

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
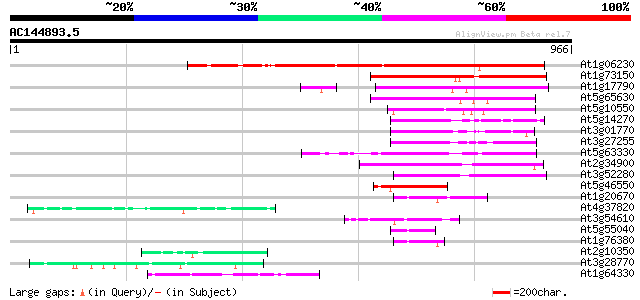
Score E
Sequences producing significant alignments: (bits) Value
At1g06230 Ring3-like bromodomain protein 494 e-139
At1g73150 unknown protein 254 2e-67
At1g17790 unknown protein (At1g17790) 242 6e-64
At5g65630 unknown protein 212 9e-55
At5g10550 bromodomain protein - like 208 1e-53
At5g14270 kinase - like protein 149 1e-35
At3g01770 unknown protein 144 3e-34
At3g27255 unknown protein 143 4e-34
At5g63330 unknown protein 140 3e-33
At2g34900 RING3 protein-like 123 4e-28
At3g52280 putative protein 114 3e-25
At5g46550 unknown protein 109 7e-24
At1g20670 hypothetical protein 56 1e-07
At4g37820 unknown protein 55 2e-07
At3g54610 histone acetyltransferase (HAT1) 52 2e-06
At5g55040 putative protein 49 1e-05
At1g76380 unknown protein 49 2e-05
At2g10350 hypothetical protein 47 5e-05
At3g28770 hypothetical protein 47 7e-05
At1g64330 unknown protein 46 9e-05
>At1g06230 Ring3-like bromodomain protein
Length = 766
Score = 494 bits (1271), Expect = e-139
Identities = 294/632 (46%), Positives = 401/632 (62%), Gaps = 43/632 (6%)
Query: 306 DVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKLEDMTSQ 365
D N + DS ++++ E S QP+ +D S + + ++ +
Sbjct: 93 DKNLTEAPSENLPGDDSDKVIDKPLVEAFSQAQPQ-----DDASLAAMDKSEEVPSQIPK 147
Query: 366 QQD--NSILEDGNS-SQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDDLHINQQA 422
QD N+++ D NS +P +L E + V D N + SQ++S +
Sbjct: 148 AQDDVNTVVVDENSIKEPPKSLAQEDVTTVIV------DKNPIEAPSQTLSLEDGDTLVV 201
Query: 423 EPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLEDDGPSSPIHRHKAVPS 482
+ + + V E+D +H A DNL ++ ++P +E D + S
Sbjct: 202 DKNPIEVSSEED-----VHVIDA--DNL-----IKEAHPENFVERDTTDAQQPAGLTSDS 249
Query: 483 TEYRHSENVTEEPNLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVG 542
+ ++ E + + RI+I++AST+KQ+K+E+ KLE +L+ VR +VK+IE K+G +G
Sbjct: 250 AHATAAGSMPMEEDADGRIRIHVASTTKQQKEEIRKKLEDQLNVVRGMVKKIEDKEGEIG 309
Query: 543 GYGNSNVVLGGGITNGGGAQRAHSEAGSVGVSRQ---PTRPLHQLSFPMFQNSRRASEGV 599
Y +S V++ GI NGGG R S S G+ R+ RP++QLS + +N++ +E V
Sbjct: 310 AYNDSRVLINTGINNGGG--RILSGFASAGLPREVIRAPRPVNQLSISVLENTQGVNEHV 367
Query: 600 EKEKRMPKANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSC 659
EKEKR PKANQFY NSEFLL DK PPAESNKKSK + KKQGG D+G G G+K FK+C
Sbjct: 368 EKEKRTPKANQFYRNSEFLLG-DKLPPAESNKKSKSSSKKQGG-DVGHGFGAGTKVFKNC 425
Query: 660 SSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAE 719
S+LLE+LM+H++ WVFN PVDV GLGL DY+TII +PMDLGT+K+ L KN YKSP+EFAE
Sbjct: 426 SALLERLMKHKHGWVFNAPVDVKGLGLLDYYTIIEHPMDLGTIKSALMKNLYKSPREFAE 485
Query: 720 DVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRY--GMDYGAPSPLS 777
DVRLTFHNAMTYNP+GQDVH MA L +IFE+RWA+IE+DYNREMR+ G + P+P
Sbjct: 486 DVRLTFHNAMTYNPEGQDVHLMAVTLLQIFEERWAVIEADYNREMRFVTGYEMNLPTPTM 545
Query: 778 R-RVPAFTPPPPLDMRRILDRQEPFARTPRS------MNNTPSSRTPAPKKPKAKDPNKR 830
R R+ PPPP+++R +DR + R P + + TPS RTPA KKPKA +PNKR
Sbjct: 546 RSRLGPTMPPPPINVRNTIDRADWSNRQPTTTPGRTPTSATPSGRTPALKKPKANEPNKR 605
Query: 831 DMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDR 890
DMTY+EKQKLS L +LP +KLDA+VQ++ K+N + +D+EIEVD D+VD E LWELDR
Sbjct: 606 DMTYEEKQKLSGHLQNLPPDKLDAIVQIVNKRNTAVKLRDEEIEVDIDSVDPETLWELDR 665
Query: 891 FVLNYKKSLSKNKRKAEQA-RERAEALQNSVQ 921
FV NYKK LSK KRKAE A + RAEA +NS Q
Sbjct: 666 FVTNYKKGLSKKKRKAELAIQARAEAERNSQQ 697
>At1g73150 unknown protein
Length = 461
Score = 254 bits (649), Expect = 2e-67
Identities = 149/322 (46%), Positives = 195/322 (60%), Gaps = 27/322 (8%)
Query: 622 DKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTPVDV 681
+ F P + K N K+GG + + KSC++LL KLM+H+ W+FNTPVDV
Sbjct: 86 NNFAPVPNKKLKTANGGKKGGVHGAAADKGTVQILKSCNNLLTKLMKHKSGWIFNTPVDV 145
Query: 682 DGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAM 741
LGLHDY II PMDLGTVKTRL+K+ YKSP EFAEDVRLTF+NAM YNP G DV+ M
Sbjct: 146 VTLGLHDYHNIIKEPMDLGTVKTRLSKSLYKSPLEFAEDVRLTFNNAMLYNPVGHDVYHM 205
Query: 742 AEQLSKIFEDRWAIIESDYNREMR-----YGMDYGAP----------SPLSRRVPAFTPP 786
AE L +FE++W +E+ Y +R +D+ AP PL P+ +PP
Sbjct: 206 AEILLNLFEEKWVPLETQYELLIRKQQPVRDIDFHAPVSTNTHNVEALPLPAPTPSLSPP 265
Query: 787 PPLDM--RRILDRQEPFARTPRSMNN--TPSSRTPAPKKPKAKDPNKRDMTYDEKQKLST 842
PP + R L+R E SM N P+ P+K + RD+T+DEK++LS
Sbjct: 266 PPPKVVENRTLERAE-------SMTNPVKPAVLPVVPEKLVEEASANRDLTFDEKRQLSE 318
Query: 843 SLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKN 902
L LP +KL+AVVQ+++K+ L+QQDDEIE+D D++D E LWEL RFV YK+SLSK
Sbjct: 319 DLQDLPYDKLEAVVQIIKKRTPELSQQDDEIELDIDSLDLETLWELFRFVTEYKESLSKK 378
Query: 903 KRKAEQARER-AEALQNSVQSS 923
K + ER AE+ NSV S
Sbjct: 379 KEEQGLDSERDAESFHNSVHES 400
Score = 42.0 bits (97), Expect = 0.002
Identities = 30/91 (32%), Positives = 48/91 (51%), Gaps = 4/91 (4%)
Query: 497 LEDR---IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGG 553
LED +KI+L+S SK E + + KL++EL+ VRSL+KR+E + N +
Sbjct: 41 LEDNHQMMKISLSSISKLEVRNLKRKLQAELEEVRSLIKRLEPQGNNFAPVPNKKLKTAN 100
Query: 554 GITNGGGAQRAHSEAGSVGVSRQPTRPLHQL 584
G GG A ++ G+V + + L +L
Sbjct: 101 G-GKKGGVHGAAADKGTVQILKSCNNLLTKL 130
>At1g17790 unknown protein (At1g17790)
Length = 487
Score = 242 bits (618), Expect = 6e-64
Identities = 136/321 (42%), Positives = 190/321 (58%), Gaps = 23/321 (7%)
Query: 631 KKSKLNWKKQGGGDMGXGLQMGS-KFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDY 689
+ K+ GG G G G+ + FK+C+SLL KLM+H+ AWVFN PVD GLGLHDY
Sbjct: 107 RSKKVKTGNGGGKKSGHGADKGTVQIFKNCNSLLTKLMKHKSAWVFNVPVDAKGLGLHDY 166
Query: 690 FTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
I+ PMDLGTVKT+L K+ YKSP +FAEDVRLTF+NA+ YNP G DV+ AE L +F
Sbjct: 167 HNIVKEPMDLGTVKTKLGKSLYKSPLDFAEDVRLTFNNAILYNPIGHDVYRFAELLLNMF 226
Query: 750 EDRWAIIESDYN---------REMRYGMDYGAPSPLSRRVPAFT----------PPPPLD 790
ED+W IE Y+ R++ + + +P+ +PA PPPP
Sbjct: 227 EDKWVSIEMQYDNLHRKFKPTRDIEFPAPAPSIAPIVEPLPAIVPSPSPSSPPPPPPPPV 286
Query: 791 MRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDP--NKRDMTYDEKQKLSTSLHSLP 848
+L+ + ++ P + AP+K + ++ N RD+T +EK++LS L LP
Sbjct: 287 AAPVLENRTWEREESMTIPVEPEAVITAPEKAEEEEAPVNNRDLTLEEKRRLSEELQDLP 346
Query: 849 VEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
+KL+ VVQ+++K N L+Q+DDEIE+D D++D LWEL RFV YK+SLSK
Sbjct: 347 YDKLETVVQIIKKSNPELSQKDDEIELDIDSLDINTLWELYRFVTGYKESLSKKNEAHGF 406
Query: 909 ARER-AEALQNSVQSSRLPIS 928
ER AE++ NS+Q +S
Sbjct: 407 GSERDAESVHNSIQEPTTLVS 427
Score = 45.1 bits (105), Expect = 2e-04
Identities = 29/72 (40%), Positives = 39/72 (53%), Gaps = 9/72 (12%)
Query: 501 IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIE---------VKQGYVGGYGNSNVVL 551
+KI+L+S SK E + + KL+SELD VRSL+KR + K G VG
Sbjct: 56 LKISLSSISKLEVRNLKRKLKSELDEVRSLIKRFDPEANPGGSMAKSGVVGRSKKVKTGN 115
Query: 552 GGGITNGGGAQR 563
GGG +G GA +
Sbjct: 116 GGGKKSGHGADK 127
>At5g65630 unknown protein
Length = 590
Score = 212 bits (539), Expect = 9e-55
Identities = 133/345 (38%), Positives = 179/345 (51%), Gaps = 60/345 (17%)
Query: 621 NDKFPPAESNKK--SKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTP 678
ND P + KK S L Q G ++ + +CS +L KLM+H++AWVFNTP
Sbjct: 133 NDLGPKKKKQKKNVSGLKRSNQFGPSDPESEKLLAGMLNTCSQILVKLMKHKWAWVFNTP 192
Query: 679 VDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDV 738
VDV GLGLHDY ++ PMDLGTVK L+K +Y SP +FA DVRLTF NAMTYNPKGQDV
Sbjct: 193 VDVVGLGLHDYHQVVKKPMDLGTVKLNLDKGFYVSPIDFATDVRLTFDNAMTYNPKGQDV 252
Query: 739 HAMAEQLSKIFEDRWAIIESDYN-REMRYGMDYGAPSP---------------------- 775
+ MA++L F+ + + ++++ P P
Sbjct: 253 YFMADKLLDHFDGMFNPAFKKFEAQQLKLTGSSSRPEPDFKPDFKQRQWNQNPPMVANPR 312
Query: 776 -------LSRRVPAFTPPPPLDMRRILD-----RQEPFARTPRSMNNTPSSRTPAPK--- 820
+++++ + PP P ++++ P P P + P P+
Sbjct: 313 KGTEQISIAKKLDSVKPPQPTLPPQLVEPSRVQSPSPPPPPPVIQPELPQPQPPPPQLEI 372
Query: 821 --------------------KPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMR 860
KPKAKDPNKR MT +EK KL +L LP EKL ++Q++R
Sbjct: 373 EVEAPPDVSEVSKGRKGKLPKPKAKDPNKRLMTMEEKSKLGMNLQDLPPEKLGQLLQILR 432
Query: 861 KKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRK 905
K+N L Q DEIE+D +AVD E LWELDRFV NYKK SK KR+
Sbjct: 433 KRNGHLAQDGDEIELDIEAVDNETLWELDRFVTNYKKMASKIKRQ 477
>At5g10550 bromodomain protein - like
Length = 678
Score = 208 bits (530), Expect = 1e-53
Identities = 128/308 (41%), Positives = 166/308 (53%), Gaps = 54/308 (17%)
Query: 651 MGSKFFKS----CSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRL 706
M K KS C +L KLM+H+++WVF PVDV GLGLHDY I+ PMDLGTVK L
Sbjct: 241 MSEKVLKSMMTTCGQILVKLMKHKWSWVFLNPVDVVGLGLHDYHRIVDKPMDLGTVKMNL 300
Query: 707 NKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRW--AIIESDYNREM 764
K Y+SP +FA DVRLTF NAM+YNPKGQDV+ MAE+L F D W ++ +E+
Sbjct: 301 EKGLYRSPIDFASDVRLTFTNAMSYNPKGQDVYLMAEKLLSQF-DVWFNPTLKRFEAQEV 359
Query: 765 RYGMDYGAPSPLSRR-------------------------------VPAFTPPPPLDMRR 793
+ P P + +P PPP +++ R
Sbjct: 360 KVMGSSSRPGPEDNQRVWNQNNVAENARKGPEQISIAKKLDSVKPLLPTLPPPPVIEITR 419
Query: 794 -------ILDRQEPFARTPRSMNNTPSS---------RTPAPKKPKAKDPNKRDMTYDEK 837
+ P + P+ +N +S R KPKAKDPNKR+MT DEK
Sbjct: 420 DPSPPPSPVQPPPPPSPPPQPVNQVEASLEVRETNKGRKGKLPKPKAKDPNKREMTMDEK 479
Query: 838 QKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKK 897
KL +L LP EKL ++Q++RK+ L Q DEIE+D +A+D E LWELDRFV NY+K
Sbjct: 480 GKLGVNLQELPPEKLGQLIQILRKRTRDLPQDGDEIELDIEALDNETLWELDRFVTNYRK 539
Query: 898 SLSKNKRK 905
SK KR+
Sbjct: 540 MASKIKRQ 547
>At5g14270 kinase - like protein
Length = 688
Score = 149 bits (375), Expect = 1e-35
Identities = 100/267 (37%), Positives = 129/267 (47%), Gaps = 32/267 (11%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +LL++LM HQY WVFNTPVDV L + DYF +I +PMDLGTVK +L Y P E
Sbjct: 139 KQCEALLKRLMSHQYGWVFNTPVDVVKLNILDYFNVIEHPMDLGTVKNKLTSGTYSCPSE 198
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NAMTYNP G DV+ MA+ L K FE RW +E +
Sbjct: 199 FAADVRLTFSNAMTYNPPGNDVYVMADTLRKFFEVRWKTLEKKLS--------------- 243
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
T P LD + E P M +T A DP KR MT ++
Sbjct: 244 --GTKVHTEPSNLDAHK-----EKHIVIPVPM--AKKRKTTAVDCENVVDPAKRVMTDED 294
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ---QDDEIEVDFDAVDAEILWELDRFVL 893
+ KL L SL E ++ +R N N+ DDEIE+D + + L++L +
Sbjct: 295 RLKLGKDLESL-TEFPAQLINFLRDHN--SNEGGIGDDEIEIDINDLSDHALFQLRDLLD 351
Query: 894 NYKKSLSKNKRKAEQARERAEALQNSV 920
+ + + K E E L SV
Sbjct: 352 EHLREIQNKKSSVEPC--EIELLHGSV 376
>At3g01770 unknown protein
Length = 620
Score = 144 bits (362), Expect = 3e-34
Identities = 98/254 (38%), Positives = 132/254 (51%), Gaps = 36/254 (14%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C SLL++LM Q+ W+FNTPVDV L + DYFTII +PMDLGTVK++L Y SP E
Sbjct: 131 KQCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSE 190
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
F+ DVRLTF NAMTYNP +V+ A+ LSK FE RW IE + PS L
Sbjct: 191 FSADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKKSSGTK------SEPSNL 244
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ P EP A+ R MN A K+ +P KR MT ++
Sbjct: 245 ATLAHKDIAIP-----------EPVAKK-RKMN--------AVKRNSLLEPAKRVMTDED 284
Query: 837 KQKLSTSLHSL---PVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWEL----D 889
+ KL L SL PV+ ++ + K+ DDEIE+D + + + L++L D
Sbjct: 285 RVKLGRDLGSLTEFPVQIINFLRDHSSKEE---RSGDDEIEIDINDLSHDALFQLRDLFD 341
Query: 890 RFVLNYKKSLSKNK 903
F+ +K S +
Sbjct: 342 EFLRENQKKDSNGE 355
>At3g27255 unknown protein
Length = 503
Score = 143 bits (361), Expect = 4e-34
Identities = 95/252 (37%), Positives = 123/252 (48%), Gaps = 24/252 (9%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +LL KL H ++WVF PVDV L + DY T I +PMDLGTVK L Y SP E
Sbjct: 178 KQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKNLASGVYSSPHE 237
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NAMTYNP G DVH M + LSK+FE RW I+ P
Sbjct: 238 FAADVRLTFTNAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK------------LPPCS 285
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ +PA T P D R+ P + R M + P P KP MT E
Sbjct: 286 MQTLPAVTLEPN-DERKAAISVPPAKK--RKMASPVRESVPEPVKPL--------MTEVE 334
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ-QDDEIEVDFDAVDAEILWELDRFVLNY 895
+ +L L SL E ++ ++K N + +DEIE+D D + E+L L + Y
Sbjct: 335 RHRLGRQLESLLDELPAHIIDFLKKHNSNGGEIAEDEIEIDIDVLSDEVLVTLRNLLDEY 394
Query: 896 KKSLSKNKRKAE 907
++ + E
Sbjct: 395 IQNKEAKQTNVE 406
>At5g63330 unknown protein
Length = 477
Score = 140 bits (353), Expect = 3e-33
Identities = 116/409 (28%), Positives = 179/409 (43%), Gaps = 83/409 (20%)
Query: 503 INLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGGGITNGGGAQ 562
++L+ S+ E++ + KL+ EL VR L K+I + + V+L
Sbjct: 61 LSLSKMSRSERKNLVHKLKMELQQVRDLSKKI-------ASFSSDTVLL----------- 102
Query: 563 RAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNSEFLLAND 622
S S P RP + +F F S+
Sbjct: 103 ---SPYNDHSCSDGPRRPPPE-NFATFVGSQ---------------------------GK 131
Query: 623 KFPPAESNKKSKLNWKKQGGGDMGXGLQMG-SKFFKSCSSLLEKLMEHQYAWVFNTPVDV 681
K PP S+K+ K+G + + K C +LL +L H+ W F TPVD
Sbjct: 132 KRPPVRSDKQRN----KKGPSRLNVPTSYTVASVMKECETLLNRLWSHKSGWPFRTPVDP 187
Query: 682 DGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAM 741
L + DYF +I +PMDLGT+++RL K Y SP +FA DVRLTF N++ YNP G H M
Sbjct: 188 VMLNIPDYFNVIKHPMDLGTIRSRLCKGEYSSPLDFAADVRLTFSNSIAYNPPGNQFHTM 247
Query: 742 AEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPP-PLDMRRILDRQEP 800
A+ +SK FE W IE +++P PP PL L+ + P
Sbjct: 248 AQGISKYFESGWKSIE--------------------KKIPMSKPPVIPLTSSASLESEIP 287
Query: 801 FARTPRSMNNTPSSRTPAPKKPKAK-DPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMM 859
F P + A K + +P K MT EK+KL L +L + + ++
Sbjct: 288 FEVAPM------RKKEAAMNDNKLRVEPAKLVMTDGEKKKLGQDLMALEEDFPQKIADLL 341
Query: 860 RKKNL*LNQQ-DDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAE 907
R+++ Q + EIE+D +A+ EIL+ + + + +Y + K+ K+E
Sbjct: 342 REQSGSDGQSGEGEIEIDIEALSDEILFMVRKLLDDYLREKKKSMEKSE 390
>At2g34900 RING3 protein-like
Length = 386
Score = 123 bits (309), Expect = 4e-28
Identities = 92/343 (26%), Positives = 154/343 (44%), Gaps = 40/343 (11%)
Query: 602 EKRMPKANQFYHNSEFLLANDKFPPAESNKK---SKLNWKK--QGGGDMGXGLQMGSK-F 655
E+++ + FY + + KK S+ N K G + G + S
Sbjct: 51 EQKVVEVEHFYSTKDGAAQTNTSKSNSGGKKIAISQPNNSKGNSAGKEKSKGKHVSSPDL 110
Query: 656 FKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPK 715
+ +++ ++ +H++AW F PVDV GLGLHDY+ +I PMDLGT+K ++ + Y + +
Sbjct: 111 MRQFATMFRQIAQHKWAWPFLEPVDVKGLGLHDYYKVIEKPMDLGTIKKKMESSEYSNVR 170
Query: 716 EFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSP 775
E DVRL F NAM YN + +DV+ MAE L + FE++W +I E + +D A
Sbjct: 171 EIYADVRLVFKNAMRYNEEKEDVYVMAESLLEKFEEKWLLIMPKLVEEEKKQVDEEAEKH 230
Query: 776 LSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNK-RDMTY 834
+++ L + A R ++N +K + + R ++
Sbjct: 231 ANKQ---------------LTMEAAQAEMARDLSNELYEIDLQLEKLRESVVQRCRKLST 275
Query: 835 DEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLN 894
EK+ LS +L L E L ++M+ + N E+E+D D LW L FV
Sbjct: 276 QEKKGLSAALGRLSPEDLSKALKMVSESNPSFPAGAPEVELDIDVQTDVTLWRLKVFVQE 335
Query: 895 YKKSLSK------------------NKRKAEQARERAEALQNS 919
K+ +K NK A++ RE ++A+ +
Sbjct: 336 ALKAANKSSGGTNAQNNNNTGTGEINKNNAKRRREISDAINKA 378
>At3g52280 putative protein
Length = 369
Score = 114 bits (284), Expect = 3e-25
Identities = 73/269 (27%), Positives = 129/269 (47%), Gaps = 20/269 (7%)
Query: 661 SLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRL---NKNWYKSPKEF 717
++ ++ +H+ AW F PV+V+GLGLHDYF +I PMD T+K ++ + YK +
Sbjct: 100 TIFRQITQHKCAWPFMHPVNVEGLGLHDYFEVIDKPMDFSTIKNQMEAKDGTGYKHVMQI 159
Query: 718 AEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLS 777
D+RL F NAM YN + DV++MA++L + FE++WA E + + +
Sbjct: 160 YADMRLVFENAMNYNEETSDVYSMAKKLLEKFEEKWAHFLPKVQEEEKIREEEEKQAA-- 217
Query: 778 RRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNK-RDMTYDE 836
+L ++ +T R + N +K K + R +T +E
Sbjct: 218 -------------KEALLAKEASHIKTTRELGNEICHANDELEKLMRKVVERCRKITIEE 264
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYK 896
K+ + +L L + L V+ ++ + N + +E+ ++ D +D LW L FV +
Sbjct: 265 KRNIGLALLKLSPDDLQKVLGIVAQANPSFQPRAEEVSIEMDILDEPTLWRLKFFVKDAL 324
Query: 897 KSLSKNKRKAE-QARERAEALQNSVQSSR 924
+ K K++ E + RE + A + V R
Sbjct: 325 DNAMKKKKEEETKTRELSGAQKKEVSKKR 353
>At5g46550 unknown protein
Length = 494
Score = 109 bits (273), Expect = 7e-24
Identities = 60/133 (45%), Positives = 83/133 (62%), Gaps = 10/133 (7%)
Query: 627 AESNKKSKLNWKKQGGGDMGXGLQMGSK------FFKSCSSLLEKLMEHQYAWVFNTPVD 680
++S++KSK K+GG +Q K + C +LL LMEH+ W+F PVD
Sbjct: 39 SQSSEKSK----KRGGPKELDEVQPKKKQRLDCDWSSQCLALLRFLMEHRGGWLFKEPVD 94
Query: 681 VDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHA 740
+ + DYF +I PMDLGTVK++L KN Y + EFA DVRLTF NAM YNP +VH
Sbjct: 95 PVKMEIPDYFNVIQKPMDLGTVKSKLLKNVYSNADEFAADVRLTFANAMHYNPLWNEVHT 154
Query: 741 MAEQLSKIFEDRW 753
+A+++++IFE RW
Sbjct: 155 IAKEINEIFEVRW 167
>At1g20670 hypothetical protein
Length = 652
Score = 55.8 bits (133), Expect = 1e-07
Identities = 47/165 (28%), Positives = 67/165 (40%), Gaps = 8/165 (4%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+L++L + V++ PVD + L DYF II NPMD T++ +L+ Y + ++F DV
Sbjct: 183 ILDRLQKKDTYGVYSDPVDPEELP--DYFEIIKNPMDFSTLRNKLDSGAYSTLEQFERDV 240
Query: 722 RLTFHNAMTYNPKG----QDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLS 777
L NAM YN + A+ E K FE+ +SD P
Sbjct: 241 FLICTNAMEYNSADTVYYRQARAIQELAKKDFENLRQ--DSDDEEPQSQQQQQQQPKVAR 298
Query: 778 RRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKP 822
R P P P + R A P +N S K P
Sbjct: 299 RGRPPKKHPEPSSIDRTASEISADALIPGDSSNKFSGAYNLRKAP 343
>At4g37820 unknown protein
Length = 532
Score = 54.7 bits (130), Expect = 2e-07
Identities = 83/444 (18%), Positives = 175/444 (38%), Gaps = 64/444 (14%)
Query: 31 INPNIEETV-----LKDENIPTTIVTSD--TDNNNGTTLENIDEAPEFAVLGDGNLAKAE 83
+ P IEET ++DE D TD NG E ++ E + + N +AE
Sbjct: 69 LRPRIEETKDVKDEVEDEEGSKNEGGGDVSTDKENGD--EIVEREEEEKAVEENNEKEAE 126
Query: 84 GSSRLEDGSSIQQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADGNSPQ 143
G+ E+G+ + +V+E G ++ E+N AS ++ G +
Sbjct: 127 GTGN-EEGNEDSNNGESEKVVDESE----GGNEISNEEAREINYKGDDASSEVMHGTEEK 181
Query: 144 --QKVENQDSAHPQASLRIG---DGNSPQQQLVEPNLAQPQASLRAGDGNSPQRQLEEEN 198
+KVE + + ++ + D + P+ +++E ++ + + SL + S + +E
Sbjct: 182 SNEKVEVEGESKSNSTENVSVHEDESGPKNEVLEGSVIK-EVSLNTTENGSDDGEQQETK 240
Query: 199 SAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLVQPQASSIIED 258
S + S+TG+ D N ++ NL A+ E
Sbjct: 241 S---ELDSKTGEKGF-------------------SDSNG---ELPETNLSTSNATETTES 275
Query: 259 GNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQT-----HVSSRTEDVNSPRPQ 313
S + +S + D+ + Q E+ + ++ SS ++ +P+
Sbjct: 276 SGSDESGSSGKSTGYQQTKNEEDEKEKVQSSEEESKVKESGKNEKDASSSQDESKEEKPE 335
Query: 314 NSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKLEDMTSQQQDNSILE 373
+++ S + E+ + E +S E+ +PE +K ++ +S Q++N I E
Sbjct: 336 RKKKEESSSQ---GEGKEEEPEKREKEDSSSQEESKEEEPE--NKEKEASSSQEENEIKE 390
Query: 374 DGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDDLHINQQAEPSNLNVRQED 433
++ + E SS + N +E + ++ + + I Q + N ++ D
Sbjct: 391 ------TEIKEKEESSSQEGNENKETEKKSSESQRKENTNSEKKIEQVESTDSSNTQKGD 444
Query: 434 DRPSSPIHRQGAISDNLHSHQQVE 457
++ + R+ S N S+++ E
Sbjct: 445 EQKTDESKRE---SGNDTSNKETE 465
>At3g54610 histone acetyltransferase (HAT1)
Length = 568
Score = 52.0 bits (123), Expect = 2e-06
Identities = 51/213 (23%), Positives = 88/213 (40%), Gaps = 32/213 (15%)
Query: 577 PTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNSEFLLANDKFPPAESNKKSKLN 636
P P LS M + R+A + E+ + + Y EFL N+ P + K ++
Sbjct: 372 PKLPYTDLS-SMIRQQRKAID--ERIRELSNCQNVYPKIEFL-KNEAGIPRKIIKVEEIR 427
Query: 637 WKKQGGGDMGXGLQMGSKFFKSCS--------------SLLEKLMEHQYAWVFNTPVDVD 682
++ G K F + +LL+ + +H AW F PVD
Sbjct: 428 GLREAGWTPDQWGHTRFKLFNGSADMVTNQKQLNALMRALLKTMQDHADAWPFKEPVD-- 485
Query: 683 GLGLHDYFTIITNPMDLGTVKTRL-NKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAM 741
+ DY+ II +P+DL + R+ ++ +Y + F D R F+N TYN +
Sbjct: 486 SRDVPDYYDIIKDPIDLKVIAKRVESEQYYVTLDMFVADARRMFNNCRTYNSPDTIYYKC 545
Query: 742 AEQLSKIFEDRWAIIESDYNREMRYGMDYGAPS 774
A +L E+ ++ +++ G+ GA S
Sbjct: 546 ATRL-----------ETHFHSKVQAGLQSGAKS 567
>At5g55040 putative protein
Length = 145
Score = 49.3 bits (116), Expect = 1e-05
Identities = 29/76 (38%), Positives = 40/76 (52%), Gaps = 2/76 (2%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
KS +L+KL + V+ PVD + L DY +I +PMD TV+ +L Y + +E
Sbjct: 49 KSLELILDKLQKKDIYGVYAEPVDPEELP--DYHDMIEHPMDFSTVRKKLANGSYSTLEE 106
Query: 717 FAEDVRLTFHNAMTYN 732
DV L NAM YN
Sbjct: 107 LESDVLLICSNAMQYN 122
>At1g76380 unknown protein
Length = 579
Score = 48.5 bits (114), Expect = 2e-05
Identities = 30/92 (32%), Positives = 47/92 (50%), Gaps = 6/92 (6%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+L+++ + V++ P D + L DY+ II NPMD T++ +L Y + ++F +DV
Sbjct: 152 ILDRVQKKDTYGVYSDPADPEELP--DYYEIIKNPMDFTTLRKKLESGAYTTLEQFEQDV 209
Query: 722 RLTFHNAMTYNPKG----QDVHAMAEQLSKIF 749
L NAM YN + AM E K F
Sbjct: 210 FLICTNAMEYNSADTVYYRQARAMLELAKKDF 241
>At2g10350 hypothetical protein
Length = 863
Score = 47.0 bits (110), Expect = 5e-05
Identities = 60/233 (25%), Positives = 92/233 (38%), Gaps = 31/233 (13%)
Query: 227 PASLRIGDGNSLRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSR-TDDGDS 285
P + + +S + H QP+ + + D N K +D S + P +++ TDD
Sbjct: 322 PIQSQTPETSSQSHTLSKHVFGQPEVAQQVPDDNL--DKAQDSSDSTPLINTEITDD--- 376
Query: 286 AQQQVEDQNLAQTHVSSRTEDVNSPRP--------QNSTQKDGDSPQILENSRPEDASLT 337
V L H S +D N P QN+ ++D DS +DAS
Sbjct: 377 -PMDVFVTPLQSEH--SNNDDANEGNPVYDIDVKYQNANEEDVDSQM-------QDASER 426
Query: 338 QPEVNSRLE---DRSFPQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPV 394
P ++S L+ + ++ + E N+ ED+ SQ +D S S L LQ
Sbjct: 427 VPAIHSGLDLSKEHNYEEQESNANDEDVDSQMKDASERVPATHSGLDLPKEHNSEELQTN 486
Query: 395 ANSTSEDLNLAQPLSQ---SVSD-DLHINQQAEPSNLNVRQEDDRPSSPIHRQ 443
AN T D + L + S SD DL I Q +E E + P+ Q
Sbjct: 487 ANKTDVDGKMQDSLDREPASHSDIDLSIEQSSEEQQAKTPPEVNFDGDPMKDQ 539
>At3g28770 hypothetical protein
Length = 2081
Score = 46.6 bits (109), Expect = 7e-05
Identities = 87/463 (18%), Positives = 171/463 (36%), Gaps = 66/463 (14%)
Query: 35 IEETVLKDENIPTTIVTSDTDNNNGTTLENIDEAP-EFAVLGDGNLAKAEGSSRLEDGSS 93
IE+ + E++ + + + D ++ T EN+ EA V N + EG + S
Sbjct: 289 IEKNLESKEDVKSEVEAAKNDGSSMT--ENLGEAQGNNGVSTIDNEKEVEGQGESIEDSD 346
Query: 94 IQQQLGDQNLVEEHA-------SSRTG-----DRNSPQQQLEELNSAQPHASFQLADG-- 139
I++ L + V+ SS TG RN+ E +NS + D
Sbjct: 347 IEKNLESKEDVKSEVEAAKNAGSSMTGKLEEAQRNNGVSTNETMNSENKGSGESTNDKMV 406
Query: 140 ----NSPQQKVENQDSAHPQASLRI---------GDGNSPQQQLVEPNLAQPQ----ASL 182
N K EN++ H + G+ S + + +E + + AS+
Sbjct: 407 NATTNDEDHKKENKEETHENNGESVKGENLENKAGNEESMKGENLENKVGNEELKGNASV 466
Query: 183 RAGDGNSPQRQLEEENSAQPQA---SSRTGDGNLPHQQ------------LEDQYSAQPP 227
A N ++ + E S + + T G + Q LE++ +P
Sbjct: 467 EAKTNNESSKEEKREESQRSNEVYMNKETTKGENVNIQGESIGDSTKDNSLENKEDVKPK 526
Query: 228 ASLRIGDGNSLRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQ 287
DGNS +++ H Q ED N ++Q V T+DGD +
Sbjct: 527 VDANESDGNSTKER---HQEAQVNNGVSTEDKNLDNIGADEQKKNDKSVEVTTNDGDHTK 583
Query: 288 QQVED------QNLAQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEV 341
++ E+ +++ ++ ++ ED + S ++ LE R + +
Sbjct: 584 EKREETQGNNGESVKNENLENK-EDKKELKDDESVGAKTNNETSLEEKREQTQKGHDNSI 642
Query: 342 NSRLEDRSFPQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRL------EGSSLQPVA 395
NS++ D + N + E +++ +E ++ ++ ++ +G +
Sbjct: 643 NSKIVDNKGGNADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSEKGEEGKENN 702
Query: 396 NSTSEDLNLAQPLSQSVS-DDLHINQQAEPSNLNVRQEDDRPS 437
+ ED L SQ+ S DD ++ + E + + + D S
Sbjct: 703 KDSMEDKKLENKESQTDSKDDKSVDDKQEEAQIYGGESKDDKS 745
Score = 39.7 bits (91), Expect = 0.008
Identities = 81/551 (14%), Positives = 211/551 (37%), Gaps = 74/551 (13%)
Query: 10 REIQRYSEGKVYRRKVFKGI--KINPNIEETVLKDENIPTTIVTSDTDNNNGTTLENIDE 67
+E ++ +G+ K K + K N + T +DE + + D +++ E
Sbjct: 780 KESEKVEKGEKKESKDAKSVETKDNKKLSSTENRDEAKERSGEDNKEDKEESKDYQSV-E 838
Query: 68 APEFAVLG--DGNLAKAEGSSRLEDGSSIQQQLG-DQNLVEEHASSRTGDRNSPQQQLEE 124
A E G D N+ E S L+D S++ + ++++ ++ + D++S ++ +
Sbjct: 839 AKEKNENGGVDTNVGNKEDSKDLKDDRSVEVKANKEESMKKKREEVQRNDKSSTKEVRDF 898
Query: 125 LNSAQPHASFQLADGNSPQ---------QKVENQDSAHPQASLRIGDGNSPQQQLVEPNL 175
N+ Q G S + K EN+D+ + + + D +++ N+
Sbjct: 899 ANNMD--IDVQKGSGESVKYKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNM 956
Query: 176 AQPQASLRAGDGNSPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDG 235
+ + + N ++Q + + +S+ + N +++ ++ +
Sbjct: 957 KKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDS----------A 1006
Query: 236 NSLRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNL 295
+ R+K E + + S E+ ++K++D+ + R S +++ E ++L
Sbjct: 1007 SKNREKKE----YEEKKSKTKEEAKKEKKKSQDKKREEKDSEERK----SKKEKEESRDL 1058
Query: 296 AQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPEL 355
T++ S +K+ + ED + E ED+ +
Sbjct: 1059 KAKKKEEETKEKKESENHKSKKKE-------DKKEHEDNKSMKKE-----EDKKEKKKHE 1106
Query: 356 NSKLEDMTSQQQDNSILEDGNSSQPQ---------LNLRLEGSSLQPVANSTSEDLNLAQ 406
SK ++D LED NS++ + +++L +E+ + +
Sbjct: 1107 ESKSRKKEEDKKDMEKLEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETK 1166
Query: 407 PLSQSVSDDLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLE 466
+ S S ++++ + S+ + +++ ++ S+ + E +E
Sbjct: 1167 EIESSKSQKNEVDKKEKKSSKDQQKKKEKEMKE-------SEEKKLKKNEEDRKKQTSVE 1219
Query: 467 DDGPSSPIHRHKAVPSTEYRHS-----------ENVTEEPNLEDRIKINLASTSKQEKQE 515
++ + K P + +++ E+ ++E + + + + S + K E
Sbjct: 1220 ENKKQKETKKEKNKPKDDKKNTTKQSGGKKESMESESKEAENQQKSQATTQADSDESKNE 1279
Query: 516 MCLKLESELDA 526
+ ++ +S+ D+
Sbjct: 1280 ILMQADSQADS 1290
>At1g64330 unknown protein
Length = 555
Score = 46.2 bits (108), Expect = 9e-05
Identities = 65/303 (21%), Positives = 124/303 (40%), Gaps = 40/303 (13%)
Query: 238 LRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQ 297
+R+KV G + +SS + +K++ + + + A ++ D +
Sbjct: 84 IRKKVHGKG--ENDSSSSSSSDSDSDKKSKRNGRGENEIELLKKQMEDANLEIADLKMKL 141
Query: 298 THVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQP--EVNSRLEDRSFPQPEL 355
E V S Q +K +S +I N R E LT E+N +LE + +L
Sbjct: 142 ATTDEHKEAVESEH-QEILKKLKESDEICGNLRVETEKLTSENKELNEKLEVAGETESDL 200
Query: 356 NSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDD 415
N KLED+ ++ DG L E +S ST E++N Q +
Sbjct: 201 NQKLEDVKKER-------DG--------LEAELASKAKDHESTLEEVNRLQGQKNETEAE 245
Query: 416 LHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLEDDGPSSPIH 475
L +Q +P+ LN Q +D + + ++ A + H+Q+ N L E+
Sbjct: 246 LEREKQEKPALLN--QINDVQKALLEQEAAYNTLSQEHKQI-----NGLFEE-------- 290
Query: 476 RHKAVP--STEYRHSENVTEE-PNLEDRIKINLASTSKQ--EKQEMCLKLESELDAVRSL 530
R + + +Y+ + + EE + + + + T K ++ + LE ++++R+
Sbjct: 291 REATIKKLTDDYKQAREMLEEYMSKMEETERRMQETGKDVASRESAIVDLEETVESLRNE 350
Query: 531 VKR 533
V+R
Sbjct: 351 VER 353
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,115,537
Number of Sequences: 26719
Number of extensions: 1123310
Number of successful extensions: 4155
Number of sequences better than 10.0: 205
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 175
Number of HSP's that attempted gapping in prelim test: 3822
Number of HSP's gapped (non-prelim): 431
length of query: 966
length of database: 11,318,596
effective HSP length: 109
effective length of query: 857
effective length of database: 8,406,225
effective search space: 7204134825
effective search space used: 7204134825
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC144893.5


