

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.8 - phase: 0
(361 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
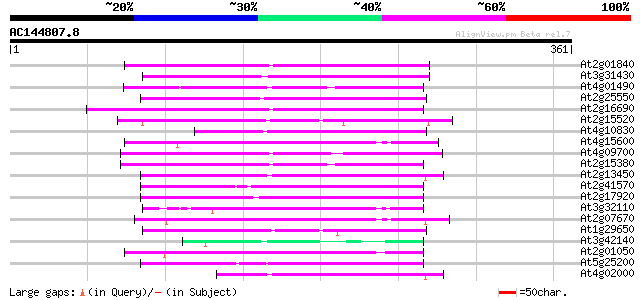
Score E
Sequences producing significant alignments: (bits) Value
At2g01840 putative non-LTR retroelement reverse transcriptase 69 3e-12
At3g31430 hypothetical protein 69 4e-12
At4g01490 putative transposon protein 67 2e-11
At2g25550 putative non-LTR retroelement reverse transcriptase 67 2e-11
At2g16690 putative Ta11-like non-LTR retroelement protein 64 1e-10
At2g15520 putative Ta11-like non-LTR retroelement protein 64 2e-10
At4g10830 putative protein 63 2e-10
At4g15600 hypothetical protein 63 3e-10
At4g09700 putative protein 62 5e-10
At2g15380 putative non-LTR retroelement reverse transcriptase 62 5e-10
At2g13450 T10F5.3 62 6e-10
At2g41570 putative Ta11-like non-LTR retroelement protein 60 2e-09
At2g17920 putative Ta11-like non-LTR retroelement protein 60 2e-09
At3g32110 non-LTR reverse transcriptase, putative 60 2e-09
At2g07670 putative non-LTR retrolelement reverse transcriptase 59 5e-09
At1g29650 reverse transcriptase, putative 59 5e-09
At3g42140 putative protein 57 1e-08
At2g01050 hypothetical protein 57 2e-08
At5g25200 putative protein 55 5e-08
At4g02000 putative zinc finger protein 55 5e-08
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 69.3 bits (168), Expect = 3e-12
Identities = 45/197 (22%), Positives = 86/197 (42%), Gaps = 3/197 (1%)
Query: 75 IQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQED 134
+Q +++ + +LG+ + + + I + W R I ++ F F +E
Sbjct: 28 VQRAVDDNRFCLLGRPLMPRNQNLRQILTTVPRTWGLVGFVRGRIIQNRRFQFIFPSEES 87
Query: 135 TKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGS 194
++R PW F + L+V W D+D L F P W+Q+R +P Q ++ I
Sbjct: 88 LDTVLRRGPWSFADRMLVVERWTPDMDPLVLNF--IPFWIQVREIPLQFLNLEVIDNIAG 145
Query: 195 SIGTVLASELYEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFC 253
S+G A + + ++ ++I++ V+ P++ + + F YE L FC
Sbjct: 146 SLGERKAVDFDPFTTTRVEFVRIQIKWDVNHPLRFQRNYQFSLGVNTVLSFYYERLRGFC 205
Query: 254 FACGLIGHTEANCKLHD 270
CG + H +C L +
Sbjct: 206 DVCGRMTHDAGSCVLQN 222
>At3g31430 hypothetical protein
Length = 336
Score = 68.9 bits (167), Expect = 4e-12
Identities = 43/186 (23%), Positives = 82/186 (43%), Gaps = 5/186 (2%)
Query: 86 ILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWF 145
+ G+ V + + SI ++ IW + + FHF +E + ++R PW
Sbjct: 134 LFGRPVMPRRQNLRSIVASMPRIWGQSGLVHGRIMEGRQFHFIFTLEESLETVLRRGPWA 193
Query: 146 FRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL- 204
F + +++ W I F P WVQ+RG+P Q + + IG ++G VL ++
Sbjct: 194 FNDWMILLQRWEPQIPL----FPFIPFWVQIRGIPFQFLNRGVVEHIGRALGQVLDTDFN 249
Query: 205 YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEA 264
E + ++ ++ ++ P++ + + FRYE L FC CG++ H
Sbjct: 250 VEVVARMDFARVLLHWDITHPLRFQRHFQFTAGVNTLLRFRYERLRGFCEVCGMLTHDFG 309
Query: 265 NCKLHD 270
C + +
Sbjct: 310 ACLIQN 315
>At4g01490 putative transposon protein
Length = 577
Score = 66.6 bits (161), Expect = 2e-11
Identities = 51/194 (26%), Positives = 82/194 (41%), Gaps = 9/194 (4%)
Query: 74 DIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRV-EHIGDKLFHFFMDEQ 132
D E ++E +++G + + A S+ LSN W N KG +G F F + +
Sbjct: 28 DNSELIQENSLTLMGILTNPSAQRLWSLIPFLSNRW-NLKGKATGSDLGRGCFQFRFEYE 86
Query: 133 EDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKI 192
ED ++++ P+ F +I+ W+ I P W++L+GLP K+M I
Sbjct: 87 EDLQKVLDNRPYHFGQWMVILQRWKPVISPSFPS--EIPFWIELQGLPMHYWIKEMLYAI 144
Query: 193 GSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQF 252
G G V+ E+ KIKV + PI + + Y+NL
Sbjct: 145 GKEAGEVVDHEI-----SPAAAKIKVLINGLQPITKETVVEFPDGSEALVYLEYKNLKSH 199
Query: 253 CFACGLIGHTEANC 266
C C + H EA+C
Sbjct: 200 CHHCQRLSHAEADC 213
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 66.6 bits (161), Expect = 2e-11
Identities = 42/184 (22%), Positives = 76/184 (40%), Gaps = 2/184 (1%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
S++ V+ + + ++ + +W P I F +E ++R PW
Sbjct: 38 SLVVTTVNPRKQNLRALIGQMPRVWGFPDSCVGRIIDKGRVQFKFQSEEAMNLVLRRGPW 97
Query: 145 FFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL 204
F + L +H W ++ E + P WVQ+ G+P T M +G+ +G V +
Sbjct: 98 SFNDWMLSIHRWYPNLS--EAEMKIIPFWVQITGIPLLFLTNAMARCVGNRLGHVADVDF 155
Query: 205 YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEA 264
E + +++++N + P++ A I FR+E L FC CG + H
Sbjct: 156 DENSNHTGFVRVRINWNLDDPLRFQRNFQFADGENTVIKFRFERLRNFCTKCGSLKHDIK 215
Query: 265 NCKL 268
C L
Sbjct: 216 VCTL 219
>At2g16690 putative Ta11-like non-LTR retroelement protein
Length = 240
Score = 64.3 bits (155), Expect = 1e-10
Identities = 48/218 (22%), Positives = 90/218 (41%), Gaps = 3/218 (1%)
Query: 50 NDPTEETVTEDTAETDSFIYYDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIW 109
N ++ + E T DS + + +E S++G+ ++ ++ + IW
Sbjct: 2 NMELDKAIFEMTINDDSPLILSNQPQLSSIERNSCSLIGRFLNPSHQRMSNWILDMPRIW 61
Query: 110 CNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRH 169
R + + F FF +D I++ W +I+ W L
Sbjct: 62 RIYSRVRGVALSQERFQFFYKSNDDLLEILKTGVWTQDEWCVIMDRWVEKPPEEYLMI-- 119
Query: 170 APIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL-YEYPDKKLIIKIKVNLAVSTPIKA 228
PIW++LR +P T++ KI +G V+ EL E + ++++VNL V P++
Sbjct: 120 LPIWIRLRNIPVNYYTEETIKKIAGCVGQVVKVELDLEKSQAQDYVRVQVNLDVRNPLRN 179
Query: 229 GIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEANC 266
+ + F YE + + CF C + H +A+C
Sbjct: 180 SKSVQVPSGEVVSVTFDYERIRKRCFFCQRLTHDKADC 217
>At2g15520 putative Ta11-like non-LTR retroelement protein
Length = 483
Score = 63.5 bits (153), Expect = 2e-10
Identities = 47/223 (21%), Positives = 99/223 (44%), Gaps = 11/223 (4%)
Query: 70 YDDSDIQERLEECQN--SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHF 127
++ D+ E L +N S++G++++ ++ + W + + ++ F F
Sbjct: 17 FEMPDLPEYLSSERNKFSLIGRVLNPACQPMKNLLRNMPRKWQKEGKVKGIALNNERFQF 76
Query: 128 FMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQ 187
D + D + ++ F + ++V W + L R+ +WVQ+ +P T +
Sbjct: 77 IFDSEHDLEEVLAKGVHTFNDWSIVVDRWYENPPDDYL--RYMLLWVQIWNIPVNYNTAK 134
Query: 188 MGIKIGSSIGTVLASELYEYPDKKLI---IKIKVNLAVSTPIKAGIYIGSAKDGAHWIDF 244
K+G IG V E+ P+ K I +++KV VS P++ + G+ ++F
Sbjct: 135 AITKLGDLIGEV--KEVVFNPELKQIREFVRVKVLFDVSRPLRRSKVVNFRNGGSATVNF 192
Query: 245 RYENLPQFCFACGLIGHTEANCKL--HDTRTNAESRNKNVLGP 285
YE + + C+ C + H + C + + A +R K ++ P
Sbjct: 193 HYERIQKRCYECQRLTHEKDVCPILVKARQDLATARRKGIIIP 235
>At4g10830 putative protein
Length = 1294
Score = 63.2 bits (152), Expect = 2e-10
Identities = 38/149 (25%), Positives = 65/149 (43%), Gaps = 2/149 (1%)
Query: 120 IGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGL 179
+G+ F +E +++ PW F + L VH W +I E + P WVQ++G+
Sbjct: 73 VGNGRVQFKFRNEESMNLVLQRGPWSFNDWMLSVHRWFPNITEG--EMKIIPFWVQIQGI 130
Query: 180 PTQCRTKQMGIKIGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGA 239
P T M +GS +G V + E ++ +++K+ ++ ++
Sbjct: 131 PILYLTNAMARVVGSRLGYVTDVDFDENANQMGFVRVKLAWNFDDHLRFQRNFQFQENEN 190
Query: 240 HWIDFRYENLPQFCFACGLIGHTEANCKL 268
I FR+E L FC CG + H C L
Sbjct: 191 TIIKFRFERLRNFCTKCGSLKHDVKECNL 219
>At4g15600 hypothetical protein
Length = 655
Score = 62.8 bits (151), Expect = 3e-10
Identities = 42/207 (20%), Positives = 89/207 (42%), Gaps = 10/207 (4%)
Query: 75 IQERLEECQNSILGKIVSEKAIHRNSIQNALS----NIWCNPKGFRVEHIGDKLFHFFMD 130
I++ + + N + + + K + R++ LS ++W G V + + F +
Sbjct: 80 IEKEVLDAMNGLYKQCMLVKILGRHTTIEVLSRKLRDLWRPTGGMSVLDLPRQFFMVRFE 139
Query: 131 EQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGI 190
+ED + G PW S L+V W + + P+WV++ LP ++ +
Sbjct: 140 VEEDYMMALTGGPWRVLGSILMVQAWSPEFNPLRDVIETTPVWVRVANLPVTFYHNEILL 199
Query: 191 KIGSSIGTVLASELYEY-PDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENL 249
I + +G + +L ++ ++ V + + P+K + + + +++ YE L
Sbjct: 200 GIAAGLGKPIKVDLTTLRKERGRFARVCVEVNLKNPLKGTLVVNGER---YFVS--YEGL 254
Query: 250 PQFCFACGLIGHTEANCKLHDTRTNAE 276
C CG+ GHT NC + T ++
Sbjct: 255 QTICSLCGIYGHTVNNCPMGRTTQGSQ 281
>At4g09700 putative protein
Length = 371
Score = 62.0 bits (149), Expect = 5e-10
Identities = 43/207 (20%), Positives = 88/207 (41%), Gaps = 9/207 (4%)
Query: 72 DSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDE 131
+S+ + E + ++LG++ + + ++ + W +G + F F +
Sbjct: 30 ESENSALIAENKFTLLGRVTNPQIQRPRALVEYMVQYWNLENRVIGRELGPERFFFRFET 89
Query: 132 QEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIK 191
+ D + +++ P+ F+ I+ W + F P W+++ +P T+
Sbjct: 90 EADLQLVLKKAPYHFKKWMFILQRWEPIVSEAFPAF--IPFWIKVNDIPMHHCTELTLNT 147
Query: 192 IGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQ 251
IG ++G V+ +L + +I+VN+ P++ I I + YE L +
Sbjct: 148 IGQALGPVIDKDLSDG-------RIRVNINGLEPLEMKIPIRLPTGEVTTVFLEYEKLEK 200
Query: 252 FCFACGLIGHTEANCKLHDTRTNAESR 278
CF+C + H E C L +N ESR
Sbjct: 201 HCFSCFSLSHEEKACPLKPLVSNKESR 227
>At2g15380 putative non-LTR retroelement reverse transcriptase
Length = 1311
Score = 62.0 bits (149), Expect = 5e-10
Identities = 47/195 (24%), Positives = 79/195 (40%), Gaps = 7/195 (3%)
Query: 72 DSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDE 131
D D + +EE +++G++ + ++ LSN W +G +F D
Sbjct: 30 DLDTSDLIEENALTLIGRLTNPSGQRLWALFLFLSNRWTLRGKATGSDLGQGVFQLKFDF 89
Query: 132 QEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIK 191
ED ++++ P+ F +I+ W I P W++L+GLP +M
Sbjct: 90 SEDLQQVLDNRPYHFDQWMVILQKWEPVISPSFPCL--IPFWIELQGLPKHYWKPEMLKS 147
Query: 192 IGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQ 251
IG +GTV+ E+ +KIKV L PI + + Y+NL
Sbjct: 148 IGEELGTVMDQEI-----TSSTVKIKVLLDGLQPITKETIVDFPDGREAVVYLDYKNLKN 202
Query: 252 FCFACGLIGHTEANC 266
C C + H E +C
Sbjct: 203 HCRHCHRLTHEEKHC 217
>At2g13450 T10F5.3
Length = 394
Score = 61.6 bits (148), Expect = 6e-10
Identities = 36/202 (17%), Positives = 86/202 (41%), Gaps = 9/202 (4%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
S++ + ++ ++ + +++ +AL W + D F + D ++R PW
Sbjct: 38 SLIARPLNPRSQNLHAVISALPRAWGLTNRVHGRVLNDTFVQFIFQSEIDLLSVLRREPW 97
Query: 145 FFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL 204
+ N ++ W ++ L +WVQ+RG+P ++ ++I +G ++ +
Sbjct: 98 LYNNWFVTAQRWEVNLTFHLLT--SIELWVQMRGIPLLYVCEETALEIAHELGEIITLDF 155
Query: 205 YEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTE 263
++ ++ I++++ ++ ++ + I I F+YE L + C +C + H
Sbjct: 156 HDSTTTQIAYIRVRIRFGITDRLRFFLRIIFDSGETALISFQYERLRRICSSCFRMTHHR 215
Query: 264 ANC------KLHDTRTNAESRN 279
+C LH RN
Sbjct: 216 NSCLYRQIESLHRVTNTTAQRN 237
>At2g41570 putative Ta11-like non-LTR retroelement protein
Length = 418
Score = 60.1 bits (144), Expect = 2e-09
Identities = 46/184 (25%), Positives = 83/184 (45%), Gaps = 5/184 (2%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
SILG+ ++ ++ + IW R + + F FF ++D I++ W
Sbjct: 37 SILGRFLNPSNQRMSNWILDMPRIWRLYSRVRGVALSQERFQFFFKSEDDLLEILKTGVW 96
Query: 145 FFRNSWLIVHPWRRDIDARSLEF-RHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASE 203
++ W +V R I+ + E+ P+W++LR +P T+ KI S +G VL E
Sbjct: 97 T-QDEWCVV--MERWIEKSTEEYLMFLPVWMRLRNIPVNYYTQDTIKKIASCVGKVLKVE 153
Query: 204 L-YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHT 262
L E + I+++V + V P++ I + F YE + + CF C + H
Sbjct: 154 LDLEKSQAQDYIRVQVIIDVRNPLRNFKEIQLPTGEIVSVTFDYERIRKRCFLCQRLTHE 213
Query: 263 EANC 266
+ +C
Sbjct: 214 KGDC 217
>At2g17920 putative Ta11-like non-LTR retroelement protein
Length = 336
Score = 60.1 bits (144), Expect = 2e-09
Identities = 40/184 (21%), Positives = 80/184 (42%), Gaps = 5/184 (2%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
SI+ + ++ + + +I AL W I D F + D + R PW
Sbjct: 38 SIIARPLNPRVQNLQAIITALPRAWGLTAHVHGRIIDDTYVQFLFQSEMDLLSVQRREPW 97
Query: 145 FFRNSWLIVHPWRRDIDARSLEF-RHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASE 203
F N ++ W+ A +L F +WVQ+RG+P +++ ++I IG +++ +
Sbjct: 98 LFNNWFVASQRWQ---PAPALNFVTTIDLWVQMRGIPFLYVSEETALEIAQEIGAIISLD 154
Query: 204 LYEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHT 262
++ ++ I+++V + ++ ++ I + I F+YE L + C C H
Sbjct: 155 FHDTTSTQIAYIRVRVRVGITDSLRFFQRITFESGESALIRFQYERLRRICSNCFRFTHN 214
Query: 263 EANC 266
C
Sbjct: 215 RNYC 218
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 59.7 bits (143), Expect = 2e-09
Identities = 44/185 (23%), Positives = 83/185 (44%), Gaps = 17/185 (9%)
Query: 86 ILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFM---DEQEDTKRIIRGN 142
+LG+ VS A+ + L +W P G ++ D HFFM +++E+ + G
Sbjct: 102 VLGRKVSLAAV-----SHRLREMW-KPSG--AMYVLDLPRHFFMVRFEKEEEFLTALTGG 153
Query: 143 PWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLAS 202
PW S L+V W + + E PIWV++ +P K + + + +G L
Sbjct: 154 PWRAFGSCLLVKAWSPEFNPLKDEIVTTPIWVRIADMPVSFYHKSILMSVAQGLGRPLKV 213
Query: 203 EL-YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGH 261
+L + ++ ++ V + + P++ + + + +++ YE L C CG+ GH
Sbjct: 214 DLTMLHFERGRFARVCVEVDLKRPLQGALLVNGDR---YYV--AYEGLTNICSKCGIYGH 268
Query: 262 TEANC 266
NC
Sbjct: 269 LVHNC 273
>At2g07670 putative non-LTR retrolelement reverse transcriptase
Length = 913
Score = 58.5 bits (140), Expect = 5e-09
Identities = 42/210 (20%), Positives = 90/210 (42%), Gaps = 12/210 (5%)
Query: 81 ECQNSILGKIVSEKAIHRN----SIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTK 136
+ NS+ + + K + RN ++ L +W V + + F + +++
Sbjct: 46 DVMNSMWKQCMVVKVLGRNISIANLNRRLREMWKPQGAMFVVDLPCQFFMIRFEREDEYL 105
Query: 137 RIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSI 196
+ G PW S+L+V W + D E PIWV+L +P + + I S+
Sbjct: 106 SALTGGPWRAFGSYLLVQAWSPEFDPLRDEITTTPIWVRLMNIPLSLYHTSILMGIAGSL 165
Query: 197 GTVLASELYE-YPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFA 255
G + ++ + ++ ++ + + ++ P+K + + + +++ YE L C
Sbjct: 166 GKPVKVDMTTLHVERARFARMCIEVDLAKPLKGTLLLNGER---YFVS--YEGLANICSR 220
Query: 256 CGLIGHTEANC--KLHDTRTNAESRNKNVL 283
CG+ GH C + + NA S++ V+
Sbjct: 221 CGMYGHLIHTCPQTIAEKEVNAASQSAPVV 250
>At1g29650 reverse transcriptase, putative
Length = 1557
Score = 58.5 bits (140), Expect = 5e-09
Identities = 41/186 (22%), Positives = 77/186 (41%), Gaps = 7/186 (3%)
Query: 86 ILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWF 145
++G++++ + + W + + D+ F F D + D + ++
Sbjct: 39 LIGRVLNPSCQPMKHLIRNMPRKWQKEGKVKGVALSDERFQFIFDSKHDLEDVLAKGVHS 98
Query: 146 FRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELY 205
F +IV W L F P+WVQ+ +P T++ K+G IG V E+
Sbjct: 99 FNEWSIIVDRWYEHPPDDYLRFM--PLWVQIWNIPVNYYTEKAITKLGDLIGEV--KEVV 154
Query: 206 EYPD---KKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHT 262
PD + +++KV VS P++ I + + F YE + + C+ C + H
Sbjct: 155 FNPDLHQAQKFVRVKVLFDVSRPLRRSKVINFKHGESATVKFHYERIQKRCYECQRLTHE 214
Query: 263 EANCKL 268
+ C L
Sbjct: 215 KDMCPL 220
>At3g42140 putative protein
Length = 273
Score = 57.4 bits (137), Expect = 1e-08
Identities = 40/160 (25%), Positives = 62/160 (38%), Gaps = 39/160 (24%)
Query: 112 PKGFRVEHIGDKL----FHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRR-DIDARSLE 166
PK E +G L F +E I+R PW F + ++ W + DA E
Sbjct: 45 PKNVDEEVVGRILEIHKIEFLFQSEESMFSILRRGPWSFNDWMCVIQRWTKLHSDA---E 101
Query: 167 FRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPI 226
F+ P W+Q+RG+P + T ++ IG +G L + NL +
Sbjct: 102 FKRIPFWIQIRGIPLRFLTARIITSIGERMGLFL----------------ETNLGRDVSV 145
Query: 227 KAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEANC 266
+ F+YE L FC CG++ H + C
Sbjct: 146 ---------------LKFQYEKLKNFCTTCGMLSHDASEC 170
>At2g01050 hypothetical protein
Length = 515
Score = 56.6 bits (135), Expect = 2e-08
Identities = 40/197 (20%), Positives = 80/197 (40%), Gaps = 10/197 (5%)
Query: 75 IQERLEECQNSILGKIVSEKAIHR----NSIQNALSNIWCNPKGFRVEHIGDKLFHFFMD 130
I E + E N + K + K + + + L +W V + + F +
Sbjct: 65 IGEEVLEAMNGLWKKCMIVKVLGSQIPISVLNRKLRELWKPSGVMTVMDLPRQFFMIRFE 124
Query: 131 EQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGI 190
+E+ + G PW ++L+V W D + P+WV+L +P + + +
Sbjct: 125 LEEEYMAALTGGPWRVLGNYLLVQDWSSRFDPLRDDIVTTPVWVRLSNIPYNYYHRCLLM 184
Query: 191 KIGSSIGTVLASELYEYP-DKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENL 249
+I +G L ++ DK ++ + + ++ P+K + I + YE L
Sbjct: 185 EIARGLGRPLKVDMNTINFDKGRFARVCIEVNLAKPLKGTVLINGDR-----YFVAYEGL 239
Query: 250 PQFCFACGLIGHTEANC 266
+ C +CG+ GH +C
Sbjct: 240 SKICSSCGIYGHLVHSC 256
>At5g25200 putative protein
Length = 367
Score = 55.5 bits (132), Expect = 5e-08
Identities = 38/183 (20%), Positives = 79/183 (42%), Gaps = 3/183 (1%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
S++ + ++ + + NS+ AL W + F + D + R PW
Sbjct: 38 SLIARPLNPRVQNLNSVVVALPRSWGLTTQVHGRVLDATYVQFLFANEIDLMMVQRREPW 97
Query: 145 FFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL 204
F N+W + + A +L +WVQ+RG+P +++ ++I +G +++ +
Sbjct: 98 LF-NNWFVAATRWQVAPAHNL-VTTIDLWVQIRGIPLPYVSEETVLEIAQDLGEIISLDF 155
Query: 205 YEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTE 263
+E ++ I+++V ++ ++ I I F+YE L + C +C H
Sbjct: 156 HEATSPQIAFIRVRVRFGITDRLRFFQRIIFDSGETATIRFQYERLRRLCSSCFRFTHNR 215
Query: 264 ANC 266
A C
Sbjct: 216 AYC 218
>At4g02000 putative zinc finger protein
Length = 314
Score = 55.5 bits (132), Expect = 5e-08
Identities = 33/153 (21%), Positives = 68/153 (43%), Gaps = 9/153 (5%)
Query: 134 DTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIG 193
D ++R PW + N ++ H W ++ L +WVQ+RG+P ++ ++I
Sbjct: 7 DLLSVLRREPWLYNNWFVTTHRWEVNLTFHLLT--SIELWVQMRGIPLLYVCEETALEIA 64
Query: 194 SSIGTVLASELYEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQF 252
+G +L + ++ ++ I++++ ++ ++ I A I F+YE L +
Sbjct: 65 HELGKILTLDFHDSTTTQIAYIRVRIRFGITDRLRFFQRIIFDFGEAALISFQYERLRRI 124
Query: 253 CFACGLIGHTEANC------KLHDTRTNAESRN 279
C +C + H +C LH + RN
Sbjct: 125 CSSCFRMTHHRNSCPYRQIEPLHRVTNSTAQRN 157
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,877,633
Number of Sequences: 26719
Number of extensions: 382894
Number of successful extensions: 1257
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1179
Number of HSP's gapped (non-prelim): 58
length of query: 361
length of database: 11,318,596
effective HSP length: 101
effective length of query: 260
effective length of database: 8,619,977
effective search space: 2241194020
effective search space used: 2241194020
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144807.8


