

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144766.6 - phase: 0
(381 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
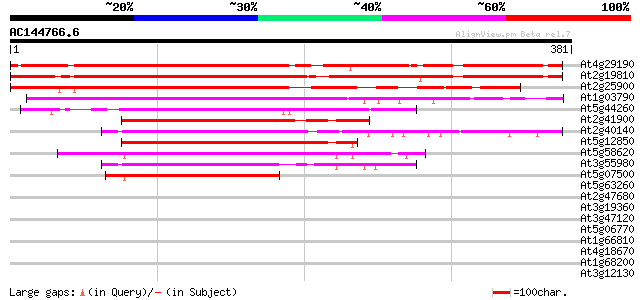
Score E
Sequences producing significant alignments: (bits) Value
At4g29190 unknown protein 393 e-110
At2g19810 putative CCCH-type zinc finger protein 387 e-108
At2g25900 Cys3His zinc finger like protein ATCTH (ATCTH) 312 2e-85
At1g03790 unknown protein 216 2e-56
At5g44260 zinc finger like protein 211 4e-55
At2g41900 CCCH-type zinc finger like protein 192 3e-49
At2g40140 putative CCCH-type zinc finger protein 192 3e-49
At5g12850 zinc finger transcription factor -like protein 191 6e-49
At5g58620 unknown protein 190 1e-48
At3g55980 unknown protein 182 2e-46
At5g07500 zinc finger transcription factor 175 3e-44
At5g63260 unknown protein 38 0.011
At2g47680 putative ATP-dependent RNA helicase A 38 0.011
At3g19360 unknown protein 37 0.014
At3g47120 putative RNA-binding protein 37 0.018
At5g06770 unknown protein 35 0.068
At1g66810 hypothetical protein 35 0.068
At4g18670 extensin-like protein 35 0.089
At1g68200 putative zinc finger protein 35 0.089
At3g12130 unknown protein 34 0.12
>At4g29190 unknown protein
Length = 356
Score = 393 bits (1010), Expect = e-110
Identities = 210/382 (54%), Positives = 259/382 (66%), Gaps = 34/382 (8%)
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNN-PTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENN 59
MM+GE RT PTV +PPWP L T+E FSP+ ++ D S M EAL+ Q Y+ N
Sbjct: 1 MMIGET-RRTYPTVEIPPWPVLEELTTSEFFSPVMNSPDCS---MLEALAGLQRYLPSNE 56
Query: 60 -DSDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPR 118
D +S ++ +D+YS DHFRM++FK+RRCARGRSHDWTECP++HPGEKARRRDPR
Sbjct: 57 PDPESYPDLLGPDSPIDAYSCDHFRMYDFKVRRCARGRSHDWTECPYAHPGEKARRRDPR 116
Query: 119 KYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHT 178
KY+YSGT+CPDFRKG CKKGDSCEFAHGVFECWLHP+RYRTQPCKDG +C R +CFFAH+
Sbjct: 117 KYHYSGTACPDFRKGGCKKGDSCEFAHGVFECWLHPARYRTQPCKDGGNCLRKICFFAHS 176
Query: 179 TEQLRAPTQQSPRSVPSVDSYD-GSPLRLAFESSCVKTLQFMSSPGSVSPPVE---SPPM 234
+QLR +SP VDS+D SP+R + Q SP S SPP+
Sbjct: 177 PDQLRFLHTRSP---DRVDSFDVSSPIR-------ARAFQLSISPVSGSPPMSPRADSES 226
Query: 235 SPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFC 294
SPMT+SL RS+GS S+N++V S RNLQ ++KS P + N G FGSPRG ++ PGF
Sbjct: 227 SPMTQSLSRSLGSCSINDVVPSFRNLQFNSVKSFPRN-NPLFG---FGSPRGSILGPGFQ 282
Query: 295 SLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMER-VESGRDIRVKMFEKLSKENSFNGSGM 353
SLP+TPT R + D+W+ EEEPVMER VESGR++R KM EKL KEN +
Sbjct: 283 SLPTTPT------RPGNLDIWEYGLEEEPVMERVVESGRELREKMREKLHKENCMDRVAQ 336
Query: 354 GSGSGLGEVVEDPDVGWVSELV 375
LGE PDVGWVS+L+
Sbjct: 337 DPDQNLGEA---PDVGWVSDLL 355
>At2g19810 putative CCCH-type zinc finger protein
Length = 359
Score = 387 bits (993), Expect = e-108
Identities = 208/381 (54%), Positives = 258/381 (67%), Gaps = 29/381 (7%)
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNNPT-AEIFSPLTSNDDYSQFYMQEALSAFQHYVNENN 59
MM+GE NPTVH+PPWP + T ++I+ S D S M EAL+ Q Y+ N
Sbjct: 1 MMIGESHRGFNPTVHIPPWPLSEDLTVSDIYG---SPDGGSS--MMEALAELQRYLPSNE 55
Query: 60 -DSDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPR 118
D DSD ++ +D+Y+ DHFRM+EFK+RRCARGRSHDWTECP++HPGEKARRRDPR
Sbjct: 56 PDPDSDPDLSGPDSPIDAYTCDHFRMYEFKVRRCARGRSHDWTECPYAHPGEKARRRDPR 115
Query: 119 KYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHT 178
K++YSGT+CP+FRKG CK+GD+CEF+HGVFECWLHP+RYRTQPCKDG +CRR VCFFAH+
Sbjct: 116 KFHYSGTACPEFRKGCCKRGDACEFSHGVFECWLHPARYRTQPCKDGGNCRRRVCFFAHS 175
Query: 179 TEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMT 238
+Q+R QSP V S D + +R AF QF SP S SPPV S +
Sbjct: 176 PDQIRVLPNQSPDRVDSFDVLSPT-IRRAF--------QFSISPSSNSPPVSPRGDSDSS 226
Query: 239 RS-LGRSVGSSSVNEMVASLRNLQLGTMK-SLPSSWNVQMG--SPRFGSPRGPVIRPGFC 294
S L RS+GS+ N++VASLRNLQL +K SL SS+N Q+G FGSPRG V+ PGF
Sbjct: 227 CSLLSRSLGSNLGNDVVASLRNLQLNKVKSSLSSSYNNQIGGYGSGFGSPRGSVLGPGFR 286
Query: 295 SLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMG 354
SLP+TPT R ++W+ EEEP MERVESGR++R ++FEKLSKEN
Sbjct: 287 SLPTTPT------RPGFMNIWENGLEEEPAMERVESGRELRAQLFEKLSKENCMGRIEPD 340
Query: 355 SGSGLGEVVEDPDVGWVSELV 375
G G+ PDVGWVS+LV
Sbjct: 341 PDQGAGDT---PDVGWVSDLV 358
>At2g25900 Cys3His zinc finger like protein ATCTH (ATCTH)
Length = 315
Score = 312 bits (799), Expect = 2e-85
Identities = 170/357 (47%), Positives = 217/357 (60%), Gaps = 54/357 (15%)
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSP-----LTSNDDYSQF---YMQEALSAFQ 52
MM+GE +R +PT+H+P W +N+PTA I SP L S +DY Y+ S F+
Sbjct: 1 MMIGENKNRPHPTIHIPQWDQINDPTATISSPFSSVNLNSVNDYPHSPSPYLDSFASLFR 60
Query: 53 HYVNENNDSDSDSEIFPTHESV-DSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEK 111
+ + +DSDS + DS+S+D FR++EFKIRRCARGRSHDWTECPF+HPGEK
Sbjct: 61 YLPSNELTNDSDSSSGDESSPLTDSFSSDEFRIYEFKIRRCARGRSHDWTECPFAHPGEK 120
Query: 112 ARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRP 171
ARRRDPRK++YSGT+CP+FRKGSC++GDSCEF+HGVFECWLHPSRYRTQPCKDGTSCRR
Sbjct: 121 ARRRDPRKFHYSGTACPEFRKGSCRRGDSCEFSHGVFECWLHPSRYRTQPCKDGTSCRRR 180
Query: 172 VCFFAHTTEQLRA-PTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVE 230
+CFFAHTTEQLR P P L F S + + S SPP E
Sbjct: 181 ICFFAHTTEQLRVLPCSLDP--------------DLGFFSGLATSPTSILVSPSFSPPSE 226
Query: 231 SPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIR 290
SPP+SP S E++AS+R +QL G + SP +R
Sbjct: 227 SPPLSP------------STGELIASMRKMQLNG------------GGCSWSSPMRSAVR 262
Query: 291 PGFCSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENS 347
F S P Q + R+ F++ EE P ME VESG+++R +M+ +LS+ENS
Sbjct: 263 LPFSS-SLRPIQAATWPRIREFEI-----EEAPAMEFVESGKELRAEMYARLSRENS 313
>At1g03790 unknown protein
Length = 393
Score = 216 bits (549), Expect = 2e-56
Identities = 143/402 (35%), Positives = 196/402 (48%), Gaps = 58/402 (14%)
Query: 12 PTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSEIFPTH 71
PTV +PP L + + + + + +H + +++ + E
Sbjct: 13 PTVEIPPRKLLLSSKSFPSDSSSPRSPRKHNWNKSNKITSEHEEDNEDNNRENKEYCYDS 72
Query: 72 ESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFR 131
+S D Y++DHFRMFEFKIRRC R RSHDWT+CPF+HPGEKARRRDPR++ YSG CP+FR
Sbjct: 73 DSDDPYASDHFRMFEFKIRRCTRSRSHDWTDCPFAHPGEKARRRDPRRFQYSGEVCPEFR 132
Query: 132 K-GSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSP 190
+ G C +GD CEFAHGVFECWLHP RYRT+ CKDG C+R VCFFAH+ QLR ++
Sbjct: 133 RGGDCSRGDDCEFAHGVFECWLHPIRYRTEACKDGKHCKRKVCFFAHSPRQLRVLPPENV 192
Query: 191 RSVPSVDSYDG-SPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTR---------- 239
V + S +P L SS TL S S SP + SPPMSP +
Sbjct: 193 SGVSASPSPAAKNPCCLFCSSSPTSTLLGNLSHLSRSPSL-SPPMSPANKAAAFSRLRNR 251
Query: 240 -----SLGRSVGSSS----VNEMVASLRNLQLG----TMKSLPSSWNVQMGSPRFGSPRG 286
S + GS + ++E+V SL ++ L S P + V + F S G
Sbjct: 252 AASAVSAAAAAGSMNYKDVLSELVNSLDSMSLAEALQASSSSPVTTPVSAAAAAFASSCG 311
Query: 287 ------------PVIRPGFCSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDI 334
P F PSTP+ + + + N F D M + +
Sbjct: 312 LSNQRLHLQQQQPSSPLQFALSPSTPSYLTNSPQANFFS--DDFTPRRRQMNDFTAMTAV 369
Query: 335 RVKMFEKLSKENSFNGSGMGSGSGLGEVVEDPDVGWVSELVS 376
R E + E+ G DPD+GWV++L++
Sbjct: 370 R----ENTNIEDGSCG--------------DPDLGWVNDLLT 393
>At5g44260 zinc finger like protein
Length = 381
Score = 211 bits (538), Expect = 4e-55
Identities = 126/284 (44%), Positives = 161/284 (56%), Gaps = 41/284 (14%)
Query: 8 HRTNPTVHVPPWPTLNNPTA--EIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDS 65
H + PTV +PP L++ + + SPL DY + ++ DSDS
Sbjct: 10 HISRPTVDIPPRKLLSSAKSPSSVSSPLR---DYKE--------------QKDYCYDSDS 52
Query: 66 EIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGT 125
E D Y+ DHFRM+EFKIRRC R RSHDWT+CPFSHPGEKARRRDPR+++Y+G
Sbjct: 53 E--------DPYAGDHFRMYEFKIRRCTRSRSHDWTDCPFSHPGEKARRRDPRRFHYTGE 104
Query: 126 SCPDF-RKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRA 184
CP+F R G C +GD C FAHGVFECWLHPSRYRT+ CKDG C+R VCFFAH+ QLR
Sbjct: 105 VCPEFSRHGDCSRGDECGFAHGVFECWLHPSRYRTEACKDGKHCKRKVCFFAHSPRQLRV 164
Query: 185 --PTQQ---------SPRSVP-SVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESP 232
P+ + SP S P SV S + L S TL +S S SPP+
Sbjct: 165 LPPSPENHISGGCGGSPSSSPASVLSNKNNRCCLFCSHSPTSTLLNLSRSPSSSPPLSPA 224
Query: 233 PMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQM 276
+ L R ++ +NE+++SL +L L + SS V M
Sbjct: 225 DKADAFSRLSRR-RTAVLNELISSLDSLSLTEALAASSSSPVTM 267
>At2g41900 CCCH-type zinc finger like protein
Length = 716
Score = 192 bits (487), Expect = 3e-49
Identities = 91/168 (54%), Positives = 111/168 (65%), Gaps = 12/168 (7%)
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
Y+ D FRM+ FK+R C+R SHDWTECPF HPGE ARRRDPRK++YS CPDFRKG+C+
Sbjct: 257 YATDEFRMYSFKVRPCSRAYSHDWTECPFVHPGENARRRDPRKFHYSCVPCPDFRKGACR 316
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
+GD CE+AHGVFECWLHP++YRT+ CKDGT C R VCFFAHT E+LR + +VP
Sbjct: 317 RGDMCEYAHGVFECWLHPAQYRTRLCKDGTGCARRVCFFAHTPEELRPLYASTGSAVP-- 374
Query: 197 DSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRS 244
SP A ++ + L PGS S P+SP G S
Sbjct: 375 -----SPRSNADYAAALSLL-----PGSPSGVSVMSPLSPSAAGNGMS 412
>At2g40140 putative CCCH-type zinc finger protein
Length = 597
Score = 192 bits (487), Expect = 3e-49
Identities = 136/365 (37%), Positives = 178/365 (48%), Gaps = 61/365 (16%)
Query: 63 SDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNY 122
+D+ + +E V Y D FRMF FK++ C+R SHDWTECPF HPGE ARRRDPRKY Y
Sbjct: 198 ADASLPDINEGV--YGTDDFRMFSFKVKPCSRAYSHDWTECPFVHPGENARRRDPRKYPY 255
Query: 123 SGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQL 182
+ CP+FRKGSC KGDSCE+AHGVFE WLHP++YRT+ CKD T C R VCFFAH ++L
Sbjct: 256 TCVPCPEFRKGSCPKGDSCEYAHGVFESWLHPAQYRTRLCKDETGCARRVCFFAHRRDEL 315
Query: 183 RAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLG 242
R + ++ S S + SP E S + L SSP + SP P+SP L
Sbjct: 316 RPVNASTGSAMVSPRSSNQSP-----EMSVMSPLTLGSSPMN-SPMANGVPLSPRNGGLW 369
Query: 243 -------------------RSVGSSSVNEMVASLR-----NLQLGTMK---------SLP 269
+S S+ +M LR N +LG +K S
Sbjct: 370 QNRVNSLTPPPLQLNGSRLKSTLSARDMDMEMELRFRGLDNRRLGDLKPSNLEETFGSYD 429
Query: 270 SSWNVQMGSPRFGS-----PRGPVIRP---GFCSLPSTPTQVPSRGRVNHFDLWDQSCEE 321
S+ +Q+ SP S P PV +P GF S + V + R + F S +
Sbjct: 430 SASVMQLQSPSRHSQMNHYPSSPVRQPPPHGFESSAAMAAAVMN-ARSSAFAKRSLSFKP 488
Query: 322 EPVMERVESGRDIRVKM--------FEKLSKENSFNGSGMGSGS---GLGEVVEDPDVGW 370
PV V K+ KL + SF G + S + ++PDV W
Sbjct: 489 APVASNVSDWGSPNGKLEWGMQRDELNKLRRSASFGIHGNNNNSVSRPARDYSDEPDVSW 548
Query: 371 VSELV 375
V+ LV
Sbjct: 549 VNSLV 553
>At5g12850 zinc finger transcription factor -like protein
Length = 706
Score = 191 bits (485), Expect = 6e-49
Identities = 90/162 (55%), Positives = 108/162 (66%), Gaps = 7/162 (4%)
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
YS D FRMF FKIR C+R SHDWTECPF+HPGE ARRRDPRK++Y+ CPDF+KGSCK
Sbjct: 253 YSTDEFRMFSFKIRPCSRAYSHDWTECPFAHPGENARRRDPRKFHYTCVPCPDFKKGSCK 312
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
+GD CE+AHGVFECWLHP++YRT+ CKDG C R VCFFAH E+LR + +PS
Sbjct: 313 QGDMCEYAHGVFECWLHPAQYRTRLCKDGMGCNRRVCFFAHANEELRPLYPSTGSGLPSP 372
Query: 197 DSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVES--PPMSP 236
+ +S + L PGS S S PP+SP
Sbjct: 373 RASSAVSASTMDMASVLNML-----PGSPSAAQHSFTPPISP 409
>At5g58620 unknown protein
Length = 607
Score = 190 bits (483), Expect = 1e-48
Identities = 111/273 (40%), Positives = 147/273 (53%), Gaps = 27/273 (9%)
Query: 33 LTSNDDYSQFYMQEALSA-FQHYVNENNDSDSDSEIFPTHESVDS-----YSNDHFRMFE 86
L NDD ++ QE + V + S+ + +P ++ Y D FRM+
Sbjct: 155 LKGNDDLNEVNGQEESEPEVEVEVEVSPPRGSERKEYPVDPTLPDIKNGVYGTDEFRMYA 214
Query: 87 FKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHG 146
FKI+ C+R SHDWTECPF HPGE ARRRDPRKY+YS CP+FRKGSC +GD+CE+AHG
Sbjct: 215 FKIKPCSRAYSHDWTECPFVHPGENARRRDPRKYHYSCVPCPEFRKGSCSRGDTCEYAHG 274
Query: 147 VFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV-DSYDGSPLR 205
+FECWLHP++YRT+ CKD T+C R VCFFAH E+LR + VPS S+
Sbjct: 275 IFECWLHPAQYRTRLCKDETNCSRRVCFFAHKPEELRPLYPSTGSGVPSPRSSFSSCNSS 334
Query: 206 LAFESSCVKTLQFMS------SPGSVSPPVES-------PPMSPMTRSLGRSVGSSSVNE 252
AF+ + L + SP VS P+ P ++P L S S++N
Sbjct: 335 TAFDMGPISPLPIGATTTPPLSPNGVSSPIGGGKTWMNWPNITPPALQLPGSRLKSALNA 394
Query: 253 MVASLRNLQLGTMKSL--PSSW-NVQMGSPRFG 282
M+SL P++W N M SP G
Sbjct: 395 REIDFSE----EMQSLTSPTTWNNTPMSSPFSG 423
>At3g55980 unknown protein
Length = 580
Score = 182 bits (463), Expect = 2e-46
Identities = 102/222 (45%), Positives = 135/222 (59%), Gaps = 25/222 (11%)
Query: 63 SDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNY 122
+D+ + +E V Y +D FRM+ FK++ C+R SHDWTEC F HPGE ARRRDPRKY Y
Sbjct: 195 ADASLPDINEGV--YGSDEFRMYSFKVKPCSRAYSHDWTECAFVHPGENARRRDPRKYPY 252
Query: 123 SGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQL 182
+ CP+FRKGSC KGDSCE+AHGVFE WLHP++Y+T+ CKD T C R VCFFAH E++
Sbjct: 253 TCVPCPEFRKGSCPKGDSCEYAHGVFESWLHPAQYKTRLCKDETGCARKVCFFAHKREEM 312
Query: 183 RAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMS--SPGSVSPPVESPPMSPMTR- 239
R V++ GS A S +L+ M SP + S V +PP+SPM
Sbjct: 313 R-----------PVNASTGS----AVAQSPFSSLEMMPGLSPLAYSSGVSTPPVSPMANG 357
Query: 240 --SLGRSVGS--SSVNEMVASLRNLQLGT-MKSLPSSWNVQM 276
S R+ GS + VN + L G+ +KS S+ ++ M
Sbjct: 358 VPSSPRNGGSWQNRVNTLTPPALQLNGGSRLKSTLSARDIDM 399
>At5g07500 zinc finger transcription factor
Length = 245
Score = 175 bits (444), Expect = 3e-44
Identities = 75/120 (62%), Positives = 91/120 (75%), Gaps = 2/120 (1%)
Query: 66 EIFPTHESVDS--YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYS 123
EI P+ ++D Y +D FRM+ +KI+RC R RSHDWTECP++H GEKA RRDPR+Y Y
Sbjct: 35 EIDPSIPNIDDAIYGSDEFRMYAYKIKRCPRTRSHDWTECPYAHRGEKATRRDPRRYTYC 94
Query: 124 GTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLR 183
+CP FR G+C +GDSCEFAHGVFE WLHP+RYRT+ C G C+R VCFFAH EQLR
Sbjct: 95 AVACPAFRNGACHRGDSCEFAHGVFEYWLHPARYRTRACNAGNLCQRKVCFFAHAPEQLR 154
>At5g63260 unknown protein
Length = 435
Score = 37.7 bits (86), Expect = 0.011
Identities = 61/247 (24%), Positives = 81/247 (32%), Gaps = 80/247 (32%)
Query: 131 RKGSCKKGDSCEFAHGV-----------------------FECWLHPSRYRTQPCKDGTS 167
R GSCK G SC+F H V EC + +RT CK G S
Sbjct: 112 RTGSCKYGSSCKFNHPVRRKLQIGRERVRERDEDVENPKLMECKYY---FRTGGCKYGES 168
Query: 168 CRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLR--------------LAFESSCV 213
CR F+H E SP SVP + ++ G P+R F S C
Sbjct: 169 CR-----FSHMKE------HNSPASVPEL-NFLGLPIRPGEKECPFYMRNGSCKFGSDCK 216
Query: 214 KTLQFMSSPGSVSPPV----ESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLP 269
++ G V P+ SP S S SS M + GT +P
Sbjct: 217 FNHPDPTAIGGVDSPLYRGNNGGSFSPKAPSQASSTSWSSTRHMNGT------GTAPFIP 270
Query: 270 -------------SSWNVQMGSPRFGSPRGPVIRPGFCSLPSTPTQVPSRGRVNHFDLWD 316
S WN S + R P + P + ++ + S + H
Sbjct: 271 SMFPHSRGVTPQASDWNGYQASSAYPPERSP-LAPSSYQVNNSLAETSSFSQYQH----Q 325
Query: 317 QSCEEEP 323
S EE P
Sbjct: 326 MSVEEFP 332
Score = 36.2 bits (82), Expect = 0.031
Identities = 29/100 (29%), Positives = 36/100 (36%), Gaps = 34/100 (34%)
Query: 101 TECPFSHP--------GEKARRRDPRKYNYSGTSCPD-FRKGSCKKGDSCEFAH------ 145
+ C F+HP E+ R RD N C FR G CK G+SC F+H
Sbjct: 120 SSCKFNHPVRRKLQIGRERVRERDEDVENPKLMECKYYFRTGGCKYGESCRFSHMKEHNS 179
Query: 146 ----------------GVFECWLHPSRYRTQPCKDGTSCR 169
G EC P R CK G+ C+
Sbjct: 180 PASVPELNFLGLPIRPGEKEC---PFYMRNGSCKFGSDCK 216
>At2g47680 putative ATP-dependent RNA helicase A
Length = 1015
Score = 37.7 bits (86), Expect = 0.011
Identities = 31/117 (26%), Positives = 44/117 (37%), Gaps = 14/117 (11%)
Query: 101 TECPFSHPGEKARRRDPRKYNYSGTS--CPDFRKGSCKKGDSCEFAHGVFE----CWLHP 154
T+C S +P Y G + C F G C +G C F H + C
Sbjct: 705 TQCTPSESDNGHGALEPEDYVEYGEAPVCVYFLNGYCNRGGQCTFTHTLQSTRPACKFFA 764
Query: 155 SRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESS 211
S +Q C++G S C F+H + P+ +P D SPL F +S
Sbjct: 765 S---SQGCRNGES-----CLFSHAMRRRTTSYLPPPQCLPEEDGSSTSPLLDLFPTS 813
>At3g19360 unknown protein
Length = 386
Score = 37.4 bits (85), Expect = 0.014
Identities = 14/37 (37%), Positives = 21/37 (55%)
Query: 122 YSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYR 158
Y C FR G+C+ G+ C FAHG+ + PS ++
Sbjct: 105 YKTRMCAKFRAGTCRNGELCNFAHGIEDLRQPPSNWQ 141
>At3g47120 putative RNA-binding protein
Length = 352
Score = 37.0 bits (84), Expect = 0.018
Identities = 24/90 (26%), Positives = 33/90 (36%), Gaps = 12/90 (13%)
Query: 68 FPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRR------------ 115
+ HE D + R R RG C FSH ++A
Sbjct: 115 YKKHEEEDEETRRQNREARGVCRAFQRGECTRGDSCKFSHDEKRAANTGWGHEEDRSSKW 174
Query: 116 DPRKYNYSGTSCPDFRKGSCKKGDSCEFAH 145
D K C F++G C +GDSC+F+H
Sbjct: 175 DHDKNREGRGVCRAFQRGECTRGDSCKFSH 204
>At5g06770 unknown protein
Length = 240
Score = 35.0 bits (79), Expect = 0.068
Identities = 14/29 (48%), Positives = 16/29 (54%)
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFE 149
NY C + KG+C GD C FAHG E
Sbjct: 205 NYKTKICDRYSKGNCTYGDRCHFAHGESE 233
>At1g66810 hypothetical protein
Length = 310
Score = 35.0 bits (79), Expect = 0.068
Identities = 24/59 (40%), Positives = 33/59 (55%), Gaps = 9/59 (15%)
Query: 133 GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCK---DGTSC-RRPVCFFAHT-TEQLR 183
G+C GD+C+FAHG+ E HP RY+T+ C+ G C C F H+ T+Q R
Sbjct: 245 GACCYGDNCQFAHGIDELRPVIRHP-RYKTEVCRMMVTGAMCPYGHRCHFRHSLTDQER 302
>At4g18670 extensin-like protein
Length = 839
Score = 34.7 bits (78), Expect = 0.089
Identities = 39/141 (27%), Positives = 48/141 (33%), Gaps = 6/141 (4%)
Query: 165 GTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGS 224
G S R PV + T + SP PS SP + + SP S
Sbjct: 401 GRSTRPPVVVPSPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSPPS 460
Query: 225 VSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSP 284
+P SPP SP T + G S SS + + P+ SP SP
Sbjct: 461 TTPSPGSPPTSPTTPTPGGSPPSSPTTPTPGG----SPPSSPTTPTPGGSPPSSPTTPSP 516
Query: 285 RGPVIRPGFCSLPSTPTQVPS 305
G P PS P VPS
Sbjct: 517 GGSPPSPSIS--PSPPITVPS 535
>At1g68200 putative zinc finger protein
Length = 308
Score = 34.7 bits (78), Expect = 0.089
Identities = 21/56 (37%), Positives = 30/56 (53%), Gaps = 8/56 (14%)
Query: 133 GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCK---DGTSC-RRPVCFFAHTTEQ 181
G+C GD C+FAHG+ E HP RY+T+ C+ G +C C F H+ +
Sbjct: 235 GTCPYGDHCQFAHGIKELRPVIRHP-RYKTEVCRMVLAGDNCPYGHRCHFRHSLSE 289
Score = 28.5 bits (62), Expect = 6.4
Identities = 13/26 (50%), Positives = 15/26 (57%)
Query: 282 GSPRGPVIRPGFCSLPSTPTQVPSRG 307
G PRG V +PG C ST +V RG
Sbjct: 177 GKPRGTVTKPGTCGQVSTTQKVYVRG 202
>At3g12130 unknown protein
Length = 248
Score = 34.3 bits (77), Expect = 0.12
Identities = 14/29 (48%), Positives = 16/29 (54%)
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFE 149
N+ C F KG+C GD C FAHG E
Sbjct: 213 NFKTKICERFSKGNCTFGDRCHFAHGEAE 241
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,999,507
Number of Sequences: 26719
Number of extensions: 474171
Number of successful extensions: 1906
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 1760
Number of HSP's gapped (non-prelim): 133
length of query: 381
length of database: 11,318,596
effective HSP length: 101
effective length of query: 280
effective length of database: 8,619,977
effective search space: 2413593560
effective search space used: 2413593560
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144766.6


