

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.2 - phase: 0
(355 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
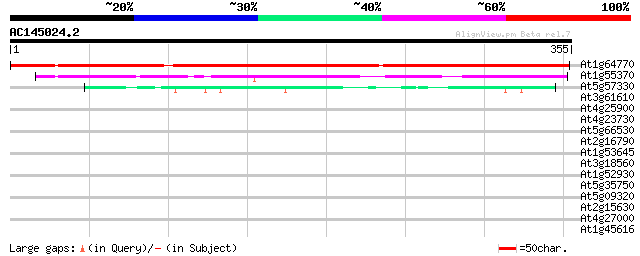
Score E
Sequences producing significant alignments: (bits) Value
At1g64770 unknown protein 410 e-115
At1g55370 unknown protein 102 2e-22
At5g57330 apospory-associated protein C 44 2e-04
At3g61610 unknown protein 39 0.003
At4g25900 possible apospory-associated like protein 38 0.010
At4g23730 unknown protein 37 0.016
At5g66530 apospory-associated protein C-like 33 0.31
At2g16790 gluconokinase like protein 29 3.4
At1g53645 unknown protein 29 3.4
At3g18560 unknown protein 29 4.5
At1g52930 unknown protein 28 5.8
At5g35750 histidine kinase (AHK2) 28 7.6
At5g09320 unknown protein 28 7.6
At2g15630 putative salt-inducible protein 28 7.6
At4g27000 putative DNA binding protein 28 10.0
At1g45616 disease resistance protein, putative 28 10.0
>At1g64770 unknown protein
Length = 348
Score = 410 bits (1053), Expect = e-115
Identities = 213/356 (59%), Positives = 264/356 (73%), Gaps = 10/356 (2%)
Query: 1 MASFLSLSL-PKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRN 59
MAS +S SL PK +++S +A T ++T E L DKFGRKGIKF ESN+IP+VEL VRN
Sbjct: 1 MASLISFSLLPKPKAVRSSISAPQTQTINT-EKLEDKFGRKGIKFSESNNIPMVELKVRN 59
Query: 60 GSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGA 119
GSSL L L DAHV SYKPKV+WKD+G EE+LYT+ +E+ +GG+G+V+ +P
Sbjct: 60 GSSLKLSLSDAHVLSYKPKVYWKDEGFEEVLYTVDGDES-----RGGVGVVIVNGEEPKG 114
Query: 120 KELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKP 179
+ S +W+VKD D DAIDALQ+EL T D+TYIVSLYPVSMATA++VKN KP
Sbjct: 115 GSSVISGCDWSVKDTDSDAIDALQIELSCTAGVLDITYIVSLYPVSMATALVVKNNGRKP 174
Query: 180 VTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA 239
VTL I+S+ RFK+R GA I+GL+ CSYC +PPL SPF++L+PSEAMK+ES FG+
Sbjct: 175 VTLKPGIMSYLRFKKRSGAGIQGLKGCSYCPNPPLSSPFELLSPSEAMKAESSGW--FGS 232
Query: 240 EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVI 299
E KPG W + ITLLE KMSR++ APP ER KA YNTPPSK+E IDQGR +F+R+I
Sbjct: 233 EEGEKPGIWAVEDSVITLLEKKMSRIYGAPPAERLKAVYNTPPSKFETIDQGRGLFFRMI 292
Query: 300 RMGFEDIYVSSPGSMSDKYGRD-YFVCTGPASILEPVTVNPGEEFRGAQVIEHDNL 354
R+GFE++YV SPGSM DKYG+ YFVCTGP S+L PV V GE +RGA VIEHDNL
Sbjct: 293 RIGFEEMYVGSPGSMWDKYGKQHYFVCTGPTSMLVPVDVASGETWRGAMVIEHDNL 348
>At1g55370 unknown protein
Length = 354
Score = 102 bits (255), Expect = 2e-22
Identities = 86/341 (25%), Positives = 152/341 (44%), Gaps = 41/341 (12%)
Query: 17 ASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYK 76
AS++A+ P+ E L +F G F + I L + NGSS ++ L +TSYK
Sbjct: 50 ASASASPAFPIDV-EYLRREFSGHGATFEDIGETCIARLKLDNGSSANVMLTRGMITSYK 108
Query: 77 PKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDY 136
+V+ G ELL+T E +GG+ + E+ +W ++ +
Sbjct: 109 VRVW--HGGKVELLHTWVEQEEEEVVIRGGVSSAFRS---SDSDEIS----DWRLQGISG 159
Query: 137 DAIDALQVELISTNRFF---DMTYIVSLYPVSMATAVIVKNKSPKPVTLTN-AILSHFRF 192
D+ D +Q+EL +++ ++ I+SL +++ + + NK P+ L +++S+
Sbjct: 160 DSKDCVQMELRRSDKKIKEIELKQIISLRENTLSIELSMTNKGISPIKLEGCSLVSYLTV 219
Query: 193 KRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQG 252
GL+ + P F ++ G + E KPG ++
Sbjct: 220 STPEATYAVGLEGSDFVETTPFLPRFGVVQ---------------GEKEEEKPGFGGEEE 264
Query: 253 VPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG 312
L +MSR++ PK T +ID+GR V R GFE++Y+ SPG
Sbjct: 265 SNYKQLNREMSRIYTCAPKSFT------------VIDRGRRNSVVVGREGFEEVYMYSPG 312
Query: 313 SMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDN 353
S + Y + +VC GP+S+L P+++ G +RG + + N
Sbjct: 313 SRLESYTKSAYVCIGPSSLLSPISLESGCVWRGVLHLHNPN 353
>At5g57330 apospory-associated protein C
Length = 312
Score = 43.5 bits (101), Expect = 2e-04
Identities = 70/327 (21%), Positives = 126/327 (38%), Gaps = 81/327 (24%)
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA---- 103
N + + L G S + L +HVTS WK++ EELL+ + ++K
Sbjct: 15 NGLDKIVLRESRGRSAEVYLYGSHVTS------WKNENGEELLHL---SSKAIFKPPKPI 65
Query: 104 KGGIGLVLNEVLQPGAKEL--LPSTLEWTVK----DVDYDAIDALQVELISTNRFFDMTY 157
+GGI L + G E W V+ + ++ + V+LI D+
Sbjct: 66 RGGIPLCFPQFSNFGTLESHGFARNRIWEVEANPPPLPLNSCSSAFVDLILRPTEDDLKI 125
Query: 158 IVSLYPVSMATAVIVK------------NKSPKPVTLTNAILSHFRFKRRGGAAIKGLQT 205
+ + + A+ + N KP T T A ++F ++GL+T
Sbjct: 126 WPNNFEFRLRIALGTEGELTLTSRIRNTNSDGKPFTFTFAYHTYFSVSDISEVRVEGLET 185
Query: 206 CSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRV 265
Y + +K + +T+QG IT E+++ ++
Sbjct: 186 LDYLDN---------------LKDRER---------------FTEQGDAITF-ESEVDKI 214
Query: 266 FAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG-----SMSDKYGR 320
Y + P+K I+D ++ + + + G D V +P ++SD
Sbjct: 215 ------------YLSTPTKIAILDHEKKRTFVIRKDGLADAVVWNPWDKKSKTISDLGDE 262
Query: 321 DY--FVCTGPASILEPVTVNPGEEFRG 345
DY +C A+I P+T+ PGEE++G
Sbjct: 263 DYKHMLCVEAAAIERPITLKPGEEWKG 289
>At3g61610 unknown protein
Length = 317
Score = 39.3 bits (90), Expect = 0.003
Identities = 28/108 (25%), Positives = 47/108 (42%), Gaps = 20/108 (18%)
Query: 249 TQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYV 308
T+QG IT E++M R + PK ++D R+ Y + + G D V
Sbjct: 204 TEQGDAITF-ESEMDRTYLRSPKV------------VAVLDHERKRTYVIGKEGLPDTVV 250
Query: 309 SSPGSMSDKYGRDY-------FVCTGPASILEPVTVNPGEEFRGAQVI 349
+P K D+ +C A++ P+T+ PGEE+ G ++
Sbjct: 251 WNPWEKKSKTMADFGDEEYKSMLCVDGAAVERPITLKPGEEWTGRLIL 298
>At4g25900 possible apospory-associated like protein
Length = 318
Score = 37.7 bits (86), Expect = 0.010
Identities = 24/72 (33%), Positives = 39/72 (53%), Gaps = 4/72 (5%)
Query: 278 YNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGS--MSDKYGRDY--FVCTGPASILE 333
Y + P + I+D ++ V + G D V +P +SD DY FV A++ +
Sbjct: 234 YLSTPDQLRIVDHKKKKTIVVHKEGQVDAVVWNPWDKKVSDLGVEDYKRFVTVESAAVAK 293
Query: 334 PVTVNPGEEFRG 345
P+TVNPG+E++G
Sbjct: 294 PITVNPGKEWKG 305
>At4g23730 unknown protein
Length = 306
Score = 37.0 bits (84), Expect = 0.016
Identities = 67/312 (21%), Positives = 118/312 (37%), Gaps = 75/312 (24%)
Query: 60 GSSLSLRLPDAHVTSYKPKVFWKDDGLEELLY-TIPANETGLYKAKGGIGLVLNEVLQPG 118
G+S+ + L V S WK D +ELL+ + AN + +GGI + + G
Sbjct: 34 GASVKISLHGGQVLS------WKTDKGDELLFNSTKANLKPPHPVRGGIPICFPQFGTRG 87
Query: 119 AKEL--LPSTLEWTVKD-----VDYDAIDALQVELI------STNRF----FDMTYIVSL 161
+ E W V++ +D+ V+L+ T R F+ VSL
Sbjct: 88 SLEQHGFARNKMWLVENNPPALPSFDSTGKAYVDLVLKSSDEDTMRIWPYSFEFHLRVSL 147
Query: 162 YPVSMATAVI-VKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQI 220
T + V+N + KP + + A ++F ++GL+T Y +
Sbjct: 148 ALDGNLTLISRVRNINSKPFSFSIAYHTYFSISDISEVRLEGLETLDYLDN--------- 198
Query: 221 LTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNT 280
+ +R +T+QG +T E+++ RV Y
Sbjct: 199 -------MHDRER--------------FTEQGDALTF-ESEIDRV------------YLN 224
Query: 281 PPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRD-------YFVCTGPASILE 333
I D R+ + + + G D+ V +P + D + +C A+I +
Sbjct: 225 SKDVVAIFDHERKRTFLIKKEGLPDVVVWNPWEKKARALTDLGDDEYRHMLCVDGAAIEK 284
Query: 334 PVTVNPGEEFRG 345
P+T+ PGEE+ G
Sbjct: 285 PITLKPGEEWTG 296
>At5g66530 apospory-associated protein C-like
Length = 307
Score = 32.7 bits (73), Expect = 0.31
Identities = 44/185 (23%), Positives = 80/185 (42%), Gaps = 20/185 (10%)
Query: 41 GIKFLESN-SIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIP-ANET 98
G++ E ++P + LT S + L +TS WK ++LL+ P A
Sbjct: 35 GVRVAEGEGNLPKLVLTSPQNSEAEIYLFGGCITS------WKVASGKDLLFVRPDAVFN 88
Query: 99 GLYKAKGGIGLVLNEVLQPGAKEL--LPSTLEWTVKD---VDYDAIDALQVELISTNRF- 152
+ GGI + PG + ++W+V D D +A L+++ +R
Sbjct: 89 KIKPISGGIPHCFPQ-FGPGLIQQHGFGRNMDWSVVDSQNADDNAAVTLELKDGPYSRAM 147
Query: 153 ----FDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSY 208
F Y V + S++T + + N KP + + A+ ++FR GA+++GL+ C
Sbjct: 148 WDFAFQALYKVIVGADSLSTELKITNTDDKPFSFSTALHTYFR-ASSAGASVRGLKGCKT 206
Query: 209 CSHPP 213
+ P
Sbjct: 207 LNKDP 211
>At2g16790 gluconokinase like protein
Length = 189
Score = 29.3 bits (64), Expect = 3.4
Identities = 25/88 (28%), Positives = 41/88 (46%), Gaps = 8/88 (9%)
Query: 214 LDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKER 273
LD +L S K R I G++P+ KPGS+T V LLE + A K
Sbjct: 79 LDGETVVLACSSLRKQ--YREILRGSDPDYKPGSYTSCKVTFVLLEGNAEVIAARLQKRA 136
Query: 274 TKAFYNTP----PSKYEII--DQGREIF 295
++ + P S+++++ D+ +IF
Sbjct: 137 SEEEHFMPLTLLQSQFDLLQADECEKIF 164
>At1g53645 unknown protein
Length = 523
Score = 29.3 bits (64), Expect = 3.4
Identities = 22/94 (23%), Positives = 40/94 (42%), Gaps = 11/94 (11%)
Query: 195 RGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFG------AEPEMKPGSW 248
RG ++ + + PP SP P + +S+SQ+ S ++P+ +P
Sbjct: 112 RGRGSVGSDTVSPFAAEPPRQSP----PPPQQQQSQSQQQRSQPQQQQPRSQPQQQPND- 166
Query: 249 TQQGVPITLLENKMSRVFAAPPKERTKAFYNTPP 282
QG P+ + +M ++PP +K PP
Sbjct: 167 ESQGSPVFVKLQEMQDATSSPPPPESKPGQADPP 200
>At3g18560 unknown protein
Length = 195
Score = 28.9 bits (63), Expect = 4.5
Identities = 24/79 (30%), Positives = 36/79 (45%), Gaps = 12/79 (15%)
Query: 28 STPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLE 87
STPE ++ R + FL S ++ L R+ S +S +L KPK K LE
Sbjct: 4 STPEHVSSAHKRISVSFLVS----LMVLCARHASRVSKKL--------KPKKTRKQTHLE 51
Query: 88 ELLYTIPANETGLYKAKGG 106
+ L + +N G +GG
Sbjct: 52 DYLESPKSNGNGSEDGRGG 70
>At1g52930 unknown protein
Length = 320
Score = 28.5 bits (62), Expect = 5.8
Identities = 23/100 (23%), Positives = 42/100 (42%), Gaps = 15/100 (15%)
Query: 257 LLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSD 316
LL+ +++VF P + R Y+ + I+D+ + F + +S P + SD
Sbjct: 178 LLKEMLTQVFGIPKEHRKSKPYHDHVFVFSIVDE---------HIWFRNYQISVPHNESD 228
Query: 317 KYGRD-----YFVCTGPASILEPVTVNPGEEFRGAQVIEH 351
K + + GP L P+ + G F G + E+
Sbjct: 229 KIAKGGLDKMTLIEVGPRFCLNPIKIFAG-SFGGPTLYEN 267
>At5g35750 histidine kinase (AHK2)
Length = 1176
Score = 28.1 bits (61), Expect = 7.6
Identities = 11/29 (37%), Positives = 16/29 (54%)
Query: 229 SESQRLISFGAEPEMKPGSWTQQGVPITL 257
S+S ++ SF E PG WT G+ +L
Sbjct: 86 SDSDQVTSFECHKESSPGMWTNYGITCSL 114
>At5g09320 unknown protein
Length = 712
Score = 28.1 bits (61), Expect = 7.6
Identities = 39/171 (22%), Positives = 64/171 (36%), Gaps = 12/171 (7%)
Query: 116 QPGAKELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVS----LYPVSMATAVI 171
+PGA + LP + T+K L+ R+ + +V L+ ++
Sbjct: 178 EPGADQFLPVLIYVTIKANP----PQFHSNLLYIQRYRRQSKLVGEAGYLFTNILSAESF 233
Query: 172 VKNKSPKPVTLTNAILSHFRFKRRGG-AAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSE 230
+ N K +++ A F K + A + G + SY + P T E M
Sbjct: 234 ISNIDAKSLSMDEA---DFETKMKSAHARLSGPGSQSYQTDHGAALPTAHNTKRENMLLH 290
Query: 231 SQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTP 281
++ SF E + ++ PIT LENK + E TK F P
Sbjct: 291 TKSTDSFSGTNETLSETPIKKADPITDLENKGAATLLNDRSEATKIFQEYP 341
>At2g15630 putative salt-inducible protein
Length = 627
Score = 28.1 bits (61), Expect = 7.6
Identities = 25/82 (30%), Positives = 35/82 (42%), Gaps = 14/82 (17%)
Query: 114 VLQPGAKELLPSTLEWTVKDVDYDAIDALQVELIST---------NRFFDMTYIVSLYPV 164
VL P E+L ++ + + D L L+ST N F+ + LY +
Sbjct: 41 VLPPITSEILLESIRSSQWHIVEHVADKLTPSLVSTTLLSLVKTPNLAFNFVNHIDLYRL 100
Query: 165 S-----MATAVIVKNKSPKPVT 181
+A AVI K SPKPVT
Sbjct: 101 DFQTQCLAIAVISKLSSPKPVT 122
>At4g27000 putative DNA binding protein
Length = 415
Score = 27.7 bits (60), Expect = 10.0
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 2/70 (2%)
Query: 272 ERTKAFYNTPPSKYEIIDQ--GREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYFVCTGPA 329
E KA Y++ + D+ GR Y +R E + + M+ +Y + TGPA
Sbjct: 191 ETFKAVYSSVKGAKVVNDRTTGRSKGYGFVRFADESEQIRAMTEMNGQYCSSRPMRTGPA 250
Query: 330 SILEPVTVNP 339
+ +P+T+ P
Sbjct: 251 ANKKPLTMQP 260
>At1g45616 disease resistance protein, putative
Length = 994
Score = 27.7 bits (60), Expect = 10.0
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 3/74 (4%)
Query: 12 LNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELT--VRNGSSL-SLRLP 68
L+L + S A + PLST +D +++LE + I+E +RN +L S+ L
Sbjct: 470 LSLKRLVSLALSGIPLSTTNITSDSEFSSHLEYLELSGCNIIEFPEFIRNQRNLSSIDLS 529
Query: 69 DAHVTSYKPKVFWK 82
+ ++ P W+
Sbjct: 530 NNNIKGQVPNWLWR 543
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,183,938
Number of Sequences: 26719
Number of extensions: 353139
Number of successful extensions: 834
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 816
Number of HSP's gapped (non-prelim): 21
length of query: 355
length of database: 11,318,596
effective HSP length: 100
effective length of query: 255
effective length of database: 8,646,696
effective search space: 2204907480
effective search space used: 2204907480
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145024.2


