

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
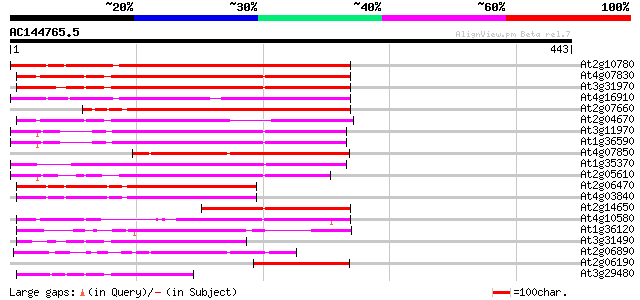
Query= AC144765.5 - phase: 0 /pseudo
(443 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 229 2e-60
At4g07830 putative reverse transcriptase 223 2e-58
At3g31970 hypothetical protein 223 2e-58
At4g16910 retrotransposon like protein 208 4e-54
At2g07660 putative retroelement pol polyprotein 201 7e-52
At2g04670 putative retroelement pol polyprotein 179 3e-45
At3g11970 hypothetical protein 159 3e-39
At1g36590 hypothetical protein 154 7e-38
At4g07850 putative polyprotein 153 2e-37
At1g35370 hypothetical protein 151 6e-37
At2g05610 putative retroelement pol polyprotein 147 1e-35
At2g06470 putative retroelement pol polyprotein 146 3e-35
At4g03840 putative transposon protein 141 8e-34
At2g14650 putative retroelement pol polyprotein 141 8e-34
At4g10580 putative reverse-transcriptase -like protein 134 8e-32
At1g36120 putative reverse transcriptase gb|AAD22339.1 131 9e-31
At3g31490 hypothetical protein 99 4e-21
At2g06890 putative retroelement integrase 88 8e-18
At2g06190 putative Ty3-gypsy-like retroelement pol polyprotein 85 9e-17
At3g29480 hypothetical protein 84 2e-16
>At2g10780 pseudogene
Length = 1611
Score = 229 bits (585), Expect = 2e-60
Identities = 125/271 (46%), Positives = 171/271 (62%), Gaps = 8/271 (2%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKI 60
N + YHP KA VADALSR+ + A + +L+ L V LS + LG+
Sbjct: 1050 NLDIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH-VNALSKEVEPLGLGAA 1107
Query: 61 N-SDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLI 119
+ +D L IR AQ+ D + Q +++++ G + GR+C+P++ LK+ I
Sbjct: 1108 DQADLLSRIRLAQERDEEIKGWA----QNNKTEYQTSNNGTIVVNGRVCVPNDRALKEEI 1163
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
L E H+S SIH G+ KMY+DLK+ + W G+KKDVAR+V C TCQ K EHQ P+GLL
Sbjct: 1164 LREAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQ 1223
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRG-HDSIWVVVDRLTKSAHFIPINISYPVAQLAEIY 238
L +PEWKWD I+MDFV+ LP + H+++WVVVDRLTKSAHF+ I+ +AE Y
Sbjct: 1224 NLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKY 1283
Query: 239 IQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
I IV+LHG+P SIVSDRD RFTS+FW++ Q
Sbjct: 1284 IDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQ 1314
>At4g07830 putative reverse transcriptase
Length = 611
Score = 223 bits (568), Expect = 2e-58
Identities = 121/264 (45%), Positives = 166/264 (62%), Gaps = 8/264 (3%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP KA VV DALSRK + A + E L +C ++ + + L + +D L
Sbjct: 67 YHPGKANVVTDALSRK--RVGAAPGQSVETLVIEIGALRLCAVAREPLGLEAVD-QTDLL 123
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
++ AQ+ D + ++ +++AE +++ G + GR+C+P ++EL++ IL E H
Sbjct: 124 SRVQLAQEKD----EGLIAASKAEGFEYQFAANGTILVHGRVCVPKDKELRQEILSEAHA 179
Query: 126 SNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPE 185
S SIH GATKMY+DLK+ + W G+K+DV +V C CQ K+EHQ LL L +PE
Sbjct: 180 SMFSIHPGATKMYRDLKRYYQWVGMKRDVGNWVEECDVCQLVKIEHQVSGSLLQSLPIPE 239
Query: 186 WKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKL 245
WKWD I+MDFV LP SR D+IWV+VDRLTKSAHF+ I + A LA+ Y+ IVKL
Sbjct: 240 WKWDFITMDFVVGLP-VSRTKDAIWVIVDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKL 298
Query: 246 HGVPSSIVSDRDPRFTSRFWRSLQ 269
HGVP SIVSDRD +FTS FWR+ Q
Sbjct: 299 HGVPVSIVSDRDSKFTSAFWRAFQ 322
>At3g31970 hypothetical protein
Length = 1329
Score = 223 bits (567), Expect = 2e-58
Identities = 123/265 (46%), Positives = 167/265 (62%), Gaps = 14/265 (5%)
Query: 6 YHPDKAKVVADALSRKTLHMSA-LMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDF 64
YH KA VVADALSRK + S +V E L +C ++ + + L + +D
Sbjct: 804 YHSGKANVVADALSRKRVGGSVEALVSEIGALR-------LCVMAQEPLGLEAVD-RADL 855
Query: 65 LGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGH 124
L +R AQ+ D + ++ +++A+ S+++ G + GR+C+P +EEL++ IL E H
Sbjct: 856 LTRVRLAQEKD----EGLIAASKADGSEYQFAANGTILVHGRVCVPKDEELRREILSEAH 911
Query: 125 KSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVP 184
S SIH ATKMY+DLK+ + W G+K+DVA +V C CQ K EHQ P GLL L +
Sbjct: 912 ASMFSIHPRATKMYRDLKRYYQWVGMKRDVANWVTECDVCQLVKAEHQVPGGLLQSLPIL 971
Query: 185 EWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVK 244
EWKWD I+MDFV LP SR D+IWV+VDRLTKSAHF+ I + LA+ Y+ IV+
Sbjct: 972 EWKWDFITMDFVVGLP-VSRTKDAIWVIVDRLTKSAHFLAIRKTDGAVLLAKKYVSEIVE 1030
Query: 245 LHGVPSSIVSDRDPRFTSRFWRSLQ 269
LHGVP SIVSDRD +FTS FWR+ Q
Sbjct: 1031 LHGVPVSIVSDRDSKFTSAFWRAFQ 1055
>At4g16910 retrotransposon like protein
Length = 687
Score = 208 bits (530), Expect = 4e-54
Identities = 119/271 (43%), Positives = 164/271 (59%), Gaps = 16/271 (5%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKI 60
+ + YHP KA V DALSR + A + L+ L L LS + LG+
Sbjct: 142 DLNIAYHPGKANQVVDALSRLRTVVEAER-NQVNLVNMMGTLHLNA-LSKEVEPLGLRAA 199
Query: 61 N-SDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLI 119
N +D L IR AQ+ D + Q +++++ G + GR+C P+++ LK+ I
Sbjct: 200 NQADLLSRIRSAQERDEEIKGWA----QNNKTEYQSSNNGTIVVNGRVCGPNDKALKEEI 255
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
L+E H+S SIH G+ KMY+DLK+ + W G+KKDVAR+V +K EHQ P+G+L
Sbjct: 256 LKEAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWV--------AKEEHQVPSGMLQ 307
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRG-HDSIWVVVDRLTKSAHFIPINISYPVAQLAEIY 238
L +PEWKWD I MDFV+ LP + H+++WVVVDRLTKSAHF+ I+ +AE Y
Sbjct: 308 NLPIPEWKWDHIMMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDAAEIIAEKY 367
Query: 239 IQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
I IV+LHG+P SIVSDRD RFTS+FW+ Q
Sbjct: 368 IDEIVRLHGIPVSIVSDRDTRFTSKFWKPFQ 398
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 201 bits (511), Expect = 7e-52
Identities = 104/213 (48%), Positives = 143/213 (66%), Gaps = 7/213 (3%)
Query: 58 LKINSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKK 117
LKI +L IR AQ+ D + + ++N+ E ++ G + GR+C+P++ LK+
Sbjct: 500 LKIWRSYL--IRLAQERDEE-IKGWTLNNKTE---YQTSNNGTIVVNGRVCVPNDRALKE 553
Query: 118 LILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGL 177
IL E H+S SIH G+ KMY+DLK+ + W G++KDVAR+V C TCQ K EHQ P+GL
Sbjct: 554 EILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMRKDVARWVAKCPTCQLVKAEHQVPSGL 613
Query: 178 LTPLDVPEWKWDSISMDFVSSLPNTSRG-HDSIWVVVDRLTKSAHFIPINISYPVAQLAE 236
L L + EWKWD I+MDFV+ LP + H+++WVVVDRLTKSAHF+ I+ +AE
Sbjct: 614 LQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAE 673
Query: 237 IYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
YI I++LHG+P SIVSDRD RFTS+FW + Q
Sbjct: 674 KYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQ 706
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 179 bits (454), Expect = 3e-45
Identities = 107/266 (40%), Positives = 148/266 (55%), Gaps = 39/266 (14%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP KA VVADALSRK + A + E L +C ++ + + L + +D L
Sbjct: 913 YHPGKANVVADALSRK--RVGAAPGQSVEALVSEIGALRLCVVAREPLGLEAVD-RADLL 969
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
R AQ+ D + ++ +++AE S+++ G + GR+C+P +EEL++ IL E H
Sbjct: 970 TRARLAQEKD----EGLIAASKAEGSEYQFAANGTIFVYGRVCVPKDEELRREILSEAHA 1025
Query: 126 SNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPE 185
S SIH GATKMY+DLK+ + W G+K+DVA +V C CQ K EHQ P
Sbjct: 1026 SMFSIHPGATKMYRDLKRYYQWVGMKRDVANWVAECDVCQLVKAEHQVP----------- 1074
Query: 186 WKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKL 245
D+IWV++DRLTKSAHF+ I + A LA+ Y+ IVKL
Sbjct: 1075 ---------------------DAIWVIMDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKL 1113
Query: 246 HGVPSSIVSDRDPRFTSRFWRSLQMR 271
HGVP SIVSDRD +FT FWR+ Q +
Sbjct: 1114 HGVPVSIVSDRDSKFTFAFWRAFQAK 1139
>At3g11970 hypothetical protein
Length = 1499
Score = 159 bits (402), Expect = 3e-39
Identities = 96/270 (35%), Positives = 140/270 (51%), Gaps = 34/270 (12%)
Query: 1 NFGLNYHPDKAKVVADALSR----KTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLG 56
++ + Y K VVADALSR + LHM A+ V E +LL+ +
Sbjct: 991 DYEIQYRQGKENVVADALSRVEGSEVLHM-AMTVVECDLLKDIQ---------------- 1033
Query: 57 MLKINSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELK 116
G ++Q D+ + + + + Q +LR + +I +P N+ +K
Sbjct: 1034 --------AGYANDSQLQDI----ITALQRDPDSKKYFSWSQNILRRKSKIVVPANDNIK 1081
Query: 117 KLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAG 176
IL H S + H G +Q +K LF+W G+ KD+ ++ +C TCQ+ K + G
Sbjct: 1082 NTILLWLHGSGVGGHSGRDVTHQRVKGLFYWKGMIKDIQAYIRSCGTCQQCKSDPAASPG 1141
Query: 177 LLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAE 236
LL PL +P+ W +SMDF+ LP S G I VVVDRL+K+AHFI ++ Y +A
Sbjct: 1142 LLQPLPIPDTIWSEVSMDFIEGLP-VSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAH 1200
Query: 237 IYIQNIVKLHGVPSSIVSDRDPRFTSRFWR 266
Y+ N+ KLHG P+SIVSDRD FTS FWR
Sbjct: 1201 AYLDNVFKLHGCPTSIVSDRDVVFTSEFWR 1230
>At1g36590 hypothetical protein
Length = 1499
Score = 154 bits (390), Expect = 7e-38
Identities = 95/270 (35%), Positives = 140/270 (51%), Gaps = 34/270 (12%)
Query: 1 NFGLNYHPDKAKVVADALSR----KTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLG 56
++ + Y K VVADALSR + LHM A+ V E +LL+ +
Sbjct: 991 DYEIQYRQGKENVVADALSRVEGSEVLHM-AMTVVECDLLKDIQ---------------- 1033
Query: 57 MLKINSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELK 116
G ++Q D+ + + + + Q +LR + +I +P N+ +K
Sbjct: 1034 --------AGYANDSQLQDI----ITALQRDPDSKKYFSWSQNILRRKSKIVVPANDNIK 1081
Query: 117 KLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAG 176
IL H S + H G +Q +K LF+ G+ KD+ ++ +C TCQ+ K + G
Sbjct: 1082 NTILLWLHGSGVGGHSGRDVTHQRVKGLFYSKGMIKDIQAYIRSCGTCQQCKSDPAASPG 1141
Query: 177 LLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAE 236
LL PL +P+ W +SMDF+ LP S G I VVVDRL+K+AHFI ++ Y +A+
Sbjct: 1142 LLQPLPIPDTIWSEVSMDFIEGLP-VSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAQ 1200
Query: 237 IYIQNIVKLHGVPSSIVSDRDPRFTSRFWR 266
Y+ N+ KLHG P+SIVSDRD FTS FWR
Sbjct: 1201 AYLDNVFKLHGCPTSIVSDRDVVFTSEFWR 1230
>At4g07850 putative polyprotein
Length = 1138
Score = 153 bits (387), Expect = 2e-37
Identities = 75/171 (43%), Positives = 108/171 (62%), Gaps = 2/171 (1%)
Query: 98 QGVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARF 157
+G L + R+CIP N L++L + E H L H G +K + ++ F W +K+DV R
Sbjct: 742 EGFLFYDNRLCIP-NSSLRELFIREAHGGGLMGHFGVSKTLKVMQDHFHWPHMKRDVERM 800
Query: 158 VYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLT 217
C TC+++K + Q P GL TPL +P W+ ISMDFV LP T G DSI+VVVDR +
Sbjct: 801 CERCTTCKQAKAKSQ-PHGLCTPLPIPLHPWNDISMDFVVGLPRTRTGKDSIFVVVDRFS 859
Query: 218 KSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
K AHFIP + + +A ++ + +V+LHG+P +IVSDRD +F S FW++L
Sbjct: 860 KMAHFIPCHKTDDAMHIANLFFREVVRLHGMPKTIVSDRDTKFLSYFWKTL 910
>At1g35370 hypothetical protein
Length = 1447
Score = 151 bits (382), Expect = 6e-37
Identities = 88/267 (32%), Positives = 138/267 (50%), Gaps = 28/267 (10%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKI 60
++ + Y K +VADALSR S+ + + + +
Sbjct: 962 DYEIQYRQGKENLVADALSRVE--------------------------GSEVLHMALSIV 995
Query: 61 NSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVD-EQGVLRFRGRICIPDNEELKKLI 119
DFL I+ A + D D++ Q ++ Q +LR + +I +P++ E+ +
Sbjct: 996 ECDFLKEIQVAYESDGVLKDIISALQQHPDAKKHYSWSQDILRRKSKIVVPNDVEITNKL 1055
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
L+ H S + G +Q +K LF+W G+ KD+ F+ +C TCQ+ K ++ GLL
Sbjct: 1056 LQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIRSCGTCQQCKSDNAAYPGLLQ 1115
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYI 239
PL +P+ W +SMDF+ LPN S G I VVVDRL+K+AHF+ + Y +A+ ++
Sbjct: 1116 PLPIPDKIWCDVSMDFIEGLPN-SGGKSVIMVVVDRLSKAAHFVALAHPYSALTVAQAFL 1174
Query: 240 QNIVKLHGVPSSIVSDRDPRFTSRFWR 266
N+ K HG P+SIVSDRD FTS FW+
Sbjct: 1175 DNVYKHHGCPTSIVSDRDVLFTSDFWK 1201
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 147 bits (370), Expect = 1e-35
Identities = 95/257 (36%), Positives = 134/257 (51%), Gaps = 34/257 (13%)
Query: 1 NFGLNYHPDKAKVVADALSR----KTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLG 56
++ + Y K +VADALSR + LHM AL V E +LL++ +Q G
Sbjct: 382 DYEIQYKQGKENLVADALSRVEGSEVLHM-ALSVVECDLLKE--------------IQAG 426
Query: 57 MLKINSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELK 116
+ + D G I QQ QA+ QGVLR + +I +P+N +K
Sbjct: 427 YVT-DGDIQGIITILQQ-------------QADSKKHYTWSQGVLRRKNKIVVPNNSGIK 472
Query: 117 KLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAG 176
IL H S + H G +Q +K LF+W + KD+ F+ +C TCQ+ K ++ G
Sbjct: 473 DTILRWLHCSGMGGHSGKEVTHQRVKGLFYWKSMVKDIQAFIRSCGTCQQCKSDNAASPG 532
Query: 177 LLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAE 236
LL PL +P+ W +SMDF+ LP S G I VVVDRL+K+AHFI + Y +A+
Sbjct: 533 LLQPLPIPDRIWSDVSMDFIDGLP-LSNGKTVIMVVVDRLSKAAHFIALAHPYSAMTVAQ 591
Query: 237 IYIQNIVKLHGVPSSIV 253
Y+ N+ KLHG PSSIV
Sbjct: 592 AYLDNVFKLHGCPSSIV 608
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 146 bits (368), Expect = 3e-35
Identities = 83/191 (43%), Positives = 116/191 (60%), Gaps = 7/191 (3%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKIN-SDF 64
YHP KA VADALSR+ + A + +L+ L V LS + LG+ + +D
Sbjct: 505 YHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH-VNALSKEVESLGLGAADQADL 562
Query: 65 LGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGH 124
L IR AQ+ D + + ++N+ E ++ G + GR+C+P++ LK+ IL E H
Sbjct: 563 LSRIRLAQERDEE-IKGWALNNKTE---YQTSNNGTIVVNGRVCVPNDRALKEEILREAH 618
Query: 125 KSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVP 184
+S SIH G+ KMY+DLK+ + W G+KKDVAR+V C TCQ K EHQ P+GLL L +P
Sbjct: 619 QSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPIP 678
Query: 185 EWKWDSISMDF 195
EWKWD I+MDF
Sbjct: 679 EWKWDHITMDF 689
>At4g03840 putative transposon protein
Length = 973
Score = 141 bits (355), Expect = 8e-34
Identities = 81/191 (42%), Positives = 115/191 (59%), Gaps = 7/191 (3%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKIN-SDF 64
YH KA VADALSR+ + A + +L+ L V LS + LG+ + ++
Sbjct: 505 YHAGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH-VNALSKEVEPLGLGAADQANL 562
Query: 65 LGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGH 124
L IR AQ+ D + + ++N+ E ++ G + GR+C+P+N LK+ IL E H
Sbjct: 563 LSRIRLAQERDEE-IKGWALNNKTE---YQTSNNGTIVVNGRVCVPNNRALKEEILREAH 618
Query: 125 KSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVP 184
+S SIH G+ K+Y+DLK+ + W G+KKDVAR+V C TCQ K EHQ P+GLL L +P
Sbjct: 619 QSKFSIHPGSNKIYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPIP 678
Query: 185 EWKWDSISMDF 195
EWKWD I+MDF
Sbjct: 679 EWKWDHITMDF 689
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 141 bits (355), Expect = 8e-34
Identities = 70/118 (59%), Positives = 84/118 (70%), Gaps = 1/118 (0%)
Query: 152 KDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWV 211
+DVA +V C CQ K EHQ P G+L L +PEWKWD I++DFV LP SR D+IWV
Sbjct: 941 EDVANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLP-VSRTKDAIWV 999
Query: 212 VVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
+VDRLTKSAHF+ I + A LA+ Y+ IVKLHGVP SIVSDRD +FTS FWR+ Q
Sbjct: 1000 IVDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQ 1057
Score = 32.7 bits (73), Expect = 0.41
Identities = 27/87 (31%), Positives = 44/87 (50%), Gaps = 7/87 (8%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP KA VVADALS K + A + E L +C ++ + + L + +D L
Sbjct: 862 YHPGKANVVADALSHK--RVGAAPGQSVEALVSEIGALRLCAVAREPLGLEAVD-RADLL 918
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESD 92
+R AQ+ D + ++ + +AE S+
Sbjct: 919 TRVRLAQEKD----EGLIAAYKAEGSE 941
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 134 bits (338), Expect = 8e-32
Identities = 96/266 (36%), Positives = 129/266 (48%), Gaps = 60/266 (22%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP KA VV DALSRK + A + + E+L +C ++ + + L + +D L
Sbjct: 887 YHPGKANVVVDALSRK--RVGAALGQSVEVLVSEIGALRLCAVAREPLGLEAVD-RADLL 943
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
+R AQ KK +EG
Sbjct: 944 TRVRLAQ-------------------------------------------KK---DEG-- 955
Query: 126 SNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPE 185
L ATKMY+DLK+ + W G+K DVA +V C CQ K EHQ G+L L +PE
Sbjct: 956 ------LRATKMYRDLKRYYQWVGMKMDVANWVAECDVCQLVKAEHQVLGGMLQSLPIPE 1009
Query: 186 WKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKL 245
WKWD I+MD V L SR D+IWV+VDRLTKSAHF+ I + A LA+ ++ IVKL
Sbjct: 1010 WKWDFITMDLVVGL-RVSRTKDAIWVIVDRLTKSAHFLAIRKTDGAAVLAKKFVSEIVKL 1068
Query: 246 HGVPSSI--VSDRDPRFTSRFWRSLQ 269
HGVP ++ DR + + R L+
Sbjct: 1069 HGVPLNMKEAQDRQRSYADKRRRELE 1094
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 131 bits (329), Expect = 9e-31
Identities = 94/267 (35%), Positives = 124/267 (46%), Gaps = 80/267 (29%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP K VVADALSRK + + QSV+ + +I
Sbjct: 803 YHPGKTNVVADALSRKRVGAAP----------------------GQSVEALVSEI----- 835
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDE--QGVLRFRGRICIPDNEELKKLILEEG 123
G++R L VV+ + + + VD G + R+C+P +EEL++ IL E
Sbjct: 836 GALR-----------LCVVAREPLKLE-AVDRAANGTILVHERVCLPKDEELRREILSEA 883
Query: 124 HKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDV 183
H S SIH GATKMY+DLK+ + W G+K+DVA +V C CQ K EHQ P GLL L +
Sbjct: 884 HASMFSIHPGATKMYRDLKRHYQWVGMKRDVANWVTECDVCQLVKAEHQVPGGLLQSLPI 943
Query: 184 PEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIV 243
EWK D + LP Y+ IV
Sbjct: 944 SEWK----KTDGAAVLPKK-----------------------------------YVSEIV 964
Query: 244 KLHGVPSSIVSDRDPRFTSRFWRSLQM 270
KLHGVP SI+S RD +FTS FWR+ Q+
Sbjct: 965 KLHGVPVSILSHRDSKFTSAFWRAFQV 991
>At3g31490 hypothetical protein
Length = 285
Score = 99.4 bits (246), Expect = 4e-21
Identities = 62/182 (34%), Positives = 96/182 (52%), Gaps = 23/182 (12%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YHP KA V +AL V E L +C ++ + ++L ++ +D L
Sbjct: 127 YHPGKANVSVEAL-----------VSEISALR-------LCAVAREPLELEVVG-RADLL 167
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
+R AQ+ D + ++ +++ E S+ + G + +IC P +E+L+ IL E H
Sbjct: 168 SRVRLAQEKD----EGLIAASKTEGSENQFAANGTILVHAQICGPKDEKLRWKILSEVHA 223
Query: 126 SNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPE 185
S SI+ ATKMY+DLK+ + W G+K+DVA ++ C CQ K EHQ P LL L +PE
Sbjct: 224 SMFSIYPRATKMYRDLKRYYQWVGMKRDVANWIAECDVCQLVKAEHQVPGALLQSLPIPE 283
Query: 186 WK 187
K
Sbjct: 284 KK 285
>At2g06890 putative retroelement integrase
Length = 1215
Score = 88.2 bits (217), Expect = 8e-18
Identities = 75/223 (33%), Positives = 103/223 (45%), Gaps = 35/223 (15%)
Query: 4 LNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSD 63
+ Y K VVADALS++ +S L VK L F + V E + D
Sbjct: 813 IKYKKGKDNVVADALSQRYTLLSTLNVK----LMGFEQIKEVYET------------DHD 856
Query: 64 FLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEG 123
F +E + KF A F+ D+ L + R+C+P N L+ L + E
Sbjct: 857 F----QEVYKACEKF---------ASGRYFRQDK--FLFYENRLCVP-NCSLRDLFVREA 900
Query: 124 HKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDV 183
H L H G K + + + F W +K DV R C TC+++K + Q P GL TPL +
Sbjct: 901 HGGGLMGHFGIAKTLEVMTEHFRWPHMKCDVKRICGRCNTCKQAKSKIQ-PNGLYTPLPI 959
Query: 184 PEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPIN 226
P+ W+ ISMDFV LP T G DSI+VV F PI+
Sbjct: 960 PKHPWNDISMDFVMGLPRT--GKDSIFVVYSPFQIVYGFNPIS 1000
>At2g06190 putative Ty3-gypsy-like retroelement pol polyprotein
Length = 280
Score = 84.7 bits (208), Expect = 9e-17
Identities = 38/76 (50%), Positives = 52/76 (68%)
Query: 193 MDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSI 252
MDFV LP T RG DS++VVVDR +K HFI + + +A+++ + +V+LHGVP SI
Sbjct: 1 MDFVLGLPRTQRGVDSVFVVVDRFSKMTHFIACKKTADASNIAKLFFKEVVRLHGVPKSI 60
Query: 253 VSDRDPRFTSRFWRSL 268
SDRD +F S FW +L
Sbjct: 61 TSDRDTKFLSHFWSTL 76
>At3g29480 hypothetical protein
Length = 718
Score = 84.0 bits (206), Expect = 2e-16
Identities = 53/140 (37%), Positives = 83/140 (58%), Gaps = 7/140 (5%)
Query: 6 YHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFL 65
YH KA VVADALSRK + ++ E L+ + R L L C ++ + + L + D L
Sbjct: 586 YHHGKASVVADALSRKRVGVAPGQSVE-ALVSEIRALRL-CAVAREPLGLEAVD-RVDLL 642
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
+R AQ+ D + ++ +++AE S ++ G + GR+C+ ++EEL++ IL E H
Sbjct: 643 SRVRLAQEKD----EGLIAASKAEGSKYQFAANGTILVHGRVCVLNDEELRREILSEAHA 698
Query: 126 SNLSIHLGATKMYQDLKKLF 145
S SIH GATKMY+DLK+ +
Sbjct: 699 SMFSIHPGATKMYRDLKEYY 718
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.148 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,461,992
Number of Sequences: 26719
Number of extensions: 327506
Number of successful extensions: 1331
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1256
Number of HSP's gapped (non-prelim): 55
length of query: 443
length of database: 11,318,596
effective HSP length: 102
effective length of query: 341
effective length of database: 8,593,258
effective search space: 2930300978
effective search space used: 2930300978
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144765.5


