

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
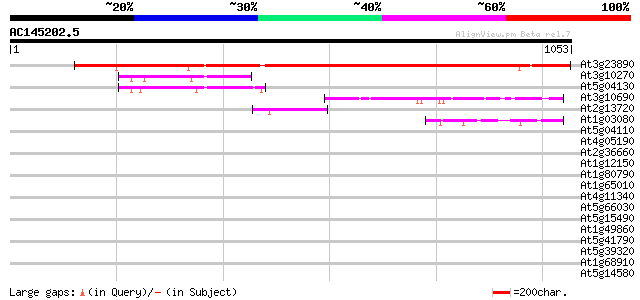
Query= AC145202.5 - phase: 0 /pseudo
(1053 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g23890 topoisomerase II 825 0.0
At3g10270 putative DNA gyrase subunit B 89 1e-17
At5g04130 DNA gyrase subunit B - like protein 83 7e-16
At3g10690 putative DNA gyrase subunit A 57 7e-08
At2g13720 unknown protein 55 3e-07
At1g03080 unknown protein 44 6e-04
At5g04110 DNA gyrase subunit B - like protein 36 0.10
At4g05190 kinesin - like protein 35 0.29
At2g36660 putative poly(A) binding protein 33 0.65
At1g12150 hypothetical protein 33 0.65
At1g80790 hypothetical protein 33 0.85
At1g65010 hypothetical protein 33 1.1
At4g11340 putative protein 32 1.5
At5g66030 Golgi-localized protein GRIP 32 1.9
At5g15490 UDP-glucose dehydrogenase-like protein 32 1.9
At1g49860 glutathione S-transferase, putative 32 2.5
At5g41790 myosin heavy chain-like protein 31 4.2
At5g39320 UDP-glucose dehydrogenase-like protein 31 4.2
At1g68910 unknown protein 31 4.2
At5g14580 polynucleotide phosphorylase 30 5.5
>At3g23890 topoisomerase II
Length = 1473
Score = 825 bits (2132), Expect = 0.0
Identities = 475/977 (48%), Positives = 626/977 (63%), Gaps = 57/977 (5%)
Query: 122 VSLCDNGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIP 180
++ C+ + WT V FKPDL KF+M LE DV+ALM KRV D+A CLG++V V LNG IP
Sbjct: 206 ITKCNKSENWTKVTFKPDLKKFNMTELEDDVVALMSKRVFDIAGCLGKSVKVELNGKQIP 265
Query: 181 VKSFKDYANFFLDRAKPF----PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKG 236
VKSF DY + +L A PLPR K+ DR E+C+SLS+G+FQQVSFVNSIATIKG
Sbjct: 266 VKSFTDYVDLYLSAANKSRTEDPLPRLTEKVNDRWEVCVSLSEGQFQQVSFVNSIATIKG 325
Query: 237 GTHVDYITKQITTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTT 295
GTHVDY+T QIT +I V KK K+ NV A VKN LWVFVNALIDNPAF+SQTKE LT
Sbjct: 326 GTHVDYVTSQITNHIVAAVNKKNKNANVKAHNVKNHLWVFVNALIDNPAFDSQTKETLTL 385
Query: 296 KPARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-----------------DAK 337
+ + G K L V KS +++NLL FK + DA
Sbjct: 386 RQSSFGSKCELSEDFLKKVGKSGVVENLLSWADFKQNKDLKKSDGAKTGRVLVEKLDDAA 445
Query: 338 LAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSR-KLFKK 396
AG +S CTLILT+G K LA++G S V+ + GVFPL KLLN S ++
Sbjct: 446 EAGGKNSRLCTLILTEGDSAKSLALAGRS-VLGNNYCGVFPLRGKLLNVREASTTQITNN 504
Query: 397 KEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKV 456
KEI+NL ILGL +N KY N LRYGQ+M+M +QD DG+H KGLLINF +SFWPSLL+V
Sbjct: 505 KEIENLKKILGLKQNMKYENVNSLRYGQMMIMTDQDHDGSHIKGLLINFIHSFWPSLLQV 564
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKG--KELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
F+ F PI+K + +KG K LSF SM +YE+WK+ L A W+IKY +GL
Sbjct: 565 PSFLVEFITPIVKAT-------RKGTKKVLSFYSMPEYEEWKESLKGNATGWDIKYYKGL 617
Query: 515 --SSAQEVREYFQDLGKHRAYFVWDDK-DETSIKLAFSKKA-EEWEAWIGNSQPGTCNYE 570
S+A+E +EYF +LG H+ FVW+D+ D +I+LAFSKK E + W+ + PG +
Sbjct: 618 GTSTAEEGKEYFSNLGLHKKDFVWEDEQDGEAIELAFSKKKIEARKNWLSSYVPGNHLDQ 677
Query: 571 RKV-INYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLT 629
R+ + Y +FVNKEL+ SM +LQRSIPS+VDGL GQRK LF +F+ K+ +V +L
Sbjct: 678 RQPKVTYSDFVNKELILFSMADLQRSIPSMVDGLKPGQRKILFVAFKKIARKEMKVAQLV 737
Query: 630 AYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYI 689
YV+ S H+G+QS+AS IIGMAQD+VGSNNINLL+P GQFGTR GGKD AS+R ++
Sbjct: 738 GYVSLLSAYHHGEQSLASAIIGMAQDYVGSNNINLLLPNGQFGTRTSGGKDSASARYIFT 797
Query: 690 KLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYH 749
KL+ VTR+LFP DDD LL+YLNEDG+ I+P WY+PIIP VLVNG +GT +S+ IP Y+
Sbjct: 798 KLSPVTRILFPKDDDLLLDYLNEDGQRIEPTWYMPIIPTVLVNGAEGIGTGWSTFIPNYN 857
Query: 750 PYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIG--YYTVDGLVERINEQTFRIK 807
P EI+ NVRR LN E MV M PWY+GF GTIEK+A G YT+ GL E ++E T RI
Sbjct: 858 PREIVANVRRLLNGESMVPMDPWYRGFKGTIEKTASKEGGCTYTITGLYEEVDETTIRIT 917
Query: 808 ELPIRMWTEDYKKFLEKI-TARG--LIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELR 864
ELPIR W +DYK FL+ + T G + + D + V+F + + EE + E
Sbjct: 918 ELPIRRWNDDYKNFLQSLKTDNGAPFFQDVKAYNDEKSVDFDL-ILSEENMLAARQEGFL 976
Query: 865 KKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLR 924
KKF LT TI+TSNM+LFD +G IKKY TPEQILEEF LR +YY +RK+ +VKN L
Sbjct: 977 KKFKLTTTIATSNMHLFDKKGVIKKYVTPEQILEEFFDLRFEYYEKRKETVVKNMEIELL 1036
Query: 925 SLRIKQKFISNIVNEKFNPINRRAELLIEIL---------KKGKSAEPHVAGAIDYSSKE 975
L K +FI +++ + R+ ++E L +K +S E +AGA+D + E
Sbjct: 1037 KLENKARFILAVLSGEIIVNKRKKADIVEDLRQKGFTPFPRKAESVEAAIAGAVDDDAAE 1096
Query: 976 QEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTS 1035
+ E+ V S +Y+YL ++ + T+E +++L A+ D+ + ++ T+
Sbjct: 1097 EP--EEILVDPESSSSYIPGSEYDYLLAMAIASLTIEKVEELLADRDKMIIAVADMKKTT 1154
Query: 1036 PDLMWLNDLELFEKEFD 1052
P +WL+DLE +KE +
Sbjct: 1155 PKSLWLSDLESLDKELE 1171
>At3g10270 putative DNA gyrase subunit B
Length = 657
Score = 89.0 bits (219), Expect = 1e-17
Identities = 75/289 (25%), Positives = 127/289 (42%), Gaps = 43/289 (14%)
Query: 204 HAKLGDRLEICLSLSDGKFQQVS---------FVNSIATIKGGTHVDYITKQITTYI--- 251
H LG R EI S D Q S + NSI TI GGTH++ + +T +
Sbjct: 257 HDVLGFRKEINGSTVDVSLQWCSDAYSDTMLGYANSIRTIDGGTHIEGVKASLTRTLNSL 316
Query: 252 --KKEVLKKKHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLKFFPGSM 309
K +V+K+K ++++ + V+ L V+ + NP F QTK L R +
Sbjct: 317 AKKLKVIKEKDISLSGEHVREGLTCIVSVKVPNPEFEGQTKTRLGNPEVRKIVDQSVQEY 376
Query: 310 LNDVQK--SQILDNLLPRIPFKHRTHVDAKLA----------------------GRPDSG 345
L + + +L++++ + ++ + AK A D
Sbjct: 377 LTEYLELHPDVLESIISKSLNAYKAALAAKRARELVRSKSVLKSSSLPGKLADCSSTDPA 436
Query: 346 KCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATR-DSRKLFKKKEIQNLMA 404
+ + + +G A G D + PL K+LN R D ++K +EIQNL+
Sbjct: 437 ESEIFIVEGDSAGGSAKQGR----DRRFQAILPLRGKILNIERKDEAAMYKNEEIQNLIL 492
Query: 405 ILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSL 453
LGL + N + LRY +++++ + D DGAH + LL+ FF+ + +L
Sbjct: 493 GLGLGVKGEDFNKENLRYHKIIILTDADVDGAHIRTLLLTFFFRYQRAL 541
>At5g04130 DNA gyrase subunit B - like protein
Length = 732
Score = 83.2 bits (204), Expect = 7e-16
Identities = 84/323 (26%), Positives = 138/323 (42%), Gaps = 51/323 (15%)
Query: 204 HAKLGDRLEICLSLSDGKFQQVS---------FVNSIATIKGGTHVDYI----TKQITTY 250
H LG R EI + D Q S + NSI TI GGTH++ + T+ + T
Sbjct: 332 HDVLGFRREINGATVDVALQWCSDAYSDTMLGYANSIRTIDGGTHIEGVKASLTRTLNTL 391
Query: 251 IKK-EVLKKKHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLKFFPGSM 309
KK + +K+K ++++ + V+ L V+ + NP F QTK L R +
Sbjct: 392 AKKSKTVKEKDISLSGEHVREGLTCIVSVKVPNPEFEGQTKTRLGNPEVRKIVDQSVQEY 451
Query: 310 LNDVQK--SQILDNLLPRIPFKHRTHVDAKLAGRPDSGKCTL------------------ 349
L + + IL++++ + ++ + AK A K L
Sbjct: 452 LTEFLELHPDILESIISKSLNAYKAALAAKRARELVRSKSVLKSSSLPGKLADCSSTDPE 511
Query: 350 ----ILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATR-DSRKLFKKKEIQNLMA 404
+ +G A G D + PL K+LN R D ++K +EIQNL+
Sbjct: 512 VSEIFIVEGDSAGGSAKQGR----DRRFQAILPLRGKILNIERKDEAAMYKNEEIQNLIL 567
Query: 405 ILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFT 464
LGL + + LRY +++++ + D DGAH + LL+ FF+ + +L + V
Sbjct: 568 GLGLGVKGEDFKKENLRYHKIIILTDADVDGAHIRTLLLTFFFRYQRALFDAG-CIYVGV 626
Query: 465 FPIIKVS-------MYNASDSKK 480
P+ KV Y+ +D KK
Sbjct: 627 PPLFKVERGKNAQYCYDDADLKK 649
>At3g10690 putative DNA gyrase subunit A
Length = 917
Score = 56.6 bits (135), Expect = 7e-08
Identities = 104/477 (21%), Positives = 196/477 (40%), Gaps = 59/477 (12%)
Query: 592 LQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTE--HSYCHYGQQSIASTI 649
L R++P V DGL R+ LF E+ ++ + + V E + +G ++ ++
Sbjct: 123 LGRALPDVRDGLKPVHRRILFAMHELGMSSKKPYKKCARVVGEVLGKFHPHGDTAVYDSL 182
Query: 650 IGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTR--LLFPVDDDKLL 707
+ MAQ F S L+ +G FG+ D A+ R +L+ + LL +D D +
Sbjct: 183 VRMAQSF--SLRCPLIQGHGNFGSID--ADPPAAMRYTECRLDPLAEAVLLSDLDQDTVD 238
Query: 708 EYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNN---- 763
N D +P +P +L+NG + +++IP ++ E++ + ++N
Sbjct: 239 FVANFDNSQKEPAVLPARLPALLLNGASGIAVGMATNIPPHNLGELVDVLCALIHNPEAT 298
Query: 764 -EEMVSMKP--------WYKGFGGTIEKSAKGIGYYTVDGL--VERINEQTFR----IKE 808
+E++ P G G ++ G G V G VE ++ +T R I E
Sbjct: 299 LQELLEYMPAPDFPTGGIIMGNLGVLDAYRTGRGRVVVRGKAEVELLDPKTKRNAVIITE 358
Query: 809 LPIR----MWTEDYKKFLEKITARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELR 864
+P + + + +E T G I R + D + + +K+ + + L
Sbjct: 359 IPYQTNKATLVQKIAELVENKTLEG-ISDIRDESDRNGMRVVIELKRGGDPALVLN-NLY 416
Query: 865 KKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLR 924
+ L + S + + + D E K+ +++L+ F R RR ++ + + Q
Sbjct: 417 RHTALQSSFSCNMVGICDGEPKLMGL---KELLQAFIDFRCSVVERRARFKLSHAQQ--- 470
Query: 925 SLRIKQKFISNIVNEKFNPINRRAELLIEILKKG---KSAEPHVAGAIDYSSKEQEAGEQ 981
++ I IV N + +IE++ K SA + S K+ EA +
Sbjct: 471 ----RKHIIEGIVVGLDN-----VDEVIELITKASSHSSATAALQSEYGLSEKQAEAILE 521
Query: 982 ESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDL 1038
++ + ++E D SSL E + KLE L + N LK +E + +L
Sbjct: 522 ITLRRLTALERKKFTDES--SSL------TEQITKLEQLLSTRTNILKLIEQEAIEL 570
>At2g13720 unknown protein
Length = 223
Score = 54.7 bits (130), Expect = 3e-07
Identities = 41/148 (27%), Positives = 75/148 (49%), Gaps = 9/148 (6%)
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKGKELS----FDSMQQYEDWKKELGNTANDWEIKYCQ 512
S F++V P +KVS S K L+ S + K + N N W IK+ +
Sbjct: 39 SSFITVLVTPYVKVSRLTFSSHKYFIVLTNLSYVTSTLNTRNGKMKDRNYLNRWTIKFYK 98
Query: 513 GLSSAQEVRE--YFQDLGKHRAYFVWDDKDETSIKLAFSKKAEEWEAWIGNSQPGT-CNY 569
G+ Q +RE ++ +++ +F+WD+ E +++ AF ++ + WI PGT +
Sbjct: 99 GVW-VQVMREKNVLSNVDQNKRFFIWDEGSEAALENAFGTSEDKQKKWIEKYVPGTFLDS 157
Query: 570 ERKVINYRNFVNKEL-LFSSMKNLQRSI 596
+++ I Y +FV K+L L+ + N ++ I
Sbjct: 158 KKQQIMYSDFVTKKLKLYFTTSNQRKHI 185
>At1g03080 unknown protein
Length = 1744
Score = 43.5 bits (101), Expect = 6e-04
Identities = 63/289 (21%), Positives = 124/289 (42%), Gaps = 63/289 (21%)
Query: 780 IEKSAKGIGYYTVDGLVERINEQTFRI----KELPIRMWT---EDYKKFLEKITARGLIE 832
+E+S + + + +DGL+E++ Q+ + KEL R+WT E+ +F+E TA ++
Sbjct: 449 LERSNQNL-HSELDGLLEKLGNQSHELTEKQKELG-RLWTCVQEENLRFMEAETAFQTLQ 506
Query: 833 SFRQKGDYEIVNFKVNVK----------------QEEIATIEKDEELRKKFNLTCTISTS 876
+ E+ + ++ QEE+ + + + NL+ S
Sbjct: 507 QLHSQSQEELSTLALELQNRSQILKDMEARNNGLQEEVQEAKDQSKSLNELNLSSAASIK 566
Query: 877 NMYLFDAEGKIKKYNTPEQILEEFCTLRL--KYYVRRKQYLVKNFTQLLRSLRIKQKFIS 934
++ + ++ K Q LE LR+ + ++++ Y +K
Sbjct: 567 SL-----QEEVSKLRETIQKLEAEVELRVDQRNALQQEIYCLK----------------- 604
Query: 935 NIVNEKFNPINRRAELLIEILK----KGKSAEPHVAGAIDYSSKEQEAGEQESVSQSVSI 990
E+ + I ++ + ++E ++ +S V + +SK +E E+ES+ ++ I
Sbjct: 605 ----EELSQIGKKHQSMVEQVELVGLHPESFGSSVKELQEENSKLKEIRERESIEKTALI 660
Query: 991 EGATLGDYEYLSSLPFENFTLE-SLKKLEAELDEKENELKTLEHTSPDL 1038
E E + L +N LE S+ L AEL+ +LKTLE S L
Sbjct: 661 E-----KLEMMEKLVQKNLLLENSISDLNAELETIRGKLKTLEEASMSL 704
>At5g04110 DNA gyrase subunit B - like protein
Length = 257
Score = 36.2 bits (82), Expect = 0.10
Identities = 18/73 (24%), Positives = 36/73 (48%), Gaps = 5/73 (6%)
Query: 225 VSFVNSIATIKGGTHVDYITKQITTYI-----KKEVLKKKHVNVNADTVKNQLWVFVNAL 279
+ + N I T+ GGT++D + IT + K ++++ K + + V L V+ +
Sbjct: 87 LGYANGIRTMDGGTYIDGVKASITRTLNSLVEKSKLVEDKDIIFTEEHVMEGLTCIVSVI 146
Query: 280 IDNPAFNSQTKEM 292
+ P F QT+ +
Sbjct: 147 VPKPEFEGQTQRL 159
>At4g05190 kinesin - like protein
Length = 777
Score = 34.7 bits (78), Expect = 0.29
Identities = 58/229 (25%), Positives = 95/229 (41%), Gaps = 36/229 (15%)
Query: 823 EKITARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMY--L 880
EK++ IE+ R++ D +V K+ V E K+E++ K +T S +MY L
Sbjct: 153 EKLSKLDAIENHRREKDCRVVAEKLQVSLREELDKVKEEKMAAKQKVT---SLEDMYKRL 209
Query: 881 FDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQL----------LRSLRIKQ 930
+ +++YNT Q E + K +++N T L L S R+ Q
Sbjct: 210 QEYNTSLQQYNTKLQTDLEVAREAHTRAEKEKSSILENLTTLRGHSKSLQDQLASSRVSQ 269
Query: 931 KFISNIVNEKFNPINRRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGE----QESVSQ 986
V +K + + L E+ + + HV ++ AGE +ESV +
Sbjct: 270 ---DEAVKQKDSLLMEVNNLQSELQQVRDDRDRHVV------QSQKLAGEILMYKESVGK 320
Query: 987 S---VSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLE 1032
S + I A G E SL E +K LE EL + +LK ++
Sbjct: 321 SSHELDILIAKSGSLEETCSL-----QKERIKMLEQELAFAKEKLKMVD 364
>At2g36660 putative poly(A) binding protein
Length = 609
Score = 33.5 bits (75), Expect = 0.65
Identities = 28/117 (23%), Positives = 57/117 (47%), Gaps = 7/117 (5%)
Query: 830 LIESFRQKGDYEIVNFKV-NVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIK 888
L E F++K + + + KV N+ + + +EELRK F+ C TS + D +GK K
Sbjct: 286 LREQFKEKHEEQKMIAKVSNIYVKNVNVAVTEEELRKHFS-QCGTITSTKLMCDEKGKSK 344
Query: 889 K-----YNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEK 940
++TPE+ ++ T + + + Y+ + R ++++ +F + + K
Sbjct: 345 GFGFVCFSTPEEAIDAVKTFHGQMFHGKPLYVAIAQKKEDRKMQLQVQFGNRVEARK 401
>At1g12150 hypothetical protein
Length = 548
Score = 33.5 bits (75), Expect = 0.65
Identities = 30/100 (30%), Positives = 47/100 (47%), Gaps = 14/100 (14%)
Query: 946 RRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYL---- 1001
+R EL+I +++ KSAE V + S++QE+ +Q+ S I+ T+ ++E L
Sbjct: 411 KRLELVIREVEEAKSAEEKVREEMKMISQKQESKKQDEESSGSKIK-ITIQEFESLKRGA 469
Query: 1002 --------SSLPFENFTLESLKKLEAELDEK-ENELKTLE 1032
L LE + K AE D K E LK +E
Sbjct: 470 GETEAAIEKKLATIAAELEEINKRRAEADNKLEANLKAIE 509
>At1g80790 hypothetical protein
Length = 634
Score = 33.1 bits (74), Expect = 0.85
Identities = 46/239 (19%), Positives = 100/239 (41%), Gaps = 23/239 (9%)
Query: 823 EKITARGLIESFRQKGDYEIVNFKVNVKQ--EEIATIEKDEELRKKFNLTCTISTSNMYL 880
E ++ G + SF EI N + V +IA +D + N T+S + L
Sbjct: 230 EYLSKVGKLRSFSDITKEEIQNKSIVVDDLANKIAMTNEDLNKLQYMNNEKTLSLRRV-L 288
Query: 881 FDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRI------KQKFIS 934
+ + + Y + ++E ++ R K+ L + +L+I K++ ++
Sbjct: 289 IEKDELDRVYKQETKKMQELSREKINRIFREKERLTNELEAKMNNLKIWSKQLDKKQALT 348
Query: 935 NIVNEKFNPINRRAELLIEILK----KGKSAEPHVAGAIDYSSKEQEAGEQESVSQSVSI 990
+ +K + ++++++ L+ + K + V +D +++E + + +
Sbjct: 349 ELERQKLDEDKKKSDVMNSSLQLASLEQKKTDDRVLRLVDEHKRKKEETLNKILQLEKEL 408
Query: 991 EGATLGDYEY------LSSLPFENFTLESLKK----LEAELDEKENELKTLEHTSPDLM 1039
+ E L + E+ E +KK ++ EL+EK +EL+ LE T+ LM
Sbjct: 409 DSKQKLQMEIQELKGKLKVMKHEDEDDEGIKKKMKKMKEELEEKCSELQDLEDTNSALM 467
>At1g65010 hypothetical protein
Length = 1318
Score = 32.7 bits (73), Expect = 1.1
Identities = 52/211 (24%), Positives = 87/211 (40%), Gaps = 26/211 (12%)
Query: 832 ESFRQKGDYEIVNFKVN--VKQEEI-ATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIK 888
+S QK E+ NF + +K+ E+ A + ++EEL+ K T+ T + + I
Sbjct: 954 DSLAQKKIEELSNFNASLLIKENELQAVVCENEELKSK--QVSTLKTIDELSDLKQSLIH 1011
Query: 889 KYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRA 948
K + + E +LK ++ T L ++L KQ + + +E + A
Sbjct: 1012 KEKELQAAIVE--NEKLKAEAALSLQRIEELTNLKQTLIDKQNELQGVFHENEELKAKEA 1069
Query: 949 ELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFEN 1008
L +I + EQ E+ES Q V+ E L + L++
Sbjct: 1070 SSLKKI--------------DELLHLEQSWLEKESEFQRVTQENLELKTQDALAAKK--- 1112
Query: 1009 FTLESLKKLEAELDEKENELKTLEHTSPDLM 1039
+E L KL+ L EKE ELK E + + M
Sbjct: 1113 --IEELSKLKESLLEKETELKCREAAALEKM 1141
>At4g11340 putative protein
Length = 273
Score = 32.3 bits (72), Expect = 1.5
Identities = 37/175 (21%), Positives = 74/175 (42%), Gaps = 31/175 (17%)
Query: 356 YVKD-LAMSGLSAVVDPDLYGVFPLSSKLLNATRDSR---KLFKKKEIQNLMAILGLVRN 411
++KD L +G++ V D D P+ LL +DSR +F + ++ + LV
Sbjct: 35 FLKDALIKNGINVVTDEDAPRGKPIDENLLKLIKDSRIAVVIFSENYPESTWCLDELVEI 94
Query: 412 KKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFPIIKVS 471
+K + K L + F+ +K+ S F + ++++
Sbjct: 95 EKQMDLKMLDSCPI--------------------FFEVETCHVKLQVARSTFNYNLLQL- 133
Query: 472 MYNASDSKKGKELS----FDSMQQYEDWKKELGNTANDWEIKYCQGLSSAQEVRE 522
+ KK +++S D+ +++E W+K L + A+ + Y +G + A V E
Sbjct: 134 --EHDERKKARQISKKAWEDAEKRFEGWRKALISVASRLGLTYKKGSNQATFVNE 186
>At5g66030 Golgi-localized protein GRIP
Length = 788
Score = 32.0 bits (71), Expect = 1.9
Identities = 41/189 (21%), Positives = 86/189 (44%), Gaps = 19/189 (10%)
Query: 851 QEEIATIEKDEELRKKFNLTCTISTSNMY--LFDAEGKIKKYNTPEQILEEFCTLRLKYY 908
QE++A++ ++ ++ K+ + + ++ +A+ K ++Y++ +E+ +K
Sbjct: 88 QEQVASLSREIDVEKQTRVAAEQALEHLREAYSEADAKSQEYSSKFSQVEQKLDQEIKER 147
Query: 909 VRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIEILKKGKSAEPHVAGA 968
+ L FT+L + R KQ+ I I EK + ++ R + E ++ S
Sbjct: 148 DEKYADLDAKFTRLHK--RAKQR-IQEIQKEK-DDLDARFREVNETAERASS-------- 195
Query: 969 IDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENEL 1028
+SS +QE E Q + E D E N +++++L L KEN++
Sbjct: 196 -QHSSMQQEL---ERTRQQAN-EALKAMDAERQQLRSANNKLRDTIEELRGSLQPKENKI 250
Query: 1029 KTLEHTSPD 1037
+TL+ + D
Sbjct: 251 ETLQQSLLD 259
>At5g15490 UDP-glucose dehydrogenase-like protein
Length = 480
Score = 32.0 bits (71), Expect = 1.9
Identities = 27/117 (23%), Positives = 47/117 (40%), Gaps = 18/117 (15%)
Query: 409 VRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVS 457
V Y+ K R G + ++ G+ F+ ++N Y +W ++K++
Sbjct: 244 VSEVSYAVGKDSRIGPKFLNSSVGFGGSCFQKDILNLVYICECNGLPEVAEYWKQVIKIN 303
Query: 458 KFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
+ I SM+N +KK L F +KK+ G+T I C+GL
Sbjct: 304 DYQKTRFVNRIVSSMFNTVSNKKIAVLGF-------AFKKDTGDTRETPAIDVCKGL 353
>At1g49860 glutathione S-transferase, putative
Length = 254
Score = 31.6 bits (70), Expect = 2.5
Identities = 18/67 (26%), Positives = 36/67 (52%), Gaps = 1/67 (1%)
Query: 981 QESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDL-M 1039
+E +S+ ++I LG+ YL+ F L L ++ L+ E ELK L ++ P++
Sbjct: 140 KEKLSEVLNIYETRLGESPYLAGESFSLADLHHLAPIDYLLNTDEEELKNLIYSRPNVAA 199
Query: 1040 WLNDLEL 1046
W+ +++
Sbjct: 200 WVEKMKM 206
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 30.8 bits (68), Expect = 4.2
Identities = 25/116 (21%), Positives = 48/116 (40%), Gaps = 3/116 (2%)
Query: 924 RSLRIKQKFISNIVNEKFNPINRRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQES 983
+SL +K IS+++ + I L E+ +K K E + ++ + +
Sbjct: 9 KSLSLKVSEISDVIQQGQTTIQELISELGEMKEKYKEKESEHSSLVELHKTHERESSSQV 68
Query: 984 VSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLM 1039
IE + ++ SL N E K L ++ E NE++ ++T +LM
Sbjct: 69 KELEAHIESSEKLVADFTQSL---NNAEEEKKLLSQKIAELSNEIQEAQNTMQELM 121
>At5g39320 UDP-glucose dehydrogenase-like protein
Length = 480
Score = 30.8 bits (68), Expect = 4.2
Identities = 33/148 (22%), Positives = 57/148 (38%), Gaps = 31/148 (20%)
Query: 421 RYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVSKFMSVFTFPIIK 469
R G + A+ G+ F+ ++N Y +W ++K++ + I
Sbjct: 256 RIGSKFLNASVGFGGSCFQKDILNLVYICQCNGLPEVAEYWKQVIKINDYQKNRFVNRIV 315
Query: 470 VSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGLSSAQEVREYFQDLGK 529
SM+N +KK L F +KK+ G+T I C+GL LG
Sbjct: 316 SSMFNTVSNKKVAILGF-------AFKKDTGDTRETPAIDVCKGL------------LGD 356
Query: 530 HRAYFVWDDK-DETSIKLAFSKKAEEWE 556
++D + E I+ S K +W+
Sbjct: 357 KAQISIYDPQVTEEQIQRDLSMKKFDWD 384
>At1g68910 unknown protein
Length = 534
Score = 30.8 bits (68), Expect = 4.2
Identities = 45/217 (20%), Positives = 87/217 (39%), Gaps = 43/217 (19%)
Query: 752 EIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPI 811
E+ K + F NEE + +K Y E + R++E
Sbjct: 197 ELEKKLMEFQQNEEQLKLKLHYT-------------------------EEVSSRMEEASE 231
Query: 812 RMWTEDYKKFLEKITARGLIESFRQK--GDYEIVNFKVNVKQEEIATIEKDEELRKKF-N 868
+W +FLE + ++ ++ G +I+ F +N + +++ EL+ K +
Sbjct: 232 FIWG----RFLEADNSSEVLTGISKELVGRLQILQFSLN------GSAQRESELKSKLED 281
Query: 869 LTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRI 928
T + ++ + EG I + + +I+ E TLR YV+ + +KN L+S+
Sbjct: 282 CTVQLEAKDLLVQKLEGTISENS---EIVSEVLTLR--EYVKSAEQKLKNTDLELKSVNA 336
Query: 929 KQKFISNIVNEKFNPINRRAELLIEILKKGKSAEPHV 965
++ I + E N E L E + +S E +
Sbjct: 337 SKQEILVHLAEMENANESVKENLFEAESRAESGEAKI 373
>At5g14580 polynucleotide phosphorylase
Length = 991
Score = 30.4 bits (67), Expect = 5.5
Identities = 17/50 (34%), Positives = 27/50 (54%)
Query: 357 VKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRKLFKKKEIQNLMAIL 406
V DLA + + +V YG F L N +D RK+F+++ Q ++IL
Sbjct: 293 VADLAATRIESVFTDPSYGKFERGEALDNIGKDVRKVFEEEGDQESLSIL 342
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.139 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,112,536
Number of Sequences: 26719
Number of extensions: 1008069
Number of successful extensions: 2768
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 2731
Number of HSP's gapped (non-prelim): 41
length of query: 1053
length of database: 11,318,596
effective HSP length: 109
effective length of query: 944
effective length of database: 8,406,225
effective search space: 7935476400
effective search space used: 7935476400
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC145202.5


