

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
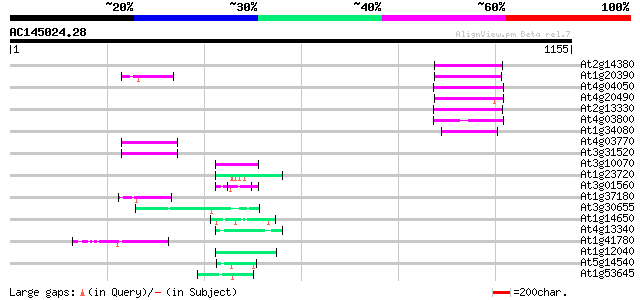
Query= AC145024.28 - phase: 0
(1155 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g14380 putative retroelement pol polyprotein 80 7e-15
At1g20390 hypothetical protein 79 1e-14
At4g04050 putative transposon protein 75 3e-13
At4g20490 putative protein 73 8e-13
At2g13330 F14O4.9 71 4e-12
At4g03800 65 2e-10
At1g34080 putative protein 58 4e-08
At4g03770 hypothetical protein 56 1e-07
At3g31520 hypothetical protein 52 3e-06
At3g10070 unknown protein 51 4e-06
At1g23720 unknown protein 50 7e-06
At3g01560 unknown protein 50 1e-05
At1g37180 hypothetical protein 49 2e-05
At3g30655 hypothetical protein, 3' partial 48 3e-05
At1g14650 splicing factor, putative 47 5e-05
At4g13340 extensin-like protein 47 8e-05
At1g41780 hypothetical protein 47 8e-05
At1g12040 leucine-rich repeat/extensin 1 (LRX1) 47 8e-05
At5g14540 unknown protein 46 1e-04
At1g53645 unknown protein 46 1e-04
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 80.1 bits (196), Expect = 7e-15
Identities = 45/140 (32%), Positives = 73/140 (52%)
Query: 875 LTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLR 934
LTF+ EDL HN L+I + ++ +LIDTGS++NV+ K L ++ + ++
Sbjct: 330 LTFTSEDLFGVDLPHNDPLVIELHIGESEVTRILIDTGSSVNVVFKDVLQKMKVHDRHIK 389
Query: 935 LSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVT 994
S + FDG G + LPI +G F V++ ++ +LG PWIHD A+
Sbjct: 390 PSVRPLTGFDGNTMMTNGTIKLPIYLGGAATWHKFVVVDKPTIYNIILGTPWIHDMQAIP 449
Query: 995 STLHQKLKFVSRGKLITVSG 1014
S+ HQ +K + + T+ G
Sbjct: 450 SSYHQCIKIPTSIGIETIRG 469
>At1g20390 hypothetical protein
Length = 1791
Score = 79.3 bits (194), Expect = 1e-14
Identities = 43/138 (31%), Positives = 73/138 (52%)
Query: 875 LTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLR 934
++F+D DL HN L++ ++ ++ VLIDTGS+++++ K L + ++ ++
Sbjct: 553 ISFTDVDLEGLDTPHNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIK 612
Query: 935 LSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVT 994
+ FDG G + LPI VG V F V+ A ++ +LG PWIH A+
Sbjct: 613 PVSKPLAGFDGDFVMTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIP 672
Query: 995 STLHQKLKFVSRGKLITV 1012
ST HQ +KF + + T+
Sbjct: 673 STYHQCVKFPTHNGIFTL 690
Score = 49.7 bits (117), Expect = 1e-05
Identities = 36/117 (30%), Positives = 50/117 (41%), Gaps = 14/117 (11%)
Query: 230 EKYQGDTCPQNHLTMYISKMIAYKNNVPLLIHCFQ--------DSLTGPAHTWFMGLK-- 279
E Y G P+ LT + + N L I F + LTGPAH WF LK
Sbjct: 175 ESYNGRNDPKEFLTSFNVAI----NRAELTIDNFDAGRCQIFIEHLTGPAHNWFSRLKPN 230
Query: 280 GVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPP 336
+ +F QL +F++ Y S +L S++Q KES + + RF I P
Sbjct: 231 SIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFVDRFKLVVTNITVP 287
>At4g04050 putative transposon protein
Length = 681
Score = 74.7 bits (182), Expect = 3e-13
Identities = 43/145 (29%), Positives = 74/145 (50%)
Query: 873 NSLTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAP 932
+ +TF + + H+ AL+I + LS+++IDTGS+++V+ ++ + ++
Sbjct: 519 HQITFWESETTDLNKPHDDALVIRIDVGNYELSHIMIDTGSSVDVLFYDAFKRMGHLDSE 578
Query: 933 LRLSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGA 992
L+ K + F G G + LP +F V+ +A F+ +LGRPW+H A
Sbjct: 579 LQGRKTPLTGFAGDTTFSLGTIQLPTIARGVRRLTSFLVVNKKAPFNAILGRPWLHAMKA 638
Query: 993 VTSTLHQKLKFVSRGKLITVSGESA 1017
V ST HQ +KF S + VS +A
Sbjct: 639 VPSTYHQCIKFPSDKGIAVVSAANA 663
>At4g20490 putative protein
Length = 853
Score = 73.2 bits (178), Expect = 8e-13
Identities = 43/157 (27%), Positives = 75/157 (47%), Gaps = 14/157 (8%)
Query: 875 LTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLR 934
+ F+ ED + HN AL+I ++ + +LIDTGS++N+M TL ++ SE+ +
Sbjct: 463 IIFTSEDASGIQHPHNDALVIKLVMEDFDVERILIDTGSSVNLMFLKTLFKMGISESKIM 522
Query: 935 LSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVT 994
+ +DG + GE+ L + VG + F V++ + ++ +LG PWI+ A+
Sbjct: 523 PKIRPMTGYDGEAKMSIGEIKLQVQVGGITQKTKFVVIDSEPIYNAILGSPWIYSMKAIP 582
Query: 995 ST--------------LHQKLKFVSRGKLITVSGESA 1017
ST L + + G LI + G A
Sbjct: 583 STNLHTIRQPEGLTDMLRDREEAAKEGNLIAIKGRRA 619
Score = 42.0 bits (97), Expect = 0.002
Identities = 24/66 (36%), Positives = 37/66 (55%), Gaps = 2/66 (3%)
Query: 263 FQDSLTGPAHTWFMGLKG--VTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFK 320
F + LTG A T F L+ + F QL+ AF++QY+ S +L S+TQ++ E+
Sbjct: 225 FVEGLTGNALTCFSRLEANSIDNFTQLSTAFLKQYRVFIQPGASSSDLWSMTQENGETLN 284
Query: 321 EYAQRF 326
+Y RF
Sbjct: 285 DYLGRF 290
>At2g13330 F14O4.9
Length = 889
Score = 70.9 bits (172), Expect = 4e-12
Identities = 41/142 (28%), Positives = 71/142 (49%)
Query: 873 NSLTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAP 932
+ +TF + ++ H+ AL+I + LS +++DTGS+++V+ + + ++
Sbjct: 45 DGITFWESEITDLDKPHDDALVIRIDVGNYELSCIMVDTGSSVDVLFYDAFKRTGHLDSK 104
Query: 933 LRLSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGA 992
L+ K + F G G + LP F V++ +A F+ +LGRPW+H A
Sbjct: 105 LQGRKTPLTGFAGDTTFSIGTIQLPTIARGVRQLTNFLVVDKKAPFNAILGRPWLHVMKA 164
Query: 993 VTSTLHQKLKFVSRGKLITVSG 1014
V ST HQ +KF S + V G
Sbjct: 165 VPSTYHQCIKFPSYKGIAVVYG 186
>At4g03800
Length = 637
Score = 65.1 bits (157), Expect = 2e-10
Identities = 41/144 (28%), Positives = 69/144 (47%), Gaps = 14/144 (9%)
Query: 872 CNSLTFSDEDLPAEGNKHNQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEA 931
C+S++F +E+ H+ AL+I++ +S +L+DTGS+++++ L
Sbjct: 182 CSSVSFDEEETRHIERPHDDALIITLDVANFKISRILVDTGSSVDLIFLGPPSPLV---- 237
Query: 932 PLRLSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAG 991
AF G + LP+ G V F V + A+++ +LG PWI
Sbjct: 238 ----------AFTSESAMSLGTIKLPVLAGKMSKIVDFVVFDKPATYNIILGTPWIFQMK 287
Query: 992 AVTSTLHQKLKFVSRGKLITVSGE 1015
AV ST HQ LKF + + T+ G+
Sbjct: 288 AVPSTYHQWLKFPTSNGVETIWGD 311
>At1g34080 putative protein
Length = 811
Score = 57.8 bits (138), Expect = 4e-08
Identities = 32/114 (28%), Positives = 63/114 (55%)
Query: 890 NQALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVRAFDGTRRS 949
+ AL+IS+ + +LIDTGS+++++ TL ++ S+ ++ + + +F
Sbjct: 474 DDALVISLDVGNCEVQRILIDTGSSVDLIFLDTLVRMGISKKDIKGAPSPLVSFTIETSM 533
Query: 950 VYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKF 1003
G + LP++ + F V + A+++ +LG PW+++ AV ST HQ +KF
Sbjct: 534 SLGTITLPVTAQGVVKMIEFTVFDRPAAYNIILGTPWLYEMKAVPSTYHQCVKF 587
Score = 30.0 bits (66), Expect = 8.0
Identities = 15/53 (28%), Positives = 28/53 (52%)
Query: 281 VTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQI 333
+ F+QL+ AF++QY+Y + +L +L+Q E + Y F + Q+
Sbjct: 237 IDNFKQLSTAFIKQYEYFIKSDVTEAQLWNLSQAADEPLRTYNTAFKEIMVQV 289
>At4g03770 hypothetical protein
Length = 464
Score = 55.8 bits (133), Expect = 1e-07
Identities = 37/122 (30%), Positives = 60/122 (48%), Gaps = 6/122 (4%)
Query: 230 EKYQGDTCPQNHLTMYISKMIAYK----NNVPLLIHCFQDSLTGPAHTWFMGLKG--VTT 283
E + G + P+ HL ++I + K L H F + L GPA WF LKG V +
Sbjct: 237 EYFNGSSDPKGHLKLFIISVARAKFRPEERDAGLCHLFVEHLKGPALNWFTRLKGNSVDS 296
Query: 284 FEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPPLDERELS 343
F++L+ F++QY + S +L SL+Q+ E +++ +F A++ D LS
Sbjct: 297 FQELSTLFLKQYSVLIDPSTSDADLWSLSQQPNEPLRDFLAKFRSTLAKVEGINDLAALS 356
Query: 344 DL 345
L
Sbjct: 357 TL 358
>At3g31520 hypothetical protein
Length = 528
Score = 51.6 bits (122), Expect = 3e-06
Identities = 35/122 (28%), Positives = 58/122 (46%), Gaps = 6/122 (4%)
Query: 230 EKYQGDTCPQNHLTMYISKMIAYK----NNVPLLIHCFQDSLTGPAHTWFMGLKG--VTT 283
E + G + P+ HL +I + K L H F + L GPA WF L+G + +
Sbjct: 245 EYFNGSSDPKRHLKSFIISVARTKFRPEERDAGLCHLFVEHLKGPALCWFSRLEGNSMDS 304
Query: 284 FEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPPLDERELS 343
F++L+ F++QY L S +L SL+Q+ E +++ + A++ D LS
Sbjct: 305 FQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKLRSTLAKVEGINDVAALS 364
Query: 344 DL 345
L
Sbjct: 365 AL 366
>At3g10070 unknown protein
Length = 539
Score = 50.8 bits (120), Expect = 4e-06
Identities = 28/88 (31%), Positives = 38/88 (42%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P P+ S + + Q N + S PQG P PP Y+P N + Q+P+ PQQ
Sbjct: 255 PSPISSPNVQSTGASQQSLQAINQPWLSSTPQGKPPLPPPSYRPQVNSPSMQQRPHIPQQ 314
Query: 485 PYQQRPYNPPQQQPRPQAPYNQQNQKQQ 512
P QQ + Q + Q Q QQ
Sbjct: 315 HISTSAATPQPQQQQSQQQHQPQEQLQQ 342
>At1g23720 unknown protein
Length = 895
Score = 50.1 bits (118), Expect = 7e-06
Identities = 46/164 (28%), Positives = 62/164 (37%), Gaps = 30/164 (18%)
Query: 425 PQPVYSNHP--FVANITPQ--MTAPQNPNYQSPRP-----QGPAPYYPPLYQPLYN---- 471
P+PVY + P +V N P + P Y+SP P P PYY P +P Y
Sbjct: 325 PKPVYKSPPPPYVYNSPPPPYYSPSPKPAYKSPPPPYVYSSPPPPYYSPSPKPTYKSPPP 384
Query: 472 --LQQLSQQPYYPQQP---YQQRP----YN---PPQQQPRPQAPYNQQNQKQQFDPLPMT 519
+ PYY P Y+ P YN PP P P+ Y + P
Sbjct: 385 PYVYSSPPPPYYSPSPKPVYKSPPPPYIYNSPPPPYYSPSPKPSYKSPPPPYVYSSPPPP 444
Query: 520 YGALLPSLLAQNLVQTIPPPRI-PDPLPRWYRPDLHCIYHQGAP 562
Y + P L ++ PPP + P P +Y P +Y P
Sbjct: 445 YYSPSPKL----TYKSSPPPYVYSSPPPPYYSPSPKVVYKSPPP 484
Score = 47.4 bits (111), Expect = 5e-05
Identities = 43/162 (26%), Positives = 63/162 (38%), Gaps = 26/162 (16%)
Query: 425 PQPVYSNHP----FVANITPQMTAPQNPNYQSPRP-----QGPAPYYPPLYQPLYN---- 471
P+PVY + P + + P + P Y+SP P P PYY P +P+Y
Sbjct: 225 PKPVYKSPPPPYVYSSPPPPYYSPSPKPAYKSPPPPYVYSSPPPPYYSPSPKPIYKSPPP 284
Query: 472 --LQQLSQQPYY---PQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQF--DPLPMTYGALL 524
+ PYY P+ Y+ P PP P PY + K + P P Y +
Sbjct: 285 PYVYNSPPPPYYSPSPKPAYKSPP--PPYVYSFPPPPYYSPSPKPVYKSPPPPYVYNSPP 342
Query: 525 PSLLAQN---LVQTIPPPRI-PDPLPRWYRPDLHCIYHQGAP 562
P + + ++ PPP + P P +Y P Y P
Sbjct: 343 PPYYSPSPKPAYKSPPPPYVYSSPPPPYYSPSPKPTYKSPPP 384
Score = 45.1 bits (105), Expect = 2e-04
Identities = 41/155 (26%), Positives = 56/155 (35%), Gaps = 28/155 (18%)
Query: 425 PQPVYSNHPFVANITPQMT----APQNPNYQSPRPQ-----GPAPYY-----PPLYQPLY 470
P P YS+ P + +P +P P Y SP P+ P PY PP Y P
Sbjct: 543 PPPCYSHSPKIEYKSPPTPYVYHSPPPPPYYSPSPKPAYKSSPPPYVYSSPPPPYYSP-- 600
Query: 471 NLQQLSQQPYY--PQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLL 528
+ +P Y P PY PP P P+ Y + P Y + P
Sbjct: 601 -----APKPVYKSPPPPYVYNSPPPPYYSPSPKPTYKSPPPPYVYSSPPPPYYSPTP--- 652
Query: 529 AQNLVQTIPPPRI-PDPLPRWYRPDLHCIYHQGAP 562
+ ++ PPP + P P +Y P Y P
Sbjct: 653 -KPTYKSPPPPYVYSSPPPPYYSPSPKPTYKSPPP 686
Score = 43.5 bits (101), Expect = 7e-04
Identities = 41/166 (24%), Positives = 57/166 (33%), Gaps = 32/166 (19%)
Query: 425 PQPVYSNHP----FVANITPQMTAPQNPNYQSPRP-----QGPAPYYPPLYQPLYN---- 471
P+P Y + P + + P + P Y+SP P P PYY P +P Y
Sbjct: 652 PKPTYKSPPPPYVYSSPPPPYYSPSPKPTYKSPPPPYVYSSPPPPYYSPAPKPTYKSPPP 711
Query: 472 --LQQLSQQPYY----------PQQPY-QQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPM 518
+ PYY P PY P PP P P+ Y + P
Sbjct: 712 PYVYSSPPPPYYSPSPKPTYKSPPPPYVYSSPPPPPYYSPSPKVEYKSPPPPYVYSSPPP 771
Query: 519 TYGALLPSLLAQNLVQTIPPPRI--PDPLPRWYRPDLHCIYHQGAP 562
Y + P + ++ PPP + P P +Y P Y P
Sbjct: 772 PYYSPSPKV----EYKSPPPPYVYSSPPPPPYYSPSPKVEYKSPPP 813
Score = 43.5 bits (101), Expect = 7e-04
Identities = 42/152 (27%), Positives = 58/152 (37%), Gaps = 23/152 (15%)
Query: 425 PQPVYSNHPFVANITPQ----MTAPQNPNYQ-SPRP--QGPAPYY------PPLYQPLYN 471
P P YS P V +P ++P P Y SP+P + P P Y PP Y P
Sbjct: 92 PPPYYSPSPKVDYKSPPPPYVYSSPPPPYYSPSPKPTYKSPPPPYVYNSPPPPYYSPSPK 151
Query: 472 LQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQN 531
++ S P PY PP P P+ Y ++ P Y + P +
Sbjct: 152 VEYKSPPP-----PYVYSSPPPPYYSPSPKVDYKSPPPPYVYNSPPPPYYSPSP----KP 202
Query: 532 LVQTIPPPRI-PDPLPRWYRPDLHCIYHQGAP 562
++ PPP I P P +Y P +Y P
Sbjct: 203 TYKSPPPPYIYSSPPPPYYSPSPKPVYKSPPP 234
Score = 37.0 bits (84), Expect = 0.065
Identities = 42/169 (24%), Positives = 58/169 (33%), Gaps = 35/169 (20%)
Query: 420 VAHGGPQP--VYSNHPFVANITPQMTAPQNPNYQSPRP-----QGPAPYYPPLYQPLYNL 472
V + P P VYS+ P P + P+Y+SP P P PYY P + +Y
Sbjct: 477 VVYKSPPPPYVYSSPP-----PPYYSPSPKPSYKSPPPPYVYNSPPPPYYSPSPKVIYKS 531
Query: 473 QQLSQ-------QPYY----------PQQPY-QQRPYNPPQQQPRPQAPYNQQNQKQQFD 514
P Y P PY P PP P P+ Y +
Sbjct: 532 PPHPHVCVCPPPPPCYSHSPKIEYKSPPTPYVYHSPPPPPYYSPSPKPAYKSSPPPYVYS 591
Query: 515 PLPMTYGALLPSLLAQNLVQTIPPPRI-PDPLPRWYRPDLHCIYHQGAP 562
P Y + P + + ++ PPP + P P +Y P Y P
Sbjct: 592 SPPPPYYSPAP----KPVYKSPPPPYVYNSPPPPYYSPSPKPTYKSPPP 636
Score = 36.6 bits (83), Expect = 0.085
Identities = 36/137 (26%), Positives = 50/137 (36%), Gaps = 22/137 (16%)
Query: 443 TAPQNPNYQSPRP----QGPAPYY------PPL-YQPLYNLQQLSQQPYYPQQPYQQRPY 491
++P P Y SP P + P P Y PPL Y P + S P PY
Sbjct: 12 SSPPPPLYDSPTPKVDYKSPPPPYVYSSPPPPLSYSPSPKVDYKS-----PPPPYVYSSP 66
Query: 492 NPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQN----LVQTIPPPRI--PDPL 545
PP P P+ Y + P Y + P + ++ V + PPP P P
Sbjct: 67 PPPYYSPSPKVEYKSPPPPYVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYSPSPK 126
Query: 546 PRWYRPDLHCIYHQGAP 562
P + P +Y+ P
Sbjct: 127 PTYKSPPPPYVYNSPPP 143
Score = 36.6 bits (83), Expect = 0.085
Identities = 29/95 (30%), Positives = 36/95 (37%), Gaps = 20/95 (21%)
Query: 425 PQPVYSNHPFVANITPQ----MTAPQNPNYQSPRPQ-----GPAPYY------PPLYQPL 469
P P YS P V +P ++P P Y SP P+ P PY P Y P
Sbjct: 796 PPPYYSPSPKVEYKSPPPPYVYSSPPPPTYYSPSPKVEYKSPPPPYVYNSPPPPAYYSPS 855
Query: 470 YNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPY 504
++ S P PY PP P P+A Y
Sbjct: 856 PKIEYKSPPP-----PYVYSSPPPPSYSPSPKAEY 885
Score = 36.6 bits (83), Expect = 0.085
Identities = 37/144 (25%), Positives = 52/144 (35%), Gaps = 23/144 (15%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYY-----PPLYQPLYNLQQLSQQP 479
P VYS+ P + +P +Y+SP P PY PP Y P ++ S P
Sbjct: 33 PPYVYSSPPPPLSYSPSPKV----DYKSP----PPPYVYSSPPPPYYSPSPKVEYKSPPP 84
Query: 480 YYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPP 539
PY PP P P+ Y + P Y + P + ++ PPP
Sbjct: 85 -----PYVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYSPSP----KPTYKSPPPP 135
Query: 540 RI-PDPLPRWYRPDLHCIYHQGAP 562
+ P P +Y P Y P
Sbjct: 136 YVYNSPPPPYYSPSPKVEYKSPPP 159
Score = 35.4 bits (80), Expect = 0.19
Identities = 37/142 (26%), Positives = 52/142 (36%), Gaps = 23/142 (16%)
Query: 425 PQPVYSNHPFVANITPQ---MTAPQNPNYQSPRPQ-----GPAPYY------PPLYQPLY 470
P P YS P V +P + + P Y SP P+ P PY PP Y P
Sbjct: 745 PPPYYSPSPKVEYKSPPPPYVYSSPPPPYYSPSPKVEYKSPPPPYVYSSPPPPPYYSPSP 804
Query: 471 NLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFD-PLPMTYGALLPSLLA 529
++ S P Y P P P P+ Y ++ P P Y + P +
Sbjct: 805 KVEYKSPPPPY----VYSSPPPPTYYSPSPKVEYKSPPPPYVYNSPPPPAYYSPSPKIEY 860
Query: 530 QN----LVQTIPPPRIPDPLPR 547
++ V + PPP P P+
Sbjct: 861 KSPPPPYVYSSPPPPSYSPSPK 882
>At3g01560 unknown protein
Length = 511
Score = 49.7 bits (117), Expect = 1e-05
Identities = 35/85 (41%), Positives = 37/85 (43%), Gaps = 13/85 (15%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSP-----RPQGPAPY--YPPLYQPLYNLQQLSQ 477
PQP SN P PQ P P+YQSP PQ P P Y P QP Y +Q
Sbjct: 293 PQPPPSNPPPYQ--APQTQTPHQPSYQSPPQQPQYPQQPPPSSGYNPEEQPPYQMQSYPP 350
Query: 478 QPYYPQQPYQQRP----YNPPQQQP 498
P Q P P YNPPQ QP
Sbjct: 351 NPPRQQPPAGSTPSQQFYNPPQPQP 375
Score = 49.3 bits (116), Expect = 1e-05
Identities = 27/69 (39%), Positives = 32/69 (46%), Gaps = 6/69 (8%)
Query: 448 PNYQSPRPQGPAPYYPPL----YQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAP 503
P++ P P P PY P +QP Y Q QQP YPQQP YNP +Q P
Sbjct: 290 PSHPQPPPSNPPPYQAPQTQTPHQPSY--QSPPQQPQYPQQPPPSSGYNPEEQPPYQMQS 347
Query: 504 YNQQNQKQQ 512
Y +QQ
Sbjct: 348 YPPNPPRQQ 356
Score = 38.1 bits (87), Expect = 0.029
Identities = 32/96 (33%), Positives = 39/96 (40%), Gaps = 12/96 (12%)
Query: 426 QPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQP 485
QP S P + Q ++ Q P P P P PP YQ Q QP Y P
Sbjct: 265 QPPSSQLP--PQLPTQFSSQQEPYCPPPSHPQPPPSNPPPYQAPQT--QTPHQPSYQSPP 320
Query: 486 YQQRPYNPPQQQPRPQAPYNQQN----QKQQFDPLP 517
Q+P P QQP P + YN + Q Q + P P
Sbjct: 321 --QQPQYP--QQPPPSSGYNPEEQPPYQMQSYPPNP 352
Score = 34.7 bits (78), Expect = 0.32
Identities = 32/128 (25%), Positives = 46/128 (35%), Gaps = 22/128 (17%)
Query: 440 PQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPR 499
P + PQ P+ + Q P+ PP ++ QQ +PY P + P PP P
Sbjct: 249 PLTSFPQPPSSTAAPSQPPSSQLPPQLPTQFSSQQ---EPYCPPPSH---PQPPPSNPPP 302
Query: 500 PQAP---------YNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIP-------PPRIPD 543
QAP Y Q+ Q+ P P +Q+ P PP
Sbjct: 303 YQAPQTQTPHQPSYQSPPQQPQYPQQPPPSSGYNPEEQPPYQMQSYPPNPPRQQPPAGST 362
Query: 544 PLPRWYRP 551
P ++Y P
Sbjct: 363 PSQQFYNP 370
>At1g37180 hypothetical protein
Length = 661
Score = 48.9 bits (115), Expect = 2e-05
Identities = 39/120 (32%), Positives = 59/120 (48%), Gaps = 17/120 (14%)
Query: 224 FKMPVFEKYQGDTCPQNHLTMYISKMIAYKNNVPL--------LIHCFQDSLTGPAHTWF 275
FK+PV Y G P HLT + ++I + VPL L F ++L+GPA TWF
Sbjct: 314 FKLPV---YNGKGDPNEHLTSF--QVIVRR--VPLEPYEEDAGLCKLFSENLSGPALTWF 366
Query: 276 MGLK--GVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQI 333
L+ + F+QL+ AF++QY Y + L +L+Q + + Y F + QI
Sbjct: 367 TQLEEGSIDNFKQLSTAFIKQYGYFIKSDITEAHLWNLSQSADDPLRTYIIVFKEIMVQI 426
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 48.1 bits (113), Expect = 3e-05
Identities = 69/278 (24%), Positives = 97/278 (34%), Gaps = 43/278 (15%)
Query: 260 IHCFQDSLTGPAHTWFMGL--KGVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKE 317
+ F SL AH W L +T ++ +AF+ ++ N+ A R E+ TQK E
Sbjct: 88 LRLFPFSLGDKAHLWEKTLPQNSITAWDDCKKAFLAKFFSNSRTARLRNEISGFTQKQNE 147
Query: 318 SFKEYAQRFIQKAAQIRPP---LDERELSDLFYETLSPCYSEKMIVCASQKFTDL-VETG 373
SF E +RF K Q + P + L Y + P + ++ F VE G
Sbjct: 148 SFGEAWERF--KGYQTKCPHHGFKQASLLSTLYRGVLPKIRILLDTASNGNFLKKDVEEG 205
Query: 374 MRIEEWARKGAAVSGSSSGGSSGVSSSGNKKFGNGYPKRN----------AQEVGMVAHG 423
+ E + S SS N K N + V ++
Sbjct: 206 WELVENFAQSDGNYNEDYDRSIRTSSDSNDKHDREMKALNDKLEKIVQMQQKHVHFISED 265
Query: 424 GPQPVYSNH----PFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQP 479
P PV ++ + Q + NY P P NL S
Sbjct: 266 EPFPVQEGEKDQWAEISYVQNQGGYNKGYNYYRPNP---------------NLSYRSTNV 310
Query: 480 YYPQ-QPYQQRPYNPPQQQPRPQAPYNQQN---QKQQF 513
PQ Q Y Q+ Q QP+P PYNQ KQQF
Sbjct: 311 ANPQDQVYAQQ--QQQQNQPKPFVPYNQNQGFIPKQQF 346
>At1g14650 splicing factor, putative
Length = 785
Score = 47.4 bits (111), Expect = 5e-05
Identities = 45/155 (29%), Positives = 59/155 (38%), Gaps = 25/155 (16%)
Query: 413 NAQEVGMVAHG------GPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYP--- 463
N +E G +G GP + P V + P + P N PRP P+ YP
Sbjct: 524 NGEEQGDGVYGDPNSFPGPAALPPPRPGVPIVRP-LPPPPNLALNLPRPP-PSAQYPGAP 581
Query: 464 -----PLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPM 518
P+ QP++ QL+ P P P Q PPQ QP P +Q +P
Sbjct: 582 RPLGVPMMQPMHQQHQLTM-PGPPGHP-QMMMNRPPQMQPGMHVPPPPGSQFAHHMQIPR 639
Query: 519 TYGALLPSLLAQ-------NLVQTIPPPRIPDPLP 546
YG L PS + + PP P PLP
Sbjct: 640 PYGQLPPSAMGMMQPPPMPGMAPPPPPEEAPPPLP 674
>At4g13340 extensin-like protein
Length = 760
Score = 46.6 bits (109), Expect = 8e-05
Identities = 39/138 (28%), Positives = 45/138 (32%), Gaps = 18/138 (13%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P PVYS P P P P Y P P P P PP+Y P P Y
Sbjct: 445 PPPVYSPPP------PPPPPPPPPVYSPPPPPPPPPPPPPVYSPPPPSPPPPPPPVYSPP 498
Query: 485 PYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDP 544
P P PP P P Y+ P P+ P PPP P P
Sbjct: 499 PPPPPPPPPPVYSPPPPPVYSSPPPPPSPAPTPVYCTRPPP-----------PPPHSPPP 547
Query: 545 LPRWYRPDLHCIYHQGAP 562
P++ P Y+ P
Sbjct: 548 -PQFSPPPPEPYYYSSPP 564
Score = 42.7 bits (99), Expect = 0.001
Identities = 38/122 (31%), Positives = 42/122 (34%), Gaps = 16/122 (13%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P PVYS P P P P Y SP P P P PP+Y P P Y
Sbjct: 476 PPPVYSPPP------PSPPPPPPPVY-SPPPPPPPPPPPPVYSP-------PPPPVYSSP 521
Query: 485 PYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMT--YGALLPSLLAQNLVQTIPPPRIP 542
P P P RP P QF P P Y + P + + PPP P
Sbjct: 522 PPPPSPAPTPVYCTRPPPPPPHSPPPPQFSPPPPEPYYYSSPPPPHSSPPPHSPPPPHSP 581
Query: 543 DP 544
P
Sbjct: 582 PP 583
Score = 38.5 bits (88), Expect = 0.022
Identities = 37/132 (28%), Positives = 48/132 (36%), Gaps = 23/132 (17%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYY--------PPLYQPLYNLQQLS 476
P P++S P + + P +P P Y P P P P PP P+Y S
Sbjct: 413 PAPIFSTPPTLTS--PPPPSPPPPVYSPPPPPPPPPPVYSPPPPPPPPPPPPVY-----S 465
Query: 477 QQPYYPQQPYQQRPYNPPQQQPRPQAP--YNQQNQKQQFDPLPMTYGALLPSLLAQNLVQ 534
P P P Y+PP P P P Y+ P P Y P + +
Sbjct: 466 PPPPPPPPPPPPPVYSPPPPSPPPPPPPVYSPPPPPPP-PPPPPVYSPPPPPVYS----- 519
Query: 535 TIPPPRIPDPLP 546
+ PPP P P P
Sbjct: 520 SPPPPPSPAPTP 531
Score = 35.0 bits (79), Expect = 0.25
Identities = 34/143 (23%), Positives = 48/143 (32%), Gaps = 9/143 (6%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQ----QLSQQPY 480
P PVYS+ P + P P P P + PP +P Y S P+
Sbjct: 514 PPPVYSSPPPPPSPAPTPVYCTRPPPPPPHSPPPPQFSPPPPEPYYYSSPPPPHSSPPPH 573
Query: 481 YPQQPYQ-QRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPP 539
P P+ P P P P P + + P P P ++ PPP
Sbjct: 574 SPPPPHSPPPPIYPYLSPPPPPTPVSSPPPTPVYSPPPPPPCIEPPP--PPPCIEYSPPP 631
Query: 540 RIPDPLPRWYRPDLHCIYHQGAP 562
P P+ + P +Y+ P
Sbjct: 632 --PPPVVHYSSPPPPPVYYSSPP 652
>At1g41780 hypothetical protein
Length = 246
Score = 46.6 bits (109), Expect = 8e-05
Identities = 52/214 (24%), Positives = 96/214 (44%), Gaps = 32/214 (14%)
Query: 129 EMPFAQGQQTSAHFQMSQPIPQATMT----QAGPTVHVGPQHEEQIYHSDSI-MGDDKAI 183
E+ +QGQ ++ P+ MT +A P V + P+ E +D I + DD+
Sbjct: 49 EVATSQGQHSAE--------PEQNMTNRVEEADPQVPI-PRAESD--EADEIKVSDDE-- 95
Query: 184 DWEERFGALEKKMSNMRGKEAVVQSIYDLCLVPDVNI-----PPKFKMPVFEKYQGDTCP 238
D EE E +S++ + +++ ++ ++ P K ++ E + G + P
Sbjct: 96 DSEENIRWAEDAVSSVPNIDCILEVYHNTPFTRKISNAIISDPEKLRI---EYFSGSSDP 152
Query: 239 QNHLTMYISKMIAYK----NNVPLLIHCFQDSLTGPAHTWFMGLKG--VTTFEQLAEAFM 292
+ HL + + K + L H F + L GP WF L+G V +F++L+ F+
Sbjct: 153 KGHLKSFKISVARAKFKPEESDAGLCHLFVEHLKGPVLDWFSRLEGNSVDSFQELSTLFL 212
Query: 293 QQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRF 326
+QY S +L SL+Q+ E ++ +F
Sbjct: 213 KQYSVLIDPGTSDVDLWSLSQQPNEPLSDFLTKF 246
>At1g12040 leucine-rich repeat/extensin 1 (LRX1)
Length = 744
Score = 46.6 bits (109), Expect = 8e-05
Identities = 42/134 (31%), Positives = 49/134 (36%), Gaps = 9/134 (6%)
Query: 425 PQPVYSNHPFVANITP--QMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYP 482
P P S P V P Q P +P Y P Q P P P Y P+ N YYP
Sbjct: 532 PCPESSPPPPVVYYAPVTQSPPPPSPVYYPPVTQSPPPPSPVYYPPVTNSPPPPSPVYYP 591
Query: 483 QQPYQQRPYNP---PQQQPRPQAP---YNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTI 536
Y P +P PQ P P P Y P P+ Y + PS + V
Sbjct: 592 PVTYSPPPPSPVYYPQVTPSPPPPSPLYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVYYP 651
Query: 537 P-PPRIPDPLPRWY 549
P P P P P +Y
Sbjct: 652 PVTPSPPPPSPVYY 665
Score = 33.9 bits (76), Expect = 0.55
Identities = 37/139 (26%), Positives = 46/139 (32%), Gaps = 25/139 (17%)
Query: 425 PQPVYSNHP--FVANITPQMTAPQNPNYQSPRPQGPAPYYPP-----LYQPLYNLQQLSQ 477
P VYS+ P +V + P +P P P P P P Y P+
Sbjct: 497 PPYVYSSPPPPYVYSSPPPPYVYSSPPPPPPSPPPPCPESSPPPPVVYYAPVTQSPPPPS 556
Query: 478 QPYYPQQPYQQRP-----YNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNL 532
YYP P Y P P P +P P+TY PS +
Sbjct: 557 PVYYPPVTQSPPPPSPVYYPPVTNSPPPPSPVYYP---------PVTYSPPPPSPVYYPQ 607
Query: 533 VQTIPPPRIPDPLPRWYRP 551
V PPP P P +Y P
Sbjct: 608 VTPSPPP----PSPLYYPP 622
Score = 33.1 bits (74), Expect = 0.94
Identities = 35/144 (24%), Positives = 44/144 (30%), Gaps = 12/144 (8%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P PVY +P V P P +P Y P P P P Y P+ YYP
Sbjct: 600 PSPVY--YPQVTPSPP----PPSPLYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVYYPPV 653
Query: 485 PYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRI--- 541
P +P Q+P P A Q PPP
Sbjct: 654 TPSPPPPSPVYYPSETQSPPPPTEYYYSPSQSPPPTKACKEGHPPQATPSYEPPPEYSYS 713
Query: 542 ---PDPLPRWYRPDLHCIYHQGAP 562
P P P Y P + + + +P
Sbjct: 714 SSPPPPSPTSYFPPMPSVSYDASP 737
>At5g14540 unknown protein
Length = 530
Score = 46.2 bits (108), Expect = 1e-04
Identities = 35/98 (35%), Positives = 39/98 (39%), Gaps = 18/98 (18%)
Query: 426 QPVYSNHPFVANITPQMTAPQNPNYQSPR------PQG---PAPYYPPLYQPLYNLQQLS 476
QP S H P + PQ PN SP+ P G P P P YQP Q L
Sbjct: 251 QPPASQHGLSP---PSLQLPQLPNQFSPQQEPYFPPSGQSQPPPTIQPPYQPPPPTQSLH 307
Query: 477 QQPYYPQQPYQQRPYNPPQQQPRP------QAPYNQQN 508
Q PY P Q P PP Q P + PY QQ+
Sbjct: 308 QPPYQPPPQQPQYPQQPPPQLQHPSGYNPEEPPYPQQS 345
Score = 35.4 bits (80), Expect = 0.19
Identities = 27/76 (35%), Positives = 30/76 (38%), Gaps = 18/76 (23%)
Query: 448 PNYQSPRPQGPA---PYYPPLYQPLYNLQQLSQ----------QPYYPQQPY-----QQR 489
P YQ P P PY PP QP Y Q Q +P YPQQ Y +Q
Sbjct: 295 PPYQPPPPTQSLHQPPYQPPPQQPQYPQQPPPQLQHPSGYNPEEPPYPQQSYPPNPPRQP 354
Query: 490 PYNPPQQQPRPQAPYN 505
P +PP Q YN
Sbjct: 355 PSHPPPGSAPSQQYYN 370
>At1g53645 unknown protein
Length = 523
Score = 45.8 bits (107), Expect = 1e-04
Identities = 37/128 (28%), Positives = 52/128 (39%), Gaps = 18/128 (14%)
Query: 388 GSSSGGSSGVSSSGNKKFGNG----YPKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMT 443
G SG G ++G +FG P R G HG +P+ S+ +I+P T
Sbjct: 46 GRGSGEDGGFPAAGRGQFGVNREPVVPGREPSSAGGYGHGRGRPIQSD-----SISPAFT 100
Query: 444 A---PQNPNYQSPRPQ------GPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPP 494
+ +P+ R P PP P QQ SQ QP QQ+P + P
Sbjct: 101 SFVKSDSPSIGRGRGSVGSDTVSPFAAEPPRQSPPPPQQQQSQSQQQRSQPQQQQPRSQP 160
Query: 495 QQQPRPQA 502
QQQP ++
Sbjct: 161 QQQPNDES 168
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,370,508
Number of Sequences: 26719
Number of extensions: 1323173
Number of successful extensions: 6516
Number of sequences better than 10.0: 244
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 184
Number of HSP's that attempted gapping in prelim test: 4937
Number of HSP's gapped (non-prelim): 917
length of query: 1155
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1045
effective length of database: 8,379,506
effective search space: 8756583770
effective search space used: 8756583770
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC145024.28


