

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
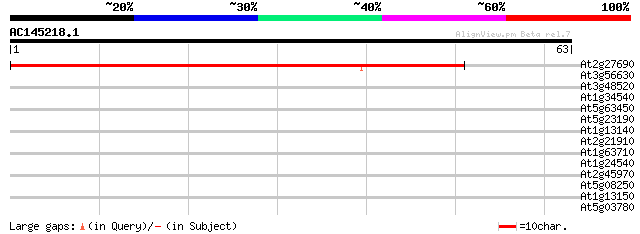
Query= AC145218.1 - phase: 0
(63 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g27690 putative cytochrome P450 47 2e-06
At3g56630 cytochrome P450-like protein 37 0.001
At3g48520 cytochrome P450-like protein 37 0.002
At1g34540 cytochrome p450, putative 37 0.002
At5g63450 cytochrome P450-like protein 34 0.016
At5g23190 cytochrome P450-like protein 30 0.29
At1g13140 cytochrome P450 monooxygenase like protein 30 0.29
At2g21910 putative cytochrome P450 28 0.65
At1g63710 cytochrome P450 like protein 28 0.65
At1g24540 putative cytochrome P450 28 1.1
At2g45970 putative cytochrome P450 27 1.9
At5g08250 cytochrome P450-like protein 27 2.5
At1g13150 putative cytochrome P450 monooxygenase 27 2.5
At5g03780 myb -like protein 25 9.4
>At2g27690 putative cytochrome P450
Length = 495
Score = 47.0 bits (110), Expect = 2e-06
Identities = 25/52 (48%), Positives = 36/52 (69%), Gaps = 1/52 (1%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDE-KIFDLRDIMRSSS 51
+ASLEL SV+VR +A EI+ EI TRL+P + SF+ + + DL+D+ R S
Sbjct: 127 LASLELGSVSVRVFAHEIVKTEIETRLLPILTSFSDNPGSVLDLQDVFRRFS 178
>At3g56630 cytochrome P450-like protein
Length = 499
Score = 37.4 bits (85), Expect = 0.001
Identities = 16/46 (34%), Positives = 30/46 (64%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A + K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLAEAATNGKLIDLQDIL 174
>At3g48520 cytochrome P450-like protein
Length = 506
Score = 37.0 bits (84), Expect = 0.002
Identities = 15/48 (31%), Positives = 32/48 (66%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E ++ ++RS+A E++ +E+ RL+P + + A DL+D+++
Sbjct: 133 LASHEFSTRSLRSFAFEVLKDEVENRLVPVLSTAADVGTTVDLQDVLK 180
>At1g34540 cytochrome p450, putative
Length = 498
Score = 37.0 bits (84), Expect = 0.002
Identities = 16/46 (34%), Positives = 29/46 (62%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLVEAATTGKLIDLQDIL 174
>At5g63450 cytochrome P450-like protein
Length = 510
Score = 33.9 bits (76), Expect = 0.016
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 135 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 182
>At5g23190 cytochrome P450-like protein
Length = 559
Score = 29.6 bits (65), Expect = 0.29
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS+E S R + + E +H RL+P + + + DL+D++
Sbjct: 162 ASIEFHSAKFRQLTTQSLFELVHKRLLPVLETSVKSSSPIDLQDVL 207
>At1g13140 cytochrome P450 monooxygenase like protein
Length = 519
Score = 29.6 bits (65), Expect = 0.29
Identities = 12/43 (27%), Positives = 25/43 (57%)
Query: 5 ELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
E+ S ++ + + + +L+ M SFAR ++ FDL+D++
Sbjct: 148 EMHSTRFVEHSFQTTQDLVRKKLLKVMESFARSQEAFDLQDVL 190
>At2g21910 putative cytochrome P450
Length = 510
Score = 28.5 bits (62), Expect = 0.65
Identities = 12/37 (32%), Positives = 22/37 (59%)
Query: 12 RSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+S++L I +I L+P + FA +FDL+D+ +
Sbjct: 140 QSFSLRTITCKIKNGLVPVLDHFAEANTVFDLQDVFQ 176
>At1g63710 cytochrome P450 like protein
Length = 523
Score = 28.5 bits (62), Expect = 0.65
Identities = 14/46 (30%), Positives = 25/46 (53%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R ++ I RL+P + S + DL+D++
Sbjct: 134 AALEFTTRTLRQAMARWVDRAIKNRLVPILESARSRAEPIDLQDVL 179
>At1g24540 putative cytochrome P450
Length = 522
Score = 27.7 bits (60), Expect = 1.1
Identities = 10/47 (21%), Positives = 26/47 (55%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
+A E+ S + + + + +L+P M + + +++FDL+D++
Sbjct: 141 VAKTEMHSSRFLEHTFTTMRDLVDQKLVPLMENLSTSKRVFDLQDLL 187
>At2g45970 putative cytochrome P450
Length = 537
Score = 26.9 bits (58), Expect = 1.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R +N I R +P + + + DL+D++
Sbjct: 134 AALEFTTRTLRQAMARWVNRAIKLRFLPILENARLGSEPIDLQDLL 179
>At5g08250 cytochrome P450-like protein
Length = 550
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/46 (30%), Positives = 25/46 (53%), Gaps = 4/46 (8%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS+E S R + ++E +H RL+P + + + DL+DI+
Sbjct: 163 ASVEFHSAKFRQLTSQSLHELVHNRLLPVLETSGK----IDLQDIL 204
>At1g13150 putative cytochrome P450 monooxygenase
Length = 529
Score = 26.6 bits (57), Expect = 2.5
Identities = 11/42 (26%), Positives = 23/42 (54%)
Query: 5 ELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDI 46
E+ S ++ + + +L+ M SFA+ ++ FDL+D+
Sbjct: 141 EMHSTGFVEHSFQTTQHLVRKKLLKVMESFAKSQEAFDLQDV 182
>At5g03780 myb -like protein
Length = 420
Score = 24.6 bits (52), Expect = 9.4
Identities = 19/62 (30%), Positives = 28/62 (44%), Gaps = 13/62 (20%)
Query: 5 ELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIF-------DLRDIMRSSSKYMSKN 57
E+ V V +A E + +P+ EK+F DL+D RS K M+KN
Sbjct: 356 EMLKVGVEKFAAEA------NKNMPWRKILEMGEKVFHETRTPADLKDKWRSMVKIMNKN 409
Query: 58 ER 59
E+
Sbjct: 410 EQ 411
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.135 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,068,212
Number of Sequences: 26719
Number of extensions: 29448
Number of successful extensions: 105
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 93
Number of HSP's gapped (non-prelim): 14
length of query: 63
length of database: 11,318,596
effective HSP length: 39
effective length of query: 24
effective length of database: 10,276,555
effective search space: 246637320
effective search space used: 246637320
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC145218.1


