

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
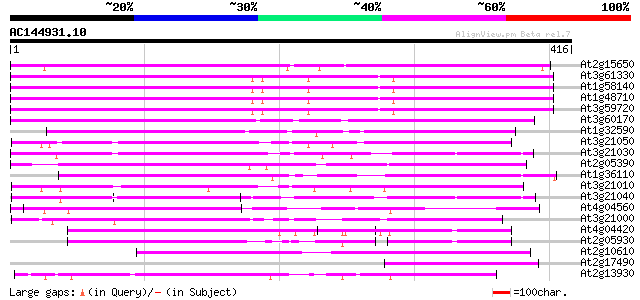
Score E
Sequences producing significant alignments: (bits) Value
At2g15650 putative retroelement pol polyprotein 232 2e-61
At3g61330 copia-type polyprotein 204 1e-52
At1g58140 hypothetical protein 202 3e-52
At1g48710 hypothetical protein 202 3e-52
At3g59720 copia-type reverse transcriptase-like protein 201 8e-52
At3g60170 putative protein 171 9e-43
At1g32590 hypothetical protein, 5' partial 161 5e-40
At3g21050 hypothetical protein 151 6e-37
At3g21030 hypothetical protein 150 1e-36
At2g05390 putative retroelement pol polyprotein 142 3e-34
At1g36110 hypothetical protein 135 3e-32
At3g21010 unknown protein 130 2e-30
At3g21040 hypothetical protein 122 4e-28
At4g04560 putative transposon protein 117 2e-26
At3g21000 unknown protein 113 2e-25
At4g04420 putative transposon protein 106 3e-23
At2g05930 copia-like retroelement pol polyprotein 104 8e-23
At2g10610 pseudogene 103 2e-22
At2g17490 putative retroelement pol polyprotein 93 2e-19
At2g13930 putative retroelement pol polyprotein 91 9e-19
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 232 bits (592), Expect = 2e-61
Identities = 139/426 (32%), Positives = 228/426 (52%), Gaps = 28/426 (6%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLP--NNPTLAQIRNHSDE-VAKEGRALAIIHAALHDDI 57
M T R + LW VVE+G P+ P A+ + +E V + AL I+ A+ D I
Sbjct: 24 MATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTNDTMALQILQTAVTDQI 83
Query: 58 FIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKV 117
F +I +KEAW+ L +EYQGS + + +K+ +LRRE+E LKM + + ++ F+D++ +
Sbjct: 84 FSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLKMYDNDNIKTFTDKLIVL 143
Query: 118 ITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRR 177
Q+ GE ++ ++++KIL+ LP F++ +S LE+ ++ +T+ EL+ L+A E R
Sbjct: 144 EIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDALTMSELLGILKAQEARV 203
Query: 178 SLRMEENVEGAFLASNKGKNQSFKSFG-------EKKFPPCPHCKKDTHLDKFCWYRP-- 228
+ R E EGAF +KG+ FK +KK+ C K H ++ C +P
Sbjct: 204 TAREESTKEGAFYVRSKGRESGFKQDNTNNRVNQDKKW--CGFHKSSKHTEEECREKPKN 261
Query: 229 ---------GVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACS 279
+KC C ++GH C++K N++ + E ++ LF AS + +
Sbjct: 262 DDHGKNKRSNIKCYKCGKIGHYANECRSK-NKERAHVTLEEEDVNEDHMLFSASEEESTT 320
Query: 280 SKE-TWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKY 338
+E WL+DSGCTNHMT +F +++S + NG V GKG + V T G +
Sbjct: 321 LREDVWLVDSGCTNHMTKEERYFSNINKSIKVPIRVRNGDIVMTAGKGDITVMTRHGKRI 380
Query: 339 ILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP---RG 395
I +V VP + ++LLSV Q++ Y + F++ +C I D G ++M +EM KSF
Sbjct: 381 IKNVFLVPGLEKNLLSVPQIISSGYWVRFQDKRCIIQDANGKEIMNIEMTDKSFKIKLSS 440
Query: 396 VEEDIL 401
VEE+ +
Sbjct: 441 VEEEAM 446
>At3g61330 copia-type polyprotein
Length = 1352
Score = 204 bits (518), Expect = 1e-52
Identities = 132/433 (30%), Positives = 218/433 (49%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIAEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKAHYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y KE W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQKENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
Score = 29.6 bits (65), Expect = 3.2
Identities = 41/227 (18%), Positives = 87/227 (38%), Gaps = 35/227 (15%)
Query: 20 PPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNL-----------EIAK 68
P P +PT +QI S E R++ ++ + + + L I K
Sbjct: 788 PTTPPTSPTSSQIEESSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQKAIEK 847
Query: 69 EAWNKLMEE----------YQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVI 118
+ W M+E ++ + K + ++ ++A K + E R + ++K
Sbjct: 848 KTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGY 907
Query: 119 TQIRLLGEDLSD------QRVVEKILVCLPEMFEAKISSLEENKNF------SEITVPEL 166
+Q +G D + + ++++ L + KI ++ F E+ + +
Sbjct: 908 SQ--RVGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQP 965
Query: 167 VNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPH 213
+ E+ + LR+++ + G A + K F EK F CP+
Sbjct: 966 QGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPY 1012
>At1g58140 hypothetical protein
Length = 1320
Score = 202 bits (514), Expect = 3e-52
Identities = 131/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>At1g48710 hypothetical protein
Length = 1352
Score = 202 bits (514), Expect = 3e-52
Identities = 131/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIIEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
Score = 29.6 bits (65), Expect = 3.2
Identities = 41/225 (18%), Positives = 84/225 (37%), Gaps = 31/225 (13%)
Query: 20 PPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNL-----------EIAK 68
P P +PT +QI S E R++ ++ + + + L I K
Sbjct: 788 PTTPPTSPTSSQIEESSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEAIEK 847
Query: 69 EAWNKLMEE----------YQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVI 118
+ W M+E ++ + K + ++ ++A K + E R + ++K
Sbjct: 848 KTWRNAMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERYKARLVAKGY 907
Query: 119 TQIRLLGEDLSDQRVVE----KILVCLPEMFEAKISSLEENKNF------SEITVPELVN 168
Q + D V ++++ L + KI ++ F E+ + +
Sbjct: 908 IQRAGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQG 967
Query: 169 ALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPH 213
+ E+ + LR+++ + G A + K F EK F CP+
Sbjct: 968 YIVKGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPY 1012
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 201 bits (510), Expect = 8e-52
Identities = 130/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ +++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFKEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDKESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
Score = 30.0 bits (66), Expect = 2.5
Identities = 41/225 (18%), Positives = 85/225 (37%), Gaps = 31/225 (13%)
Query: 20 PPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNL-----------EIAK 68
P P +PT +QI S E R++ ++ + + + L I K
Sbjct: 788 PTTPPTSPTSSQIEESSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEAIEK 847
Query: 69 EAWNKLMEE----------YQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVI 118
+ W M+E ++ + K + ++ ++A K + E R + ++K
Sbjct: 848 KTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGY 907
Query: 119 TQIRLLGEDLSDQRVVE----KILVCLPEMFEAKISSLEENKNF------SEITVPELVN 168
+Q + D V ++++ L + KI ++ F E+ + +
Sbjct: 908 SQRAGIDYDEIFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQG 967
Query: 169 ALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPH 213
+ E+ + LR+++ + G A + K F EK F CP+
Sbjct: 968 YIVKGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPY 1012
>At3g60170 putative protein
Length = 1339
Score = 171 bits (432), Expect = 9e-43
Identities = 112/390 (28%), Positives = 192/390 (48%), Gaps = 11/390 (2%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVA-KEGRALAIIHAALHDDIFI 59
M+ +LR++ LW +VE+G + P R+ +E K+ + + A+ +I
Sbjct: 26 MENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAVEEAKLKDLKVKNFLFQAIDREILE 85
Query: 60 KILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVIT 119
IL+ +K W + ++YQGS + K+ ++ LR+EFE L MKE E + F R V+
Sbjct: 86 TILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMKEGEKIDTFLGRTLTVVN 145
Query: 120 QIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSL 179
+++ GE + +V KIL L F + S+EE+ + S +++ EL +L EQR +
Sbjct: 146 KMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTLSIDELHGSLLVHEQRLNG 205
Query: 180 RMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLG 239
++E E A +++ + + G + + + R V+C C+ LG
Sbjct: 206 HVQE--EQALKVTHEERPSQGRGRGVFRGSR----GRGRGRGRSGTNRAIVECYKCHNLG 259
Query: 240 HVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVS 299
H + C + E+ A E +E+E L + E W +DSGC+NHMT +
Sbjct: 260 HFQYECP----EWEKNANYAELEEEEELLLMAYVEQNQANRDEVWFLDSGCSNHMTGSKE 315
Query: 300 FFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMM 359
+F EL+E F V GN + V GKG V V+ + I +V +VPE+ +LLS+GQ+
Sbjct: 316 WFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQVIPEVYYVPELRNNLLSLGQLQ 375
Query: 360 EKNYSLHFKNMKCTIFDPYGSKLMIVEMRG 389
E+ ++ ++ C ++ P +M M G
Sbjct: 376 ERGLAILIRDGTCKVYHPSKGAIMETNMSG 405
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 161 bits (408), Expect = 5e-40
Identities = 105/350 (30%), Positives = 175/350 (50%), Gaps = 13/350 (3%)
Query: 28 TLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKM 87
T AQ +++ K+ + + A++ I IL E +K+ W + +YQG++R +
Sbjct: 11 TGAQRTELAEKTVKDHKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSA 70
Query: 88 KVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEA 147
++ LRR FE L+MK ET+ + R+ ++ +R LGED+ D +VVEKIL L E F
Sbjct: 71 QLQRLRRSFEVLEMKIGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTY 130
Query: 148 KISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKK 207
+ ++EE+ N E+TV L ++L EQ +L + E A + + + G
Sbjct: 131 VVCAIEESNNIKELTVDGLQSSLMVHEQ--NLSRHDVEERVLKAETQWRPDGGRGRGGS- 187
Query: 208 FPPCPHCKKDTHLDKFCWY--RPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQED 265
P + + + Y R V+C C+++GH + C + E++A VE E+
Sbjct: 188 --PSRGRGRGGYQGRGRGYVNRDTVECFKCHKMGHYKAECPS----WEKEANYVE--MEE 239
Query: 266 EEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGK 325
+ L + K+ W +DSGC+NHM +F ELD F V G+ + + V+GK
Sbjct: 240 DLLLMAHVEQIGDEEKQIWFLDSGCSNHMCGTREWFLELDSGFKQNVRLGDDRRMAVEGK 299
Query: 326 GVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIF 375
G + +E I+ I DV FVP + +L SVGQ+ +K + C ++
Sbjct: 300 GKLRLEVDGRIQVISDVYFVPGLKNNLFSVGQLQQKGLRFIIEGDVCEVW 349
>At3g21050 hypothetical protein
Length = 420
Score = 151 bits (382), Expect = 6e-37
Identities = 116/386 (30%), Positives = 187/386 (48%), Gaps = 36/386 (9%)
Query: 2 KTYLRAQSLWDVVEKGNNPPP--LPNNPT------LAQIRNHSDEVAKEGRALAIIHAAL 53
K L +WDVV+ G +P P +P LAQ RN V K+ +AL I+ ++L
Sbjct: 23 KKRLVENGVWDVVQNGVSPNPTKIPQLAATIQAVDLAQWRNR---VIKDNKALKIMQSSL 79
Query: 54 HDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDR 113
D +F K +++ AKE W+ L + TK+ K+ L ++FE L M E E + + R
Sbjct: 80 PDSVFRKTISIASAKELWDLLKK----GNDTKEAKLRRLEKQFEKLMMYEGEPMDLYLKR 135
Query: 114 ISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQAS 173
+ ++ + +LG +SD +V+ K+L L ++ I L+E ++T+ +L+ A +
Sbjct: 136 VEEITERFEVLGNPISDDKVITKLLTSLSWPYDDSIPVLKEFMTLPDLTLRDLLKAFELF 195
Query: 174 EQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL---DKFCWYRPGV 230
+E + K N K+ E+ PC C K+ H D + +RP V
Sbjct: 196 GSHPETMPQELM--------KFINILRKAHSERM--PCGICVKNNHNQEEDLYYNHRPRV 245
Query: 231 -KCRACNQL---GHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLI 286
NQ GH + C N N Q+ + R+ Q+ E + + + W++
Sbjct: 246 VNLSGANQFRGQGHYARDCSNTRNLQQAEKRI----QKPEHLMLGVTVGGITFDEGMWMV 301
Query: 287 DSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVP 346
+ T+HMT FF LD S+ ++V +G+ V +GKG V + T G K I +VLFVP
Sbjct: 302 HTTTTDHMTPYEKFFTTLDRSYRARVGLADGKVVMAEGKGDVMIMTREGKKRIKNVLFVP 361
Query: 347 EINQSLLSVGQMMEKNYSLHFKNMKC 372
IN++ LSV QM ++ S+ F KC
Sbjct: 362 GINKNALSVAQMTDQGCSVTFGGGKC 387
>At3g21030 hypothetical protein
Length = 411
Score = 150 bits (380), Expect = 1e-36
Identities = 111/404 (27%), Positives = 195/404 (47%), Gaps = 47/404 (11%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIR-----NHSDEVAKEGRALAIIHAALHD 55
+KT L +WDVV+ G +P P N A I+ + V K+ +AL I+ ++L D
Sbjct: 22 VKTRLVENGVWDVVQNGVSPNPTTNPELAATIKAVDLAQWRNLVVKDNKALKIMQSSLPD 81
Query: 56 DIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRIS 115
+F K +++ AKE W+ L + TK+ K+ L ++FE L M E E ++ + R+
Sbjct: 82 SVFRKTISIASAKELWDLLKK----GNDTKEAKLRRLEKQFENLMMYEGERMKLYLKRLE 137
Query: 116 KVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQ 175
K+I ++ +LG +SD +V+ K+L LP ++ + L+E E+T +L+ A +
Sbjct: 138 KIIEELFVLGNPISDDKVIAKLLTSLPRSYDDSVPVLKEFMTLPELTHRDLLKAFEMFGS 197
Query: 176 RRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGV----- 230
+E ++ + +++ E+ + C C+ H + C+Y P V
Sbjct: 198 DPKTMPQELMKFIMILR--------RAWSERMW--CSACETYNHNQEDCYYNPRVVYLGG 247
Query: 231 ---KCRACNQLGHVEKVCKNKTNQQ--EQQARVVEHHQEDEEQLFKASCYLACSSKETWL 285
+C C + GH + C N N Q +++ V + DE+ W+
Sbjct: 248 GRGQCFQCGERGHFARDCCNTRNLQPIDRKREVKDDFTYDEDM---------------WM 292
Query: 286 IDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKY-ILDVLF 344
+ + NHM+ FF LD ++ + V NG V V+G G V + G++ I DVLF
Sbjct: 293 LYTTSENHMSPYEKFFTSLDRTYKAGVQLPNGNVVMVEGIGDVKF-MMEGVRMTIKDVLF 351
Query: 345 VPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMR 388
VP IN+++LS +M ++ YS+ + + I D G K+ + +R
Sbjct: 352 VPWINKNILSFSRMKKRGYSMEVDDGEWIIRDRTG-KVFVKTLR 394
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 142 bits (358), Expect = 3e-34
Identities = 101/406 (24%), Positives = 189/406 (45%), Gaps = 49/406 (12%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+ LR +W+ ++ G SD++ K A A++ ++ + ++
Sbjct: 1 MEATLRVHKVWETIDPG------------------SDDMEKNDMARALLFQSVPESTILQ 42
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
+ + +K W + G+ER K+ K+ L EF+ L MK+ ET+ EF RIS++ T+
Sbjct: 43 VGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNMKDNETIDEFVGRISEISTK 102
Query: 121 IRLLGEDLSDQRVVEKILVCLP-EMFEAKISSLEENKNFSEITVPELVNALQASEQR--- 176
LGE++ + ++V+K L LP + + I++LE+ + + ++V ++ E R
Sbjct: 103 SESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNTTGFEDIVGRMKTYEDRVCD 162
Query: 177 -------RSLRMEENVEGAF-----------LASNKGKNQSFKSFGEKKFPPCPHCKKDT 218
+ M N E ++ +S +G+ +K C C K
Sbjct: 163 EDDSPEEQGKLMYANSESSYDTRGGRGRGRGRSSGRGRGGYGYQQRDKSKVICYRCDKTG 222
Query: 219 HLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLAC 278
H C R +L ++ +N + E ++ ++ E+ K + AC
Sbjct: 223 HYASECLDR-------LLKLIKAQEQQQNNEDDDEIESLMMHEVVYLNERSVKPKEFEAC 275
Query: 279 SSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGI-K 337
S +W +D+G +NHMT N+ +F +L+E KV FG+ + +KGKG + + T GI K
Sbjct: 276 SD-NSWYLDNGASNHMTGNLQWFSKLNEMITGKVRFGDDSRIDIKGKGSIVLITKGGIRK 334
Query: 338 YILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM 383
+ DV F+P++ +++S+GQ E + K+ + T+ D G L+
Sbjct: 335 TLTDVYFIPDLKSNIISLGQATEAGCDVRMKDDQLTLHDREGCLLL 380
>At1g36110 hypothetical protein
Length = 745
Score = 135 bits (341), Expect = 3e-32
Identities = 97/386 (25%), Positives = 182/386 (47%), Gaps = 51/386 (13%)
Query: 37 DEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREF 96
D+ K A +I ++ + + +++ L AK W+ + Y G++R K+ ++ L EF
Sbjct: 52 DDGYKNTMARGLIFQSIPESLTLQVGTLATAKLVWDSIKTRYVGADRVKEARLQTLMAEF 111
Query: 97 EALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLP-EMFEAKISSLEEN 155
E +KMKE+E + F+ R++++ T+ LG ++ ++V+K L LP + I+SLE+
Sbjct: 112 EKMKMKESEKIDVFAGRLAELATRSDALGSNIETSKLVKKFLNALPLRKYIHIIASLEQV 171
Query: 156 KNFSEITVPELVNALQASEQR-RSLRMEENVEGAFLASN------------KGKNQSFKS 202
+ + + ++V ++ E+R +E+ +G + +N +G+ F
Sbjct: 172 LDLNNTSFEDIVGRIKVYEERVWDGEEQEDDQGKLMYANTDTQDSWYASRGRGQGGGFNG 231
Query: 203 FGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHH 262
G + +DT V C C++LGH C + + +
Sbjct: 232 RGRGR----GRGSRDT---------SKVTCYRCDKLGHYASNCPDSVKPSVFETEL---- 274
Query: 263 QEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKV 322
W +D+G +NHMT N ++F ++DES KV FG+ + +
Sbjct: 275 ----------------DGDNVWYLDNGASNHMTGNRAYFSKIDESITGKVRFGDDSCIDI 318
Query: 323 KGKGVVAVETLLGIKYIL-DVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSK 381
KGKG + + G K +L +V F+P++ +++S+GQ E ++ K T++D G K
Sbjct: 319 KGKGSILFMSKNGEKKVLAEVYFIPDVKSNIISLGQATEAGCNIRMKENYLTLYDRDG-K 377
Query: 382 LMIVEMRGKSFPRGVEEDILA--CFS 405
L++ MR K+ V ++ A CFS
Sbjct: 378 LLVKAMRSKNRLYKVIMEVKASKCFS 403
>At3g21010 unknown protein
Length = 438
Score = 130 bits (326), Expect = 2e-30
Identities = 115/411 (27%), Positives = 185/411 (44%), Gaps = 52/411 (12%)
Query: 2 KTYLRAQSLWDVVEKGNNPPP--LPNNPTLAQIRNHS---DEVAKEGRALAIIHAALHDD 56
KT L + LWDVVE G P P +P Q S D K+ +AL I+ ++L D
Sbjct: 25 KTTLTEKGLWDVVENGVPPDPSKVPELAATIQAEEFSKWRDLAEKDMKALQILQSSLTDS 84
Query: 57 IFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISK 116
F K L+ AK+ W+ L + ++ K+ L ++FE L M E E + + DR+ +
Sbjct: 85 AFRKTLSASSAKDLWDSLEKG-----NNEQAKLRRLEKQFEELSMNEGEPINPYIDRVIE 139
Query: 117 VITQIRLLGEDLSDQRVVEKILVCLPEMFE---AKISSLEENKNFSEITVPELVNALQA- 172
++ Q+R SD +V+EK+L L ++ + L + KN S + EL +
Sbjct: 140 IVEQLRRFKIAKSDYQVIEKVLATLSGSYDDVAPVLGDLVDLKNMSLKSFVELFYVYDSI 199
Query: 173 SEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFC-------- 224
+E+ L ++ N +S E++ C C K+ H + C
Sbjct: 200 TEESIHLMLKAN--------------RLRSMAEEE--SCVLCNKNNHKQEDCSSSIPKAG 243
Query: 225 ------WYRPGV-KCRACNQLGHVEKVCKNKTNQ---QEQQARVVEHHQEDEEQLFKASC 274
+P KC C + GH K C K Q E Q +VE + + + L A+
Sbjct: 244 KSSGARQSKPKKGKCFQCGERGHKAKDCNKKIQQPAVTEDQIEIVEEKEIEVDFLLLAAL 303
Query: 275 YL--ACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
L + W+I TNHMT F LD ++ +KV +G + V+G G V ++
Sbjct: 304 NLNEFTYDENMWMIYGHATNHMTPYEKLFTTLDRTYKAKVGLVDGTIIMVEGIGDVKIKM 363
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYS-LHFKNMKCTIFDPYGSK 381
G K I +V+FVP +N+++LSV QM + S ++ +C + D G +
Sbjct: 364 KEGKQKTINNVIFVPGLNRNVLSVQQMKSRGCSFINGARGECIVHDRTGKE 414
>At3g21040 hypothetical protein
Length = 534
Score = 122 bits (306), Expect = 4e-28
Identities = 88/314 (28%), Positives = 145/314 (46%), Gaps = 22/314 (7%)
Query: 78 YQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKI 137
Y + K K+L L ++FE L M + E++ + +R+ I Q R G LSD V+ K+
Sbjct: 227 YDKNNPIKDAKLLRLEKQFEKLMMYDGESMDSYLERVLDTIEQFRASGNPLSDDDVIAKL 286
Query: 138 LVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKN 197
L L ++ I LEE N ++T +L+ L+ + M + ++ + K K
Sbjct: 287 LTSLSWPYDDAIPVLEELMNLPDMTPHDLIGVLEMFGSHPEI-MPQGLKDFVKSLRKAKA 345
Query: 198 QSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQAR 257
+ C CK H + C+Y P V + GH + C N N Q
Sbjct: 346 EQLW---------CGVCKMYNHYQEDCYYNPRVVNLGGGERGHCARDCCNTRNLQPGATL 396
Query: 258 VVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNG 317
+ + +++ + ++ W++ + NHMT FF LD ++ + V
Sbjct: 397 QAQKREVNDDFTY---------DEDMWMLYTTTENHMTPYEKFFTSLDRTYRAGVQLAGV 447
Query: 318 QHVKVKGKGVVAVETLLGIKY-ILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFD 376
V V+G G V T+ G+K I DVLFVP IN+++LS +M ++ YSL + +CTI D
Sbjct: 448 NVVMVEGIGDVKF-TMKGVKMTIKDVLFVPRINKNVLSFSRMKKRGYSLEMEGGECTIRD 506
Query: 377 PYGSKLMIVEMRGK 390
G K+ + +R K
Sbjct: 507 EKG-KIFVQTLREK 519
Score = 87.0 bits (214), Expect = 2e-17
Identities = 48/175 (27%), Positives = 95/175 (53%), Gaps = 8/175 (4%)
Query: 2 KTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHS-----DEVAKEGRALAIIHAALHDD 56
+T L +WDVV+ G +P P + A I+ + + ++ +AL I+ +AL D
Sbjct: 23 ETRLVENGVWDVVQNGVSPNPTTDPEVAATIKVEDLTQWRNRMIEDNKALKIMQSALPDS 82
Query: 57 IFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISK 116
+F K +++ +KE W+ L + G++ + K+ L ++ E L M E E ++ + R+ K
Sbjct: 83 VFRKTISVASSKELWDLLKK---GNDNEEAKKLRRLEKQLENLTMYEGERMKSYLKRVEK 139
Query: 117 VITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQ 171
+I + + G +SD++V+ K+L LP ++ I+ L+E ++T +L+ A +
Sbjct: 140 IIEEFFVWGNPISDEKVIAKLLTSLPRPYDDSITVLKEFMTLPDLTHRDLLKAFE 194
>At4g04560 putative transposon protein
Length = 590
Score = 117 bits (292), Expect = 2e-26
Identities = 93/399 (23%), Positives = 166/399 (41%), Gaps = 114/399 (28%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLP--NNPTLAQIRNHSDEVA-KEGRALAIIHAALHDDI 57
MKT + + LW++V++G PP ++P Q + + + K+ AL I+ A+ D I
Sbjct: 22 MKTIFQTKKLWEIVDEGVPKPPAEGDHSPEAVQQKTRCEAASLKDLTALQILQTAVSDSI 81
Query: 58 FIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKV 117
F +I A L E+E LKMKE++ + F ++ ++
Sbjct: 82 FPRIAPASSA----------------------LGKPWEYENLKMKESDNINTFMTKLIEM 119
Query: 118 ITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRR 177
Q+R+ GE+ SD ++V+KIL+ LP+ F+ ++ +++ K+ + ++ + + +
Sbjct: 120 GNQLRVHGEEKSDYQIVQKILISLPKRFDIIVAMMKQTKDLTSLSAGKWCDVCE------ 173
Query: 178 SLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQ 237
R N ++ NKG E+ +C CN+
Sbjct: 174 --RKNHNESDCWMKKNKGVLSQQVGNNER------------------------RCFVCNK 207
Query: 238 LGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSK---ETWLIDSGCTNHM 294
GH+ K C+ + ++ + +E +DE+ + ++ SS ETWL+DSGCTNHM
Sbjct: 208 PGHLAKNCRLRRTERVDLS--LEETNDDEDHMLFSAVEEETSSNVNDETWLVDSGCTNHM 265
Query: 295 TNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLS 354
T V +F LD+S
Sbjct: 266 TKEVKYFITLDQSV---------------------------------------------- 279
Query: 355 VGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP 393
K Y + F++ +C I DP G +++ ++M KSFP
Sbjct: 280 ------KGYRVLFEDNRCLINDPQGRRILDMKMMQKSFP 312
Score = 72.8 bits (177), Expect = 3e-13
Identities = 43/169 (25%), Positives = 88/169 (51%), Gaps = 7/169 (4%)
Query: 11 WDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKE-------GRALAIIHAALHDDIFIKILN 63
W VE G PP + ++ ++ +A E +AL++I +L + F ++
Sbjct: 372 WFAVEDGWMPPTTKDAKRDIVSKSRTEWIADEKTAANHNSQALSVIFGSLLRNKFTQVQG 431
Query: 64 LEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRL 123
AKE W L ++ + K+ ++ L EFE L M+ E+V +F+ ++S + + +
Sbjct: 432 CLSAKEVWEILQVSFECTNNVKRTRLDMLASEFENLTMEAEESVDDFNGKLSSITQEAVV 491
Query: 124 LGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQA 172
LG+ D+++V+K L LP+ F++ S+++ + N ++ ++V +QA
Sbjct: 492 LGKTYKDKKMVKKFLRSLPDKFQSHKSAIDVSLNSDQLKFDQVVGMMQA 540
>At3g21000 unknown protein
Length = 405
Score = 113 bits (282), Expect = 2e-25
Identities = 100/373 (26%), Positives = 169/373 (44%), Gaps = 44/373 (11%)
Query: 2 KTYLRAQSLWDVVEKGNNPPPLPNNPTLA------QIRNHSDEVAKEGRALAIIHAALHD 55
K+ L Q LWDVV G P NP LA ++ D V K+ +AL I+ ++L D
Sbjct: 25 KSTLIEQGLWDVVVNGVPQDP-SKNPELAATIQPEELSKWRDFVVKDAKALQILQSSLTD 83
Query: 56 DIFIKILNLEIAKEAWNKLME--EYQGSERTKKMKVLNLRREFEALKMKETETVREFSDR 113
+F K L+ AK+ W+ L + E R +++ + L ++ E LKM + E+ + D+
Sbjct: 84 SVFRKTLSASSAKDVWDLLRKGNEQATIRRLEQVTIRRLEKQLEDLKMVDKESGSSYLDK 143
Query: 114 ISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQAS 173
+++ ++ + SD + + + L F+ S LEE + ++T LV
Sbjct: 144 ALEILERLGRAKLEKSDYEICKNVFTTLSGSFDGLDSMLEELIDVHKMTSKSLVEYFYYR 203
Query: 174 EQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCR 233
S EE + G K+ KS EK C C K+ H + C +R
Sbjct: 204 VHESS--TEEAIFGLL------KDLRLKSKSEKW---CGLCYKNNHNQEDCKFR------ 246
Query: 234 ACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNH 293
T+++E++ +V ++ + A Y + W+I +
Sbjct: 247 -------------IHTDKEEKEDEIVVDYRLETVPNLGAKTY----DDDIWIIHKMAPIN 289
Query: 294 MTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLG-IKYILDVLFVPEINQSL 352
MT V +F LD +F + V +G + V+GKG V + G K I +V+FVP +N+++
Sbjct: 290 MTPYVKYFTTLDRTFKATVGTVDGTVLLVEGKGDVKIRMKEGKKKTIRNVIFVPGLNRNV 349
Query: 353 LSVGQMMEKNYSL 365
LS G+M+ K YS+
Sbjct: 350 LSFGKMVSKRYSI 362
>At4g04420 putative transposon protein
Length = 1008
Score = 106 bits (264), Expect = 3e-23
Identities = 76/268 (28%), Positives = 124/268 (45%), Gaps = 40/268 (14%)
Query: 44 RALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKE 103
RAL++I ++ + F +I N E AKEAW+KL + Y+G+ K+ ++ L +FE L M+E
Sbjct: 80 RALSLIFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLSMEE 139
Query: 104 TETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITV 163
TE + EFS +IS + ++ LG+ D+++V+K+L CLP FE+K +++ + + I
Sbjct: 140 TENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDSIDF 199
Query: 164 PELVNALQASEQRRSLRMEENVEGAFLASNKGKNQ----------------------SFK 201
E+V LQA E + +G LA++ KN+ K
Sbjct: 200 EEVVGMLQAYELEITSGKGGYSKGLALAASAKKNEIQELKDTMSMMAKDFSRAMRRVEKK 259
Query: 202 SFGEKKFPP-------------CPHCKKDTHLDKFC--WYRPGVKCRACNQLGHVEKVC- 245
FG + C C+ H+ C R +KC CN LGH + C
Sbjct: 260 GFGRNQGTDRYRDRSSKRDEIQCHECQGYGHIKAECPSLKRKDLKCSECNGLGHTKFDCV 319
Query: 246 --KNKTNQQEQQARVVEHHQEDEEQLFK 271
K+K ++ + + D E K
Sbjct: 320 GSKSKPDKSCSSESESDSNDGDSEDYIK 347
Score = 68.2 bits (165), Expect = 8e-12
Identities = 41/155 (26%), Positives = 76/155 (48%), Gaps = 14/155 (9%)
Query: 229 GVKCRACNQLGHVEKVCKN---KTNQQEQQARVVEHHQEDEEQLFKAS---CYLACSS-- 280
G +C C + GH+++ C + N+ ++Q ++ + + + C++A +S
Sbjct: 543 GYECYYCGRHGHIQRYCYRYAARLNKLKRQGKLYPYQGRTSKMYVRREDLYCHVAYTSIE 602
Query: 281 ---KETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIK 337
K+ W DSG + HM + S + SKV FG G K+KGKG + T
Sbjct: 603 EGIKKPWYFDSGASRHMVGSQSNLENYTSVKESKVTFGGGDKRKIKGKGDL---TKAEKP 659
Query: 338 YILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKC 372
+ +V FV + +L+SV Q+ ++ ++ F ++KC
Sbjct: 660 QLTNVYFVEGLTANLISVSQLCDEGLTVSFNSVKC 694
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 104 bits (260), Expect = 8e-23
Identities = 70/231 (30%), Positives = 115/231 (49%), Gaps = 34/231 (14%)
Query: 44 RALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKE 103
RAL++I +++ + F +I N E AKEAW+KL + Y+G+ K+ ++ L +FE L M+E
Sbjct: 92 RALSLIFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTMEE 151
Query: 104 TETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITV 163
TE + EFS +IS + ++ LG+ D+++V+K+L CLP FE+K +++ + + + I
Sbjct: 152 TENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTNSIDF 211
Query: 164 PELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKF 223
E+V QA E + S KG +G K CP K
Sbjct: 212 EEVVGMFQAYELE-------------ITSGKG------GYGHIK-AECPSLK-------- 243
Query: 224 CWYRPGVKCRACNQLGHVEKVC---KNKTNQQEQQARVVEHHQEDEEQLFK 271
R +KC C LGH++ C K+K ++ + + D E K
Sbjct: 244 ---RKDLKCSECKGLGHIKFDCVGSKSKPDRSCSSESESDSNDGDSEDYIK 291
Score = 57.4 bits (137), Expect = 1e-08
Identities = 31/92 (33%), Positives = 49/92 (52%), Gaps = 3/92 (3%)
Query: 281 KETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYIL 340
K+ W DSG + HMT + S + SKV FG G K+KGKG + T +
Sbjct: 514 KKPWYFDSGASRHMTGSQSNLENYTSVKESKVTFGGGDKGKIKGKGDL---TKAEKPQLT 570
Query: 341 DVLFVPEINQSLLSVGQMMEKNYSLHFKNMKC 372
+V FV + +L+SV Q+ ++ ++ F ++KC
Sbjct: 571 NVYFVEGLTANLISVSQLCDEGLTVSFNSVKC 602
>At2g10610 pseudogene
Length = 531
Score = 103 bits (257), Expect = 2e-22
Identities = 72/294 (24%), Positives = 138/294 (46%), Gaps = 23/294 (7%)
Query: 95 EFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLP-EMFEAKISSLE 153
EF LKMK++ + +F+ +S++ + LG + + R+V+K L +P + + +++LE
Sbjct: 3 EFHRLKMKDSHKIDDFAGMLSEISVKSAALGVIIEESRLVKKFLKSVPRKRYIHIVAALE 62
Query: 154 ENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPH 213
+ + + IT ++V L+ E+R + + + L + + +S GE++ P
Sbjct: 63 QVLDLNAITFDDIVGRLKVYEERVYDEDDPEEDQSKLLYSSMDSSKRESEGEEEEDPIGE 122
Query: 214 CKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKAS 273
+D G + NK Q + + E +E+ +S
Sbjct: 123 EDED---------------------GMETQENDNKETQVADELMMHEVVYLNEQSCMPSS 161
Query: 274 CYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETL 333
+ W +D+G +NHMT + +F ++D S K+ FG+ + +KGKG +A L
Sbjct: 162 FETNVDGENVWYLDNGASNHMTGDRRYFDKMDHSITGKIRFGDDSRIDIKGKGTIAFIDL 221
Query: 334 LG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVE 386
G + +LDV F+PE+ +++S+GQ E + K+ T+ D G L+ E
Sbjct: 222 NGKPRVMLDVYFIPELKSNIISLGQATESGCEIKLKDGYLTMHDQEGKLLVKAE 275
>At2g17490 putative retroelement pol polyprotein
Length = 822
Score = 93.2 bits (230), Expect = 2e-19
Identities = 51/114 (44%), Positives = 66/114 (57%)
Query: 279 SSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKY 338
S TWLIDSGCTNHMT N F +++ F + GNG + +GKG + V T +
Sbjct: 25 SGGNTWLIDSGCTNHMTPNEKLFTKINRDFKVPIRVGNGAVMMSEGKGDIEVMTRKDKRG 84
Query: 339 ILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSF 392
I DVL VP++ ++LLSV QM+ Y + KN CTI D K+ VEM KSF
Sbjct: 85 IRDVLLVPKLGKNLLSVPQMIINGYQVTLKNNYCTIHDSARKKIGEVEMVNKSF 138
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 91.3 bits (225), Expect = 9e-19
Identities = 93/390 (23%), Positives = 164/390 (41%), Gaps = 61/390 (15%)
Query: 4 YLRAQSLWDVVEKGNNPPPLPN----NPTLAQIRNHSDEVAKEGR---ALAIIHAALHDD 56
+L L DVV G P PL +P + R+ +DEVA+ R A +I + D
Sbjct: 3 HLSVIGLKDVV-LGTTPKPLTEEEEEDPEKRKKRD-ADEVARLERCDKAKNVIFLNVADK 60
Query: 57 IFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISK 116
+ KI + A EAW L + ++ + F KM+E + + E D K
Sbjct: 61 VLRKIELCKTAAEAWETLDRLFMIRSLPHRVYT---QLSFYTFKMQENKKIDENIDDFLK 117
Query: 117 VITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQR 176
++ + L D++D+ +L LP ++ + +++ + + ++ + +++ A + E+
Sbjct: 118 IVADLNHLQIDVTDEVQAILLLSSLPARYDGLVETMKYSNSREKLRLDDVMVAARDKERE 177
Query: 177 RSLRMEENVEGAFLAS----------NKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWY 226
S VEG F NKGKN+S + K + CW
Sbjct: 178 LSQNNRPVVEGHFARGRPDGKNNNQGNKGKNRSRSKSADGK--------------RVCWI 223
Query: 227 RPGVKCRACNQLGHVEKVC-----KNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSK 281
C + GH +K C +NK+ QQ E + F + L + +
Sbjct: 224 --------CGKEGHFKKQCYKWIERNKSKQQGSDNG--ESSLAKSTEAFNPAMVLLATDE 273
Query: 282 ---------ETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
W++D+GC+ HMT +FK+ E V GN + VKG G + +
Sbjct: 274 TLVVTDSIANEWVLDTGCSFHMTPRKDWFKDFKELSSGYVKMGNDTYSPVKGIGSIKIRN 333
Query: 333 LLGIKYIL-DVLFVPEINQSLLSVGQMMEK 361
G + IL DV ++P + ++L+S+G + ++
Sbjct: 334 SDGSQVILTDVRYMPNMTRNLISLGTLEDR 363
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,505,944
Number of Sequences: 26719
Number of extensions: 411030
Number of successful extensions: 1865
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 68
Number of HSP's that attempted gapping in prelim test: 1580
Number of HSP's gapped (non-prelim): 222
length of query: 416
length of database: 11,318,596
effective HSP length: 102
effective length of query: 314
effective length of database: 8,593,258
effective search space: 2698283012
effective search space used: 2698283012
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144931.10


