Q++ SK+K K+ KKG
Sbjct: 407 QAKGSKEKGKATKKG 421
>At1g76720 putative translation initiation factor IF-2
Length = 1224
Score = 37.0 bits (84), Expect = 0.020
Identities = 48/226 (21%), Positives = 91/226 (40%), Gaps = 31/226 (13%)
Query: 28 GEDAVSNDLDPHVTVNT------ETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
G+D+V+++ T +T ET +N N++ T + ++
Sbjct: 230 GDDSVADETSQAKTPDTKSVEVIETGKIKKKKKNKNKVARTLEEEDDLDKLLAELGETPA 289
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPV---KNKNAKVGSSSGVNASLVKNSKIGKDK-QA 137
+ S EV E+ +AQ GPV +N K G V + K K K+K +
Sbjct: 290 AERPASSTPEV------EKVQAQPGPVAPVENAGEKEGEKETVETAAAKKKKKKKEKDKE 343
Query: 138 SPAVSNGTSALDSRPR-------QPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGES 190
A + TS+++++ +P++ + + + KK PK +++ E +
Sbjct: 344 KKAAAAATSSVEAKEEKQEESVTEPLQPKKKDAKGKAAEKKIPKHVRE------MQEALA 397
Query: 191 SLYPFSLRKGESSSLRLRKRFRQRRRRR--VACKQNPRRVKKPRLK 234
RK + +LRK +RRR+ A + +R +K + K
Sbjct: 398 RRQEAEERKKKEEEEKLRKEEEERRRQEELEAQAEEAKRKRKEKEK 443
>At5g49400 putative protein
Length = 280
Score = 36.2 bits (82), Expect = 0.035
Identities = 39/156 (25%), Positives = 68/156 (43%), Gaps = 22/156 (14%)
Query: 75 SNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNK---NAKVGSSSGVNASLVKNSKI 131
S D++DGS+ ++E + + ++ +K K+K +K S S AS+ +
Sbjct: 144 SVDDLDGSD---DDDEEERPDATNGKAEVEKRSKKSKRKHRSKSDSESDSEASVFETDSD 200
Query: 132 GKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESS 191
G ++S S+ + + D R R+ R+++ SKKK K K+ ESS
Sbjct: 201 GSSGESSSEYSSSSDSEDERRRR---------RKAKKSKKKQKQRKERRRRYSSSSSESS 251
Query: 192 LYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRR 227
+ ES+S R RR+++ N RR
Sbjct: 252 -------ESESASDSDSDEDRSRRKKKSKRHSNKRR 280
>At4g31880 unknown protein
Length = 873
Score = 36.2 bits (82), Expect = 0.035
Identities = 38/167 (22%), Positives = 65/167 (38%), Gaps = 24/167 (14%)
Query: 25 SNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSNL 84
++ + S D E + SE ++ S+ + + S+ + G +
Sbjct: 725 TSQDDKTASKSKDSKEASREEEASSEEESEEEEPPKTVGKSGSSRSKKDISSVSKSGKSK 784
Query: 85 TVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNG 144
SK+KE K +T S+++ GPVK+ AK SK GK K S + S
Sbjct: 785 ASSKKKEEPSKATTS-SKSKSGPVKSVPAK--------------SKTGKGKAKSGSASTP 829
Query: 145 TSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESS 191
S S+++ +S+ + K+P+ K K QG S
Sbjct: 830 ASKA---------KESASESESEETPKEPEPATKAKSGKSQGSQSKS 867
>At4g00990 unknown protein
Length = 454
Score = 36.2 bits (82), Expect = 0.035
Identities = 25/91 (27%), Positives = 44/91 (47%), Gaps = 4/91 (4%)
Query: 94 IKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPR 153
+ V T ++ + PVK +N KV A L K G+ K+AS + +D +
Sbjct: 167 VNVLTHTAKVEIPPVKYQNIKVHQKKYAEAMLQKQQYSGQVKEASELENKSMKEVD-ESK 225
Query: 154 QPIKNRSSNDRQSQLSKKKPKSLKKGPLDKV 184
+ +K++++N+ QS S + S G +KV
Sbjct: 226 KDLKDKAANEEQSNNSSRPSGS---GEAEKV 253
>At4g00250 unknown protein (At4g00250)
Length = 319
Score = 35.8 bits (81), Expect = 0.045
Identities = 38/136 (27%), Positives = 62/136 (44%), Gaps = 14/136 (10%)
Query: 59 LESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
LES AT +S+ E+E S++ + S+ S+E + K V+ S+ K P N+K SS
Sbjct: 4 LESPATASSS--EVESSSEEIFKSS---SEESKPKDPVTVPSSKTLKSPSAAVNSKTDSS 58
Query: 119 --SGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSL 176
S + ++ K + SPAV +G ++ + +S D + KK+ +
Sbjct: 59 DDSEKQSFVLTRRKKKEGAAESPAVKSG-------KKRAGEGSTSRDMHVKRVKKEDDNK 111
Query: 177 KKGPLDKVQGEGESSL 192
K P E E SL
Sbjct: 112 KANPQRVWSEEDEISL 127
>At1g36440 hypothetical protein
Length = 409
Score = 35.8 bits (81), Expect = 0.045
Identities = 38/166 (22%), Positives = 67/166 (39%), Gaps = 15/166 (9%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAM---KEIEGSNDN 78
+ + G DA + V DG+ + GN+ M KE+ +
Sbjct: 222 ENEAEEGSDANGGEKGSEVGEKETKIEKDGSEDE----SDVGGGNNEMEMDKELAEDESD 277
Query: 79 VDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGV--NASLVKNSKIGKDKQ 136
V G N + +KE+ + ++ K V+NK+ GS+ + + ++++ K+ K
Sbjct: 278 VGGGNKEMEMDKEIDVGGGNKEMEKDKEMVQNKSDVGGSNREMEKDKRVLQDDKMDGGKA 337
Query: 137 ASPAVSNGTSALD----SRPRQPIKNRSSNDRQSQLSKKKPKSLKK 178
A P+ S D S R+ K+ D S+ KK K +KK
Sbjct: 338 AEPSKKREMSHDDGDDPSNKRR--KSHDDGDDPSEGGVKKLKVVKK 381
>At5g20925 protein kinase (TOUSLED)
Length = 688
Score = 35.4 bits (80), Expect = 0.059
Identities = 29/138 (21%), Positives = 60/138 (43%), Gaps = 8/138 (5%)
Query: 81 GSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPA 140
G T ++ I VS + + + V+N++ ++GS + + ++ +D Q
Sbjct: 247 GKQKTTKVISDLLISVSKTERQEARTKVRNESLRLGSVGVLRTGTII-AETWEDGQM--- 302
Query: 141 VSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKG 200
L+++ RQ ++ + + +RQ +L KK+ K D G E + P + K
Sbjct: 303 ----LKDLNAQLRQLLETKEAIERQRKLLKKRQNGDKNDGTDTESGAQEEDIIPDEVYKS 358
Query: 201 ESSSLRLRKRFRQRRRRR 218
+S++ + R R R
Sbjct: 359 RLTSIKREEEAVLRERER 376
>At1g56660 hypothetical protein
Length = 522
Score = 35.0 bits (79), Expect = 0.077
Identities = 44/205 (21%), Positives = 73/205 (35%), Gaps = 27/205 (13%)
Query: 51 GNSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKE--------KEVKIKVSTEQSR 102
G S+ + + +S +KE + + DG ++ E KE +KV +
Sbjct: 55 GKSKKDKEKKKGKNVDSEVKEDKDDDKKKDGKMVSKKHEEGHGDLEVKESDVKVEEHEKE 114
Query: 103 AQKGPVKN-------------KNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALD 149
+KG K KN K SG KN K K+K+ VS L+
Sbjct: 115 HKKGKEKKHEELEEEKEGKKKKNKKEKDESGPEE---KNKKADKEKKHED-VSQEKEELE 170
Query: 150 SRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRK 209
+ K + ++ ++ KKKPK KK + E + KGE L
Sbjct: 171 EEDGKKNKKKEKDESGTEEKKKKPKKEKKQKEESKSNEDKK--VKGKKEKGEKGDLEKED 228
Query: 210 RFRQRRRRRVACKQNPRRVKKPRLK 234
+++ + + KK + K
Sbjct: 229 EEKKKEHDETDQEMKEKDSKKNKKK 253
Score = 28.5 bits (62), Expect = 7.2
Identities = 27/120 (22%), Positives = 47/120 (38%), Gaps = 29/120 (24%)
Query: 70 KEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNS 129
K+++G + + +L E++ K T+Q +K KNK
Sbjct: 210 KKVKGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSKKNKK----------------- 252
Query: 130 KIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGE 189
K+K S A + + ++P K + D ++ KK K KKG +K + E E
Sbjct: 253 ---KEKDESCA--------EEKKKKPDKEKKEKDESTEKEDKKLKG-KKGKGEKPEKEDE 300
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,636,636
Number of Sequences: 26719
Number of extensions: 358930
Number of successful extensions: 1975
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 95
Number of HSP's that attempted gapping in prelim test: 1883
Number of HSP's gapped (non-prelim): 166
length of query: 419
length of database: 11,318,596
effective HSP length: 102
effective length of query: 317
effective length of database: 8,593,258
effective search space: 2724062786
effective search space used: 2724062786
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145331.12


