

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145329.13 - phase: 2 /pseudo
(528 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
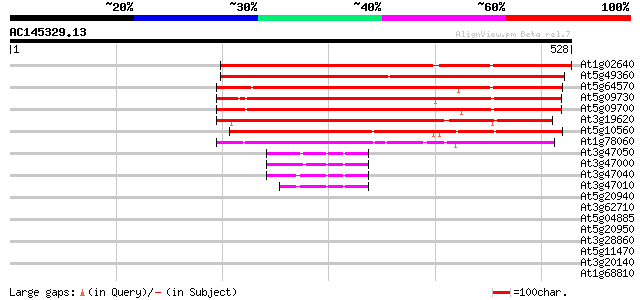
Sequences producing significant alignments: (bits) Value
At1g02640 beta-xylosidase, putative 451 e-127
At5g49360 xylosidase 402 e-112
At5g64570 beta-xylosidase 319 3e-87
At5g09730 beta-xylosidase - like protein 307 1e-83
At5g09700 beta-glucosidase - like protein 296 1e-80
At3g19620 beta-xylosidase, putative 275 3e-74
At5g10560 beta-xylosidase - like protein 274 1e-73
At1g78060 putative protein 239 3e-63
At3g47050 beta-D-glucan exohydrolase - like protein 50 3e-06
At3g47000 beta-D-glucan exohydrolase - like protein 48 2e-05
At3g47040 beta-D-glucan exohydrolase - like protein 47 2e-05
At3g47010 beta-D-glucan exohydrolase - like protein 45 1e-04
At5g20940 beta-glucosidase - like protein 41 0.002
At3g62710 beta-D-glucan exohydrolase-like protein 40 0.003
At5g04885 unknown protein 40 0.004
At5g20950 beta-D-glucan exohydrolase-like protein 37 0.027
At3g28860 P-glycoprotein, putative 30 2.5
At5g11470 putative protein 29 7.4
At3g20140 cytochrome P450, putative 29 7.4
At1g68810 putative bHLH transcription factor (bHLH030) 29 7.4
>At1g02640 beta-xylosidase, putative
Length = 768
Score = 451 bits (1159), Expect = e-127
Identities = 219/331 (66%), Positives = 264/331 (79%), Gaps = 7/331 (2%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
P+QGI YA+TIHQ+GC +V C DD+ F A++AAR ADAT+LV+GLDQSIEAE DR S
Sbjct: 443 PVQGITGYARTIHQKGCVDVHCMDDRLFDAAVEAARGADATVLVMGLDQSIEAEFKDRNS 502
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG QQ+LVS+VA A+KGP ILVLMSGGP+DI+FA+ D K+ I+WAGYPGQ GG AI
Sbjct: 503 LLLPGKQQELVSRVAKAAKGPVILVLMSGGPIDISFAEKDRKIPAIVWAGYPGQEGGTAI 562
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPVVYPFGH 377
ADILFG+A+PGGKLP+TWYPQ+YL NL MT M+MRP PGRTYRFY GPVVYPFGH
Sbjct: 563 ADILFGSANPGGKLPMTWYPQDYLTNLPMTEMSMRPVHSKRIPGRTYRFYDGPVVYPFGH 622
Query: 378 GLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
GL+YT F H ++ AP V+ + V G N +S K+IRVTHARC +LS+ +HV+V NVG
Sbjct: 623 GLSYTRFTHNIADAPKVIPIAV----RGRNGTVSGKSIRVTHARCDRLSLGVHVEVTNVG 678
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
SRDGTHT+LVFSAPP G W P+K LVAFE+VHV K+RV+VNIHVCK LSVVD++G
Sbjct: 679 SRDGTHTMLVFSAPP--GGEWAPKKQLVAFERVHVAVGEKKRVQVNIHVCKYLSVVDRAG 736
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
RRIP+G+H +HIGD H+VSLQA LG+IK
Sbjct: 737 NRRIPIGDHGIHIGDESHTVSLQASTLGVIK 767
>At5g49360 xylosidase
Length = 774
Score = 402 bits (1032), Expect = e-112
Identities = 202/328 (61%), Positives = 249/328 (75%), Gaps = 5/328 (1%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RYA+T+HQ GC+ VAC+ ++ FG A AAR ADAT+LV+GLDQSIEAET DRT
Sbjct: 447 PLQGISRYARTLHQAGCAGVACKGNQGFGAAEAAAREADATVLVMGLDQSIEAETRDRTG 506
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG+QQDLV++VA AS+GP ILVLMSGGP+D+TFAKNDP+VA I+WAGYPGQAGGAAI
Sbjct: 507 LLLPGYQQDLVTRVAQASRGPVILVLMSGGPIDVTFAKNDPRVAAIIWAGYPGQAGGAAI 566
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHG 378
A+I+FG A+PGGKLP+TWYPQ+Y+ + MT MAMR S YPGRTYRFYKGPVV+PFG G
Sbjct: 567 ANIIFGAANPGGKLPMTWYPQDYVAKVPMTVMAMRASG-NYPGRTYRFYKGPVVFPFGFG 625
Query: 379 LTYTHFVHELSSAPTV-VSVPVHGHRHGNN-TNISNKAIRVTHARCGKL-SIALHVDVKN 435
L+YT F H L+ +P +SV + N N S+ +I+V+H C + LHV+V N
Sbjct: 626 LSYTTFTHSLAKSPLAQLSVSLSNLNSANTILNSSSHSIKVSHTNCNSFPKMPLHVEVSN 685
Query: 436 VGSRDGTHTLLVFSAPP-NGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVD 494
G DGTHT+ VF+ PP NG K L+AFEKVHV A KQ V+V++ CK L VVD
Sbjct: 686 TGEFDGTHTVFVFAEPPINGIKGLGVNKQLIAFEKVHVMAGAKQTVQVDVDACKHLGVVD 745
Query: 495 KSGIRRIPMGEHSLHIGDVKHSVSLQAE 522
+ G RRIPMGEH LHIGD+KH++ +Q +
Sbjct: 746 EYGKRRIPMGEHKLHIGDLKHTILVQPQ 773
>At5g64570 beta-xylosidase
Length = 784
Score = 319 bits (817), Expect = 3e-87
Identities = 167/331 (50%), Positives = 218/331 (65%), Gaps = 8/331 (2%)
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETV 254
K PLQG+ T + GCSNVAC G A A AD ++LVIG DQSIEAE+
Sbjct: 456 KYTTPLQGLAGTVSTTYLPGCSNVACAVADVAG-ATKLAATADVSVLVIGADQSIEAESR 514
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
DR L LPG QQ+LV +VA A+KGP +LV+MSGG DITFAKNDPK+AGILW GYPG+AG
Sbjct: 515 DRVDLHLPGQQQELVIQVAKAAKGPVLLVIMSGGGFDITFAKNDPKIAGILWVGYPGEAG 574
Query: 315 GAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVY 373
G AIADI+FG +P GKLP+TWYPQ Y++ + MT M MRP K GYPGRTYRFY G VY
Sbjct: 575 GIAIADIIFGRYNPSGKLPMTWYPQSYVEKVPMTIMNMRPDKASGYPGRTYRFYTGETVY 634
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHAR----CGKLSIAL 429
FG GL+YT F H L AP++VS+ + + ++ + H G + +
Sbjct: 635 AFGDGLSYTKFSHTLVKAPSLVSLGLEENHVCRSSECQSLDAIGPHCENAVSGGGSAFEV 694
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
H+ V+N G R+G HT+ +F+ PP H P+K LV FEK+ + + + VR + +CK
Sbjct: 695 HIKVRNGGDREGIHTVFLFTTPP--AIHGSPRKHLVGFEKIRLGKREEAVVRFKVEICKD 752
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
LSVVD+ G R+I +G+H LH+GD+KHS+S++
Sbjct: 753 LSVVDEIGKRKIGLGKHLLHVGDLKHSLSIR 783
>At5g09730 beta-xylosidase - like protein
Length = 773
Score = 307 bits (786), Expect = 1e-83
Identities = 159/329 (48%), Positives = 213/329 (64%), Gaps = 8/329 (2%)
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETV 254
K PLQG+ + +Q GC NVAC D G A+D A ADA +LV+G DQSIE E
Sbjct: 447 KYTTPLQGLAETVSSTYQLGC-NVACVD-ADIGSAVDLAASADAVVLVVGADQSIEREGH 504
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
DR L LPG QQ+LV++VA A++GP +LV+MSGG DITFAKND K+ I+W GYPG+AG
Sbjct: 505 DRVDLYLPGKQQELVTRVAMAARGPVVLVIMSGGGFDITFAKNDKKITSIMWVGYPGEAG 564
Query: 315 GAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVY 373
G AIAD++FG +P G LP+TWYPQ Y++ + M+NM MRP K GYPGR+YRFY G VY
Sbjct: 565 GLAIADVIFGRHNPSGNLPMTWYPQSYVEKVPMSNMNMRPDKSKGYPGRSYRFYTGETVY 624
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPV---HGHRHGNNTNISNKAIRVTHARCGKLSIALH 430
F LTYT F H+L AP +VS+ + H R ++ +A G +H
Sbjct: 625 AFADALTYTKFDHQLIKAPRLVSLSLDENHPCRSSECQSLDAIGPHCENAVEGGSDFEVH 684
Query: 431 VDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLL 490
++VKN G R G+HT+ +F+ P H P K L+ FEK+ + + VR N++VCK L
Sbjct: 685 LNVKNTGDRAGSHTVFLFTTSPQ--VHGSPIKQLLGFEKIRLGKSEEAVVRFNVNVCKDL 742
Query: 491 SVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
SVVD++G R+I +G H LH+G +KHS+++
Sbjct: 743 SVVDETGKRKIALGHHLLHVGSLKHSLNI 771
>At5g09700 beta-glucosidase - like protein
Length = 411
Score = 296 bits (759), Expect = 1e-80
Identities = 155/330 (46%), Positives = 210/330 (62%), Gaps = 8/330 (2%)
Query: 195 KLCRPLQGIGRYAKTI-HQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAET 253
K PLQG+ R T + +GC NV C + A A ADAT+LV+G DQ+IE ET
Sbjct: 83 KYTTPLQGLERTVLTTKYHRGCFNVTCTE-ADLDSAKTLAASADATVLVMGADQTIEKET 141
Query: 254 VDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
+DR L LPG QQ+LV++VA A++GP +LV+MSGG DITFAKND K+ I+W GYPG+A
Sbjct: 142 LDRIDLNLPGKQQELVTQVAKAARGPVVLVIMSGGGFDITFAKNDEKITSIMWVGYPGEA 201
Query: 314 GGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVV 372
GG AIAD++FG +P GKLP+TWYPQ Y++ + MTNM MRP K GY GRTYRFY G V
Sbjct: 202 GGIAIADVIFGRHNPSGKLPMTWYPQSYVEKVPMTNMNMRPDKSNGYLGRTYRFYIGETV 261
Query: 373 YPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCG---KLSIAL 429
Y FG GL+YT+F H+L AP VS+ + + + + H + +
Sbjct: 262 YAFGDGLSYTNFSHQLIKAPKFVSLNLDESQSCRSPECQSLDAIGPHCEKAVGERSDFEV 321
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
+ V+NVG R+GT T+ +F+ PP H P+K L+ FEK+ + K + VR + VCK
Sbjct: 322 QLKVRNVGDREGTETVFLFTTPPE--VHGSPRKQLLGFEKIRLGKKEETVVRFKVDVCKD 379
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
L VVD+ G R++ +G H LH+G +KHS ++
Sbjct: 380 LGVVDEIGKRKLALGHHLLHVGSLKHSFNI 409
>At3g19620 beta-xylosidase, putative
Length = 781
Score = 275 bits (704), Expect = 3e-74
Identities = 143/326 (43%), Positives = 202/326 (61%), Gaps = 15/326 (4%)
Query: 195 KLCRPLQGIGRYA--KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAE 252
K P+QG+ +Y K +++ GC +V C D A+ A AD T+LV+GLDQ++EAE
Sbjct: 439 KYTSPIQGLQKYVPEKIVYEPGCKDVKCGDQTLISAAVKAVSEADVTVLVVGLDQTVEAE 498
Query: 253 TVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQ 312
+DR +L LPG+Q+ LV VA A+K +LV+MS GP+DI+FAKN + +LW GYPG+
Sbjct: 499 GLDRVNLTLPGYQEKLVRDVANAAKKTVVLVIMSAGPIDISFAKNLSTIRAVLWVGYPGE 558
Query: 313 AGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPV 371
AGG AIA ++FG +P G+LP TWYPQE+ +AMT+M MRP S G+PGR+YRFY G
Sbjct: 559 AGGDAIAQVIFGDYNPSGRLPETWYPQEFADKVAMTDMNMRPNSTSGFPGRSYRFYTGKP 618
Query: 372 VYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHV 431
+Y FG+GL+Y+ F + SAP+++ + + + N T ++ ++ C L I + +
Sbjct: 619 IYKFGYGLSYSSFSTFVLSAPSIIHIKTNPIMNLNKTT----SVDISTVNCHDLKIRIVI 674
Query: 432 DVKNVGSRDGTHTLLVFSAPPN------GGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIH 485
VKN G R G+H +LVF PP GG VP LV FE+V V ++ V+
Sbjct: 675 GVKNHGLRSGSHVVLVFWKPPKCSKSLVGGG--VPLTQLVGFERVEVGRSMTEKFTVDFD 732
Query: 486 VCKLLSVVDKSGIRRIPMGEHSLHIG 511
VCK LS+VD G R++ G H L IG
Sbjct: 733 VCKALSLVDTHGKRKLVTGHHKLVIG 758
>At5g10560 beta-xylosidase - like protein
Length = 792
Score = 274 bits (700), Expect = 1e-73
Identities = 149/323 (46%), Positives = 202/323 (62%), Gaps = 13/323 (4%)
Query: 208 KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQD 267
KT + GCS+V+C D FG A+ A+ AD I+V GLD S E E DR SL LPG Q+D
Sbjct: 472 KTSYASGCSDVSCDSDTGFGEAVAIAKGADFVIVVAGLDLSQETEDKDRVSLSLPGKQKD 531
Query: 268 LVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTAS 327
LVS VAA SK P ILVL GGPVD+TFAKNDP++ I+W GYPG+ GG A+A+I+FG +
Sbjct: 532 LVSHVAAVSKKPVILVLTGGGPVDVTFAKNDPRIGSIIWIGYPGETGGQALAEIIFGDFN 591
Query: 328 PGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVH 386
PGG+LP TWYP+ + ++AM++M MR S GYPGRTYRFY GP VY FG GL+YT F +
Sbjct: 592 PGGRLPTTWYPESF-TDVAMSDMHMRANSSRGYPGRTYRFYTGPQVYSFGTGLSYTKFEY 650
Query: 387 ELSSAPTVVSV-----PVHGHR----HGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
++ SAP +S+ H+ HG + ++ C L + V V N G
Sbjct: 651 KILSAPIRLSLSELLPQQSSHKKQLQHGEELRYLQLDDVIVNS-CESLRFNVRVHVSNTG 709
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
DG+H +++FS P + VP+K L+ +++VHV + I CK LSV + G
Sbjct: 710 EIDGSHVVMLFSKMPPVLS-GVPEKQLIGYDRVHVRSNEMMETVFVIDPCKQLSVANDVG 768
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQ 520
R IP+G H L +GD++HS+S++
Sbjct: 769 KRVIPLGSHVLFLGDLQHSLSVE 791
>At1g78060 putative protein
Length = 767
Score = 239 bits (610), Expect = 3e-63
Identities = 132/326 (40%), Positives = 197/326 (59%), Gaps = 16/326 (4%)
Query: 195 KLCRPLQGIGRYAKT-IHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAET 253
K PL + Y K ++ QGC +VAC + A+ A++AD +L++GLDQ+ E E
Sbjct: 441 KTVTPLDALRSYVKNAVYHQGCDSVAC-SNAAIDQAVAIAKNADHVVLIMGLDQTQEKED 499
Query: 254 VDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
DR L LPG QQ+L++ VA A+K P +LVL+ GGPVDI+FA N+ K+ I+WAGYPG+A
Sbjct: 500 FDRVDLSLPGKQQELITSVANAAKKPVVLVLICGGPVDISFAANNNKIGSIIWAGYPGEA 559
Query: 314 GGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVY 373
GG AI++I+FG +PGG+LPVTWYPQ ++ N+ MT+M MR S GYPGRTY+FYKGP VY
Sbjct: 560 GGIAISEIIFGDHNPGGRLPVTWYPQSFV-NIQMTDMRMR-SATGYPGRTYKFYKGPKVY 617
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVT------HARCGKLSI 427
FGHGL+Y+ + + T+ ++ ++ TN + ++R T C
Sbjct: 618 EFGHGLSYSAYSYRFK---TLAETNLYLNQSKAQTN--SDSVRYTLVSEMGKEGCDVAKT 672
Query: 428 ALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWV-PQKSLVAFEKVHVPAKTKQRVRVNIHV 486
+ V+V+N G G H +L+F+ GG +K LV F+ + + K + I +
Sbjct: 673 KVTVEVENQGEMAGKHPVLMFARHERGGEDGKRAEKQLVGFKSIVLSNGEKAEMEFEIGL 732
Query: 487 CKLLSVVDKSGIRRIPMGEHSLHIGD 512
C+ LS ++ G+ + G++ L +GD
Sbjct: 733 CEHLSRANEFGVMVLEEGKYFLTVGD 758
>At3g47050 beta-D-glucan exohydrolase - like protein
Length = 612
Score = 50.1 bits (118), Expect = 3e-06
Identities = 32/97 (32%), Positives = 56/97 (56%), Gaps = 5/97 (5%)
Query: 242 VIGLDQSIEAETV-DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPK 300
++ + +S AET+ D + L++P + ++++ VA K PT+++L SG P+ + + K
Sbjct: 484 IVAVGESPYAETMGDNSELVIPFNGSEIITTVA--EKIPTLVILFSGRPMFLEPQVLE-K 540
Query: 301 VAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWY 337
++ A PG G IAD++FG GKLP TW+
Sbjct: 541 AEALVAAWLPGTEG-QGIADVIFGDYDFRGKLPATWF 576
>At3g47000 beta-D-glucan exohydrolase - like protein
Length = 608
Score = 47.8 bits (112), Expect = 2e-05
Identities = 32/97 (32%), Positives = 55/97 (55%), Gaps = 5/97 (5%)
Query: 242 VIGLDQSIEAETV-DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPK 300
++ + + AET+ D + L +P + D+V+ VA PT+++L+SG PV + + K
Sbjct: 486 IVAVGEPPYAETMGDNSELRIPFNGTDIVTAVAEII--PTLVILISGRPVVLEPTVLE-K 542
Query: 301 VAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWY 337
++ A PG G +AD++FG GKLPV+W+
Sbjct: 543 TEALVAAWLPGTEG-QGVADVVFGDYDFKGKLPVSWF 578
>At3g47040 beta-D-glucan exohydrolase - like protein
Length = 636
Score = 47.4 bits (111), Expect = 2e-05
Identities = 32/97 (32%), Positives = 55/97 (55%), Gaps = 5/97 (5%)
Query: 242 VIGLDQSIEAETV-DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPK 300
++ + ++ AET+ D + L +P + D+V+ A A K PT++VL SG P+ + + K
Sbjct: 511 IVAVGETPYAETLGDNSELTIPLNGNDIVT--ALAEKIPTLVVLFSGRPLVLEPLVLE-K 567
Query: 301 VAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWY 337
++ A PG G + D++FG GKLPV+W+
Sbjct: 568 AEALVAAWLPGTEG-QGMTDVIFGDYDFEGKLPVSWF 603
>At3g47010 beta-D-glucan exohydrolase - like protein
Length = 609
Score = 45.1 bits (105), Expect = 1e-04
Identities = 26/83 (31%), Positives = 48/83 (57%), Gaps = 4/83 (4%)
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
D + L +P + ++++ A A K PT+++L SG P+ + + K ++ A +PG G
Sbjct: 501 DNSELTIPFNGNNIIT--AVAEKIPTLVILFSGRPMVLEPTVLE-KTEALVAAWFPGTEG 557
Query: 315 GAAIADILFGTASPGGKLPVTWY 337
++D++FG GKLPV+W+
Sbjct: 558 -QGMSDVIFGDYDFKGKLPVSWF 579
>At5g20940 beta-glucosidase - like protein
Length = 626
Score = 40.8 bits (94), Expect = 0.002
Identities = 42/153 (27%), Positives = 65/153 (42%), Gaps = 25/153 (16%)
Query: 228 PALDAARHADATILVIGLDQSIEAETV-DRTSLLLPGHQQDLVSKVAAASKGPTILVLMS 286
P + + D ++ + + AE D T+L + + V A+ K ++V++S
Sbjct: 493 PDTNFVKAGDFDYAIVAVGEKPYAEGFGDSTNLTISEPGPSTIGNVCASVK--CVVVVVS 550
Query: 287 GGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLA 346
G PV + + D VA L PG G +AD+LFG GKL TW+
Sbjct: 551 GRPVVMQISNIDALVAAWL----PGTEG-QGVADVLFGDYGFTGKLARTWF--------- 596
Query: 347 MTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHGL 379
+ P +G P + P +YPFG GL
Sbjct: 597 -KTVDQLPMNVGDP------HYDP-LYPFGFGL 621
>At3g62710 beta-D-glucan exohydrolase-like protein
Length = 650
Score = 40.0 bits (92), Expect = 0.003
Identities = 41/154 (26%), Positives = 68/154 (43%), Gaps = 23/154 (14%)
Query: 228 PALDAAR-HADATILVIGLDQSIEAETV-DRTSLLLPGHQQDLVSKVAAASKGPTILVLM 285
P D A+ HADA ++ + ++ AET D +L + D +S + +++L+
Sbjct: 516 PNQDTAKLHADAAYTIVVVGETPYAETFGDSPTLGITKPGPDTLSHTCGSGM-KCLVILV 574
Query: 286 SGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNL 345
+G P+ I + + W PG G +AD+LFG G LP TW
Sbjct: 575 TGRPLVIEPYIDMLDALAVAWL--PGTEG-QGVADVLFGDHPFTGTLPRTW--------- 622
Query: 346 AMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHGL 379
M ++ P +G + Y +YPFG+G+
Sbjct: 623 -MKHVTQLPMNVG--DKNY-----DPLYPFGYGI 648
>At5g04885 unknown protein
Length = 665
Score = 39.7 bits (91), Expect = 0.004
Identities = 34/101 (33%), Positives = 51/101 (49%), Gaps = 14/101 (13%)
Query: 242 VIGLDQSIEAETV---DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKND 298
+I + + AET D+ ++L PG ++S A K ++V++SG P+ + +
Sbjct: 505 IIAVGEPPYAETAGDSDKLTMLDPGPA--IISSTCQAVK--CVVVVISGRPLVM-----E 555
Query: 299 PKVAGI--LWAGYPGQAGGAAIADILFGTASPGGKLPVTWY 337
P VA I L A + G I D LFG GKLPVTW+
Sbjct: 556 PYVASIDALVAAWLPGTEGQGITDALFGDHGFSGKLPVTWF 596
>At5g20950 beta-D-glucan exohydrolase-like protein
Length = 624
Score = 37.0 bits (84), Expect = 0.027
Identities = 43/146 (29%), Positives = 62/146 (42%), Gaps = 27/146 (18%)
Query: 237 DATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAK 296
D I+V+G E D T+L + ++ V + K ++V++SG PV I
Sbjct: 498 DYAIVVVGEPPYAEMFG-DTTNLTISDPGPSIIGNVCGSVK--CVVVVVSGRPVVI---- 550
Query: 297 NDPKVAGI--LWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP 354
P V+ I L A + G +AD LFG GKL TW+ ++ P
Sbjct: 551 -QPYVSTIDALVAAWLPGTEGQGVADALFGDYGFTGKLARTWF----------KSVKQLP 599
Query: 355 SKIGYPGRTYRFYKGPVVYPFGHGLT 380
+G R Y +YPFG GLT
Sbjct: 600 MNVG--DRHY-----DPLYPFGFGLT 618
>At3g28860 P-glycoprotein, putative
Length = 1252
Score = 30.4 bits (67), Expect = 2.5
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 221 RDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPT 280
+D +DAAR A+A + GL + + +R + L G Q+ ++ A K PT
Sbjct: 1110 KDGATESEVIDAARAANAHGFISGLPEGYKTPVGER-GVQLSGGQKQRIAIARAVLKNPT 1168
Query: 281 ILVL 284
+L+L
Sbjct: 1169 VLLL 1172
>At5g11470 putative protein
Length = 691
Score = 28.9 bits (63), Expect = 7.4
Identities = 22/77 (28%), Positives = 37/77 (47%), Gaps = 7/77 (9%)
Query: 240 ILVIGLDQSIEAETVDRTSLLLPGHQQDLV-------SKVAAASKGPTILVLMSGGPVDI 292
I+ L+Q EA ++RTS+ +P + LV ++ L+L SG P+
Sbjct: 551 IVYSALNQQCEARMIERTSVTIPHIGEALVIFKTREVAERVIRRLDEGCLLLSSGRPLVA 610
Query: 293 TFAKNDPKVAGILWAGY 309
+FAK P L++G+
Sbjct: 611 SFAKITPPGKPSLFSGH 627
>At3g20140 cytochrome P450, putative
Length = 510
Score = 28.9 bits (63), Expect = 7.4
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 14/71 (19%)
Query: 366 FYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKL 425
F+K P V G GL P+ +S+P+ GH H +N+++K+++ ++ G L
Sbjct: 29 FFKKPKVSQDGFGLP-----------PSPLSLPIIGHLHLLFSNLTHKSLQKLSSKYGPL 77
Query: 426 SIALHVDVKNV 436
L++ + NV
Sbjct: 78 ---LYLRIFNV 85
>At1g68810 putative bHLH transcription factor (bHLH030)
Length = 368
Score = 28.9 bits (63), Expect = 7.4
Identities = 18/64 (28%), Positives = 34/64 (53%), Gaps = 10/64 (15%)
Query: 465 VAFEKVHVPAKTKQRVRVNIHVCKLLSVV------DKSGIRRIPMGEHSLHIGDVKHSVS 518
+A K H A+ ++R R+N H+ KL S++ DK+ + + E H+ ++K S
Sbjct: 172 LAASKSHSEAERRRRERINNHLAKLRSILPNTTKTDKASL----LAEVIQHVKELKRETS 227
Query: 519 LQAE 522
+ +E
Sbjct: 228 VISE 231
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.337 0.146 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,701,523
Number of Sequences: 26719
Number of extensions: 484449
Number of successful extensions: 1478
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1431
Number of HSP's gapped (non-prelim): 22
length of query: 528
length of database: 11,318,596
effective HSP length: 104
effective length of query: 424
effective length of database: 8,539,820
effective search space: 3620883680
effective search space used: 3620883680
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC145329.13


