

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.10 + phase: 0
(235 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
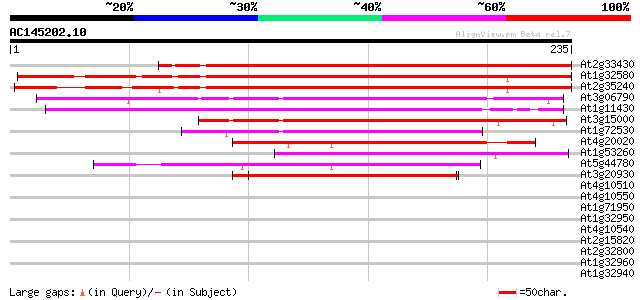
Score E
Sequences producing significant alignments: (bits) Value
At2g33430 plastid protein 293 4e-80
At1g32580 unknown protein, like plastid protein 282 1e-76
At2g35240 unknown protein 281 3e-76
At3g06790 DAG protein, putative 142 2e-34
At1g11430 unknown protein 139 2e-33
At3g15000 unknown protein 137 6e-33
At1g72530 DAG-like protein 106 1e-23
At4g20020 DAG-like protein 105 2e-23
At1g53260 DAG-like protein emb|CAA16610.1 99 2e-21
At5g44780 unknown protein 88 4e-18
At3g20930 unknown protein 84 6e-17
At4g10510 putative subtilisin-like protease 37 0.007
At4g10550 subtilisin-like protease -like protein 36 0.015
At1g71950 unknown protein 35 0.026
At1g32950 putative serine proteinase (At1g32950) 35 0.026
At4g10540 putative subtilisin-like protease 35 0.034
At2g15820 unknown protein 35 0.034
At2g32800 putative protein kinase 34 0.059
At1g32960 unknown protein 33 0.13
At1g32940 unknown protein 31 0.50
>At2g33430 plastid protein
Length = 219
Score = 293 bits (751), Expect = 4e-80
Identities = 139/173 (80%), Positives = 155/173 (89%), Gaps = 3/173 (1%)
Query: 63 GIHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMID 122
G +R+G YSPL GSN FSDRPP EMAPLFPGCDY HWLI++DKPGGEGATKQQMID
Sbjct: 50 GANRSG-STYSPLNSGSN--FSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMID 106
Query: 123 CYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDP 182
CY++TLA+V+GSEEEAKK+IYNVSCERY GFGCE+DEETS KLEG+PGVLFVLPDSYVDP
Sbjct: 107 CYIQTLAKVVGSEEEAKKRIYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDP 166
Query: 183 EHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRDNQR 235
E++DYGAELFVNGEIVQRSPERQRRVEPQ QR RPRY+DRT+Y RRR+N R
Sbjct: 167 ENKDYGAELFVNGEIVQRSPERQRRVEPQPQRAQDRPRYNDRTRYSRRRENTR 219
>At1g32580 unknown protein, like plastid protein
Length = 229
Score = 282 bits (721), Expect = 1e-76
Identities = 145/233 (62%), Positives = 177/233 (75%), Gaps = 11/233 (4%)
Query: 4 RAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGG 63
R+ AS +RL ST+ +T+ S S + S P + FI +++ F ++
Sbjct: 7 RSTASRITKRLISTSGATTPSPSY----ILSRRSTPVFSHAVGFISSLNR--FTTIRTRM 60
Query: 64 IHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDC 123
G +YSPL GSN FSDR P EMAPLFPGCDY HWLI++DKPGGE ATKQQMIDC
Sbjct: 61 DRSGG--SYSPLKSGSN--FSDRAPTEMAPLFPGCDYEHWLIVMDKPGGENATKQQMIDC 116
Query: 124 YVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPE 183
YV+TLA+++GSEEEAKKKIYNVSCERYFGFGCE+DEETSNKLEG+PGVLF+LPDSYVD E
Sbjct: 117 YVQTLAKIIGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFILPDSYVDQE 176
Query: 184 HQDYGAELFVNGEIVQRSPERQRR-VEPQAQRGDSRPRYHDRTKYVRRRDNQR 235
++DYGAELFVNGEIVQR PERQR+ +E QR + +P+YHD+T+YVRRR+N R
Sbjct: 177 NKDYGAELFVNGEIVQRPPERQRKIIELTTQRTNDKPKYHDKTRYVRRRENMR 229
>At2g35240 unknown protein
Length = 232
Score = 281 bits (718), Expect = 3e-76
Identities = 150/245 (61%), Positives = 177/245 (72%), Gaps = 30/245 (12%)
Query: 3 TRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHF- 61
+R+ AS +R FST+ + VA PS L S RF S TIF ++ +
Sbjct: 6 SRSTASCVAKRFFSTSNA-----------VASPSPLPSHLISRRF----SPTIFHAVGYI 50
Query: 62 ----------GGIHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPG 111
+ R+G +YSPL GSN FSDRPP EMAPLFPGCDY HWLI+++KPG
Sbjct: 51 PALTRFTTIRTRMDRSGG-SYSPLKSGSN--FSDRPPTEMAPLFPGCDYEHWLIVMEKPG 107
Query: 112 GEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGV 171
GE A KQQMIDCYV+TLA+++GSEEEA+KKIYNVSCERYFGFGCE+DEETSNKLEG+PGV
Sbjct: 108 GENAQKQQMIDCYVQTLAKIVGSEEEARKKIYNVSCERYFGFGCEIDEETSNKLEGLPGV 167
Query: 172 LFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRR-VEPQAQRGDSRPRYHDRTKYVRR 230
LFVLPDSYVDPE +DYGAELFVNGE+V R PERQRR VE QRG +P+YHDR + VRR
Sbjct: 168 LFVLPDSYVDPEFKDYGAELFVNGEVVPRPPERQRRMVELTNQRGSDKPKYHDRIRNVRR 227
Query: 231 RDNQR 235
R+N R
Sbjct: 228 RENMR 232
>At3g06790 DAG protein, putative
Length = 244
Score = 142 bits (357), Expect = 2e-34
Identities = 90/224 (40%), Positives = 128/224 (56%), Gaps = 8/224 (3%)
Query: 12 RRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFI-IPMSQTIFQSLHFGGIHRAGYC 70
RR ST + + S+ST ++ LS S P + S+T + +
Sbjct: 7 RRTLSTLLNKTLSSSTSYSSSFPTLSSRSRFAMPLIEKVSSSRTSLGPCYISTRPKTSGS 66
Query: 71 NYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQ 130
YSPL S ++S+RPP E L GCDY HWLI+++ + T+++MI+ YVKTL
Sbjct: 67 GYSPLNDPS-PNWSNRPPKETI-LLDGCDYEHWLIVMEFTDPK-PTEEEMINSYVKTLTS 123
Query: 131 VLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAE 190
VLG +EEAKKKIY+V Y GFG + EE S K++ +PGVL+VLPDSY+D ++DYG +
Sbjct: 124 VLGWQEEAKKKIYSVCTSTYTGFGALISEELSCKVKALPGVLWVLPDSYLDVPNKDYGGD 183
Query: 191 LFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDR--TKYVRRRD 232
L+V G+++ R + R E + R RPRY R T V RR+
Sbjct: 184 LYVEGKVIPR--PQYRFTEQRHTRPRPRPRYDRRRETMQVERRE 225
>At1g11430 unknown protein
Length = 232
Score = 139 bits (349), Expect = 2e-33
Identities = 80/217 (36%), Positives = 122/217 (55%), Gaps = 7/217 (3%)
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGGIHRAGYCNYSPL 75
S TT++S+S T L++ S+L+ R + + G R + +
Sbjct: 3 SFTTTSSSSLLLKTLLPVSHLNRFSTLSGIRVGDSWTPLLRNISTAGSRRRVAIVKAATV 62
Query: 76 VPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSE 135
++ S+ + PGCDYNHWLI+++ P ++ QMID Y+ TLA VLGS
Sbjct: 63 DSDYSSKRSNSNEQRETIMLPGCDYNHWLIVMEFPKDPAPSRDQMIDTYLNTLATVLGSM 122
Query: 136 EEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNG 195
EEAKK +Y S Y GF C +DEETS K +G+PGVL+VLPDSY+D +++DYG + ++NG
Sbjct: 123 EEAKKNMYAFSTTTYTGFQCTIDEETSEKFKGLPGVLWVLPDSYIDVKNKDYGGDKYING 182
Query: 196 EIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRD 232
EI+ P +P+ QR +++ + +Y R+RD
Sbjct: 183 EII---PCTYPTYQPK-QRNNTK---YQSKRYERKRD 212
>At3g15000 unknown protein
Length = 395
Score = 137 bits (344), Expect = 6e-33
Identities = 69/161 (42%), Positives = 107/161 (65%), Gaps = 9/161 (5%)
Query: 80 NTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAK 139
N ++S+RPP E L GCD+ HWL++++ P GE T+ ++ID Y+KTLAQ++GSE+EA+
Sbjct: 78 NPNWSNRPPKETI-LLDGCDFEHWLVVVEPPQGE-PTRDEIIDSYIKTLAQIVGSEDEAR 135
Query: 140 KKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQ 199
KIY+VS Y+ FG + E+ S+KL+ + V +VLPDSY+D ++DYG E F++G+ V
Sbjct: 136 MKIYSVSTRCYYAFGALVSEDLSHKLKELSNVRWVLPDSYLDVRNKDYGGEPFIDGKAVP 195
Query: 200 RSPE------RQRRVEPQAQRGDSRPRYHDRTK-YVRRRDN 233
P+ R + R + RPR +DR++ + RRR+N
Sbjct: 196 YDPKYHEEWIRNNARANERNRRNDRPRNNDRSRNFERRREN 236
>At1g72530 DAG-like protein
Length = 188
Score = 106 bits (264), Expect = 1e-23
Identities = 55/133 (41%), Positives = 77/133 (57%), Gaps = 8/133 (6%)
Query: 73 SPLVPGSNTSFSDRPPA-------EMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYV 125
SPL P S S S + L GCDY HWL+++ P G T+ ++ +V
Sbjct: 20 SPLSPFSGNSGSINSETTSWSELIRVPSLVEGCDYKHWLVLMKPPNGY-PTRNHIVQSFV 78
Query: 126 KTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQ 185
+TLA LGSEEEAK+ IY+VS + Y+ FGC + E + K+ +P V +VLPDS++
Sbjct: 79 ETLAMALGSEEEAKRSIYSVSTKYYYAFGCRIHEPLTYKIRSLPDVKWVLPDSFIVDGDN 138
Query: 186 DYGAELFVNGEIV 198
YG E FV+GE+V
Sbjct: 139 RYGGEPFVDGEVV 151
>At4g20020 DAG-like protein
Length = 406
Score = 105 bits (262), Expect = 2e-23
Identities = 56/129 (43%), Positives = 79/129 (60%), Gaps = 10/129 (7%)
Query: 94 LFPGCDYNHWLIIIDKPGGEGA-TKQQMIDCYVKTLAQVLG-SEEEAKKKIYNVSCERYF 151
LF GCDYNHWLI +D E + ++M+ Y +T AQ LG S EEAK+++Y S Y
Sbjct: 81 LFEGCDYNHWLITMDFSKEETPKSPEEMVAAYEETCAQGLGISVEEAKQRMYACSTTTYQ 140
Query: 152 GFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQ 211
GF + E+ S K + +PGV+F+LPDSY+DP++++YG + + NG I R P
Sbjct: 141 GFQAIMTEQESEKFKDLPGVVFILPDSYIDPQNKEYGGDKYENGVITHR--------PPP 192
Query: 212 AQRGDSRPR 220
Q G +RPR
Sbjct: 193 IQSGRARPR 201
>At1g53260 DAG-like protein emb|CAA16610.1
Length = 358
Score = 99.0 bits (245), Expect = 2e-21
Identities = 55/138 (39%), Positives = 80/138 (57%), Gaps = 15/138 (10%)
Query: 112 GEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGV 171
G + ++ID Y+KTLAQV+GSEEEA+ KIY+VS + YF FG + E+ S+K++ +P V
Sbjct: 60 GVSLARDEIIDYYIKTLAQVVGSEEEARMKIYSVSHKCYFAFGALVSEDLSHKIKELPKV 119
Query: 172 LFVLPDSYVDPEHQDYGAELFVNGEIVQRSP---------------ERQRRVEPQAQRGD 216
+VLPDSY+D +++DYG E F++G+ V P E +R P+ G
Sbjct: 120 KWVLPDSYLDGKNKDYGGEPFIDGKAVPYDPKYHEEWIRNNANATNENRRPRRPRNSDGG 179
Query: 217 SRPRYHDRTKYVRRRDNQ 234
R + T Y R NQ
Sbjct: 180 RNDRGNQDTGYRRPPPNQ 197
>At5g44780 unknown protein
Length = 723
Score = 87.8 bits (216), Expect = 4e-18
Identities = 55/166 (33%), Positives = 83/166 (49%), Gaps = 14/166 (8%)
Query: 36 LSKPSSLTSPRFIIPMSQTIFQSLHFGGIHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLF 95
LS+PS T ++P S + ++ N SP S T + P
Sbjct: 25 LSRPSDFTPVPSLLPRSV----------VKQSTAINRSPARLFSTTQYQYDPYTGEDSFM 74
Query: 96 P---GCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLG-SEEEAKKKIYNVSCERYF 151
P GCD+NHWLI ++ P ++++MI + +T A+ L S EEAKKKIY + Y
Sbjct: 75 PDNEGCDFNHWLITMNFPKDNLPSREEMISIFEQTCAKGLAISLEEAKKKIYAICTTSYQ 134
Query: 152 GFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEI 197
GF + K +PGV +++PDSY+D E++ YG + + NG I
Sbjct: 135 GFQATMTIGEVEKFRDLPGVQYIIPDSYIDVENKVYGGDKYENGVI 180
>At3g20930 unknown protein
Length = 374
Score = 84.0 bits (206), Expect = 6e-17
Identities = 41/94 (43%), Positives = 64/94 (67%)
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
LF HW++ IDKPG TK QM+D V+ L++VL +E++A+ +Y+VS + FGF
Sbjct: 167 LFDHGTVKHWMVRIDKPGVGIVTKAQMVDHCVQLLSKVLWNEKDAQMCLYHVSWQSDFGF 226
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDY 187
C+LDE ++ +L G+PGVL V+PD+ + ++DY
Sbjct: 227 CCDLDERSAVELAGVPGVLAVVPDNSFESLNKDY 260
Score = 82.0 bits (201), Expect = 2e-16
Identities = 34/88 (38%), Positives = 61/88 (68%)
Query: 101 NHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEE 160
++W++++DKP ++K M+D YV+ LA+VLG+E++A+ IY+ S + +FGF C +DE+
Sbjct: 71 SYWMVLLDKPPHWVSSKSAMVDYYVEILAKVLGNEKDAQVSIYDASFDTHFGFCCHIDED 130
Query: 161 TSNKLEGIPGVLFVLPDSYVDPEHQDYG 188
S +L +PGV+ + P+ E ++YG
Sbjct: 131 ASRQLASLPGVVSIRPEQDYSSEKKNYG 158
>At4g10510 putative subtilisin-like protease
Length = 765
Score = 37.4 bits (85), Expect = 0.007
Identities = 18/57 (31%), Positives = 31/57 (53%)
Query: 126 KTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDP 182
+ L +LGS+EEA + + + GF +L E + K+ +P V+ V+PD + P
Sbjct: 44 RMLWSLLGSKEEAHGSMVHSFRHGFSGFAAKLTESQAKKIADLPEVVHVIPDRFYKP 100
>At4g10550 subtilisin-like protease -like protein
Length = 778
Score = 36.2 bits (82), Expect = 0.015
Identities = 17/54 (31%), Positives = 30/54 (55%)
Query: 126 KTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSY 179
+ L +LGS+E+A + + GF +L E + K+ +P V+ V+PDS+
Sbjct: 56 RMLWSLLGSKEDANDSMVYSYRHGFSGFAAKLTESQAKKIADLPDVVHVIPDSF 109
>At1g71950 unknown protein
Length = 136
Score = 35.4 bits (80), Expect = 0.026
Identities = 21/64 (32%), Positives = 33/64 (50%)
Query: 113 EGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVL 172
E T ++ +++TL+ LGSEE AK + E GF +L E ++ PGV+
Sbjct: 55 EKPTDEEPKTYHLRTLSSALGSEEAAKDALIYSYKEAASGFSAKLTPEQVAEISKQPGVI 114
Query: 173 FVLP 176
V+P
Sbjct: 115 QVVP 118
>At1g32950 putative serine proteinase (At1g32950)
Length = 763
Score = 35.4 bits (80), Expect = 0.026
Identities = 16/54 (29%), Positives = 31/54 (56%)
Query: 128 LAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVD 181
L+ +LGS+++A + + + GF +L + + K+ P V+ V+PDSY +
Sbjct: 53 LSSLLGSKDDAHESMVYSYRHGFSGFAAKLTKSQAKKIADSPEVIHVIPDSYYE 106
>At4g10540 putative subtilisin-like protease
Length = 775
Score = 35.0 bits (79), Expect = 0.034
Identities = 17/54 (31%), Positives = 30/54 (55%)
Query: 126 KTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSY 179
+ L +LGS+E+A + + + GF +L + + KL +P V+ V PDS+
Sbjct: 52 RMLWSLLGSKEDAHSSMVHSYRHGFSGFAAKLTKSQAKKLADLPEVVHVTPDSF 105
>At2g15820 unknown protein
Length = 849
Score = 35.0 bits (79), Expect = 0.034
Identities = 31/88 (35%), Positives = 43/88 (48%), Gaps = 10/88 (11%)
Query: 15 FSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGGI-HRAGYCNYS 73
FS++ S ++S+S+ TV+ SSL+S II S T+F+SL F I HR+ Y S
Sbjct: 19 FSSSFSLASSSSS---TVSVTTFNISSLSSNPNIINSSSTLFRSLSFSLIRHRSSYSRRS 75
Query: 74 ------PLVPGSNTSFSDRPPAEMAPLF 95
V G+ T F PLF
Sbjct: 76 LRRLSIHTVHGNKTQFFSHSSTRTPPLF 103
>At2g32800 putative protein kinase
Length = 848
Score = 34.3 bits (77), Expect = 0.059
Identities = 24/85 (28%), Positives = 39/85 (45%), Gaps = 3/85 (3%)
Query: 8 SSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGGIHRA 67
SS K S TT+T+ +T T TT+ + S S ++ + +I+Q+ G
Sbjct: 437 SSLKSTSTSATTTTTRTTMTTTTSTTSFNASSESTPSSNYVTALEDSIYQTAETG---EN 493
Query: 68 GYCNYSPLVPGSNTSFSDRPPAEMA 92
Y NY+ S+ SF P E++
Sbjct: 494 PYFNYNSRRVMSSKSFVLDTPREIS 518
>At1g32960 unknown protein
Length = 777
Score = 33.1 bits (74), Expect = 0.13
Identities = 15/52 (28%), Positives = 29/52 (54%)
Query: 128 LAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSY 179
LA +LGS+++A + + GF +L + + K+ +P V+ V+PD +
Sbjct: 56 LASLLGSKKDADDSMVYSYRHGFSGFAAKLTKSQAKKIADLPEVVHVIPDGF 107
>At1g32940 unknown protein
Length = 774
Score = 31.2 bits (69), Expect = 0.50
Identities = 16/54 (29%), Positives = 29/54 (53%)
Query: 128 LAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVD 181
L+ +LGS+ +A + + + GF +L E + KL P V+ V+ DS+ +
Sbjct: 53 LSSLLGSKVDAHESMVYSYRHGFSGFAAKLTESQAKKLADSPEVVHVMADSFYE 106
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,543,652
Number of Sequences: 26719
Number of extensions: 251782
Number of successful extensions: 890
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 841
Number of HSP's gapped (non-prelim): 40
length of query: 235
length of database: 11,318,596
effective HSP length: 96
effective length of query: 139
effective length of database: 8,753,572
effective search space: 1216746508
effective search space used: 1216746508
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC145202.10


