

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.14 - phase: 0
(113 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
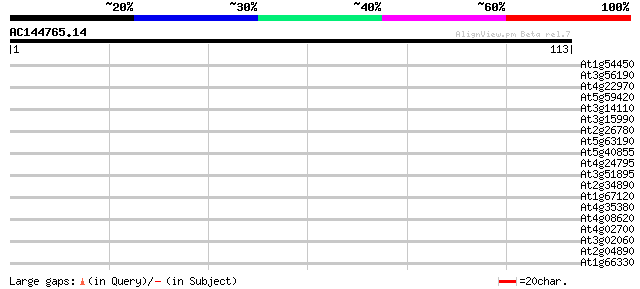
Score E
Sequences producing significant alignments: (bits) Value
At1g54450 hypothetical protein 29 0.44
At3g56190 alpha-soluble NSF attachment protein 28 0.97
At4g22970 putative protein 27 1.3
At5g59420 oxysterol-binding protein - like 27 1.7
At3g14110 unknown protein 27 1.7
At3g15990 putative sulfate transporter 27 2.2
At2g26780 unknown protein 26 2.8
At5g63190 topoisomerase-like protein 26 3.7
At5g40855 putative protein 25 4.8
At4g24795 unknown protein 25 4.8
At3g51895 sulfate transporter (ATST1) 25 4.8
At2g34890 putative CTP synthase 25 6.3
At1g67120 hypothetical protein 25 6.3
At4g35380 putative protein 25 8.2
At4g08620 putative sulfate transporter 25 8.2
At4g02700 sulfate transporter protein 25 8.2
At3g02060 helicase like protein 25 8.2
At2g04890 putative SCARECROW gene regulator 25 8.2
At1g66330 unknown protein 25 8.2
>At1g54450 hypothetical protein
Length = 535
Score = 28.9 bits (63), Expect = 0.44
Identities = 20/81 (24%), Positives = 37/81 (44%), Gaps = 4/81 (4%)
Query: 29 RLHDLNLEDANAILRE--AAEIFEGLGMVDSAAQCFTDLGDYERAGINFNFGIHVYSFMS 86
+LH + A+L E ++F+ + D C DL + +G FN ++ FM+
Sbjct: 419 QLHRMECMAQEAVLFEDILCQLFDMVKPEDEGFICLNDLKGSKLSGNVFNILFNLNKFMA 478
Query: 87 FSIKKHYDILEHKYQVYPTWS 107
F + + I + + PTW+
Sbjct: 479 FETRDPFLIRQER--ANPTWT 497
>At3g56190 alpha-soluble NSF attachment protein
Length = 289
Score = 27.7 bits (60), Expect = 0.97
Identities = 23/85 (27%), Positives = 34/85 (39%), Gaps = 4/85 (4%)
Query: 2 ATMCFERAGDSYW--GKKSKAASLRATAIRLH--DLNLEDANAILREAAEIFEGLGMVDS 57
A C ERA + + G+ + AA + D E A A +AAE F+ + S
Sbjct: 91 AASCLERAVNIFCEIGRLNMAARYYKEIAEYYESDQKFEQAIAYFEKAAEFFQNEEVTTS 150
Query: 58 AAQCFTDLGDYERAGINFNFGIHVY 82
A QC + Y + I +Y
Sbjct: 151 ANQCNLKVAQYAAQLEQYEKAIKIY 175
>At4g22970 putative protein
Length = 1773
Score = 27.3 bits (59), Expect = 1.3
Identities = 26/74 (35%), Positives = 35/74 (47%), Gaps = 7/74 (9%)
Query: 33 LNLEDANAILREA-AEIFEGL--GMVDSAAQCFTDLGDYERAGINFNFGIHVYSFMSFSI 89
L LE+ LR A+++E L MV S +C L R FN G V SF ++
Sbjct: 195 LLLEEVGGWLRVLDAKVYEKLHRAMVTSMGKCAVSL---VREAERFN-GDLVISFCDLTV 250
Query: 90 KKHYDILEHKYQVY 103
K+HY K +VY
Sbjct: 251 KEHYKSALSKDRVY 264
>At5g59420 oxysterol-binding protein - like
Length = 457
Score = 26.9 bits (58), Expect = 1.7
Identities = 23/92 (25%), Positives = 39/92 (42%), Gaps = 3/92 (3%)
Query: 11 DSYWGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYER 70
DSYW K T++ + + + +L++ AEI E ++D A +C Y R
Sbjct: 67 DSYWKMMQKYIGSDITSMVTLPVVIFEPMTMLQKMAEIMEYSHLLDQADEC---EDPYLR 123
Query: 71 AGINFNFGIHVYSFMSFSIKKHYDILEHKYQV 102
++ I VY + K IL Y++
Sbjct: 124 LVYASSWAISVYYAFQRTWKPFNPILGETYEM 155
>At3g14110 unknown protein
Length = 316
Score = 26.9 bits (58), Expect = 1.7
Identities = 19/49 (38%), Positives = 27/49 (54%), Gaps = 10/49 (20%)
Query: 27 AIRLHDLNLEDANAILREAAE---IFEGLGMVDSAAQCFTDLGDYERAG 72
AI+ H + L AI + +E I E G + A C+T+LGD E+AG
Sbjct: 262 AIQYHSMVL----AISKRESEDSGITEAYGAI---ADCYTELGDLEKAG 303
>At3g15990 putative sulfate transporter
Length = 653
Score = 26.6 bits (57), Expect = 2.2
Identities = 8/27 (29%), Positives = 14/27 (51%)
Query: 52 LGMVDSAAQCFTDLGDYERAGINFNFG 78
+ M S C+ G + R+ +N+N G
Sbjct: 377 MNMAGSCTSCYVTTGSFSRSAVNYNAG 403
>At2g26780 unknown protein
Length = 1732
Score = 26.2 bits (56), Expect = 2.8
Identities = 21/82 (25%), Positives = 35/82 (42%), Gaps = 13/82 (15%)
Query: 23 LRATAIRLHDLN-------LEDANAILREAAEIFEGLGM------VDSAAQCFTDLGDYE 69
L T RLH N LE+ + + + ++E L + ++S Q L
Sbjct: 1227 LFVTVCRLHAANIGIETEKLENLRISISKGSPMWETLDLCINIVDIESLEQLIPRLTQLV 1286
Query: 70 RAGINFNFGIHVYSFMSFSIKK 91
R G+ N + V SF+S ++K
Sbjct: 1287 RGGVGLNTRVGVASFISLLVQK 1308
>At5g63190 topoisomerase-like protein
Length = 729
Score = 25.8 bits (55), Expect = 3.7
Identities = 12/32 (37%), Positives = 17/32 (52%)
Query: 35 LEDANAILREAAEIFEGLGMVDSAAQCFTDLG 66
+EDA + + E +E G+ A QC DLG
Sbjct: 608 VEDAKDKISKLLEEYETGGVTSEACQCIRDLG 639
>At5g40855 putative protein
Length = 78
Score = 25.4 bits (54), Expect = 4.8
Identities = 9/30 (30%), Positives = 20/30 (66%)
Query: 73 INFNFGIHVYSFMSFSIKKHYDILEHKYQV 102
++F +H+ + +F + K YD+++ KYQ+
Sbjct: 30 LHFTILLHLDTCHTFLVPKRYDLVKCKYQL 59
>At4g24795 unknown protein
Length = 702
Score = 25.4 bits (54), Expect = 4.8
Identities = 11/32 (34%), Positives = 17/32 (52%)
Query: 35 LEDANAILREAAEIFEGLGMVDSAAQCFTDLG 66
+EDA + E +E G+V A +C +LG
Sbjct: 575 VEDAKDKISNLLEEYESSGLVSEACKCIHELG 606
>At3g51895 sulfate transporter (ATST1)
Length = 658
Score = 25.4 bits (54), Expect = 4.8
Identities = 9/30 (30%), Positives = 16/30 (53%)
Query: 49 FEGLGMVDSAAQCFTDLGDYERAGINFNFG 78
F + +V S C+ G + R+ +N+N G
Sbjct: 367 FGMMNIVGSFTSCYLTTGPFSRSAVNYNAG 396
>At2g34890 putative CTP synthase
Length = 597
Score = 25.0 bits (53), Expect = 6.3
Identities = 17/53 (32%), Positives = 27/53 (50%), Gaps = 5/53 (9%)
Query: 19 KAASLRATAIRLHDLNLEDANAILR-EAAEIFEGLGMVDSAAQCFTDLGDYER 70
++ LR T I++ DA I E E+F ++D ++ DLG+YER
Sbjct: 28 QSCGLRVTCIKIDPYLNYDAGTISPYEHGEVF----VLDDGSEVDLDLGNYER 76
>At1g67120 hypothetical protein
Length = 5138
Score = 25.0 bits (53), Expect = 6.3
Identities = 11/26 (42%), Positives = 16/26 (61%)
Query: 29 RLHDLNLEDANAILREAAEIFEGLGM 54
R+H L D +A+ +EAA+IF M
Sbjct: 472 RVHGLPSYDGHAVYQEAADIFSASNM 497
>At4g35380 putative protein
Length = 1711
Score = 24.6 bits (52), Expect = 8.2
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 11/69 (15%)
Query: 42 LREAAEI-FEGLGMVDSAAQCFTDLGDYER-AGINFNFG------IHVYSFMSFSIKKHY 93
L+EAA + F G V + D+ D E +G + N G +H+ S++ KKH
Sbjct: 1448 LKEAASLTFAGFMKV---LRTMDDIEDVETLSGQSVNIGDLDDDSLHIMSYVVSRTKKHI 1504
Query: 94 DILEHKYQV 102
D+L +V
Sbjct: 1505 DVLSQIVEV 1513
>At4g08620 putative sulfate transporter
Length = 649
Score = 24.6 bits (52), Expect = 8.2
Identities = 8/28 (28%), Positives = 15/28 (53%)
Query: 52 LGMVDSAAQCFTDLGDYERAGINFNFGI 79
+ +V S C+ G + R+ +NF G+
Sbjct: 374 MNVVGSMTSCYIATGSFSRSAVNFMAGV 401
>At4g02700 sulfate transporter protein
Length = 646
Score = 24.6 bits (52), Expect = 8.2
Identities = 8/30 (26%), Positives = 17/30 (56%)
Query: 49 FEGLGMVDSAAQCFTDLGDYERAGINFNFG 78
F + ++ S + C+ G + R+ +N+N G
Sbjct: 358 FGMMNILGSFSSCYLTTGPFSRSAVNYNAG 387
>At3g02060 helicase like protein
Length = 823
Score = 24.6 bits (52), Expect = 8.2
Identities = 6/15 (40%), Positives = 12/15 (80%)
Query: 90 KKHYDILEHKYQVYP 104
K+HYD++ ++ +YP
Sbjct: 333 KQHYDVISERFSLYP 347
>At2g04890 putative SCARECROW gene regulator
Length = 413
Score = 24.6 bits (52), Expect = 8.2
Identities = 12/33 (36%), Positives = 18/33 (54%)
Query: 64 DLGDYERAGINFNFGIHVYSFMSFSIKKHYDIL 96
D+ D E G+NF + +H S S++ H D L
Sbjct: 235 DVRDGEALGVNFAYMLHHLPDESVSMENHRDRL 267
>At1g66330 unknown protein
Length = 417
Score = 24.6 bits (52), Expect = 8.2
Identities = 22/76 (28%), Positives = 29/76 (37%), Gaps = 5/76 (6%)
Query: 7 ERAGDSYWGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGL--GMVDSAAQCFTD 64
+ A D +W + A LR R +EDA L I + + MVDS T
Sbjct: 190 KNAADKHWSDGALEADLRRADFRAKQRAMEDALMALEFIKNIHDMMVNKMVDSLVTSETG 249
Query: 65 LGD---YERAGINFNF 77
D E+ GI F
Sbjct: 250 TTDRISLEKNGIALGF 265
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,305,926
Number of Sequences: 26719
Number of extensions: 80718
Number of successful extensions: 278
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 265
Number of HSP's gapped (non-prelim): 19
length of query: 113
length of database: 11,318,596
effective HSP length: 89
effective length of query: 24
effective length of database: 8,940,605
effective search space: 214574520
effective search space used: 214574520
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144765.14


