

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145330.19 - phase: 0 /pseudo
(469 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
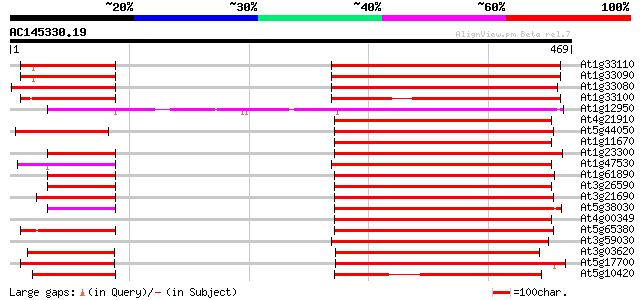
Score E
Sequences producing significant alignments: (bits) Value
At1g33110 Unknown protein (T9L6.1) 249 2e-66
At1g33090 unknown protein 248 5e-66
At1g33080 unknown protein 240 1e-63
At1g33100 unknown protein 219 3e-57
At1g12950 unknown protein 219 3e-57
At4g21910 unknown protein 206 2e-53
At5g44050 putative protein 206 3e-53
At1g11670 unknown protein 206 3e-53
At1g23300 conserved hypothetical protein 204 8e-53
At1g47530 unknown protein 204 1e-52
At1g61890 unknown protein 203 1e-52
At3g26590 integral membrane protein, putative 202 3e-52
At3g21690 integral membrane protein, putative 202 3e-52
At5g38030 putative transmembrane protein 201 9e-52
At4g00349 unknown protein 199 3e-51
At5g65380 unknown protein 198 5e-51
At3g59030 unknown protein 187 1e-47
At3g03620 unknown protein 180 1e-45
At5g17700 unknown protein 178 6e-45
At5g10420 putative protein 171 1e-42
>At1g33110 Unknown protein (T9L6.1)
Length = 494
Score = 249 bits (637), Expect = 2e-66
Identities = 116/191 (60%), Positives = 154/191 (79%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ ++ICLNING EMMI+LGF+AAASVRVSNELG G+ K AKF+ + V TSL++G LF
Sbjct: 296 DALAICLNINGLEMMIALGFLAAASVRVSNELGSGNPKGAKFATLTAVFTSLSLGIVLFF 355
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF R R++YIFT+++ VAA V +LSPLL+ SIL+NSVQPVLSGVA+GAGWQ V YVN
Sbjct: 356 VFLFLRGRVSYIFTTSEAVAAEVADLSPLLAFSILMNSVQPVLSGVAVGAGWQGYVTYVN 415
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIP+GI+LG ++ QVKG+W+GMLFG +QT VL ++T +T+WD+QV+ + +R
Sbjct: 416 LACYYLVGIPIGIILGYVVGLQVKGVWIGMLFGIFVQTCVLTVMTLRTDWDQQVSTSLRR 475
Query: 450 VNRWSKVESTD 460
+NRW ES D
Sbjct: 476 LNRWVVPESRD 486
Score = 105 bits (263), Expect = 4e-23
Identities = 50/81 (61%), Positives = 67/81 (81%), Gaps = 2/81 (2%)
Query: 10 LLKKPEEEN--EQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSR 67
LLKK E E++EL L ++VW E+K +W+VAAPAIFTRFSTFG+ IISQ+F+GH+G
Sbjct: 12 LLKKTAENGGEEKDELGLKQKVWIESKKLWIVAAPAIFTRFSTFGVSIISQSFIGHLGPI 71
Query: 68 ELAAFALVFTVLIRFANGILM 88
ELAA+++ FTVL+RF+NGIL+
Sbjct: 72 ELAAYSITFTVLLRFSNGILL 92
>At1g33090 unknown protein
Length = 494
Score = 248 bits (633), Expect = 5e-66
Identities = 115/191 (60%), Positives = 154/191 (80%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ ++IC+N+N +MMI+LGF+AA SVRVSNELG+G+ + AKF+ +V V TSL+IG LF
Sbjct: 296 DALAICINVNALQMMIALGFLAAVSVRVSNELGRGNPEGAKFATIVAVFTSLSIGLVLFF 355
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF R R++YIFT+++ VAA V +LSPLL+ SILLNSVQPVLSGVA+GAGWQ VAY+N
Sbjct: 356 VFLFLRGRISYIFTTSEAVAAEVADLSPLLAFSILLNSVQPVLSGVAVGAGWQGYVAYIN 415
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIPVG+VLG ++ QVKG+W+GMLFG +QT VL I+T +T+WD+QV+ + K
Sbjct: 416 LACYYLLGIPVGLVLGYVVGLQVKGVWIGMLFGIFVQTCVLTIMTLRTDWDQQVSTSLKN 475
Query: 450 VNRWSKVESTD 460
+NRW ES D
Sbjct: 476 INRWVVPESRD 486
Score = 101 bits (252), Expect = 8e-22
Identities = 47/81 (58%), Positives = 65/81 (80%), Gaps = 2/81 (2%)
Query: 10 LLKKPEEEN--EQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSR 67
LLKK E E +EL L ++VW E+K +WVVAAP+IFT+FST+G+ +++Q FVGHIG
Sbjct: 12 LLKKTTENGGEENDELGLKEKVWIESKKLWVVAAPSIFTKFSTYGVSLVTQGFVGHIGPT 71
Query: 68 ELAAFALVFTVLIRFANGILM 88
ELAA+++ FTVL+RF+NGIL+
Sbjct: 72 ELAAYSITFTVLLRFSNGILL 92
>At1g33080 unknown protein
Length = 494
Score = 240 bits (612), Expect = 1e-63
Identities = 110/189 (58%), Positives = 149/189 (78%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
N ++IC+NIN EMM++ GFMAAASVRVSNE+G G++ AKF+ +V V TSL+IG F
Sbjct: 296 NALAICININALEMMVAFGFMAAASVRVSNEIGSGNSNGAKFATMVVVSTSLSIGIIFFF 355
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF RER++YIFT+++ VA V +LSPLL+ SILLNS+QPVLSGVA+GAGWQ V VN
Sbjct: 356 IFLFLRERVSYIFTTSEAVATQVADLSPLLAFSILLNSIQPVLSGVAVGAGWQKYVTVVN 415
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIP G+ LG ++ QVKG+W+GM+FG +QT VL ++T +T+WD+QV+ + KR
Sbjct: 416 LACYYLVGIPSGLFLGYVVGLQVKGVWLGMIFGIFVQTCVLTVMTMRTDWDQQVSSSLKR 475
Query: 450 VNRWSKVES 458
+NRW + ES
Sbjct: 476 LNRWVEPES 484
Score = 100 bits (250), Expect = 1e-21
Identities = 47/88 (53%), Positives = 66/88 (74%), Gaps = 1/88 (1%)
Query: 2 ERDLKQNLLLKKPEEENEQEE-LSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAF 60
E ++ + LL K E E + L + ++VW E+K +WVVA PAIFTRFST G+ +ISQAF
Sbjct: 5 EGEVTETLLKKSTENRGEDRDGLGMKEKVWRESKKLWVVAGPAIFTRFSTSGLSLISQAF 64
Query: 61 VGHIGSRELAAFALVFTVLIRFANGILM 88
+GH+GS ELAA+++ TVL+RF+NGIL+
Sbjct: 65 IGHLGSTELAAYSITLTVLLRFSNGILL 92
>At1g33100 unknown protein
Length = 475
Score = 219 bits (558), Expect = 3e-57
Identities = 104/191 (54%), Positives = 142/191 (73%), Gaps = 16/191 (8%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ ++IC++IN EMMI+LGF+AA SVRVSNELG G+ K AKF+ ++ V T
Sbjct: 293 DALAICISINALEMMIALGFLAAVSVRVSNELGSGNPKGAKFATLIAVFTG--------- 343
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
R++YIFT+++ VAA V +LSPLL+ SILLNSVQPVLSGVAIGAGWQ VAYVN
Sbjct: 344 -------RISYIFTTSEAVAAEVADLSPLLAFSILLNSVQPVLSGVAIGAGWQGYVAYVN 396
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIP+G++LG ++ QVKG+W+GMLFG +QT VL ++T +T+WD+QV+ + +
Sbjct: 397 LACYYLVGIPIGVILGYVVGLQVKGVWIGMLFGIFVQTCVLTVMTLRTDWDQQVSTSLRN 456
Query: 450 VNRWSKVESTD 460
+NRW ES D
Sbjct: 457 INRWVVPESRD 467
Score = 102 bits (254), Expect = 5e-22
Identities = 45/79 (56%), Positives = 67/79 (83%), Gaps = 1/79 (1%)
Query: 10 LLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSREL 69
L+KK E E++EL + ++VW E+K +WVVAAPAIFTR+STFG+ +++QAF+GH+G EL
Sbjct: 12 LVKKTGRE-EEDELGMKEKVWIESKKLWVVAAPAIFTRYSTFGVSMVTQAFIGHLGPTEL 70
Query: 70 AAFALVFTVLIRFANGILM 88
AA+++ FT+L+RF+NGIL+
Sbjct: 71 AAYSITFTILLRFSNGILL 89
>At1g12950 unknown protein
Length = 522
Score = 219 bits (558), Expect = 3e-57
Identities = 151/469 (32%), Positives = 230/469 (48%), Gaps = 53/469 (11%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGIL---- 87
E++ +W +A PAIFT S + + ++Q F GHI + LAA ++ +V+ F+ GI+
Sbjct: 67 ESRKLWKLAGPAIFTTMSQYSLGAVTQVFAGHISTLALAAVSIENSVIAGFSFGIMLGMG 126
Query: 88 --MKIVVGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDT 145
++ + G ++ +R W S+++ + S + + T
Sbjct: 127 SALETLCGQAFGAGKVSMLGVYLQRSWVILSVTALFL-----------SLIYIFAAPILT 175
Query: 146 LRPR*KHIRSCRKHFSLVNSYYVCFYCLIHLPNIPSITKQEHHHCILG---------SFF 196
+ I + FS+ + Y I+ P + Q + G SFF
Sbjct: 176 FIGQTAAISAMAGIFSIYMIPQIFAYA-INFPTAKFLQSQSKIMVMAGISGVVLVIHSFF 234
Query: 197 ------HNHSCIPLLAFDNEISIWDCWCNDFNY-FGILDSQHWSTHICYLWLVS*NLEWF 249
H +P LA S W Y F + WS + W NL F
Sbjct: 235 TWLVMSRLHWGLPGLALVLNTSWWVIVVAQLVYIFNCTCGEAWSG---FTWEAFHNLWGF 291
Query: 250 LIFSIQRPLACCQTFPFSRRNVV---------------SICLNINGWEMMISLGFMAAAS 294
+ S+ C + V+ SIC+NI GW M++ G AA S
Sbjct: 292 VKLSLASAAMLCLEIWYFMALVLFAGYLKNAEVSVAALSICMNILGWAAMVAFGTNAAVS 351
Query: 295 VRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGE 354
VRVSNELG + AKFS+VV V+ S AIG F+ LFFR +F ++EV V E
Sbjct: 352 VRVSNELGASHPRTAKFSLVVAVILSTAIGMFIAAGLLFFRNEYPVLFVEDEEVRNVVRE 411
Query: 355 LSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKG 414
L+P+L+ I++N+VQPVLSGVA+GAGWQ+ VAYVNI CYY+ G+P G++LG + + V G
Sbjct: 412 LTPMLAFCIVINNVQPVLSGVAVGAGWQAVVAYVNIACYYLFGVPFGLLLGFKLEYGVMG 471
Query: 415 IWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWSKVESTDQET 463
IW GM+ GT +Q+IVL + KTNW+++ ++A +R+ W V + ++ET
Sbjct: 472 IWWGMVTGTFVQSIVLTWMICKTNWEKEASMAEERIKEWGGVPA-EKET 519
>At4g21910 unknown protein
Length = 509
Score = 206 bits (525), Expect = 2e-53
Identities = 96/182 (52%), Positives = 134/182 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC++I+ M+S+GF AA SVR SNELG G+ K+A FS S I L
Sbjct: 319 LSICMSISALSFMVSVGFNAAVSVRTSNELGAGNPKSAWFSTWTATFVSFVISVTEALAV 378
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
++FR+ ++YIFT + +VA AV +L P L+I+I+LN +QPVLSGVA+G GWQ+ VAYVN+G
Sbjct: 379 IWFRDYVSYIFTEDADVAKAVSDLCPFLAITIILNGIQPVLSGVAVGCGWQTYVAYVNVG 438
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG +LG +Q KGIW GM+ GTL+QT++LL +TY+T+WD++V ARKR++
Sbjct: 439 CYYVVGIPVGCILGFTFDFQAKGIWTGMIGGTLMQTLILLYVTYRTDWDKEVEKARKRLD 498
Query: 452 RW 453
W
Sbjct: 499 LW 500
Score = 37.0 bits (84), Expect = 0.023
Identities = 20/62 (32%), Positives = 34/62 (54%), Gaps = 7/62 (11%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAA-------FALVFTVLIRFAN 84
E KL++ +A PAI G+ I ++ F GH+G ELAA F+LV+ +++ +
Sbjct: 57 EMKLLFRLALPAILVYLVNSGMGISARIFAGHLGKNELAAASIGNSCFSLVYGLMLGMGS 116
Query: 85 GI 86
+
Sbjct: 117 AV 118
>At5g44050 putative protein
Length = 491
Score = 206 bits (523), Expect = 3e-53
Identities = 95/184 (51%), Positives = 141/184 (76%), Gaps = 1/184 (0%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC++ING EMM+ L F A SVRV+NELG G+ K A+F+++++V SL IG + +
Sbjct: 302 MSICMSINGLEMMVPLAFFAGTSVRVANELGAGNGKRARFAMIISVTQSLIIGIIISVLI 361
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
F +++ ++F+S++ V AV LS LLS +ILLNSVQPVLSGVA+G+GWQS VA++N+G
Sbjct: 362 YFLLDQIGWMFSSSETVLKAVNNLSILLSFAILLNSVQPVLSGVAVGSGWQSLVAFINLG 421
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVTVARKRV 450
CYY IG+P+GIV+G + + VKGIW GM+F GT++QT++L+ IT + +W+++ A+ RV
Sbjct: 422 CYYFIGLPLGIVMGWMFKFGVKGIWAGMIFGGTMVQTLILIFITMRCDWEKEAQNAKVRV 481
Query: 451 NRWS 454
N+WS
Sbjct: 482 NKWS 485
Score = 68.2 bits (165), Expect = 9e-12
Identities = 34/77 (44%), Positives = 48/77 (62%)
Query: 6 KQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIG 65
K + L K + E+E + K +W ETK +W + PAIFTR +T I +I+QAF GH+G
Sbjct: 14 KAKIPLLKDQNVAEEENGEIKKEIWLETKKLWRIVGPAIFTRVTTNLIFVITQAFAGHLG 73
Query: 66 SRELAAFALVFTVLIRF 82
ELAA ++V V+I F
Sbjct: 74 ELELAAISIVNNVIIGF 90
>At1g11670 unknown protein
Length = 503
Score = 206 bits (523), Expect = 3e-53
Identities = 98/182 (53%), Positives = 136/182 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ ++A FS VT S + F +
Sbjct: 313 LAICMSISAMSFMVSVGFNAAASVRVSNELGAGNPRSAAFSTAVTTGVSFLLSLFEAIVI 372
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++YIFT + VA AV ELSP L+I+I+LN VQPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 373 LSWRHVISYIFTDSPAVAEAVAELSPFLAITIVLNGVQPVLSGVAVGCGWQAYVAYVNIG 432
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYYI+GIP+G VLG +GIW GM+ GTL+QTI+L+I+T++T+WD++V A +R++
Sbjct: 433 CYYIVGIPIGYVLGFTYDMGARGIWTGMIGGTLMQTIILVIVTFRTDWDKEVEKASRRLD 492
Query: 452 RW 453
+W
Sbjct: 493 QW 494
Score = 37.0 bits (84), Expect = 0.023
Identities = 19/57 (33%), Positives = 34/57 (59%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ ++++ F G +GS +LAA +L + F G+++
Sbjct: 50 EMKYLFHLAAPAIFVYVINNGMSMLTRIFAGRLGSMQLAAASLGNSGFNMFTLGLML 106
>At1g23300 conserved hypothetical protein
Length = 515
Score = 204 bits (519), Expect = 8e-53
Identities = 93/191 (48%), Positives = 139/191 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NI GW +M++ GF AA SVR SNELG + AKF ++V ++TS++IG + +
Sbjct: 306 LSICMNILGWPIMVAFGFNAAVSVRESNELGAEHPRRAKFLLIVAMITSVSIGIVISVTL 365
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
+ R++ +F+ ++EV V +L+PLL+++I++N++QPVLSGVA+GAGWQ VAYVNIG
Sbjct: 366 IVLRDKYPAMFSDDEEVRVLVKQLTPLLALTIVINNIQPVLSGVAVGAGWQGIVAYVNIG 425
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+ GIP+G+VLG + VKGIW GML GT++QT VLL I Y+TNW ++ ++A R+
Sbjct: 426 CYYLCGIPIGLVLGYKMELGVKGIWTGMLTGTVVQTSVLLFIIYRTNWKKEASLAEARIK 485
Query: 452 RWSKVESTDQE 462
+W + +E
Sbjct: 486 KWGDQSNKREE 496
Score = 46.2 bits (108), Expect = 4e-05
Identities = 20/57 (35%), Positives = 36/57 (63%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E+K +W +A PAIFT F + + ++Q GH+ + LAA ++ +V+ F+ GI++
Sbjct: 44 ESKKLWWLAGPAIFTSFCQYSLGAVTQILAGHVNTLALAAVSIQNSVISGFSVGIML 100
>At1g47530 unknown protein
Length = 484
Score = 204 bits (518), Expect = 1e-52
Identities = 98/184 (53%), Positives = 131/184 (70%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+NI GW MIS+GF AA SVRVSNELG G+A AKFS++V +TS IG +
Sbjct: 295 DAISICMNIEGWTAMISIGFNAAISVRVSNELGAGNAALAKFSVIVVSITSTLIGIVCMI 354
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L ++ Y+FTS++ VAA ++ LL ++LLNS+QPVLSGVA+GAGWQ+ VAYVN
Sbjct: 355 VVLATKDSFPYLFTSSEAVAAETTRIAVLLGFTVLLNSLQPVLSGVAVGAGWQALVAYVN 414
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
I CYYIIG+P G+VLG + V+GIW GM+ G +QT++L+ I Y TNW+++ A R
Sbjct: 415 IACYYIIGLPAGLVLGFTLDLGVQGIWGGMVAGICLQTLILIGIIYFTNWNKEAEQAESR 474
Query: 450 VNRW 453
V RW
Sbjct: 475 VQRW 478
Score = 50.4 bits (119), Expect = 2e-06
Identities = 28/87 (32%), Positives = 45/87 (51%), Gaps = 5/87 (5%)
Query: 7 QNLLLKKPEEENEQEELSLGKRVW-----NETKLMWVVAAPAIFTRFSTFGIQIISQAFV 61
+ L L P E E +VW E+K +W +A PAIFT S + + ++Q F
Sbjct: 5 KTLPLLDPREPPELTGTKSASKVWAKEFGEESKRLWELAGPAIFTAISQYSLGALTQTFS 64
Query: 62 GHIGSRELAAFALVFTVLIRFANGILM 88
G +G ELAA ++ +V+ A G+++
Sbjct: 65 GRLGELELAAVSVENSVISGLAFGVML 91
>At1g61890 unknown protein
Length = 501
Score = 203 bits (517), Expect = 1e-52
Identities = 98/184 (53%), Positives = 135/184 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ +AA FS VVT S + F +
Sbjct: 310 LAICMSISAISFMVSVGFNAAASVRVSNELGAGNPRAAAFSTVVTTGVSFLLSVFEAIVV 369
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++Y FT + VA AV +LSP L+I+I+LN +QPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 370 LSWRHVISYAFTDSPAVAEAVADLSPFLAITIVLNGIQPVLSGVAVGCGWQAFVAYVNIG 429
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG VLG KGIW GM+ GTL+QTI+L+I+T +T+WD++V A R++
Sbjct: 430 CYYVVGIPVGFVLGFTYDMGAKGIWTGMIGGTLMQTIILVIVTLRTDWDKEVEKASSRLD 489
Query: 452 RWSK 455
+W +
Sbjct: 490 QWEE 493
Score = 43.5 bits (101), Expect = 2e-04
Identities = 23/57 (40%), Positives = 35/57 (61%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ I+++ F GH+GS ELAA +L + F G+L+
Sbjct: 47 EMKFLFHLAAPAIFVYVINNGMSILTRIFAGHVGSFELAAASLGNSGFNMFTYGLLL 103
>At3g26590 integral membrane protein, putative
Length = 500
Score = 202 bits (514), Expect = 3e-52
Identities = 94/182 (51%), Positives = 132/182 (71%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NI GW MI++G A SVRVSNELG + AKFS++V V+TS IG + +
Sbjct: 307 LSICMNILGWTAMIAIGMNTAVSVRVSNELGANHPRTAKFSLLVAVITSTLIGFIVSMIL 366
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L FR++ +F +++V V EL+P+L++SI++N+VQPVLSGVA+GAGWQ+ VAYVNI
Sbjct: 367 LIFRDQYPSLFVKDEKVIILVKELTPILALSIVINNVQPVLSGVAVGAGWQAVVAYVNIA 426
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+ GIP G++LG +++ V GIW GML GT++QTIVL + KTNWD + ++A R+
Sbjct: 427 CYYVFGIPFGLLLGYKLNYGVMGIWCGMLTGTVVQTIVLTWMICKTNWDTEASMAEDRIR 486
Query: 452 RW 453
W
Sbjct: 487 EW 488
Score = 49.3 bits (116), Expect = 5e-06
Identities = 23/57 (40%), Positives = 37/57 (64%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
ETK +W +A PAIFT + + + I+Q F GHI + LAA ++ +V+ F+ GI++
Sbjct: 45 ETKKLWYLAGPAIFTSVNQYSLGAITQVFAGHISTIALAAVSVENSVVAGFSFGIML 101
>At3g21690 integral membrane protein, putative
Length = 506
Score = 202 bits (514), Expect = 3e-52
Identities = 95/183 (51%), Positives = 135/183 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+ I+GW MIS+GF AA SVRVSNELG G+ K+A FS+++ + SL L +
Sbjct: 315 LSICMTISGWVFMISVGFNAAISVRVSNELGAGNPKSAAFSVIIVNIYSLITCVILAIVI 374
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ L+Y FT KEV+ AV +L PLL+++++LN +QPVLSGVA+G GWQ+ VA VN+G
Sbjct: 375 LACRDVLSYAFTEGKEVSDAVSDLCPLLAVTLVLNGIQPVLSGVAVGCGWQTFVAKVNVG 434
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYYIIGIP+G + G ++ KGIW GM+ GT+IQT +L +T++T+W ++V A KR++
Sbjct: 435 CYYIIGIPLGALFGFYFNFGAKGIWTGMIGGTVIQTFILAWVTFRTDWTKEVEEASKRLD 494
Query: 452 RWS 454
+WS
Sbjct: 495 KWS 497
Score = 42.4 bits (98), Expect = 6e-04
Identities = 24/66 (36%), Positives = 40/66 (60%)
Query: 23 LSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRF 82
L L K E+KL++ +AAPA+ + + + +Q F GH+G+ ELAA +L T + F
Sbjct: 43 LRLRKATIIESKLLFNLAAPAVIVYMINYLMSMSTQIFSGHLGNLELAAASLGNTGIQVF 102
Query: 83 ANGILM 88
A G+++
Sbjct: 103 AYGLML 108
>At5g38030 putative transmembrane protein
Length = 498
Score = 201 bits (510), Expect = 9e-52
Identities = 97/190 (51%), Positives = 136/190 (71%), Gaps = 1/190 (0%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NI GW MI++G AA SVRVSNELG + AKFS++V V+TS IG + +
Sbjct: 307 LSICMNILGWTAMIAIGMNAAVSVRVSNELGAKHPRTAKFSLLVAVITSTVIGLAISIAL 366
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L FR++ +F ++EV V +L+P+L++SI++N+VQPVLSGVA+GAGWQ+ VAYVNI
Sbjct: 367 LIFRDKYPSLFVGDEEVIIVVKDLTPILAVSIVINNVQPVLSGVAVGAGWQAVVAYVNIV 426
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+ GIP G++LG +++ V GIW GML GT++QTIVL + +TNWD + +A R+
Sbjct: 427 CYYVFGIPFGLLLGYKLNFGVMGIWCGMLTGTVVQTIVLTWMICRTNWDTEAAMAEGRIR 486
Query: 452 RWSKVESTDQ 461
W E +DQ
Sbjct: 487 EWGG-EVSDQ 495
Score = 43.5 bits (101), Expect = 2e-04
Identities = 19/57 (33%), Positives = 34/57 (59%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K +W +A PAIF + + + +Q F GHI + LAA ++ +V+ F+ G+++
Sbjct: 45 EVKKLWYLAGPAIFMSITQYSLGAATQVFAGHISTIALAAVSVENSVIAGFSFGVML 101
>At4g00349 unknown protein
Length = 514
Score = 199 bits (506), Expect = 3e-51
Identities = 93/182 (51%), Positives = 134/182 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NINGWE M+ +G AA SVRVSNELG G +AAK+S++VTV+ SL IG +
Sbjct: 321 LSICMNINGWEGMLFIGINAAISVRVSNELGSGHPRAAKYSVIVTVIESLVIGVVCAIVI 380
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ A IFT ++E+ AV +L+ LL I+++LNS+QPV+SGVA+G GWQ+ VAY+N+
Sbjct: 381 LITRDDFAVIFTESEEMRKAVADLAYLLGITMILNSLQPVISGVAVGGGWQAPVAYINLF 440
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY G+P+G +LG V+GIW+GM+ GT +QT++LL + Y TNW+++V A +R+
Sbjct: 441 CYYAFGLPLGFLLGYKTSLGVQGIWIGMICGTSLQTLILLYMIYITNWNKEVEQASERMK 500
Query: 452 RW 453
+W
Sbjct: 501 QW 502
>At5g65380 unknown protein
Length = 486
Score = 198 bits (504), Expect = 5e-51
Identities = 94/184 (51%), Positives = 133/184 (72%), Gaps = 1/184 (0%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+ INGWEMMI L F A VRV+NELG G+ K A+F+ +V+V SL IG F ++
Sbjct: 299 LSICMAINGWEMMIPLAFFAGTGVRVANELGAGNGKGARFATIVSVTQSLIIGLFFWVLI 358
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
+ ++A+IF+S+ V AV +LS LL+ ++LLNSVQPVLSGVA+G+GWQS VAY+N+G
Sbjct: 359 MLLHNQIAWIFSSSVAVLDAVNKLSLLLAFTVLLNSVQPVLSGVAVGSGWQSYVAYINLG 418
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVTVARKRV 450
CYY IG+P+G ++G V GIW GM+F GT +QT++L IT + +W+++ A R+
Sbjct: 419 CYYCIGVPLGFLMGWGFKLGVMGIWGGMIFGGTAVQTMILSFITMRCDWEKEAQKASARI 478
Query: 451 NRWS 454
N+WS
Sbjct: 479 NKWS 482
Score = 67.0 bits (162), Expect = 2e-11
Identities = 34/79 (43%), Positives = 49/79 (61%), Gaps = 1/79 (1%)
Query: 10 LLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSREL 69
LLK P E E L R+ ETK +W + PAIF+R +T+ + +I+QAF GH+G EL
Sbjct: 16 LLKSPHTAEEDGE-GLKDRILVETKKLWQIVGPAIFSRVTTYSMLVITQAFAGHLGDLEL 74
Query: 70 AAFALVFTVLIRFANGILM 88
AA ++V V + F G+L+
Sbjct: 75 AAISIVNNVTVGFNFGLLL 93
>At3g59030 unknown protein
Length = 507
Score = 187 bits (474), Expect = 1e-47
Identities = 92/181 (50%), Positives = 125/181 (68%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+ W+M LG AA SVRVSNELG G+ + A S+VV +T++ I S L +
Sbjct: 312 DAISICMYYLNWDMQFMLGLSAAISVRVSNELGAGNPRVAMLSVVVVNITTVLISSVLCV 371
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L FR L+ FTS+ EV AAV +L PLL++SI LN +QP+LSGVAIG+GWQ+ VAYVN
Sbjct: 372 IVLVFRVGLSKAFTSDAEVIAAVSDLFPLLAVSIFLNGIQPILSGVAIGSGWQAVVAYVN 431
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ YY+IG+P+G VLG V GIW GM+ G ++QT+ L+++T KTNW +V A +R
Sbjct: 432 LVTYYVIGLPIGCVLGFKTSLGVAGIWWGMIAGVILQTLTLIVLTLKTNWTSEVENAAQR 491
Query: 450 V 450
V
Sbjct: 492 V 492
Score = 37.0 bits (84), Expect = 0.023
Identities = 17/60 (28%), Positives = 33/60 (54%), Gaps = 1/60 (1%)
Query: 29 VWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
VW E+KL+W ++ +I + + ++ F GH+GS +LA ++ + A GI++
Sbjct: 49 VW-ESKLLWTLSGASIVVSVLNYMLSFVTVMFTGHLGSLQLAGASIATVGIQGLAYGIML 107
>At3g03620 unknown protein
Length = 466
Score = 180 bits (457), Expect = 1e-45
Identities = 91/171 (53%), Positives = 121/171 (70%)
Query: 273 SICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFL 332
SIC I WE+ I LGF+ AA VRV+NELGKG A A +FSI V + S +G L
Sbjct: 294 SICQYIYTWELNICLGFLGAACVRVANELGKGDAHAVRFSIKVILTISTLMGVIFSALCL 353
Query: 333 FFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGC 392
F R++Y+F+++ EV+ AV +LS +L++SILLNS+QP+LSGVA+GAG QS VA VN+
Sbjct: 354 AFCGRISYLFSNSDEVSDAVNDLSVILAVSILLNSIQPILSGVAVGAGMQSIVAVVNLAS 413
Query: 393 YYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQV 443
YY IGIP+G++L + H VKG+W GML G IQTI+L I YKT+W+ +V
Sbjct: 414 YYAIGIPLGLILTYVFHLGVKGLWSGMLAGIAIQTIILCYIIYKTDWELEV 464
Score = 62.8 bits (151), Expect = 4e-10
Identities = 28/72 (38%), Positives = 45/72 (61%)
Query: 16 EENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALV 75
+ N +E + L +VW+E MW +A P+ R ++FG I++QAF+GH LAA+AL+
Sbjct: 15 QSNNRESIYLRTKVWSEVNKMWRIALPSSLFRMTSFGSIIVAQAFIGHSSELGLAAYALL 74
Query: 76 FTVLIRFANGIL 87
+ IRF G++
Sbjct: 75 QSTFIRFLYGLM 86
>At5g17700 unknown protein
Length = 497
Score = 178 bits (451), Expect = 6e-45
Identities = 92/196 (46%), Positives = 129/196 (64%), Gaps = 4/196 (2%)
Query: 273 SICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFL 332
SIC I WEM I G M AA VRV+NELGKG A A +FSI V ++ S IG L
Sbjct: 297 SICQYIYSWEMNICFGLMGAACVRVANELGKGDADAVRFSIKVVLVVSAVIGVICSALCL 356
Query: 333 FFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGC 392
F +++Y+F+ ++ V+ AV +LS +LSISIL N +QP+LSGVAIGAG QS VA VN+
Sbjct: 357 AFGGQISYLFSDSQAVSDAVADLSIVLSISILFNIIQPILSGVAIGAGMQSMVALVNLAS 416
Query: 393 YYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNR 452
YY IG+P+G++L + ++ +KG+W GML G IQT++L + YKT+W+ +V +R+
Sbjct: 417 YYAIGVPLGVLLVYVFNFGIKGLWSGMLAGVGIQTLILCYVIYKTDWELEVKKTNERMKT 476
Query: 453 WS----KVESTDQETK 464
W+ V+ST T+
Sbjct: 477 WTLNLPAVQSTTISTR 492
Score = 69.3 bits (168), Expect = 4e-12
Identities = 32/78 (41%), Positives = 48/78 (61%)
Query: 10 LLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSREL 69
LL E E +E L L K++W+E + MW +A P+ R +FG +++QAF+GH L
Sbjct: 12 LLNGSETEQRRESLYLRKKIWSEVRKMWRIALPSTLFRVMSFGCVVVAQAFIGHSSETGL 71
Query: 70 AAFALVFTVLIRFANGIL 87
AA+AL+ + IRF GI+
Sbjct: 72 AAYALLQSTFIRFIYGIM 89
>At5g10420 putative protein
Length = 456
Score = 171 bits (432), Expect = 1e-42
Identities = 85/174 (48%), Positives = 117/174 (66%), Gaps = 26/174 (14%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+++NGWEMMI L F A VRV+NELG G+ K A+F+ +V++
Sbjct: 300 LSICMSVNGWEMMIPLAFFAGTGVRVANELGAGNGKGARFATIVSI-------------- 345
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
T ++ V AV LS LL+ ++LLNSVQPVLSGVA+G+GWQS VAY+N+G
Sbjct: 346 -----------TLSEAVLNAVDNLSVLLAFTVLLNSVQPVLSGVAVGSGWQSYVAYINLG 394
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVT 444
CYY+IG+P G+ +G I + VKGIW GM+F GT IQT++L+IIT + +WD + T
Sbjct: 395 CYYLIGLPFGLTMGWIFKFGVKGIWAGMIFGGTAIQTLILIIITTRCDWDNEET 448
Score = 62.8 bits (151), Expect = 4e-10
Identities = 27/69 (39%), Positives = 44/69 (63%)
Query: 20 QEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVL 79
+E + + +W ETK +W + P+IFT +T+ I II+QAF GH+G ELAA +++
Sbjct: 26 EEGGGMKREIWIETKKIWYIVGPSIFTGLATYSILIITQAFAGHLGDLELAAISIINNFT 85
Query: 80 IRFANGILM 88
+ F G+L+
Sbjct: 86 LGFNYGLLL 94
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.329 0.141 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,524,058
Number of Sequences: 26719
Number of extensions: 448568
Number of successful extensions: 1457
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 1336
Number of HSP's gapped (non-prelim): 108
length of query: 469
length of database: 11,318,596
effective HSP length: 103
effective length of query: 366
effective length of database: 8,566,539
effective search space: 3135353274
effective search space used: 3135353274
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC145330.19


