

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
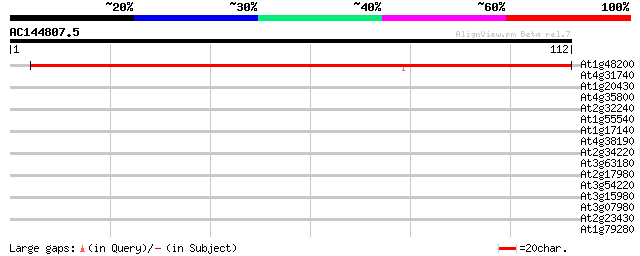
Query= AC144807.5 + phase: 0
(112 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g48200 hypothetical protein 94 1e-20
At4g31740 putative protein 31 0.088
At1g20430 Unknown protein 30 0.15
At4g35800 DNA-directed RNA polymerase (EC 2.7.7.6) II largest chain 29 0.33
At2g32240 putative myosin heavy chain 28 0.75
At1g55540 unknown protein 27 2.2
At1g17140 unknown protein 27 2.2
At4g38190 unknown protein 26 2.8
At2g34220 hypothetical protein 26 3.7
At3g63180 hypothetical protein 25 4.8
At2g17980 putative SEC1 family transport protein 25 6.3
At3g54220 SCARECROW1 25 8.2
At3g15980 putative coatomer complex subunit 25 8.2
At3g07980 putative MAP3K epsilon protein kinase 25 8.2
At2g23430 cyclin-dependent kinase inhibitor protein 25 8.2
At1g79280 hypothetical protein 25 8.2
>At1g48200 hypothetical protein
Length = 122
Score = 93.6 bits (231), Expect = 1e-20
Identities = 52/110 (47%), Positives = 69/110 (62%), Gaps = 2/110 (1%)
Query: 5 HKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTL 64
+KI A NN VIN LG F +LG RS QQK +EAL +K+SL K+NK+++ T+
Sbjct: 3 NKIAMFLSEAMNNNAVINTCLGVSFVVLGLRSDKQQKYVEALAEQKESLFKSNKAMKLTM 62
Query: 65 WDWKQQLYAEAAS--DSAPVPLARLYEIYGEAAPPQQSAPGNLFFSLLLI 112
W+WKQQL+AEAAS ++A VPL+ L IYGE + SLLL+
Sbjct: 63 WEWKQQLFAEAASAGNAAVVPLSTLKAIYGEVTTTTNQSGDYSQPSLLLV 112
>At4g31740 putative protein
Length = 171
Score = 31.2 bits (69), Expect = 0.088
Identities = 21/81 (25%), Positives = 39/81 (47%), Gaps = 15/81 (18%)
Query: 27 AVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAP----- 81
A+ +L + NQ ++ EA+EA +L + + R+ + DW ++LY ++ S P
Sbjct: 7 AIMYLLSLETINQSEV-EAVEA---ALPSADSASRSNIVDWAEKLYGQSISAVTPGVKNL 62
Query: 82 ------VPLARLYEIYGEAAP 96
+P+AR E + P
Sbjct: 63 LSSDQQLPVARTVEALTDGKP 83
>At1g20430 Unknown protein
Length = 116
Score = 30.4 bits (67), Expect = 0.15
Identities = 20/63 (31%), Positives = 33/63 (51%)
Query: 31 ILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARLYEI 90
+L RS +Q+ I LE + L K S+ + + K L +A+ DS+ V +RL +
Sbjct: 51 LLSFRSVSQKYRIHDLEEDTAVLKKEQDSLTDRMSKIKSDLLHQASIDSSGVFASRLRLL 110
Query: 91 YGE 93
+GE
Sbjct: 111 FGE 113
>At4g35800 DNA-directed RNA polymerase (EC 2.7.7.6) II largest chain
Length = 1840
Score = 29.3 bits (64), Expect = 0.33
Identities = 16/51 (31%), Positives = 30/51 (58%), Gaps = 3/51 (5%)
Query: 37 YNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARL 87
Y Q+++++A+E D + K + ++RN+L D Q LY E D+ + +L
Sbjct: 856 YIQRRLVKAME---DIMVKYDGTVRNSLGDVIQFLYGEDGMDAVWIESQKL 903
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 28.1 bits (61), Expect = 0.75
Identities = 16/54 (29%), Positives = 28/54 (51%)
Query: 21 INVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAE 74
+N+ L A G+ + Q + ALEAEK+ ++ + T+ D +QL +E
Sbjct: 1010 VNLKLNLELANHGSEANELQTKLSALEAEKEQTANELEASKTTIEDLTKQLTSE 1063
>At1g55540 unknown protein
Length = 1821
Score = 26.6 bits (57), Expect = 2.2
Identities = 20/76 (26%), Positives = 37/76 (48%), Gaps = 10/76 (13%)
Query: 37 YNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQ---QLYAEAASDSAPVPLARLYEIYGE 93
+N+QK+ LEA++ + K NK + + L + ++ +L + ++ P+AR
Sbjct: 832 WNRQKLNPELEAKRQHIMKLNKDLTHQLIELERYFNRLELDRYNEDGGHPVAR------- 884
Query: 94 AAPPQQSAPGNLFFSL 109
P +SAP SL
Sbjct: 885 RGVPNRSAPSRRVQSL 900
>At1g17140 unknown protein
Length = 344
Score = 26.6 bits (57), Expect = 2.2
Identities = 20/81 (24%), Positives = 41/81 (49%), Gaps = 7/81 (8%)
Query: 13 RASENNTVINVG-LGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQL 71
+A+E+ V V +G A++ + ++ +E++E KD+L K +R W++
Sbjct: 211 KANEDEMVSKVSRIGEELEESRAKTAHLKEKLESMEEAKDALEAEMKKLRVQTEQWRK-- 268
Query: 72 YAEAASDSAPVPLARLYEIYG 92
A+D+A L+ +E+ G
Sbjct: 269 ----AADAAAAVLSGEFEMNG 285
>At4g38190 unknown protein
Length = 1111
Score = 26.2 bits (56), Expect = 2.8
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 18/68 (26%)
Query: 8 VDIAKRASENNTV---------------INVGLGAVF---AILGARSYNQQKIIEALEAE 49
+D + R + NNTV + VG G +F A+ G N K++E E+E
Sbjct: 650 IDPSDRYANNNTVFFDGNMRALDGVQGPVYVGTGTMFRRFALYGFDPPNPDKLLEKKESE 709
Query: 50 KDSLTKTN 57
++LT ++
Sbjct: 710 TEALTTSD 717
>At2g34220 hypothetical protein
Length = 718
Score = 25.8 bits (55), Expect = 3.7
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 1/37 (2%)
Query: 47 EAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVP 83
E K S K N ++ TLWDW + Y E + VP
Sbjct: 348 EVLKSSCAKENCTLSCTLWDWLRD-YTEENLELPGVP 383
>At3g63180 hypothetical protein
Length = 975
Score = 25.4 bits (54), Expect = 4.8
Identities = 13/45 (28%), Positives = 21/45 (45%)
Query: 54 TKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARLYEIYGEAAPPQ 98
T + ++R DW Q A++A S+ +PL + PPQ
Sbjct: 354 TDRDSNLRLQNLDWSQPQQAKSAPHSSVLPLPVAVASWPSGVPPQ 398
>At2g17980 putative SEC1 family transport protein
Length = 627
Score = 25.0 bits (53), Expect = 6.3
Identities = 12/42 (28%), Positives = 24/42 (56%), Gaps = 1/42 (2%)
Query: 36 SYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAAS 77
++ K I++L A + T N + R+ + DW ++LY ++ S
Sbjct: 457 AFQYVKKIKSLNASF-AATSANSASRSNIVDWAEKLYGQSIS 497
>At3g54220 SCARECROW1
Length = 653
Score = 24.6 bits (52), Expect = 8.2
Identities = 10/25 (40%), Positives = 15/25 (60%)
Query: 75 AASDSAPVPLARLYEIYGEAAPPQQ 99
++SD +P LY+I +PPQQ
Sbjct: 195 SSSDPSPQTFEPLYQISNNPSPPQQ 219
>At3g15980 putative coatomer complex subunit
Length = 921
Score = 24.6 bits (52), Expect = 8.2
Identities = 17/56 (30%), Positives = 24/56 (42%), Gaps = 10/56 (17%)
Query: 30 AILGARSYNQQKIIEALEAEKDSLTKTN----------KSIRNTLWDWKQQLYAEA 75
A L ARSY K+ E + ++ L+K N + N DW+ L EA
Sbjct: 758 AALMARSYLPSKVSEIVALWREDLSKVNPKAAESLADPEEYSNLFEDWQVALSVEA 813
>At3g07980 putative MAP3K epsilon protein kinase
Length = 1367
Score = 24.6 bits (52), Expect = 8.2
Identities = 9/18 (50%), Positives = 15/18 (83%)
Query: 40 QKIIEALEAEKDSLTKTN 57
Q+++E++ AEK +TKTN
Sbjct: 309 QEVVESVSAEKVEVTKTN 326
>At2g23430 cyclin-dependent kinase inhibitor protein
Length = 191
Score = 24.6 bits (52), Expect = 8.2
Identities = 15/52 (28%), Positives = 29/52 (54%), Gaps = 1/52 (1%)
Query: 13 RASENNTVINVG-LGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNT 63
R+ ++++V VG G + G+ Y ++++I E +KD T+T+ R T
Sbjct: 32 RSEKSSSVSVVGDNGVSSSCSGSNEYKKKELIHLEEEDKDGDTETSTYRRGT 83
>At1g79280 hypothetical protein
Length = 2111
Score = 24.6 bits (52), Expect = 8.2
Identities = 11/33 (33%), Positives = 19/33 (57%)
Query: 47 EAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDS 79
E EKD L+K N+S+ L + K++ +D+
Sbjct: 1472 EKEKDELSKQNQSLAKQLEEAKEEAGKRTTTDA 1504
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.131 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,217,339
Number of Sequences: 26719
Number of extensions: 72969
Number of successful extensions: 248
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 236
Number of HSP's gapped (non-prelim): 16
length of query: 112
length of database: 11,318,596
effective HSP length: 88
effective length of query: 24
effective length of database: 8,967,324
effective search space: 215215776
effective search space used: 215215776
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144807.5


