

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144760.12 + phase: 0 /pseudo
(1302 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
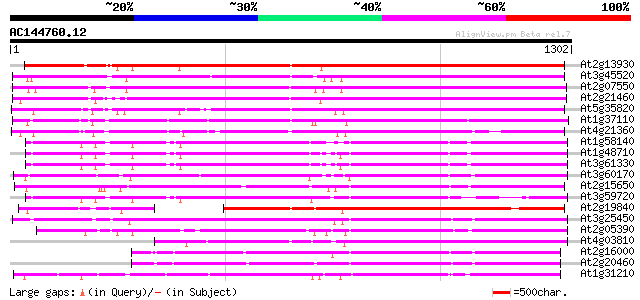
Sequences producing significant alignments: (bits) Value
At2g13930 putative retroelement pol polyprotein 984 0.0
At3g45520 copia-like polyprotein 954 0.0
At2g07550 putative retroelement pol polyprotein 953 0.0
At2g21460 putative retroelement pol polyprotein 949 0.0
At5g35820 copia-like retrotransposable element 948 0.0
At1g37110 940 0.0
At4g21360 putative transposable element 868 0.0
At1g58140 hypothetical protein 795 0.0
At1g48710 hypothetical protein 795 0.0
At3g61330 copia-type polyprotein 791 0.0
At3g60170 putative protein 778 0.0
At2g15650 putative retroelement pol polyprotein 700 0.0
At3g59720 copia-type reverse transcriptase-like protein 698 0.0
At2g19840 copia-like retroelement pol polyprotein 689 0.0
At3g25450 hypothetical protein 669 0.0
At2g05390 putative retroelement pol polyprotein 652 0.0
At4g03810 putative retrotransposon protein 647 0.0
At2g16000 putative retroelement pol polyprotein 588 e-168
At2g20460 putative retroelement pol polyprotein 582 e-166
At1g31210 putative reverse transcriptase 552 e-157
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 984 bits (2545), Expect = 0.0
Identities = 569/1319 (43%), Positives = 799/1319 (60%), Gaps = 79/1319 (5%)
Query: 34 QPLTEKK---PDSMKD---DEWSLLDR--QALGVVRLSLSRNVAFNIAKEKTTAGLMKAL 85
+PLTE++ P+ K DE + L+R +A V+ L+++ V I KT A + L
Sbjct: 19 KPLTEEEEEDPEKRKKRDADEVARLERCDKAKNVIFLNVADKVLRKIELCKTAAEAWETL 78
Query: 86 SSMYEKPPSSNKVHLMRRLFTLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALIL 145
++ ++V+ +T +M E + ++I++ + L+ + I+ DEV+A++L
Sbjct: 79 DRLFMIRSLPHRVYTQLSFYTFKMQENKKIDENIDDFLKIVADLNHLQIDVTDEVQAILL 138
Query: 146 LSSLPDSWSAIVTAVSSSSGSKKMKFDDVRDLVLSEEIRRRELGESSSSSVLHTESRGRN 205
LSSLP + +V + S+ +K++ DDV +++ + REL +++ V +RGR
Sbjct: 139 LSSLPARYDGLVETMKYSNSREKLRLDDV---MVAARDKERELSQNNRPVVEGHFARGRP 195
Query: 206 STRGNGRGKSKARRSKSKNHRSSHNSKSIECWNCGKTGHFKNQC-----RLPTKNQ-EEK 259
+ N +G RS+SK S + K + CW CGK GHFK QC R +K Q +
Sbjct: 196 DGKNNNQGNKGKNRSRSK----SADGKRV-CWICGKEGHFKKQCYKWIERNKSKQQGSDN 250
Query: 260 DEANVASTSGGGDALICSLESKE---------ESWVLDSGASFHASSQKEFFKNYVPGNL 310
E+++A ++ + + L + E WVLD+G SFH + +K++FK++ +
Sbjct: 251 GESSLAKSTEAFNPAMVLLATDETLVVTDSIANEWVLDTGCSFHMTPRKDWFKDFKELSS 310
Query: 311 GKVYLGNEQSCKVVGKGEVKIK-LNGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFH 369
G V +GN+ V G G +KI+ +GS L +VR++PN+T+NLIS+G L D G
Sbjct: 311 GYVKMGNDTYSPVKGIGSIKIRNSDGSQVILTDVRYMPNMTRNLISLGTLEDRGCWFKSQ 370
Query: 370 GDDWKISKGAMTIARGRKSGTLY-----KTAGACHLIAVATNENPNLWHKRLGHMSEKGM 424
KI KG TI +G+K TLY G H A +E LWH RLGHMS+KGM
Sbjct: 371 DGILKIVKGCSTILKGQKRDTLYILDGVTEEGESHSSAEVKDETA-LWHSRLGHMSQKGM 429
Query: 425 KVMHSKGKLPSLRSIEIDICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWG-PTTVP 483
+++ KG L E++ CEDC+ GKQ RVSF + K EKL VHSD+WG P
Sbjct: 430 EILVKKGCLRREVIKELEFCEDCVYGKQHRVSFAPAQHVTK-EKLAYVHSDLWGSPHNPA 488
Query: 484 SIGGKHYFVTFIDDHSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEY 543
S+G YF++F+DD+SRKVW+YFL+ K E FE F WK MVEN++D K+KKLRTDNG EY
Sbjct: 489 SLGNSQYFISFVDDYSRKVWIYFLRKKDEAFEKFVEWKKMVENQSDRKVKKLRTDNGLEY 548
Query: 544 EDTKFKKFCYEHGIRMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVN 603
+ F+KFC E GI +T TPQ NG+AER+NRT+ ++ RS+ +SG+ KKFWAEA +
Sbjct: 549 CNHYFEKFCKEEGIVRHKTCAYTPQQNGIAERLNRTIMDKVRSMLSRSGMEKKFWAEAAS 608
Query: 604 TSAYLINRGPSVPLEHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCI 663
T+ YLINR PS + +PEE W+G LS LR FGC+AY+H +DQG KL+P+SKK I
Sbjct: 609 TAVYLINRSPSTAINFDLPEEKWTGALPDLSSLRKFGCLAYIH-ADQG--KLNPRSKKGI 665
Query: 664 FIGYGEDEFGYRLWDDENKKMVRSKDVIFNERVMYKD---KHNTTTNDSGLSE-PVYVEM 719
F Y E GY++W E+KK V S++VIF E+VM+KD T ++S L + V +M
Sbjct: 666 FTSYPEGVKGYKVWVLEDKKCVISRNVIFREQVMFKDLKGDSQNTISESDLEDLRVNPDM 725
Query: 720 DD-------------------------VPGSPTDKSPQSGELAESSIRQPSDT--LVHPT 752
+D V SPT + E +S + T LV
Sbjct: 726 NDQEFTDQGGATQDNSNPSEATTSHNPVLNSPTHSQDEESEEEDSDAVEDLSTYQLVRDR 785
Query: 753 PVPVLRRSSRPHAPNRRYIDYMLLTDG-GEPEDYDEACQTTDASKWELAMKEEMKSLISN 811
++ + + + N Y DG EP+ Y EA D KW AMKEEM S+ N
Sbjct: 786 VRRTIKANPKYNESNMVGFAYYSEDDGKPEPKSYQEALLDPDWEKWNAAMKEEMVSMSKN 845
Query: 812 QTWELAKLPIGKKALHNKWVYRVKEDHDG--SKRYKARLVVKGFRQKEGIDYTEIFAPVV 869
TW+L P K + +WV+ K G + R+ ARLV KGF QKEG+DY EIF+PVV
Sbjct: 846 HTWDLVTKPEKVKLIGCRWVFTRKAGIPGVEAPRFIARLVAKGFTQKEGVDYNEIFSPVV 905
Query: 870 KLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKS 929
K +IR +LS+V N+ L+Q+DVKTAFLHG L EEIYM QPEGF + N VC+LK+S
Sbjct: 906 KHVSIRYLLSMVVHYNMELQQMDVKTAFLHGFLEEEIYMAQPEGFEIKRGSNKVCLLKRS 965
Query: 930 LYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYK-SSYIILLLYVDDMLVAGSNI 988
LYGLKQ+PRQW ++F+ FM + + D C +FK+ +YI LLLYVDDML+A +N
Sbjct: 966 LYGLKQSPRQWNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYIYLLLYVDDMLIASANK 1025
Query: 989 DEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAK 1048
E+ LK LS+EF+MKDLG AKKILGM+I+RD+ G+L LS Y+ +VL+ F M +AK
Sbjct: 1026 SEVNELKQLLSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYVKKVLRSFQMDNAK 1085
Query: 1049 LVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRF 1108
VSTPL HF+L + EE+ E M +PYA+ IGS+MY+M+ TRPD+ +++GV+SRF
Sbjct: 1086 PVSTPLGIHFKLKAATDKEYEEQFERMKIVPYANTIGSIMYSMIGTRPDLAYSLGVISRF 1145
Query: 1109 MSNPGKAHWEAVKWILRYLRGTTEKCLYFGKGE-IKVEGYVDADFAGEVDHRRSTTGYIF 1167
MS P K HW+AVKW+LRY+RGT +K L F K E + GY D+D+ D RRS TGY+F
Sbjct: 1146 MSKPLKDHWQAVKWVLRYMRGTEKKKLCFRKQEDFLLRGYCDSDYGSNFDTRRSITGYVF 1205
Query: 1168 TVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQ 1227
TVG ++SW S++QK+VA+S+TE EY+A+TEA KE +WL+G ELG Q+ ++SDSQ
Sbjct: 1206 TVGGNTISWKSKLQKVVAISSTEAEYMALTEAVKEALWLKGFAAELGHSQDYVEVHSDSQ 1265
Query: 1228 SAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
SAI LAKNS H RTKHI +R HFIR ++ ++ ++KI NPA++ TK V + K +
Sbjct: 1266 SAITLAKNSVHHERTKHIDIRLHFIRDIICAGLIKVVKIATECNPANIFTKTVPLAKFE 1324
>At3g45520 copia-like polyprotein
Length = 1363
Score = 954 bits (2465), Expect = 0.0
Identities = 545/1366 (39%), Positives = 792/1366 (57%), Gaps = 104/1366 (7%)
Query: 6 VKIEKFDG-ADFGFWKMQIEDYLYQKKLHQPLTE------KKPDSMKDDE---------- 48
+++EKFDG D+ WK ++ ++ L L E K+ DS K DE
Sbjct: 6 IEVEKFDGRGDYTMWKEKLLAHIDMLGLSAVLRESETPMGKERDSEKSDEDEKEEREKME 65
Query: 49 -WSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTL 107
+ R+A + LS+S V I KE + A +++AL +Y N+++L ++L++
Sbjct: 66 AFEEKKRKARSTIVLSVSDRVLRKIKKETSAAAMLEALDRLYMSKALPNRIYLKQKLYSF 125
Query: 108 RMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSK 167
+M+E +S+ +I+E + L ++ + DE +A++LL SLP + + + SSG
Sbjct: 126 KMSENLSIEGNIDEFLHIVADLENLNVLVSDEDQAILLLMSLPKPFDQLKDTLKYSSGKT 185
Query: 168 KMKFDDVRDLVLSEEIRRRELGESSSSSV--LHTESRGRNSTRGNGRGKSKARRSKSKNH 225
+ D+V + S E+ + +S L+ + + N R + K K +RSKSK+
Sbjct: 186 VLSLDEVAAAIYSRELEFGSVKKSIKGQAEGLYVKDKAENRGRSEQKDKGKGKRSKSKSK 245
Query: 226 RSSHNSKSIECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGG--------------- 270
R CW CG+ GH K+ C K Q + +N +SGG
Sbjct: 246 RG--------CWICGEDGHLKSTCPNKNKPQFKNQGSNKGESSGGKGNLVEGSVNFVESA 297
Query: 271 ----GDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGK 326
+AL + E+ W++D+G +H + ++E+ +++ G V +GN+ +V G
Sbjct: 298 GMFVSEALSSTDIHLEDEWIMDTGCIYHMTHKREWLEDFDEEAGGSVRMGNKSISRVKGV 357
Query: 327 GEVKI-KLNGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARG 385
G V+I NG L+NVR+IP++ +NL+S+G G+ +I G + G
Sbjct: 358 GTVRIVNDNGLTVTLQNVRYIPDMDRNLLSLGTFEKAGHKFESENGMLRIKSGNQVLLEG 417
Query: 386 RKSGTLY----KTAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEI 441
R+ TLY K A L N++ LWH+RL HMS+K M ++ KG L + +
Sbjct: 418 RRYDTLYILHGKPATDESLAVARANDDTVLWHRRLCHMSQKNMSLLIKKGFLDKKKVSML 477
Query: 442 DICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVP-SIGGKHYFVTFIDDHSR 500
D CEDCI G+ K++ F + KK KLE VHSD+WG TVP S+G YF++FIDD++R
Sbjct: 478 DTCEDCIYGRAKKIGFNLAQHDTKK-KLEYVHSDLWGAPTVPMSLGNCQYFISFIDDYTR 536
Query: 501 KVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRME 560
KVWVYFLK K E FE F W ++VEN++ ++K LRTDNG E+ + F FC E G +
Sbjct: 537 KVWVYFLKTKDEAFEKFVSWISLVENQSGERVKTLRTDNGLEFCNRMFDGFCEEKGFQRH 596
Query: 561 RTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHK 620
RT TPQ NGV ERMNRT+ E+ RS+ SGLPK+FWAEA +T+ LIN+ P + +
Sbjct: 597 RTCAYTPQQNGVVERMNRTIMEKVRSMLCDSGLPKRFWAEATHTAVLLINKTPCSAINFE 656
Query: 621 IPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDE 680
P++ WSGK S+LR +GCV +VH +D G KL+ ++KK + IGY GY++W E
Sbjct: 657 FPDKRWSGKAPIYSYLRRYGCVTFVH-TDGG--KLNLRAKKGVLIGYPSGVKGYKVWLIE 713
Query: 681 NKKMVRSKDVIFNERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTD-------KSPQS 733
KK V S++V F E +YKD E V E DD GS D +
Sbjct: 714 EKKCVVSRNVSFQENAVYKDLMQR-------KEQVSCEEDDHAGSYIDLDLEADKDNSSG 766
Query: 734 GELAESSIRQPS-----------------DTLVHPTPVP---VLRRSSRP-HAPNR---- 768
GE +++ + + +T VH +P+ V R R AP R
Sbjct: 767 GEQSQAQVTPATRGAVTSTPPRYETDDIEETDVHQSPLSYHLVRDRERREIRAPRRFDDE 826
Query: 769 -RYIDYMLLTDGG---EPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKK 824
Y + + T+ G EP DY EA + + KW LAM EE++S + N TW P ++
Sbjct: 827 DYYAEALYTTEDGDAVEPADYKEAVRDENWDKWRLAMNEEIESQLKNDTWTTVTRPEKQR 886
Query: 825 ALHNKWVYRVKEDHDG--SKRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVA 882
+ ++W+Y+ K+ G R+KARLV KG+ Q+EG+DY EIFAPVVK +IR +LSIVA
Sbjct: 887 IIGSRWIYKYKQGIPGVEEPRFKARLVAKGYAQREGVDYHEIFAPVVKHVSIRILLSIVA 946
Query: 883 SENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYM 942
ENL LEQLDVKTAFLHG+L E+IYM PEG KEN VC+L KSLYGLKQAPRQW
Sbjct: 947 QENLELEQLDVKTAFLHGELKEKIYMMPPEGCESLFKENEVCLLNKSLYGLKQAPRQWNE 1006
Query: 943 KFESFMHKEGFQKCNADHCCFFKRYK-SSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKE 1001
KF +M + GF++ + D C + K+ S + LL YVDDMLVA +N+ I LK +LS +
Sbjct: 1007 KFNHYMTEIGFKRSDYDSCAYTKKLSDDSTMYLLFYVDDMLVAANNMQAIDALKKELSIK 1066
Query: 1002 FDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLS 1061
F+MKDLG AKKILG++I D++ GVL LS Y+N+VL+ FNM ++K TPL +H ++
Sbjct: 1067 FEMKDLGAAKKILGIEIIIDREAGVLWLSQESYLNKVLKTFNMLESKPALTPLGAHLKMK 1126
Query: 1062 QEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVK 1121
+ E+E M +PY+SA+GS+MYAM+ TRPD+ + VGVVSRFMS P K HW VK
Sbjct: 1127 SATEEKLSTEEEYMNSVPYSSAVGSIMYAMIGTRPDLAYPVGVVSRFMSQPAKEHWLGVK 1186
Query: 1122 WILRYLRGTTE-KCLYFGKGEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRI 1180
W+LRY++GT + + Y + + GY DAD+A ++D RRS TG +FT+G ++SW S +
Sbjct: 1187 WVLRYIKGTVDTRLCYKRNSDFSICGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGL 1246
Query: 1181 QKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHS 1240
Q++VA S+TE EY+++TEA KE IWL+GLL + G+ Q+ ++ DSQSAI L+KN+ H
Sbjct: 1247 QRVVAQSSTECEYMSLTEAVKEAIWLKGLLKDFGYEQKNVEIFCDSQSAIALSKNNVHHE 1306
Query: 1241 RTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
RTKHI +++HFIR ++ D + + KI KNPAD+ TKV+ ++K +
Sbjct: 1307 RTKHIDVKFHFIREIIADGKVEVSKISTEKNPADIFTKVLPVNKFQ 1352
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 953 bits (2464), Expect = 0.0
Identities = 544/1361 (39%), Positives = 792/1361 (57%), Gaps = 89/1361 (6%)
Query: 6 VKIEKFDG-ADFGFWKMQIEDYLYQKKLHQPLTEKKP---------DSMKDDEWSLLDRQ 55
+++EKFDG D+ WK ++ ++ L+ L E + +S +D E L +
Sbjct: 6 IEVEKFDGRGDYTMWKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEKFE 65
Query: 56 AL--------GVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTL 107
AL + LS++ V I KE T A ++ AL +Y N+++ ++L++
Sbjct: 66 ALEEKKKKARSAIVLSVTDRVLRKIKKESTAAAMLLALDKLYMSKALPNRIYPKQKLYSF 125
Query: 108 RMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSK 167
+M+E +SV +I+E + T L ++ + DE +A++LL++LP ++ + + SSG
Sbjct: 126 KMSENLSVEGNIDEFLQIITDLENMNVIISDEDQAILLLTALPKAFDQLKDTLKYSSGKS 185
Query: 168 KMKFDDVRDLVLSEEIRRRELGESSSS-----SVLHTESRGRNSTRGNGRGKSKARRSKS 222
+ D+V + S+E+ ELG S L+ + + N +G +GK K ++ KS
Sbjct: 186 ILTLDEVAAAIYSKEL---ELGSVKKSIKVQAEGLYVKDKNENKGKGEQKGKGKGKKGKS 242
Query: 223 KNHRSSHNSKSIECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGG------------ 270
K K CW CG+ GHF++ C K Q ++ + +SGG
Sbjct: 243 K--------KKPGCWTCGEEGHFRSSCPNQNKPQFKQSQVVKGESSGGKGNLAEAAGYYV 294
Query: 271 GDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVK 330
+AL + E+ W+LD+G S+H + ++E+F + G V +GN+ +V G G ++
Sbjct: 295 SEALSSTEVHLEDEWILDTGCSYHMTYKREWFHEFNEDAGGSVRMGNKTVSRVRGVGTIR 354
Query: 331 IK-LNGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSG 389
+K +G L NVR+IP++ +NL+S+G GY +I G + GR+
Sbjct: 355 VKNSDGLTIVLTNVRYIPDMDRNLLSLGTFEKAGYKFESEDGILRIKAGNQVLLTGRRYD 414
Query: 390 TLY----KTAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICE 445
TLY K + L V ++ LWH+RL HMS+K M+++ KG L + +D+CE
Sbjct: 415 TLYLLNWKPVASESLAVVKRADDTVLWHQRLCHMSQKNMEILVRKGFLDKKKVSSLDVCE 474
Query: 446 DCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVP-SIGGKHYFVTFIDDHSRKVWV 504
DCI GK KR SF + K EKLE +HSD+WG VP S+G YF++ IDD +RKVWV
Sbjct: 475 DCIYGKAKRKSFSLAHHDTK-EKLEYIHSDLWGAPFVPLSLGKCQYFMSIIDDFTRKVWV 533
Query: 505 YFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVP 564
YF+K K E FE F W +VEN+TD ++K LRTDNG E+ + F FC GI RT
Sbjct: 534 YFMKTKDEAFEKFVEWVNLVENQTDRRVKTLRTDNGLEFCNKLFDGFCESIGIHRHRTCA 593
Query: 565 GTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEE 624
TPQ NGVAERMNRT+ E+ RS+ SGLPK+FWAEA +T+ LIN+ PS L +IP++
Sbjct: 594 YTPQQNGVAERMNRTIMEKVRSMLSDSGLPKRFWAEATHTTVLLINKTPSSALNFEIPDK 653
Query: 625 VWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKM 684
WSG S+LR +GCVA+VH D KL+P++KK + IGY GY++W + +K
Sbjct: 654 KWSGNPPVYSYLRRYGCVAFVHTDD---GKLEPRAKKGVLIGYPVGVKGYKVWILDERKC 710
Query: 685 VRSKDVIFNERVMYKD----KHNTTTND-----SGLSEPVYVEMDDVPGS--------PT 727
V S+++IF E +YKD + N +T + S L + E D + G P
Sbjct: 711 VVSRNIIFQENAVYKDLMQRQENVSTEEDDQTGSYLEFDLEAERDVISGGDQEMVNTIPA 770
Query: 728 DKSPQSGELAESSIRQPSDTLVHPTPVP---VLRRSSRP-HAPNR-----RYIDYMLLTD 778
+SP D+ V+ +P+ V R R AP R Y + + T+
Sbjct: 771 PESPVVSTPTTQDTNDDEDSDVNQSPLSYHLVRDRDKREIRAPRRFDDEDYYAEALYTTE 830
Query: 779 GG---EPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVK 835
G EPE+Y +A + KW+LAM EE+ S N TW + P ++ + +W+++ K
Sbjct: 831 DGEAVEPENYRKAKLDANFDKWKLAMDEEIDSQEKNNTWTIVTRPENQRIIGCRWIFKYK 890
Query: 836 EDHDG--SKRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDV 893
G R+KARLV KG+ QKEGIDY EIFAPVVK +IR +LSIVA E+L LEQLDV
Sbjct: 891 LGILGVEEPRFKARLVAKGYAQKEGIDYHEIFAPVVKHVSIRVLLSIVAQEDLELEQLDV 950
Query: 894 KTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGF 953
KTAFLHG+L E+IYM PEG+ K N VC+L K+LYGLKQAP+QW KF++FM + F
Sbjct: 951 KTAFLHGELKEKIYMSPPEGYESMFKANEVCLLNKALYGLKQAPKQWNEKFDNFMKEICF 1010
Query: 954 QKCNADHCCFFKRY-KSSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKK 1012
K D C + K S + LL+YVDD+LVA N + I LK L F+MKDLG AKK
Sbjct: 1011 VKSAYDSCAYTKVLPDGSVMYLLIYVDDILVASKNKEAITALKANLGMRFEMKDLGAAKK 1070
Query: 1013 ILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEK 1072
ILGM+I RD+ GVL LS Y+N++L+ +NM +AK TPL +HF+ + ++
Sbjct: 1071 ILGMEIIRDRTLGVLWLSQEGYLNKILETYNMAEAKPAMTPLGAHFKFQAATEQKLIRDE 1130
Query: 1073 ELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTE 1132
+ M +PY+SA+GS+MYAM+ TRPD+ + VG++SRFMS P K HW VKW+LRY++GT +
Sbjct: 1131 DFMKSVPYSSAVGSIMYAMLGTRPDLAYPVGIISRFMSQPIKEHWLGVKWVLRYIKGTLK 1190
Query: 1133 KCLYFGK-GEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEV 1191
L + K + GY DAD+A ++D RRS TG +FT+G ++SW S +Q++VA STTE
Sbjct: 1191 TRLCYKKSSSFSIVGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGLQRVVAQSTTES 1250
Query: 1192 EYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHF 1251
EY+++TEA KE IWL+GLL + G+ Q+ ++ DSQSAI L+KN+ H RTKHI ++YHF
Sbjct: 1251 EYMSLTEAVKEAIWLKGLLKDFGYEQKSVEIFCDSQSAIALSKNNVHHERTKHIDVKYHF 1310
Query: 1252 IRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLKLCSTLV 1292
IR ++ D + ++KI KNPAD+ TKV+ + K + L+
Sbjct: 1311 IREIISDGTVEVLKISTEKNPADIFTKVLAVSKFQAALNLL 1351
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 949 bits (2453), Expect = 0.0
Identities = 545/1352 (40%), Positives = 799/1352 (58%), Gaps = 94/1352 (6%)
Query: 6 VKIEKFDG-ADFGFWKMQI---------------EDYLYQKKLHQPLTEK--KPDSMKDD 47
+++EKFDG D+ WK ++ ED L +K LTE+ K + ++ +
Sbjct: 6 IEVEKFDGRGDYTMWKEKLMAHLDILGLSVALKEEDDLVEKVAEMQLTEEEEKEEVLRRE 65
Query: 48 EWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTL 107
R+A + LS++ V I KE++ A ++ L +Y N+++ ++L++
Sbjct: 66 LLEEKRRKARSAIVLSVTDRVLRKIKKEQSAAAMLGVLDKLYMSKALPNRIYQKQKLYSF 125
Query: 108 RMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSK 167
+M+E +S+ +I+E + L + + DE +A++LL SLP + + + G
Sbjct: 126 KMSENLSIEGNIDEFLRIIADLENTNVLVSDEDQAILLLMSLPKPFDQLRDTLKYGLGRV 185
Query: 168 KMKFDDVRDLVLSEEIRRRELGESSSSSVLH---------TESRGRNSTRGNGRGKSKAR 218
+ D+V + S+E+ ELG + S TE+RGR RGN K+R
Sbjct: 186 TLSLDEVVAAIYSKEL---ELGSNKKSIKGQAEGLFVKEKTETRGRTEQRGNNNNNKKSR 242
Query: 219 RSKSKNHRSSHNSKSIECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICSL 278
SKS++ + CW CG++ + EAN S +AL +
Sbjct: 243 -SKSRSKKG--------CWICGESSN----------GSSNYSEANGLYVS---EALSSTD 280
Query: 279 ESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKLN-GSV 337
E+ WV+D+G S+H + ++E+F++ G V +GN+ KV G G +++K G V
Sbjct: 281 IHLEDEWVMDTGCSYHMTYKREWFEDLNEDAGGSVRMGNKTVSKVRGIGTIRVKNEAGMV 340
Query: 338 WELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLY----K 393
L NVR+IP + +NL+S+G GY+ I G + R+ TLY +
Sbjct: 341 VRLTNVRYIPEMDRNLLSLGTFEKSGYSFKLENGTLSIIAGDSVLLTVRRCYTLYLLQWR 400
Query: 394 TAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICEDCILGKQK 453
L V ++ LWH+RLGHMS+K M ++ KG L + +++ CEDCI GK K
Sbjct: 401 PVTEESLSVVKRQDDTILWHRRLGHMSQKNMDLLLKKGLLDKKKVSKLETCEDCIYGKAK 460
Query: 454 RVSFQTSGRTPKKEKLELVHSDVWGPTTVP-SIGGKHYFVTFIDDHSRKVWVYFLKHKSE 512
R+ F + + +EKLE VHSD+WG +VP S+G YF++FIDD++RKV +YFLK K E
Sbjct: 461 RIGFNLA-QHDTREKLEYVHSDLWGAPSVPFSLGKCQYFISFIDDYTRKVRIYFLKTKDE 519
Query: 513 VFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGV 572
F+ F W +VEN+TD +IK LRTDNG E+ + F +FC + GI RT TPQ NGV
Sbjct: 520 AFDKFVEWANLVENQTDKRIKTLRTDNGLEFCNRSFDEFCSQKGILWHRTCAYTPQQNGV 579
Query: 573 AERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVK 632
AERMNRTL E+ RS+ SGLPKKFWAEA +T+A LIN+ PS L +++P++ WSGK
Sbjct: 580 AERMNRTLMEKVRSMLSDSGLPKKFWAEATHTTAILINKTPSSALNYEVPDKRWSGKSPI 639
Query: 633 LSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIF 692
S+LR FGC+A+VH D KL+P++KK I +GY GY++W E KK V S++VIF
Sbjct: 640 YSYLRRFGCIAFVHTDD---GKLNPRAKKGILVGYPIGVKGYKIWLLEEKKCVVSRNVIF 696
Query: 693 NERVMYKDKHNTTTNDSGLSE---PVYVEMD-----------DVP----GSPTDKSPQSG 734
E YKD + + +E Y+++D D P SP + SP +
Sbjct: 697 QENASYKDMMQSKDAEKDENEAPPSSYLDLDLDHEEVITSGGDDPIVEAQSPFNPSPATT 756
Query: 735 ELAESSIRQPSDTLVHPTPVPVLRRSSRP--HAPNR----RYIDYMLLT--DGG--EPED 784
+ + +D + P ++R R AP R Y+ L T D G EP D
Sbjct: 757 QTYSEGVNSETDIIQSPLSYQLVRDRDRRTIRAPVRFDDEDYLAEALYTTEDSGEIEPAD 816
Query: 785 YDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGSK-- 842
Y EA ++ + +KW+LAM EEM+S I N TW + K P +K + ++W+Y+ K G +
Sbjct: 817 YSEAKRSMNWNKWKLAMNEEMESQIKNHTWTVVKRPQHQKVIGSRWIYKFKLGIPGVEEG 876
Query: 843 RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDL 902
R+KARLV KG+ Q++GIDY EIFAPVVK +IR ++SIVA E+L LEQLDVKTAFLHG+L
Sbjct: 877 RFKARLVAKGYAQRKGIDYHEIFAPVVKHVSIRILMSIVAQEDLELEQLDVKTAFLHGEL 936
Query: 903 VEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCC 962
E+IYM PEG+ E KE+ VC+L KSLYGLKQAP+QW KF ++M + GF + D C
Sbjct: 937 KEKIYMVPPEGYEEMFKEDEVCLLNKSLYGLKQAPKQWNEKFNAYMSEIGFIRSLYDSCA 996
Query: 963 FFKRYK-SSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRD 1021
+ K S + LLLYVDDMLVA N ++I LK +LS+ FDMKDLG AK+ILGM+I R+
Sbjct: 997 YIKELSDGSRVYLLLYVDDMLVAAKNKEDISQLKEELSQRFDMKDLGAAKRILGMEIIRN 1056
Query: 1022 KQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYA 1081
+++ L LS Y+N++L+ +NM ++K V TPL +H ++ + E++++ M IPY+
Sbjct: 1057 REENTLWLSQNGYLNKILETYNMAESKHVVTPLGAHLKMRAATVEKQEQDEDYMKSIPYS 1116
Query: 1082 SAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTT-EKCLYFGKG 1140
SA+GS+MYAM+ TRPD+ + VG++SR+MS P + HW VKW+LRY++G+ K Y
Sbjct: 1117 SAVGSIMYAMIGTRPDLAYPVGIISRYMSQPAREHWLGVKWVLRYIKGSLGTKLQYKRSS 1176
Query: 1141 EIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEAS 1200
+ KV GY DAD A D RRS TG +FT+G ++SW S Q++VALSTTE EY+++TEA
Sbjct: 1177 DFKVVGYCDADHAACKDRRRSITGLVFTLGGSTISWKSGQQRVVALSTTEAEYMSLTEAV 1236
Query: 1201 KELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEV 1260
KE +W++GLL E G+ Q+ ++ DSQSAI L+KN+ H RTKHI +RY +IR ++ +
Sbjct: 1237 KEAVWMKGLLKEFGYEQKSVEIFCDSQSAIALSKNNVHHERTKHIDVRYQYIRDIIANGD 1296
Query: 1261 LTLIKIQGSKNPADMLTKVVTIDKLKLCSTLV 1292
++KI KNPAD+ TK+V ++K + TL+
Sbjct: 1297 GDVVKIDTEKNPADIFTKIVPVNKFQAALTLL 1328
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 948 bits (2451), Expect = 0.0
Identities = 547/1362 (40%), Positives = 810/1362 (59%), Gaps = 113/1362 (8%)
Query: 4 GEVKIEKFDG-ADFGFWKMQIEDYLYQKKLHQPLTEKKPDSMKDDEWSLLDR-------- 54
G ++EKFDG D+ WK ++ ++ L + L E++ ++D + D
Sbjct: 4 GRAEVEKFDGDGDYILWKEKLLAHMEMLGLLEGLGEEEEAVVEDSTTEISDGGNQDPETA 63
Query: 55 --------------QALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHL 100
+A + LSL NV + K+KT AG++K L ++ N+++L
Sbjct: 64 TSKLEDKILKEKRGKARSTIILSLGNNVLRKVIKQKTAAGMIKVLDQLFMAKSLPNRIYL 123
Query: 101 MRRLFTLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAV 160
+RL+ +M+E M++ +++N+ + + L +V + DE +A++LL SLP + + +
Sbjct: 124 KQRLYGYKMSENMTMEENVNDFFKLISDLENVKVVVPDEDQAIVLLMSLPRQFDQLKETL 183
Query: 161 SSSSGSKKMKFDDVRDLVLSEEIRRRELGES-----SSSSVLHTESRGRNSTRGNGRGKS 215
+ + +++ + S+ + ELG S ++S L + RGR+ TRG G K+
Sbjct: 184 KYCKTT--LHLEEITSAIRSKIL---ELGASGKLLKNNSDGLFVQDRGRSETRGKGPNKN 238
Query: 216 KARRSKSKNHRSSHNSKSIECWNCGKTGHFKNQC-----RLPTKNQEEKDEANV------ 264
K+R SKSK + CW CGK GHFK QC R + E+ EA+
Sbjct: 239 KSR-SKSKGAGKT-------CWICGKEGHFKKQCYVWKERNKQGSTSERGEASTVTARVT 290
Query: 265 -ASTSGGGDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKV 323
A+ AL+ E ++W+LD+G SFH + +K++ ++ GKV +GN+ +V
Sbjct: 291 DAAALVVSRALLGFAEVTPDTWILDTGCSFHMTCRKDWIIDFKETASGKVRMGNDTYSEV 350
Query: 324 VGKGEVKIKL-NGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTI 382
G G+V+IK +GS L +VR+IP ++KNLIS+G L D+G I K +T+
Sbjct: 351 KGIGDVRIKNEDGSTILLTDVRYIPEMSKNLISLGTLEDKGCWFESKKGILTIFKNDLTV 410
Query: 383 ARGRKSGTLY-----KTAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLR 437
G+K TLY AG ++I +E +LWH RLGH+ KG++V+ SKG L +
Sbjct: 411 LTGKKESTLYFLQGTTLAGEANVIDKEKDET-SLWHSRLGHIGAKGLQVLVSKGHLD--K 467
Query: 438 SIEIDICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVP-SIGGKHYFVTFID 496
+I I G K V+ K+KL+ VHSD+WG T VP SIG YF+TFID
Sbjct: 468 NIMISF------GAAKHVT---------KDKLDYVHSDLWGSTNVPFSIGKCQYFITFID 512
Query: 497 DHSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHG 556
D +R+ W+YF++ K E F F WK +EN+ D K+K L TDNG E+ + +F FC + G
Sbjct: 513 DFTRRTWIYFIRTKDEAFSKFVEWKTQIENQQDKKLKILITDNGLEFCNQEFDSFCRKEG 572
Query: 557 IRMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVP 616
+ RT TPQ NGVAERMNRT+ + R + +SGL K+FWAEA +T+ +LIN+ PS
Sbjct: 573 VIRHRTCAYTPQQNGVAERMNRTIMNKVRCMLSESGLGKQFWAEAASTAVFLINKSPSSS 632
Query: 617 LEHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRL 676
+E IPEE W+G L+ FG VAY+H SDQG KL+P++KK IF+GY + +++
Sbjct: 633 IEFDIPEEKWTGHPPDYKILKKFGSVAYIH-SDQG--KLNPRAKKGIFLGYPDGVKRFKV 689
Query: 677 WDDENKKMVRSKDVIFNERVMYKD--KHNTTTNDSGLSEP--VYVEMDDVPGSPTDKSP- 731
W E++K V S+D++F E MYK+ K++ + D L+E +E+ ++ ++S
Sbjct: 690 WLLEDRKCVVSRDIVFQENQMYKELQKNDMSEEDKQLTEVERTLIELKNLSADDENQSEG 749
Query: 732 ---QSGELAESSIRQPSDTLVHPTP---------VPVLRRSSRPHAPNRRYIDY------ 773
+ E A ++ D V T + R R +R+++
Sbjct: 750 GDNSNQEQASTTRSASKDKQVEETDSDDDCLENYLLARDRIRRQIRAPQRFVEEDDSLVG 809
Query: 774 --MLLTDGGE---PEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHN 828
+ +T+ GE PE Y+EA ++ + KW+ A EEM S+ N TW++ P GK+ +
Sbjct: 810 FALTMTEDGEVYEPETYEEAMRSPECEKWKQATIEEMDSMKKNDTWDVIDKPEGKRVIGC 869
Query: 829 KWVYRVKEDHDGSK--RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENL 886
KW+++ K G + RYKARLV KGF Q+EGIDY EIF+PVVK +IR +LSIV ++
Sbjct: 870 KWIFKRKAGIPGVEPPRYKARLVAKGFSQREGIDYQEIFSPVVKHVSIRYLLSIVVQFDM 929
Query: 887 YLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFES 946
LEQLDVKTAFLHG+L E I M QPEG+ +E VC+LKKSLYGLKQ+PRQW +F+S
Sbjct: 930 ELEQLDVKTAFLHGNLDEYILMSQPEGYEDEDSTEKVCLLKKSLYGLKQSPRQWNQRFDS 989
Query: 947 FMHKEGFQKCNADHCCFFKRYKS-SYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMK 1005
FM G+Q+ + C + ++ SYI LLLYVDDML+A N D+I+ LK L++EF+MK
Sbjct: 990 FMINSGYQRSKYNPCVYTQQLNDGSYIYLLLYVDDMLIASQNKDQIQKLKESLNREFEMK 1049
Query: 1006 DLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQS 1065
DLGPA+KILGM+ITR++++G+L LS +EY+ VL+ F M +K+ TPL +HF+L
Sbjct: 1050 DLGPARKILGMEITRNREQGILDLSQSEYVAGVLRAFGMDQSKVSQTPLGAHFKLRAANE 1109
Query: 1066 PQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILR 1125
+ E M +PY +AIGS+MY+M+ +RPD+ + VGVVSRFMS P K HW+AVKW++R
Sbjct: 1110 KTLARDAEYMKLVPYPNAIGSIMYSMIGSRPDLAYPVGVVSRFMSKPSKEHWQAVKWVMR 1169
Query: 1126 YLRGTTEKCLYFGKGE-IKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIV 1184
Y++GT + CL F K + ++ GY D+D+A ++D RRS TG++FT G ++SW S +Q++V
Sbjct: 1170 YMKGTQDTCLRFKKDDKFEIRGYCDSDYATDLDRRRSITGFVFTAGGNTISWKSGLQRVV 1229
Query: 1185 ALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKH 1244
ALSTTE EY+A+ EA KE IWL+GL E+GF Q+ + DSQSAI L+KNS H RTKH
Sbjct: 1230 ALSTTEAEYMALAEAVKEAIWLRGLAAEMGFEQDAVEVMCDSQSAIALSKNSVHHERTKH 1289
Query: 1245 IGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
I +RYHFIR + D + ++KI + NPAD+ TK V + KL+
Sbjct: 1290 IDVRYHFIREKIADGEIQVVKISTTWNPADIFTKTVPVSKLQ 1331
>At1g37110
Length = 1356
Score = 940 bits (2429), Expect = 0.0
Identities = 534/1372 (38%), Positives = 808/1372 (57%), Gaps = 106/1372 (7%)
Query: 6 VKIEKFDG-ADFGFWKMQIEDYLYQKKLHQPLTE-------------------------- 38
V+I+ F+G DF WK++I+ L L LT+
Sbjct: 8 VEIKVFNGDRDFSLWKIRIQAQLGVLGLKDTLTDFSLTKTVPLTKSEAKQESGDGESSGT 67
Query: 39 -KKPDSMKDDEWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNK 97
+ PD +K ++ QA ++ +S V + TTA L L+ Y + N+
Sbjct: 68 KEVPDPVKIEQ----SEQAKNIIINHISDVVLLKVNHYATTADLWATLNKKYMETSLPNR 123
Query: 98 VHLMRRLFTLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIV 157
++ +L++ +M M++ Q+++E + +L S+ I+ D+EV+A+++L+SLP S +
Sbjct: 124 IYTQLKLYSFKMVSTMTIDQNVDEFLRIVAELGSLEIQVDEEVQAILILNSLPASHIQLK 183
Query: 158 TAVSSSSGSKKMKFDDVRDLVLSEEIRRRELGES-----SSSSVLHTESRGRNSTRGNGR 212
+ G+K + DV S E REL E+ ++VL+T RGR R N +
Sbjct: 184 HTLKY--GNKTLTVQDVTSSAKSLE---RELAEAVDLDKGQAAVLYTTERGRPLVRNNQK 238
Query: 213 GKSKARRSKSKNHRSSHNSKS-IECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGG- 270
G RS+S NSK+ + CW C K GH K C K E + + +
Sbjct: 239 GGQGKGRSRS-------NSKTKVPCWYCKKEGHVKKDCYSRKKKMESEGQGEAGVITEKL 291
Query: 271 --GDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGE 328
+AL + + ++ W+LDSG + H +S++++F ++ + LG++ S + G+G
Sbjct: 292 VFSEALSVNEQMVKDLWILDSGCTSHMTSRRDWFISFQEKGNTTILLGDDHSVESQGQGT 351
Query: 329 VKIKLNGSVWE-LKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKIS--KGAMTIARG 385
++I +G + L+NV+++P+L +NLIS G L GY G + K+ K T RG
Sbjct: 352 IRIDTHGGTIKILENVKYVPHLRRNLISTGTLDKLGYR--HEGGEGKVRYFKNNKTALRG 409
Query: 386 RKSGTLYKTAGACHLIAVATNENPN----LWHKRLGHMSEKGMKVMHSKGKLPSLRSIEI 441
S LY G+ + + E LWH RLGHMS +KV+ KG + E+
Sbjct: 410 SLSNGLYVLDGSTVMSELCNAETDKVKTALWHSRLGHMSMNNLKVLAGKGLIDRKEINEL 469
Query: 442 DICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWG-PTTVPSIGGKHYFVTFIDDHSR 500
+ CE C++GK K+VSF G+ ++ L VH+D+WG P PSI GK YF++ IDD +R
Sbjct: 470 EFCEHCVMGKSKKVSFNV-GKHTSEDALSYVHADLWGSPNVTPSISGKQYFLSIIDDKTR 528
Query: 501 KVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRME 560
KVW+YFLK K E F+ F WK++VEN+ + K+K LRTDNG E+ +++F +C EHGI
Sbjct: 529 KVWLYFLKSKDETFDKFCEWKSLVENQVNKKVKCLRTDNGLEFCNSRFDSYCKEHGIERH 588
Query: 561 RTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHK 620
RT TPQ NGVAERMNRT+ E+ R L +SG+ + FWAEA T+AYLINR P+ + H
Sbjct: 589 RTCTYTPQQNGVAERMNRTIMEKVRCLLNKSGVEEVFWAEAAATAAYLINRSPASAINHN 648
Query: 621 IPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDE 680
+PEE+W ++ HLR FG +AYVH DQG KL P++ K F+GY GY++W E
Sbjct: 649 VPEEMWLNRKPGYKHLRKFGSIAYVH-QDQG--KLKPRALKGFFLGYPAGTKGYKVWLLE 705
Query: 681 NKKMVRSKDVIFNERVMYKD------------KHNTTTND---------SGLSEPVYVEM 719
+K V S++V+F E V+Y+D + TT+++ SG + ++
Sbjct: 706 EEKCVISRNVVFQESVVYRDLKVKEDDTDNLNQKETTSSEVEQNKFAEASGSGGVIQLQS 765
Query: 720 DDVPGSPTDKSPQSGELAESSIR-QPSDTLVHPTPVPVLR-RSSRPHAPNRRYIDYMLLT 777
D P + ++S S E E S + Q + T + R R R P R+ + +T
Sbjct: 766 DSEPITEGEQSSDSEEEVEYSEKTQETPKRTGLTTYKLARDRVRRNINPPTRFTEESSVT 825
Query: 778 DG---------GEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHN 828
EP+ Y EA ++ D KW++A +EM SL+ N TW+L P +K +
Sbjct: 826 FALVVVENCIVQEPQSYQEAMESQDCEKWDMATHDEMDSLMKNGTWDLVDKPKDRKIIGC 885
Query: 829 KWVYRVKEDHDGSK--RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENL 886
+W++++K G + R+KARLV KG+ Q+EG+DY EIFAPVVK +IR ++S+V ++L
Sbjct: 886 RWLFKLKSGIPGVEPTRFKARLVAKGYTQREGVDYQEIFAPVVKHVSIRILMSLVVDKDL 945
Query: 887 YLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFES 946
LEQ+DVKT FLHGDL EE+YM QPEGF+ + EN VC LKKSLYGLKQ+PRQW +F+
Sbjct: 946 ELEQMDVKTTFLHGDLEEELYMEQPEGFVSDSSENKVCRLKKSLYGLKQSPRQWNKRFDR 1005
Query: 947 FMHKEGFQKCNADHCCFFKRY-KSSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMK 1005
FM + F + D C + K + +I LLLYVDDML+AG++ EI +K QLS EF+MK
Sbjct: 1006 FMSSQQFIRSEHDACVYVKHVSEHDFIYLLLYVDDMLIAGASKAEINRVKEQLSTEFEMK 1065
Query: 1006 DLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQS 1065
D+G A +ILG+ I RD++ GVL+LS YI +VL RFNM AK+ + P+ +HF+L+ +
Sbjct: 1066 DMGGASRILGIDIYRDRKGGVLKLSQEIYIRKVLDRFNMSGAKMTNAPVGAHFKLA---A 1122
Query: 1066 PQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILR 1125
+ E+E +PY+SA+GS+MYAM+ TRPD+ +A+ ++SR+MS PG HWEAVKW++R
Sbjct: 1123 VREEDECVDTDVVPYSSAVGSIMYAMLGTRPDLAYAICLISRYMSKPGSMHWEAVKWVMR 1182
Query: 1126 YLRGTTEKCLYFGK-GEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIV 1184
YL+G + L F K + V GY D+++A ++D RRS +GY+FT+G +VSW + +Q +V
Sbjct: 1183 YLKGAQDLNLVFTKEKDFTVTGYCDSNYAADLDRRRSISGYVFTIGGNTVSWKASLQPVV 1242
Query: 1185 ALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKH 1244
A+STTE EY+A+ EA+KE +W++GLL ++G Q+K ++ DSQSAI L+KNS +H RTKH
Sbjct: 1243 AMSTTEAEYIALAEAAKEAMWIKGLLQDMGMQQDKVKIWCDSQSAICLSKNSVYHERTKH 1302
Query: 1245 IGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLKLCSTLVGLLE 1296
I +R+++IR ++E + ++KI S+NP D LTK + ++K K ++ L++
Sbjct: 1303 IDVRFNYIRDVVESGDVDVLKIHTSRNPVDALTKCIPVNKFKSALGVLKLMK 1354
>At4g21360 putative transposable element
Length = 1308
Score = 868 bits (2242), Expect = 0.0
Identities = 508/1354 (37%), Positives = 769/1354 (56%), Gaps = 133/1354 (9%)
Query: 5 EVKIEKFDG-ADFGFWKMQIE-------------DYLYQKKLHQPLTEKKPDSMKDDEWS 50
+V+I+ F+G DF WK++IE D+ K + +EKK +DDE
Sbjct: 6 KVEIKTFNGDRDFSLWKIRIEAQLGVLGLKPALSDFTLTKTILVVKSEKKESESEDDETD 65
Query: 51 LLDR-------------QALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNK 97
QA + ++ V + T A L L+ ++ + N+
Sbjct: 66 SKKTEEVPDPIKFEQSDQAKNFIINHITDTVLLKVQHCVTAAELWATLNKLFMETSLPNR 125
Query: 98 VHLMRRLFTLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIV 157
++ RL++ +M + +S+ Q+ +E + +L S+ I+ +EV+A+++L+SLP S+ +
Sbjct: 126 IYTQLRLYSFKMVDNLSIDQNTDEFLRIVAELGSLQIQVGEEVQAILILNSLPPSYIQLK 185
Query: 158 TAVSSSSGSKKMKFDDVRDLVLSEEIRRRELGESS------SSSVLHTESRGRNSTRGN- 210
+ G+K + V+D+V S + REL E +S+ L+T RGR T+
Sbjct: 186 HTLKY--GNKTLS---VQDVVSSAKSLERELSEQKETIRAPASTALYTAERGRPQTKNTQ 240
Query: 211 GRGKSKARRSKSKNHRSSHNSKS-IECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSG 269
G+GK + R NSKS + CW C K GH K C + E + + +
Sbjct: 241 GQGKGRGRS----------NSKSRLTCWFCKKEGHVKKDCYAGKRKLENEGQGKAGVITE 290
Query: 270 G---GDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGK 326
+AL + ++ WV+DSG ++H +S+ ++F + + LG++ + + G
Sbjct: 291 KLVYSEALSMYDQEAKDKWVIDSGCTYHMTSRMDWFSEFNENETTMILLGDDHTVESKGS 350
Query: 327 GEVKIKLNG-SVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKIS--KGAMTIA 383
G VK+ +G S+ LKNVR +PNL +NLIS G L GY G D K+ K T
Sbjct: 351 GTVKVNTHGGSIRVLKNVRFVPNLRRNLISTGTLDKLGYK--HEGGDGKVRFYKENKTAL 408
Query: 384 RGRKSGTLYKTAGACHLIA------VATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLR 437
G LY G H + +NE LWH RLGHMS MK++ KG L
Sbjct: 409 CGNLVNGLYVLDG--HTVVNENCNVEGSNEKTELWHCRLGHMSLNNMKILAEKGLLEKKD 466
Query: 438 SIEIDICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDD 497
E+ CE+C++GK K++SF G+ E L +H+D+WG K YF++ IDD
Sbjct: 467 IKELSFCENCVMGKSKKLSFNV-GKHITDEVLGYIHADLWG---------KQYFLSIIDD 516
Query: 498 HSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGI 557
SRKVW+ FLK K E FE F WK +VEN+ + K+K LRTDNG E+ + KF +FC ++GI
Sbjct: 517 KSRKVWLMFLKTKDETFERFCEWKELVENQVNKKVKILRTDNGLEFCNLKFDEFCKQNGI 576
Query: 558 RMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPL 617
RT TPQ NGVA+RMNRTL E+ R L +SGL + FWAEA T+AYL+NR P+ +
Sbjct: 577 ERHRTCTYTPQQNGVAKRMNRTLMEKVRCLLNESGLEEVFWAEAAATAAYLVNRSPASAV 636
Query: 618 EHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLW 677
+H +PEE+W K+ HLR FGC+AYVH+ DQG KL P++ K +F+GY + GY++W
Sbjct: 637 DHNVPEELWLDKKPGYKHLRRFGCIAYVHL-DQG--KLKPRALKGVFLGYPQGTKGYKVW 693
Query: 678 DDENKKMVRSKDVIFNERVMYKD-KHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGEL 736
+ +K V S++++FNE +YKD + ++ + +S+ V + ++G +
Sbjct: 694 LLDEEKCVISRNIVFNENQVYKDIRESSEQSVKDISDLEGYNEFQVSVKEHGECSKTGGV 753
Query: 737 AESSIRQPSDTLVHPTPVPVLR------------RSSRPHAPNRRYIDYMLLT------- 777
I Q SD+ T P++ R R P ++ DY
Sbjct: 754 TIEEIDQESDSENSVTQEPLIASIDLSNYQSARDRERRAPNPPQKLADYTHFALALVMAE 813
Query: 778 --DGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVK 835
+ EP+ Y +A + KW MKEE+ SL+ N TW++ + P +K + +W++++K
Sbjct: 814 EIESEEPQCYHDAKKDKHWIKWNGGMKEEIDSLLKNGTWDIVEWPKEQKVISCRWLFKLK 873
Query: 836 EDHDG--SKRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDV 893
G ++RYKARLV +GF Q++GIDY E+FAPVVK +IR ++S V +++ LEQ+DV
Sbjct: 874 PGIPGVEAQRYKARLVARGFTQQKGIDYEEVFAPVVKHISIRILMSAVVKDDMELEQMDV 933
Query: 894 KTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGF 953
KT FLHG+L + +YM QPEGF +++ VC+LKKSLYGLKQAPRQW KF +FM F
Sbjct: 934 KTTFLHGELDQVLYMEQPEGFEVNPEKDQVCLLKKSLYGLKQAPRQWNKKFHAFMLSLQF 993
Query: 954 QKCNADHCCFFKRYK-SSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKK 1012
+ D C + K ++ LLLYVDDML+A + EI LK LS +F+MKD+G A +
Sbjct: 994 ARSEHDSCVYVKEVNPGEFVYLLLYVDDMLLAAKSKSEISKLKEALSLKFEMKDMGAASR 1053
Query: 1013 ILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEK 1072
ILG+ I R++++G L+LS Y+++V+QRF M DAK+VSTP+ +HF+L+ +
Sbjct: 1054 ILGIDIIRNRKEGTLRLSQTRYVDKVIQRFRMADAKVVSTPMGAHFKLTSLIDEIGSVDP 1113
Query: 1073 ELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTE 1132
E+ +PY+SA+GS+MYAM+ T PD+ +A+G+VSRFMS PG
Sbjct: 1114 EV---VPYSSAVGSVMYAMIGTIPDVAYAMGLVSRFMSRPG------------------- 1151
Query: 1133 KCLYFGKGEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVE 1192
++V+GY D+D A ++D RRS +GY+FTVG +VSW S +Q +VALS+T+ E
Sbjct: 1152 -------ANLEVQGYCDSDHAADLDKRRSISGYVFTVGGNTVSWKSSLQHVVALSSTQAE 1204
Query: 1193 YVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFI 1252
++A+TEA KE IW++GLL ++G + + ++ DSQSAI L+KN+AFH RTKH+ ++++FI
Sbjct: 1205 FIALTEAVKEAIWIRGLLEDMGLQPKPATVWCDSQSAICLSKNNAFHDRTKHVEVKFYFI 1264
Query: 1253 RSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
R ++E + + KI S NPADMLTK + + K +
Sbjct: 1265 RDIIEAGEVKVRKIHTSVNPADMLTKCIPVKKFE 1298
>At1g58140 hypothetical protein
Length = 1320
Score = 795 bits (2054), Expect = 0.0
Identities = 479/1325 (36%), Positives = 732/1325 (55%), Gaps = 132/1325 (9%)
Query: 36 LTEKKPDSMKDDEWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSS 95
L++ + D ++D D++AL ++ L + + + + + L + Y+
Sbjct: 51 LSQTQKDGLRDSRKR--DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQV 108
Query: 96 NKVHL--MRRLF-TLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDS 152
KV L +R F L+M EG V+ + + + VT L G + DD +L SL
Sbjct: 109 KKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLK 168
Query: 153 WSAIVTAVSSSSG-----------------SKKMKFDDVRDLVLSEEIRRRELGES---- 191
+ IVT + + KK K +D+ + VL+ +I + E G+S
Sbjct: 169 FEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRR 228
Query: 192 SSSSVL------------------HTESRGRNSTRGNGRGKSKARRSKSKNHRSSHNSKS 233
V +T RG NS+RG G+G K+R KS S
Sbjct: 229 GGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKS----------S 278
Query: 234 IECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICSLESKEE-----SWVLD 288
++C+NCGK GH+ ++C+ P+ + E+ V D L+ + K+E W LD
Sbjct: 279 VKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMASYKKDEQEENHKWYLD 338
Query: 289 SGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKL-NGSVWELKNVRHIP 347
SGAS H +K F G V LG+E +V GKG + I+L NG + NV +IP
Sbjct: 339 SGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYIP 398
Query: 348 NLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLYKTA------------ 395
++ N++S+GQL ++GY D ++ ++I R ++S + K
Sbjct: 399 SMKTNILSLGQLLEKGY-------DIRLKDNNLSI-RDQESNLITKVPMSKNRMFVLNIR 450
Query: 396 -GACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIE--IDICEDCILGKQ 452
+ + E LWH R GH++ G++++ K + L I +CE C+LGKQ
Sbjct: 451 NDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQ 510
Query: 453 KRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSE 512
++SF + ++ LEL+H+DV GP S+G +YF+ FIDD SRK WVYFLK KSE
Sbjct: 511 FKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSE 570
Query: 513 VFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGV 572
VFE FK++KA VE E+ L IK +R+D GGE+ +F K+C ++GIR + TVP +PQ NGV
Sbjct: 571 VFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGV 630
Query: 573 AERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVK 632
AER NRT+ E ARS+ LPK+ WAEAV + YL+NR P+ + K P+E WSG++
Sbjct: 631 AERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPG 690
Query: 633 LSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIF 692
+SHLRVFG +A+ H+ D+ R+KLD KS+K IFIGY + GY+L++ + KK + S++++F
Sbjct: 691 VSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVF 750
Query: 693 NERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPSDTLVHPT 752
+E + + + +N+ + + E +D P PT + P S E PT
Sbjct: 751 DE----EGEWDWNSNEEDYNFFPHFE-EDKP-EPTREEPPSEE---------------PT 789
Query: 753 PVPVLRRSSRPHAPNRRYIDYMLLTDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQ 812
P SS+ + + EP D+ EA + W AM EE+KS+ N
Sbjct: 790 TPPTSPTSSQ-------------IEEKCEPMDFQEA---IEKKTWRNAMDEEIKSIQKND 833
Query: 813 TWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGIDYTEIFAPVVKL 871
TWEL LP G KA+ KWVY+ K++ G +RYKARLV KG+ Q+ GIDY E+FAPV +L
Sbjct: 834 TWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDEVFAPVARL 893
Query: 872 NTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLY 931
T+R ++S+ A + Q+DVK+AFL+GDL EE+Y+ QP+G++ +G+E+ V LKK+LY
Sbjct: 894 ETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKALY 953
Query: 932 GLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDDMLVAGSNIDEI 991
GLKQAPR W + + + ++ F KC +H + K K +I LYVDD++ G+N
Sbjct: 954 GLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIFTGNNPSMF 1013
Query: 992 KNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVS 1051
+ K +++KEF+M D+G LG+++ + + G+ ++ Y VL++F M D+ V
Sbjct: 1014 EEFKKEMTKEFEMTDIGLMSYYLGIEV-KQEDNGIF-ITQEGYAKEVLKKFKMDDSNPVC 1071
Query: 1052 TPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSN 1111
TP+ +LS+ +EE E + + S +GSL Y + CTRPDI +AVGVVSR+M +
Sbjct: 1072 TPMECGIKLSK------KEEGEGVDPTTFKSLVGSLRY-LTCTRPDILYAVGVVSRYMEH 1124
Query: 1112 PGKAHWEAVKWILRYLRGTTEKCLYFG-KGEIKVEGYVDADFAGEVDHRRSTTGYIFTVG 1170
P H++A K ILRY++GT L++ + K+ GY D+D+ G+VD R+ST+G++F +G
Sbjct: 1125 PTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIG 1184
Query: 1171 TRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEK-SALYSDSQSA 1229
+ +WMS+ Q IV LST E EYVA T IWL+ LL EL QE+ + ++ D++SA
Sbjct: 1185 DTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSA 1244
Query: 1230 IHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLKLCS 1289
I LAKN FH R+KHI RYH+IR + + + L ++ AD+ TK + +
Sbjct: 1245 IALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKREDFIKMR 1304
Query: 1290 TLVGL 1294
+L+G+
Sbjct: 1305 SLLGV 1309
>At1g48710 hypothetical protein
Length = 1352
Score = 795 bits (2053), Expect = 0.0
Identities = 480/1338 (35%), Positives = 734/1338 (53%), Gaps = 126/1338 (9%)
Query: 36 LTEKKPDSMKDDEWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSS 95
L++ + D ++D D++AL ++ L + + + + + L + Y+
Sbjct: 51 LSQTQKDGLRDSRKR--DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQV 108
Query: 96 NKVHL--MRRLF-TLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDS 152
KV L +R F L+M EG V+ + + + VT L G + DD +L SL
Sbjct: 109 KKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLK 168
Query: 153 WSAIVTAVSSSSG-----------------SKKMKFDDVRDLVLSEEIRRRELGES---- 191
+ IVT + + KK K +D+ + VL+ +I + E G+S
Sbjct: 169 FEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIIEQVLNMQITKEENGQSYQRR 228
Query: 192 SSSSVL------------------HTESRGRNSTRGNGRGKSKARRSKSKNHRSSHNSKS 233
V +T RG NS+RG G+G K+R KS S
Sbjct: 229 GGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKS----------S 278
Query: 234 IECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICSLESKEE-----SWVLD 288
++C+NCGK GH+ ++C+ P+ + E+ V D L+ + K+E W LD
Sbjct: 279 VKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMASYKKDEQEENHKWYLD 338
Query: 289 SGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKL-NGSVWELKNVRHIP 347
SGAS H +K F G V LG+E +V GKG + I+L NG + NV +IP
Sbjct: 339 SGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYIP 398
Query: 348 NLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLYKTA------------ 395
++ N++S+GQL ++GY D ++ ++I R ++S + K
Sbjct: 399 SMKTNILSLGQLLEKGY-------DIRLKDNNLSI-RDQESNLITKVPMSKNRMFVLNIR 450
Query: 396 -GACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIE--IDICEDCILGKQ 452
+ + E LWH R GH++ G++++ K + L I +CE C+LGKQ
Sbjct: 451 NDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQ 510
Query: 453 KRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSE 512
++SF + ++ LEL+H+DV GP S+G +YF+ FIDD SRK WVYFLK KSE
Sbjct: 511 FKMSFPKESSSRAQKSLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSE 570
Query: 513 VFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGV 572
VFE FK++KA VE E+ L IK +R+D GGE+ +F K+C ++GIR + TVP +PQ NGV
Sbjct: 571 VFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGV 630
Query: 573 AERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVK 632
AER NRT+ E ARS+ LPK+ WAEAV + YL+NR P+ + K P+E WSG++
Sbjct: 631 AERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKSG 690
Query: 633 LSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIF 692
+SHLRVFG +A+ H+ D+ R+KLD KS+K IFIGY + GY+L++ + KK + S++++F
Sbjct: 691 VSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVF 750
Query: 693 NERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPSDTLVHPT 752
+E + + + +N+ + + E D+ PT + P S E P+ PT
Sbjct: 751 DE----EGEWDWNSNEEDYNFFPHFEEDE--PEPTREEPPSEE--------PTTPPTSPT 796
Query: 753 PVPVLRRSSRPHAP-------------NRRYIDYMLLTDGGEPEDYDEACQTTDASKWEL 799
+ SS P N+ + L EP D+ EA + W
Sbjct: 797 SSQI-EESSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEA---IEKKTWRN 852
Query: 800 AMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEG 858
AM EE+KS+ N TWEL LP G K + KWVY+ K++ G +RYKARLV KG+ Q+ G
Sbjct: 853 AMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERYKARLVAKGYIQRAG 912
Query: 859 IDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEG 918
IDY E+FAPV +L T+R ++S+ A + Q+DVK+AFL+GDL EE+Y+ QP+G++ +G
Sbjct: 913 IDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKG 972
Query: 919 KENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYV 978
+E+ V LKK+LYGLKQAPR W + + + ++ F KC +H + K K +I LYV
Sbjct: 973 EEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYV 1032
Query: 979 DDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRV 1038
DD++ G+N + K +++KEF+M D+G LG+++ + + G+ ++ Y V
Sbjct: 1033 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEV-KQEDNGIF-ITQEGYAKEV 1090
Query: 1039 LQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDI 1098
L++F M D+ V TP+ +LS+ +EE E + + S +GSL Y + CTRPDI
Sbjct: 1091 LKKFKMDDSNPVCTPMECGIKLSK------KEEGEGVDPTTFKSLVGSLRY-LTCTRPDI 1143
Query: 1099 GHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFG-KGEIKVEGYVDADFAGEVD 1157
+AVGVVSR+M +P H++A K ILRY++GT L++ + K+ GY D+D+ G+VD
Sbjct: 1144 LYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVD 1203
Query: 1158 HRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQ 1217
R+ST+G++F +G + +WMS+ Q IV LST E EYVA T IWL+ LL EL Q
Sbjct: 1204 DRKSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQ 1263
Query: 1218 EK-SALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADML 1276
E+ + ++ D++SAI LAKN FH R+KHI RYH+IR + + + L ++ AD+
Sbjct: 1264 EEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIF 1323
Query: 1277 TKVVTIDKLKLCSTLVGL 1294
TK + + +L+G+
Sbjct: 1324 TKPLKREDFIKMRSLLGV 1341
>At3g61330 copia-type polyprotein
Length = 1352
Score = 791 bits (2043), Expect = 0.0
Identities = 478/1338 (35%), Positives = 732/1338 (53%), Gaps = 126/1338 (9%)
Query: 36 LTEKKPDSMKDDEWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSS 95
L++ + D ++D D++AL ++ L + + + + + L + Y+
Sbjct: 51 LSQTQKDGLRDSRKR--DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQV 108
Query: 96 NKVHL--MRRLF-TLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDS 152
KV L +R F L+M EG V+ + + + VT L G + DD +L SL
Sbjct: 109 KKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLK 168
Query: 153 WSAIVTAVSSSSG-----------------SKKMKFDDVRDLVLSEEIRRRELGES---- 191
+ IVT + + KK K +D+ + VL+ +I + E G+S
Sbjct: 169 FEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIAEQVLNMQITKEENGQSYQRR 228
Query: 192 SSSSVL------------------HTESRGRNSTRGNGRGKSKARRSKSKNHRSSHNSKS 233
V +T RG NS+RG G+G K+R KS S
Sbjct: 229 GGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKS----------S 278
Query: 234 IECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICSLESKEE-----SWVLD 288
++C+NCGK GH+ ++C+ P+ + E+ V D L+ + K+E W LD
Sbjct: 279 VKCYNCGKFGHYASECKAPSNKKFEEKAHYVEEKIQEEDMLLMASYKKDEQKENHKWYLD 338
Query: 289 SGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKL-NGSVWELKNVRHIP 347
SGAS H +K F G V LG+E +V GKG + I+L NG + NV +IP
Sbjct: 339 SGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYIP 398
Query: 348 NLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLYKTA------------ 395
++ N++S+GQL ++GY D ++ ++I R ++S + K
Sbjct: 399 SMKTNILSLGQLLEKGY-------DIRLKDNNLSI-RDQESNLITKVPMSKNRMFVLNIR 450
Query: 396 -GACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIE--IDICEDCILGKQ 452
+ + E LWH R GH++ G++++ K + L I +CE C+LGKQ
Sbjct: 451 NDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQ 510
Query: 453 KRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSE 512
++SF + ++ LEL+H+DV GP S+G +YF+ FIDD SRK WVYFLK KSE
Sbjct: 511 FKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSE 570
Query: 513 VFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGV 572
VFE FK++KA VE E+ L IK +R+D GGE+ +F K+C ++GIR + TVP +PQ NGV
Sbjct: 571 VFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGV 630
Query: 573 AERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVK 632
ER NRT+ E ARS+ LPK+ WAEAV + YL+NR P+ + K P+E WSG++
Sbjct: 631 VERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPG 690
Query: 633 LSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIF 692
+SHLRVFG +A+ H+ D+ R+KLD KS+K IFIGY + GY+L++ + KK + S++++F
Sbjct: 691 VSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVF 750
Query: 693 NERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPSDTLVHPT 752
+E + + + +N+ + + E D+ PT + P S E P+ PT
Sbjct: 751 DE----EGEWDWNSNEEDYNFFPHFEEDE--PEPTREEPPSEE--------PTTPPTSPT 796
Query: 753 PVPVLRRSSRPHAP-------------NRRYIDYMLLTDGGEPEDYDEACQTTDASKWEL 799
+ SS P N+ + L EP D+ +A + W
Sbjct: 797 SSQI-EESSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQKA---IEKKTWRN 852
Query: 800 AMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEG 858
AM EE+KS+ N TWEL LP G KA+ KWVY+ K++ G +RYKARLV KG+ Q+ G
Sbjct: 853 AMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRVG 912
Query: 859 IDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEG 918
IDY E+FAPV +L T+R ++S+ A + Q+DVK+AFL+GDL EE+Y+ QP+G++ +G
Sbjct: 913 IDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKG 972
Query: 919 KENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYV 978
+E+ V LKK LYGLKQAPR W + + + ++ F KC +H + K K +I LYV
Sbjct: 973 EEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYV 1032
Query: 979 DDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRV 1038
DD++ G+N + K +++KEF+M D+G LG+++ + + G+ ++ Y V
Sbjct: 1033 DDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEV-KQEDNGIF-ITQEGYAKEV 1090
Query: 1039 LQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDI 1098
L++F + D+ V TP+ +LS+ +EE E + + S +GSL Y + CTRPDI
Sbjct: 1091 LKKFKIDDSNPVCTPMECGIKLSK------KEEGEGVDPTTFKSLVGSLRY-LTCTRPDI 1143
Query: 1099 GHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFG-KGEIKVEGYVDADFAGEVD 1157
+AVGVVSR+M +P H++A K ILRY++GT L++ + K+ GY D+D+ G+VD
Sbjct: 1144 LYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVD 1203
Query: 1158 HRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQ 1217
R+ST+G++F +G + +WMS+ Q IV LST E EYVA T IWL+ LL EL Q
Sbjct: 1204 DRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQ 1263
Query: 1218 EK-SALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADML 1276
E+ + ++ D++SAI LAKN FH R+KHI RYH+IR + + + L ++ AD
Sbjct: 1264 EEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADFF 1323
Query: 1277 TKVVTIDKLKLCSTLVGL 1294
TK + + +L+G+
Sbjct: 1324 TKPLKRENFIKMRSLLGV 1341
>At3g60170 putative protein
Length = 1339
Score = 778 bits (2008), Expect = 0.0
Identities = 472/1361 (34%), Positives = 733/1361 (53%), Gaps = 112/1361 (8%)
Query: 8 IEKFDGADFGFWKMQIEDYLYQKKLHQ--------------PLTEKKPDSMKDDEWSLLD 53
I +FDG + FW M +E++L ++L + P++E + ++ +E L D
Sbjct: 12 IPRFDGY-YDFWSMTMENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAV--EEAKLKD 68
Query: 54 RQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHL--MRRLFTL-RMA 110
+ + ++ R + I + T+ + +++ Y+ + L +R+ F L M
Sbjct: 69 LKVKNFLFQAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMK 128
Query: 111 EGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSKKMK 170
EG + + V ++ + G + +L SL ++ +V ++ S+ +
Sbjct: 129 EGEKIDTFLGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTLS 188
Query: 171 FDDVRDLVLSEEIRRR-ELGESSSSSVLHTESRGRNSTRGNGRGKSKARRSKSKNH-RSS 228
D++ +L E R + E + V H E ++G GRG + R + + RS
Sbjct: 189 IDELHGSLLVHEQRLNGHVQEEQALKVTHEE----RPSQGRGRGVFRGSRGRGRGRGRSG 244
Query: 229 HNSKSIECWNCGKTGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICSL-----ESKEE 283
N +EC+ C GHF+ +C KN AN A + L+ + +++E
Sbjct: 245 TNRAIVECYKCHNLGHFQYECPEWEKN------ANYAELEEEEELLLMAYVEQNQANRDE 298
Query: 284 SWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKLNGSVWELKNV 343
W LDSG S H + KE+F G V LGN+ VVGKG VK+K+NG + V
Sbjct: 299 VWFLDSGCSNHMTGSKEWFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQVIPEV 358
Query: 344 RHIPNLTKNLISVGQLADEGYTTVFHGDDWKI---SKGAM--TIARGRKSGTLYKTAGAC 398
++P L NL+S+GQL + G + K+ SKGA+ T G + L +
Sbjct: 359 YYVPELRNNLLSLGQLQERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFLLASKPQK 418
Query: 399 HLIAVATNE----NPNLWHKRLGHMSEKGMKVMHSKGK---LPSLRSIEIDICEDCILGK 451
+ + + T E +LWH R GH++++G+K++ K LP L++ + +IC C+ GK
Sbjct: 419 NSLCLQTEEVMDKENHLWHCRFGHLNQEGLKLLAHKKMVIGLPILKATK-EICAICLTGK 477
Query: 452 QKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKS 511
Q R S +L+LVHSD+ GP T S GK Y ++FIDD +RK WVYFL KS
Sbjct: 478 QHRESMSKKTSWKSSTQLQLVHSDICGPITPISHSGKRYILSFIDDFTRKTWVYFLHEKS 537
Query: 512 EVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNG 571
E F FK +KA VE E + LRTD GGE+ +F +FC HGI + T TPQ NG
Sbjct: 538 EAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTPQQNG 597
Query: 572 VAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEV 631
VAER NRT+ RS+ + +PK FW+EA S ++ NR P+ +E PEE WSG++
Sbjct: 598 VAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWSGRKP 657
Query: 632 KLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVI 691
+ + RVFGC+ YVHI DQ R+KLD KSKKC+F+G E+ +RL+D KK+V SKDV+
Sbjct: 658 VVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVISKDVV 717
Query: 692 FNE--------------RVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELA 737
F+E V + +S + EP+ V + GS ++
Sbjct: 718 FDEDKSWDWDQADVEAKEVTLECGDEDDEKNSEVVEPIAVASPNHVGS-------DNNVS 770
Query: 738 ESSIRQPSDTLVHPTPVPVLRRSSRPHAPNRRYIDYMLLTDGGEPEDYDE---------- 787
S I PS P P PV + +R P DY + GE E+ +E
Sbjct: 771 SSPILAPSS----PAPSPVAAKVTRERRPPGWMADY----ETGEGEEIEENLSVMLLMMM 822
Query: 788 ----ACQTTDASK---WELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDG 840
Q DA K W AM+ E++S++ N TWEL LP G + KWVY+ K + DG
Sbjct: 823 TEADPIQFDDAVKDKIWREAMEHEIESIVKNNTWELTTLPKGFTPIGVKWVYKTKLNEDG 882
Query: 841 S-KRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLH 899
+YKARLV KG+ Q GIDYTE+FAPV +L+T+R++L+I + N + QLDVK+AFLH
Sbjct: 883 EVDKYKARLVAKGYAQCYGIDYTEVFAPVARLDTVRTILAISSQFNWEIFQLDVKSAFLH 942
Query: 900 GDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNAD 959
G+L EE+Y+ QPEGF+ EG+E V L+K+LYGLKQAPR WY + E++ KE F++C ++
Sbjct: 943 GELKEEVYVRQPEGFIREGEEEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCPSE 1002
Query: 960 HCCFFKRYKSSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQIT 1019
H F K + +I+ LYVDD++ GS+ K + EF+M DLG K LG+++
Sbjct: 1003 HTLFTKTRVGNILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEV- 1061
Query: 1020 RDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIP 1079
+ G + + Y VL RF M ++ V P+ +L+++++ + +E
Sbjct: 1062 -KQSDGGIFICQRRYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVDE------TM 1114
Query: 1080 YASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFGK 1139
+ +GSLMY V TRPD+ + V ++SRFMSNP +HW A K ILRYL+GT E +++ +
Sbjct: 1115 FKQLVGSLMYLTV-TRPDLMYGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRR 1173
Query: 1140 GE---IKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAV 1196
+ +K+ + D+D+AG+++ RRST+G++F + + ++ W S+ Q +VALSTTE EY+A
Sbjct: 1174 RKNRSLKLMAFTDSDYAGDLNDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAA 1233
Query: 1197 TEASKELIWLQGLLTELGFMQEKSA--LYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRS 1254
+ + +WL+ +L +LG +EKSA + D+ S I L+K+ H ++KHI +R+H++R
Sbjct: 1234 AFCACQCVWLRKVLEKLG-AEEKSATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRD 1292
Query: 1255 LLEDEVLTLIKIQGSKNPADMLTKVVTIDKLKLCSTLVGLL 1295
L+ +V+ L AD+ TK + +++ + L+G++
Sbjct: 1293 LVNGDVVKLEYCPTEDQVADIFTKPLKLEQFEKLRALLGMV 1333
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 700 bits (1807), Expect = 0.0
Identities = 445/1353 (32%), Positives = 694/1353 (50%), Gaps = 107/1353 (7%)
Query: 11 FDGADFGFWKMQIEDYLYQKKLHQPL----------TEKKPDSMKD----DEWSLLDRQA 56
FDG + FW +++ +KL + E+ P++ + +E D A
Sbjct: 12 FDGEKYDFWSIKMATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTNDTMA 71
Query: 57 LGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLM---RRLFTLRMAEGM 113
L +++ +++ + IA ++ L Y+ P V L R L+M +
Sbjct: 72 LQILQTAVTDQIFSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLKMYDND 131
Query: 114 SVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSKKMKFDD 173
++ ++L ++ QL+ G + + +L SLP + +IV+ + + + +
Sbjct: 132 NIKTFTDKLIVLEIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDALTMSE 191
Query: 174 VRDLVLSEEIRRRELGESSSSSVLHTESRGRNS--------TRGN------GRGKSKAR- 218
+ ++ ++E R ES+ + S+GR S R N G KS
Sbjct: 192 LLGILKAQEARVTAREESTKEGAFYVRSKGRESGFKQDNTNNRVNQDKKWCGFHKSSKHT 251
Query: 219 ----RSKSKNHRSSHNSKS-IECWNCGKTGHFKNQCRLPTKN-------QEEKDEANVAS 266
R K KN N +S I+C+ CGK GH+ N+CR K +E+ +E ++
Sbjct: 252 EEECREKPKNDDHGKNKRSNIKCYKCGKIGHYANECRSKNKERAHVTLEEEDVNEDHMLF 311
Query: 267 TSGGGDALICSLESKEESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGK 326
++ + S +E+ W++DSG + H + ++ +F N + + N GK
Sbjct: 312 SASEEE----STTLREDVWLVDSGCTNHMTKEERYFSNINKSIKVPIRVRNGDIVMTAGK 367
Query: 327 GEVKIKLNGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKIS----KGAMTI 382
G++ + +KNV +P L KNL+SV Q+ GY F I K M I
Sbjct: 368 GDITVMTRHGKRIIKNVFLVPGLEKNLLSVPQIISSGYWVRFQDKRCIIQDANGKEIMNI 427
Query: 383 ARGRKSGTLYKTAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEI- 441
KS + K + A + WHKRLGH+S K ++ M K + L ++
Sbjct: 428 EMTDKSFKI-KLSSVEEEAMTANVQTEETWHKRLGHVSNKRLQQMQDKELVNGLPRFKVT 486
Query: 442 -DICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSR 500
+ C+ C LGKQ R SF +T +EKLE+VH+DV GP SI G Y+V F+DD++
Sbjct: 487 KETCKACNLGKQSRKSFPKESQTKTREKLEIVHTDVCGPMQHQSIDGSRYYVLFLDDYTH 546
Query: 501 KVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRME 560
WVYFLK KSE F FK++KA+VE +++ IK LR + FC + GI +
Sbjct: 547 MCWVYFLKQKSETFATFKKFKALVEKQSNCSIKTLRP----------MEVFCEDEGINRQ 596
Query: 561 RTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHK 620
T+P +PQ NG AER NR+L E ARS+ V+ LP K WAEAV TSAYL NR PS +E
Sbjct: 597 VTLPYSPQQNGAAERKNRSLVEMARSMLVEQDLPLKLWAEAVYTSAYLQNRLPSKAIEDD 656
Query: 621 I-PEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDD 679
+ P E W G + +SHLR+FG + YVHI DQ R KLD K+K I IGY GYR++
Sbjct: 657 VTPMEKWCGHKPNVSHLRIFGSICYVHIPDQKRRKLDAKAKCGILIGYSNQTKGYRVFLL 716
Query: 680 ENKKMVRSKDVIFNERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAE- 738
E++K+ S+DV+F E DK + + + ++D+ S + S +L++
Sbjct: 717 EDEKVEVSRDVVFQE-----DKKWDWDKQEEVKKTFVMSINDIQESRDQQETSSHDLSQI 771
Query: 739 ----------------SSIRQPSDTLVHPTPVP------VLRRSSRPHAPNRRYIDYMLL 776
S + + +P +L ++ R L
Sbjct: 772 DDHANNGEGETSSHVLSQVNDQEERETSESPKKYKSMKEILEKAPRMENDEAAQGIEACL 831
Query: 777 TDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKE 836
EP+ YDEA +WE AM EE+K + N+TW+L P K + KW+Y++K
Sbjct: 832 VANEEPQTYDEA---RGDKEWEEAMNEEIKVIEKNRTWKLVDKPEKKNVISVKWIYKIKT 888
Query: 837 DHDGSK-RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKT 895
D G+ ++KARLV +GF Q+ GIDY E FAPV + +TIR++L+ A L Q+DVK+
Sbjct: 889 DASGNHVKHKARLVARGFSQEYGIDYLETFAPVSRYDTIRALLAYAAQMKWRLYQMDVKS 948
Query: 896 AFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQK 955
AFL+G+L EE+Y+ QP GF+ EGKE V L K+LYGLKQAPR WY + +S+ + GF +
Sbjct: 949 AFLNGELEEEVYVTQPPGFVIEGKEEKVLRLYKALYGLKQAPRAWYERIDSYFIQNGFAR 1008
Query: 956 CNADHCCFFKRYKSSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILG 1015
D + K+ +I+ LYVDD+++ G+N I K + EF+M DLG LG
Sbjct: 1009 SMNDAALYSKKKGEDVLIVSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLG 1068
Query: 1016 MQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELM 1075
M++ +D G+ LS +Y N+++ +F M ++K VSTPL + + ++KE
Sbjct: 1069 MEVNQD-DSGIF-LSQEKYANKLIDKFGMKESKSVSTPLTPQGKRKGVEG----DDKEFA 1122
Query: 1076 AKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTE-KC 1134
Y +G L+Y + +RPD+ +A +SR+MS+P H++ K +LRY++GT+
Sbjct: 1123 DPTKYRRIVGGLLY-LCASRPDVMYASSYLSRYMSSPSIQHYQEAKRVLRYVKGTSNFGV 1181
Query: 1135 LYFGKGEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYV 1194
L+ K ++ GY D+D+ G ++ ++STTGY+FT+G W S Q+ VA ST E EY+
Sbjct: 1182 LFTSKETPRLVGYSDSDWGGSLEDKKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYI 1241
Query: 1195 AVTEASKELIWLQGLLTELGF-MQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIR 1253
AV A+ + IWLQ L + G +E + D++SAI + +N H RTKHI ++YHF+R
Sbjct: 1242 AVCAATNQAIWLQRLFEDFGLKFKEGIPILCDNKSAIAIGRNPVQHRRTKHIEIKYHFVR 1301
Query: 1254 SLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
++ L +G AD+LTK +++ + +
Sbjct: 1302 EAEHKGLIQLEYCKGEDQLADVLTKALSVSRFE 1334
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 698 bits (1801), Expect = 0.0
Identities = 451/1331 (33%), Positives = 685/1331 (50%), Gaps = 192/1331 (14%)
Query: 36 LTEKKPDSMKDDEWSLLDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSS 95
L++ + D ++D D++AL ++ L + + + + + L + Y+
Sbjct: 51 LSQTQKDGLRDSRKR--DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQV 108
Query: 96 NKVHL--MRRLF-TLRMAEGMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDS 152
KV L +R F L+M EG V+ + + + VT L G + DD +L SL
Sbjct: 109 KKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLK 168
Query: 153 WSAIVTAVSSSSG-----------------SKKMKFDDVRDLVLSEEIRRRELGES---- 191
+ IVT + + KK K +D+ + VL+ +I + E G+S
Sbjct: 169 FEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRR 228
Query: 192 SSSSVL------------------HTESRGRNSTRGNGRGKSKARRSKSKNHRSSHNSKS 233
V +T RG NS+RG G+G K+R KS S
Sbjct: 229 GGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKS----------S 278
Query: 234 IECWNCGKTGHFKNQCRLPTKNQEEKDEAN-VASTSGGGDALICSLESKEE-----SWVL 287
++C+NCGK GH+ ++C+ P+ N++ K++AN V D L+ + K+E W L
Sbjct: 279 VKCYNCGKFGHYASECKAPS-NKKFKEKANYVEEKIQEEDMLLMASYKKDEQEENHKWYL 337
Query: 288 DSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKL-NGSVWELKNVRHI 346
DSGAS H +K F G V LG+E +V GKG + I+L NG + NV +I
Sbjct: 338 DSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYI 397
Query: 347 PNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLYKTA----------- 395
P++ N++S+GQL ++GY D ++ ++I R ++S + K
Sbjct: 398 PSMKTNILSLGQLLEKGY-------DIRLKDNNLSI-RDKESNLITKVPMSKNRMFVLNI 449
Query: 396 --GACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIE--IDICEDCILGK 451
+ + E LWH R GH++ G++++ K + L I +CE C+LG
Sbjct: 450 RNDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGN 509
Query: 452 QKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKS 511
Q ++SF + ++ LEL+H+DV GP S+G +YF+ FIDD SRK WVYFLK KS
Sbjct: 510 QFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKS 569
Query: 512 EVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNG 571
EVFE FK++KA VE E+ L IK +R+D+GGE+ +F K+C ++GIR + TVP +PQ NG
Sbjct: 570 EVFEIFKKFKAHVEKESGLVIKTMRSDSGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNG 629
Query: 572 VAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEV 631
VAER NRT+ E ARS+ LPK+ WAEAV + YL+NR P+ + K P+E WSG++
Sbjct: 630 VAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKP 689
Query: 632 KLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVI 691
+SHLRVFG +A+ H+ D+ RNKLD KS+K IFIGY + GY+L++ + KK + S++++
Sbjct: 690 GVSHLRVFGSIAHAHVPDEKRNKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIV 749
Query: 692 FNERVMY------KDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPS 745
F+E + +D + + EP E + SP S ++ ESS +
Sbjct: 750 FDEEGEWDWNSNEEDYNFFPHFEEDKPEPTREEPPSEEPTTPPTSPTSSQIEESSSER-- 807
Query: 746 DTLVHPTPVPVLRRSSRPHAPNRRYIDYMLLTDGGEPEDYDEACQTTDASKWELAMKEEM 805
TP + N+ + L EP D+ EA + W AM EE+
Sbjct: 808 ------TPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEA---IEKKTWRNAMDEEI 858
Query: 806 KSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGIDYTEI 864
KS+ N TWEL LP G KA+ KWVY+ K++ G +RYKARLV KG+ Q+ GIDY EI
Sbjct: 859 KSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDEI 918
Query: 865 FAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVC 924
FAPV +L T+R ++S+ A + Q+DVK+AFL+GDL EE+Y+ QP+G++ +G+E+ V
Sbjct: 919 FAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVL 978
Query: 925 MLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDDMLVA 984
LKK LYGLKQAPR W + + + ++ F KC +H + K K +I LYVDD++
Sbjct: 979 RLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIFT 1038
Query: 985 GSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNM 1044
G+N + K +++KEF+M D+G LG+++ + + G+ ++ Y VL++F M
Sbjct: 1039 GNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEV-KQEDNGIF-ITQEGYAKEVLKKFKM 1096
Query: 1045 GDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGV 1104
D+ S +GSL Y + CTRPDI +AVGV
Sbjct: 1097 DDSN--------------------------------PSLVGSLRY-LTCTRPDILYAVGV 1123
Query: 1105 VSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFGKGEIKVEGYVDADFAGEVDHRRSTTG 1164
VSR+M +P H++A K ILRY++GT L++
Sbjct: 1124 VSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHY--------------------------- 1156
Query: 1165 YIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEK-SALY 1223
STT + V A IWL+ LL EL QE+ + ++
Sbjct: 1157 ----------------------STTSDYKLVVCHA----IWLRNLLKELSLPQEEPTKIF 1190
Query: 1224 SDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTID 1283
D++SAI LAKN FH R+KHI RYH+IR + + + L ++ AD+ TK + +
Sbjct: 1191 VDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKRE 1250
Query: 1284 KLKLCSTLVGL 1294
+L+G+
Sbjct: 1251 DFIKMRSLLGV 1261
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 689 bits (1777), Expect = 0.0
Identities = 372/829 (44%), Positives = 522/829 (62%), Gaps = 59/829 (7%)
Query: 496 DDHSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEH 555
+D SRKVWVYFLK K E F +F WK MVE +++ K+K LRTDNG E+ + KF + C +
Sbjct: 319 NDWSRKVWVYFLKTKDEAFASFTEWKKMVETQSERKLKHLRTDNGLEFCNHKFDEVCKKE 378
Query: 556 GIRMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSV 615
GI RT TPQ NGVAER+NRT+ + RS+ +SGL KKFWA+A +T+ YLINR PS
Sbjct: 379 GIVRHRTCTYTPQQNGVAERLNRTIMNKVRSMLSESGLDKKFWAKAASTAVYLINRSPSS 438
Query: 616 PLEHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYR 675
+E+KIPEE+W+ S L+ FGCV YV+ S +G KLDP++KK +F+GY G+R
Sbjct: 439 SIENKIPEELWTSAVPNFSGLKRFGCVVYVY-SQEG--KLDPRAKKGVFVGYPNGVKGFR 495
Query: 676 LWDDENKKMVRSKDVIFNERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGE 735
+W E ++ S++V+F E VMYKD N +T SG+S + + +P + + E
Sbjct: 496 VWMIEEERCSISRNVVFREDVMYKDILNQST--SGMSFDFPLATNRIPSFECAGNRKEDE 553
Query: 736 LAESSIRQPSDTLVHPTPVPVLRRSSRPHAPNRRY------------------------- 770
++ DT P+ SS ++ R Y
Sbjct: 554 ISVQGGVSDDDTKQSSEESPISTGSSGQNSGQRTYQIARDKPKRQTKIPDKLRDYELNEE 613
Query: 771 -------IDYMLLTDGGEPE--DYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPI 821
YM+ DGG PE DY +A Q +D W A+ EE++SL+ N TW L
Sbjct: 614 VLDEIAGYAYMITEDGGNPEPNDYQKALQDSDYKMWLKAVDEEIESLLKNNTWVLVNRDQ 673
Query: 822 GKKALHNKWVYRVKEDHDGSK--RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLS 879
+K + KWV++ K G + R+KARLVVKG+ QKEGIDY EIF+PVVK +IR +LS
Sbjct: 674 FQKPIGCKWVFKRKSGIVGVEKPRFKARLVVKGYSQKEGIDYQEIFSPVVKHVSIRLLLS 733
Query: 880 IVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQ 939
+V ++ L+Q+DVKTAFLHG L E IY+ QPEG++ + + VC+LK+SLYGL+Q+PRQ
Sbjct: 734 MVTHCDMELQQMDVKTAFLHGYLDETIYIEQPEGYVHKRYPDKVCLLKRSLYGLRQSPRQ 793
Query: 940 WYMKFESFMHKEGFQKCNADHCCFFKRYKSS-YIILLLYVDDMLVAGSNIDEIKNLKIQL 998
W +F FM K G+++ D C +FK +S YI LLLYVDD+L+A + + +LK L
Sbjct: 794 WNNRFNEFMQKIGYERSKYDSCVYFKELQSGEYIYLLLYVDDILIASRDKRTVCDLKALL 853
Query: 999 SKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHF 1058
+ EF+MKDLG AKKILGM+I RD++ G + +S Y+ +VL F M AK V TP+ +HF
Sbjct: 854 NSEFEMKDLGDAKKILGMEIVRDRKAGTMSISQEGYLLKVLGNFGMDQAKPVFTPMGAHF 913
Query: 1059 RLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWE 1118
+L + + E+M +PY SA+GSLMY+M+ TRPD+ H+VG+V RFMS P K HW+
Sbjct: 914 KLKPATDEEVMRQSEVMRAVPYQSAVGSLMYSMIGTRPDLAHSVGLVCRFMSKPLKEHWQ 973
Query: 1119 AVKWILRYLRGTTE-KCLYFGKGEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWM 1177
AVKWILRY+RG+ + K Y +GE+ +EGY D+D+A + + RRST+G
Sbjct: 974 AVKWILRYIRGSIDRKLCYKNEGELILEGYCDSDYAADKEGRRSTSGV------------ 1021
Query: 1178 SRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSA 1237
K+VALS+TE EY+A+T+ +KE IWL+G ++ELGF+Q+ ++ DSQSAI LAKN+
Sbjct: 1022 ----KVVALSSTEAEYMALTDGAKEAIWLKGHVSELGFVQKTVNIHCDSQSAIALAKNAV 1077
Query: 1238 FHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKVVTIDKLK 1286
+H RTKHI ++YHFIR L+ + + ++KI NPAD+ TKV+ + K +
Sbjct: 1078 YHERTKHIDVKYHFIRDLVNNGEVQVLKIDTEDNPADIFTKVLPVSKFQ 1126
Score = 130 bits (326), Expect = 6e-30
Identities = 95/337 (28%), Positives = 168/337 (49%), Gaps = 40/337 (11%)
Query: 21 MQIEDYLYQKKLHQPLTE-------KKPDSMKDDEWSL-LDRQALGVVRLSLSRNVAFNI 72
M ++D L ++ PL E KK +++++ + D +A+ ++ +++ V NI
Sbjct: 1 MGLKDALVERAPLPPLKEEDESDPAKKKQRIEEEKARIDQDEKAMDMIFINVGDKVLRNI 60
Query: 73 AKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTLRMAEGMSVAQHINELNIVTTQLSSV 132
KT A L +Y N+V+L +++ RM + ++ ++++E + + L+++
Sbjct: 61 ENSKTAAEAWATLDKLYLVKSLPNRVYLQLKVYNYRMQDSKTLEENVDEFQKMISDLNNL 120
Query: 133 GIEFDDEVRALILLSSLPDSWSAIVTAVSSSSGSKKMKFDDVRDLVLSEEIRRRELGESS 192
I+ DEV+A+++LS+LPDS+ + + G + +K DDV S+E+ R
Sbjct: 121 QIQVPDEVQAILILSALPDSYDMLKETL--KYGREGIKLDDVISAAKSKELELR------ 172
Query: 193 SSSVLHTESRGRNSTRGNG---RGKSKARRSKSKNHRSSHNSKSIECWNCGKTGHFKNQC 249
+S G + G G RGKS+AR S +S+ K CW CGK GHFK QC
Sbjct: 173 -------DSSGGSRPVGEGLYVRGKSQARGSDGP--KSTEGKK--VCWICGKEGHFKRQC 221
Query: 250 -RLPTKNQEE--------KDEANVASTSGGGDALICSLESKEESWVLDSGASFHASSQKE 300
+ KN+ KD+A + + + +E W++D+G SFH + +KE
Sbjct: 222 YKWLEKNKANGAGETALVKDDAQDLVGLVASEVNMSEGKDDQEEWIMDTGCSFHMTPRKE 281
Query: 301 FFKNYVPGNLGKVYLGNEQSCKVVGKGEVK-IKLNGS 336
+ ++V GKV + N +V G G+VK IK +G+
Sbjct: 282 YLMDFVEAKSGKVRMANNSFSEVKGIGKVKFIKKDGT 318
>At3g25450 hypothetical protein
Length = 1343
Score = 669 bits (1726), Expect = 0.0
Identities = 427/1336 (31%), Positives = 683/1336 (50%), Gaps = 79/1336 (5%)
Query: 6 VKIEKFDGADFGFWKMQIEDYLYQKKLHQPLTEKKPDSMKDDEWSLLDRQALGVVRLSLS 65
++ + ++ W M++E L KL + D K+D A ++ S+
Sbjct: 20 IQCPMLNSVNYTVWTMRMEAVLRVHKLWGTIEPGSADEEKND-------MARALLFQSIP 72
Query: 66 RNVAFNIAKEKTTAGLMKALSSMY---EKPPSSNKVHLMRRLFTLRMAEGMSVAQHINEL 122
++ + K+KT++ + +A+ S E+ + LM L+M + ++ ++ +
Sbjct: 73 ESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMAEFDKLKMKDSETIDDYVGRI 132
Query: 123 NIVTTQLSSVGIEFDDEVRALILLSSLP-DSWSAIVTAVSSSSGSKKMKFDDVRDLVLSE 181
+ +TT+ +++G + ++ L SLP + IV A+ K F+D+ + +
Sbjct: 133 SEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAALEQVLDLKTTTFEDIAGRIKTY 192
Query: 182 EIRRRELGESSSSSVLHTESRGRNSTRGNGRGKSKARRSKSKNHRSSHNSKSIECWNCGK 241
E R + +S E +G+ T + + +KS
Sbjct: 193 EDRVWDDDDSH-------EDQGKLMTEVEEEVVDDLEEEEEEVINKEIKAKS------HV 239
Query: 242 TGHFKNQCRLPTKNQEEKDEANVASTSGGGDALICS--------LESK-EESWVLDSGAS 292
RL + ++E+D+ + A + + + + LES +W LD+GAS
Sbjct: 240 IDRLLKLIRLQEQKEKEEDDTHEAESLMMHEVVYLNEKNIRPTELESCINNAWYLDNGAS 299
Query: 293 FHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVK-IKLNGSVWELKNVRHIPNLTK 351
H + + +F GKV G++ + GKG + I G L +V +IP+L
Sbjct: 300 NHMTGNRAWFCKLDEMITGKVRFGDDSCINIKGKGSIPFISKGGERKILFDVYYIPDLKS 359
Query: 352 NLISVGQLADEGYTTVFHGDDWKIS--KGAMTIARGRKSGTLYKTAGACH---LIAVATN 406
N++S+GQ + G D + +G + I R LYK + + + T
Sbjct: 360 NILSLGQATESGCDIRMREDYLTLHDREGNLLIKAQRSRNRLYKVSLEVENSKCLQLTTT 419
Query: 407 ENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSI---EIDICEDCILGKQKRVSFQTSGRT 463
+WH RLGH+S + +K M K + + S E + C C+ GKQ R SF +
Sbjct: 420 NESTIWHARLGHISFETIKAMIKKELVIGISSSVPQEKETCGSCLFGKQARHSFPKATSY 479
Query: 464 PKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSEVFEAFKRWKAM 523
+ LEL+H D+ GP + + K Y IDDHSR +W LK KSE F FK +KA+
Sbjct: 480 RAAQVLELIHGDLCGPISPSTAAKKRYVFVLIDDHSRYMWSILLKEKSEAFGKFKEFKAL 539
Query: 524 VENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGVAERMNRTLTER 583
VE E IK RTD GGE+ +F++FC + GI T P TPQ NGV ER NRTL
Sbjct: 540 VEQECGAIIKTFRTDRGGEFLSHEFQEFCAKEGINRHLTAPYTPQQNGVVERRNRTLLGM 599
Query: 584 ARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVKLSHLRVFGCVA 643
RS+ +P W EAV S YLINR + L ++ P EV+ K+ + HLRVFGCV+
Sbjct: 600 TRSILKHMNMPNYLWGEAVRHSTYLINRVGTRSLSNQTPYEVFKHKKPNVEHLRVFGCVS 659
Query: 644 YVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIFNER--VMYKDK 701
Y + KLD +S+ +++G YRL D +++ S+DV+F+E M+++
Sbjct: 660 YAKVEVPNLKKLDDRSRMLVYLGTEPGSKAYRLLDPTKRRIFVSRDVVFDENRSWMWQES 719
Query: 702 HNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPSDTLVHPT--------- 752
+ T +SG E + + D S + E E+ I + ++
Sbjct: 720 SSETDKESGTFTITLSEFGNNGVTENDISTEPEETEEAEINGEDENIIEEAETEEHDQSQ 779
Query: 753 --PVPVLRRSSRPHAPN--RRYI-------DYMLLTDGGEPEDYDEACQTTDASKWELAM 801
P PV R + PN + Y+ +++LL EP D+ EA + +W A
Sbjct: 780 EEPQPVRRSQRQVIRPNYLKDYVLCAEIEAEHLLLAVNDEPWDFKEA---NKSKEWRDAC 836
Query: 802 KEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGID 860
KEE++S+ N+TW L LP+G KA+ KWV+++K + DGS +YKARLV KG+ Q+ G+D
Sbjct: 837 KEEIQSIEKNRTWSLVDLPVGSKAIGVKWVFKLKHNSDGSINKYKARLVAKGYVQRHGVD 896
Query: 861 YTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKE 920
+ E+FAPV ++ T+R ++++ AS + LDVKTAFLHG+L E++Y+ QPEGF + +
Sbjct: 897 FEEVFAPVARIETVRLIIALAASNGWEIHHLDVKTAFLHGELREDVYVSQPEGFTNKESK 956
Query: 921 NMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDD 980
V L K+LYGL+QAPR W K + + F+KC+ + + K+ + +++ +YVDD
Sbjct: 957 EKVYKLHKALYGLRQAPRAWNTKLNEILKELKFEKCHKEPSLYRKQEGENILVVAVYVDD 1016
Query: 981 MLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQ 1040
+LV GSN+D I N K + +F+M DLG LG+++ + K + L Y ++L+
Sbjct: 1017 LLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQSKDG--ITLKQERYAKKILE 1074
Query: 1041 RFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGH 1100
M V+TP+ + LS+ Q + +E + Y IG L Y ++ TRPD+ +
Sbjct: 1075 EAGMSKCNTVNTPMIASLELSKAQDEKRIDETD------YRRNIGCLRY-LLHTRPDLSY 1127
Query: 1101 AVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFGKGE-IKVEGYVDADFAGEVDHR 1159
VG++SR++ P ++H A+K ILRYL+GTT LYF KGE + GY D+ ++D
Sbjct: 1128 NVGILSRYLQEPRESHGAALKQILRYLQGTTSHGLYFKKGENAGLIGYSDSSHNVDLDDG 1187
Query: 1160 RSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTE-LGFMQE 1218
+ST G+IF + ++W S+ Q++V LS+ E E++A TEA+K+ IWLQ LL E +G E
Sbjct: 1188 KSTGGHIFYLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQELLAEVIGTECE 1247
Query: 1219 KSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTK 1278
K + D++SAI L KN FH R+KHI RYHFIR +E+ + + + G + AD+LTK
Sbjct: 1248 KVTIRVDNKSAIALTKNPVFHGRSKHIHRRYHFIRECVENGQIEVEHVPGVRQKADILTK 1307
Query: 1279 VVTIDKLKLCSTLVGL 1294
+ K L+G+
Sbjct: 1308 ALGKIKFLEMRELIGV 1323
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 652 bits (1683), Expect = 0.0
Identities = 421/1289 (32%), Positives = 657/1289 (50%), Gaps = 95/1289 (7%)
Query: 63 SLSRNVAFNIAKEKTTAGLMKALSSMY---EKPPSSNKVHLMRRLFTLRMAEGMSVAQHI 119
S+ + + K KT+ + +A+ + E+ + LM L M + ++ + +
Sbjct: 34 SVPESTILQVGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNMKDNETIDEFV 93
Query: 120 NELNIVTTQLSSVGIEFDDEVRALILLSSLP-DSWSAIVTAVSSSSGSKKMKFDDV---- 174
++ ++T+ S+G E ++ L SLP + I+ A+ F+D+
Sbjct: 94 GRISEISTKSESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNTTGFEDIVGRM 153
Query: 175 ---RDLVLSEEIRRRELGESS-SSSVLHTESRGRNSTRGNGRGKSKARRSKSKNHRSSHN 230
D V E+ E G+ ++S ++RG RG GRG+S R ++
Sbjct: 154 KTYEDRVCDEDDSPEEQGKLMYANSESSYDTRGG---RGRGRGRSSGRGRGGYGYQQRDK 210
Query: 231 SKSIECWNCGKTGHFKNQC--------RLPTKNQEEKDEANVASTSGGGDALIC--SLES 280
SK I C+ C KTGH+ ++C + + Q +D+ + S + S++
Sbjct: 211 SKVI-CYRCDKTGHYASECLDRLLKLIKAQEQQQNNEDDDEIESLMMHEVVYLNERSVKP 269
Query: 281 KE------ESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKLN 334
KE SW LD+GAS H + ++F GKV G++ + GKG + +
Sbjct: 270 KEFEACSDNSWYLDNGASNHMTGNLQWFSKLNEMITGKVRFGDDSRIDIKGKGSIVLITK 329
Query: 335 GSVWE-LKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKIS--KGAMTIARGRKSGTL 391
G + + L +V IP+L N+IS+GQ + G D + +G + + R L
Sbjct: 330 GGIRKTLTDVYFIPDLKSNIISLGQATEAGCDVRMKDDQLTLHDREGCLLLRATRSRNRL 389
Query: 392 YKTAGACHLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICEDCILGK 451
YK EN M K + + S +P E + C C+LGK
Sbjct: 390 YKVD--------LNVENVKCLQLEAATMVRKELVIGISN--IPK----EKETCGSCLLGK 435
Query: 452 QKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKS 511
Q R F + + LELVH D+ GP T + K Y + IDDH+R +W LK KS
Sbjct: 436 QARQPFPKATTYRASQVLELVHGDLCGPITQSTTAKKRYILVLIDDHTRYMWSMLLKEKS 495
Query: 512 EVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNG 571
E FE F+ +K VE E+ +KIK RTD GGE+ +F+ FC + GI T P TPQ NG
Sbjct: 496 EAFEKFRDFKTKVEQESGVKIKTFRTDKGGEFVSQEFQDFCAKEGINRHLTAPYTPQQNG 555
Query: 572 VAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEV 631
V ER NRTL RS+ +P W EAV S Y+INR + L+++ P EV+ ++
Sbjct: 556 VVERRNRTLLGMTRSILKHMKMPNYLWGEAVRHSTYIINRVGTRSLQNQTPYEVFKQRKP 615
Query: 632 KLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVI 691
+ HLRVFGC+ Y I KLD +SK +++G YRL D N+K+++
Sbjct: 616 NVEHLRVFGCIGYAKIEGPHLRKLDDRSKMLVYLGTEPGSKAYRLLDPTNRKIIKWN--- 672
Query: 692 FNERVMYKDKHNTTT------NDSGLSEPVYVEMDDVPGSPTDKSPQSG-------ELAE 738
N +D T + ++G+ E +E + + + G E +
Sbjct: 673 -NSDSETRDISGTFSLTLGEFGNNGIQESDDIETEKNGEESENSHEEEGENEHNEQEQID 731
Query: 739 SSIRQPSDTLVHPTPVPVLRRSSRPHAPNRRYIDYMLLTD----------GGEPEDYDEA 788
+ QPS H TP+P LRRS+R DY+L+ + EP D+ EA
Sbjct: 732 AEETQPS----HATPLPTLRRSTRQVGKPNYLDDYVLMAEIEGEQVLLAINDEPWDFKEA 787
Query: 789 CQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKAR 847
+W A KEE+ S+ N+TW L LP+ +K + KWV+++K + DGS +YKAR
Sbjct: 788 ---NKLKEWRDACKEEILSIEKNKTWSLIDLPVRRKVIGLKWVFKIKRNSDGSINKYKAR 844
Query: 848 LVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIY 907
LV KG+ Q+ GIDY E+FA V ++ TIR ++++ AS + LDVKTAFLHG+L E++Y
Sbjct: 845 LVAKGYVQRHGIDYDEVFAHVARIETIRVIIALAASNGWEVHHLDVKTAFLHGELREDVY 904
Query: 908 MHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRY 967
+ QPEGF + E V L K+LYGLKQAPR W K + + F KC+ + + ++
Sbjct: 905 VTQPEGFTNKDNEGKVYKLHKALYGLKQAPRAWNTKLNKILQELNFVKCSKEPSVYRRQE 964
Query: 968 KSSYIILLLYVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVL 1027
+ +I+ +YVDD+LV GS++D I K ++ +F+M DLG LG+++ K +L
Sbjct: 965 EKKLLIVAIYVDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLHRKNGIIL 1024
Query: 1028 QLS*AEYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSL 1087
+ Y ++++ M + V P+A+ L + Q E++ + + Y IG L
Sbjct: 1025 RQE--RYAMKIIEEAGMSNCNPVLIPMAAGLELCKAQ------EEKCITERDYRRMIGCL 1076
Query: 1088 MYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFGKG-EIKVEG 1146
Y +V TRPD+ + VGV+SR++ P ++H A+K +LRYL+GT LY +G + + G
Sbjct: 1077 RY-IVHTRPDLSYCVGVLSRYLQQPRESHGNALKQVLRYLKGTMSHGLYLKRGFKSGLVG 1135
Query: 1147 YVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWL 1206
Y D+ + ++D +ST G+IF + ++W S+ Q++VALS+ E E++A TEA+K+ IWL
Sbjct: 1136 YSDSSHSADLDDGKSTAGHIFYLHQCPITWCSQKQQVVALSSCEAEFMAATEAAKQAIWL 1195
Query: 1207 QGLLTEL-GFMQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIK 1265
Q L E+ G EK + D++SAI L KN FH R+KHI RYHFIR +E+ ++ +
Sbjct: 1196 QDLFAEVCGTTSEKVMIRVDNKSAIALTKNLVFHGRSKHIHRRYHFIRECVENNLVEVDH 1255
Query: 1266 IQGSKNPADMLTKVVTIDKLKLCSTLVGL 1294
+ G + AD+LTK + K + LVG+
Sbjct: 1256 VPGVEQRADILTKPLGRIKFREMRELVGV 1284
>At4g03810 putative retrotransposon protein
Length = 964
Score = 647 bits (1670), Expect = 0.0
Identities = 379/978 (38%), Positives = 549/978 (55%), Gaps = 40/978 (4%)
Query: 337 VWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKISKGAMTIARGRKSGTLYKTAG 396
V ELKN ++P + KN+ISV L EG+ + M L+
Sbjct: 2 VLELKNCYYVPAINKNIISVSCLDMEGFHFSIKNKCCSFDRDDMFYGSAPLDNGLHVLNQ 61
Query: 397 ACHLIAVATNE------NPN-LWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICEDCIL 449
+ + + T + NP LWH RLGH++EK ++ +HS G L S + CE C+L
Sbjct: 62 SMPIYNIRTKKFKSNDLNPTFLWHCRLGHINEKHIQKLHSDGLLNSFDYESYETCESCLL 121
Query: 450 GKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKH 509
GK + F T + L L+H+DV GP + + G YF+TF DD SR +VY +KH
Sbjct: 122 GKMTKAPF-TGHSERASDLLGLIHTDVCGPMSTSARGNYQYFITFTDDFSRYGYVYLMKH 180
Query: 510 KSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQH 569
KS+ FE FK ++ V+N+ IK LR+D GGEY F E GI + T PGTPQ
Sbjct: 181 KSKSFENFKEFQNEVQNQFGKSIKALRSDRGGEYLSQVFSDHLRECGIVSQLTPPGTPQW 240
Query: 570 NGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGK 629
NGV+ER NRTL + RS+ + LP FW A+ TSA+++NR PS +E K P E+W+GK
Sbjct: 241 NGVSERRNRTLLDMVRSMMSHTDLPSPFWGYALETSAFMLNRCPSKSVE-KTPYEIWTGK 299
Query: 630 EVKLSHLRVFGCVAYVH--ISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRS 687
LS L+++GC +Y I+D KL PKS KC F+GY ++ GY + + K+
Sbjct: 300 VPNLSFLKIWGCESYAKRLITD----KLGPKSDKCYFVGYPKETKGYYFYHPTDNKVFVV 355
Query: 688 KDVIFNERVMYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKSPQSGELAESSIRQPSDT 747
++ F ER T+ L E V DVP S + + E + +P
Sbjct: 356 RNGAFLEREFLS---KGTSGSKVLLEEVREPQGDVPTSQEEHQLDLRRVVEPILVEPE-- 410
Query: 748 LVHPTPVPVLRRSSRPHAPNRRYIDYML------LTDGGEPEDYDEACQTTDASKWELAM 801
+RRS R R+ D+++ + + EP Y+EA D+ KW A
Sbjct: 411 ---------VRRSERSRHEPDRFRDWVMDDHALFMIESDEPTSYEEALMGPDSDKWLEAA 461
Query: 802 KEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGSKR-YKARLVVKGFRQKEGID 860
K EM+S+ N+ W L LP G K + KW+++ K D DG+ + YKA LV KG++Q GID
Sbjct: 462 KSEMESMSQNKVWTLVDLPDGVKPIECKWIFKKKIDMDGNIQIYKAGLVAKGYKQVHGID 521
Query: 861 YTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKE 920
Y E ++PV L +IR +L+ A + + Q+DVKTAFL+G+L E +YM QPEGF
Sbjct: 522 YDETYSPVAMLKSIRILLATAAHYDYEIWQMDVKTAFLNGNLEEHVYMTQPEGFTVPEAA 581
Query: 921 NMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDD 980
VC L +S+YGLKQA R W ++F + + F + + C + K S+ L+LYVDD
Sbjct: 582 RKVCKLHRSIYGLKQASRSWNLRFNEAIKEFDFIRNEEEPCVYKKTSGSAVAFLVLYVDD 641
Query: 981 MLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQ 1040
+L+ G++I ++++K L F MKD+G A ILG++I RD+ ++ LS YI++VL
Sbjct: 642 ILLLGNDIPLLQSVKTWLGSCFSMKDMGEAAYILGIRIYRDRLNKIIGLSQDTYIDKVLH 701
Query: 1041 RFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGH 1100
RFNM D+K P++ LS+ Q P T +E+E M+KIPYASAIGS+MYAM+ TRPD+
Sbjct: 702 RFNMHDSKKGFIPMSHGITLSKTQCPSTHDERERMSKIPYASAIGSIMYAMLYTRPDVAC 761
Query: 1101 AVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCL-YFGKGEIKVEGYVDADFAGEVDHR 1159
A+ + SR+ S+PG++HW V+ I +YLR T +K L Y G E+ V GY DA F + D
Sbjct: 762 ALSMTSRYQSDPGESHWIVVRNIFKYLRRTKDKFLVYGGSEELVVSGYTDASFQTDKDDF 821
Query: 1160 RSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEK 1219
RS +G+ F + +VSW S Q VA STTE EY+A +EA+KE++W++ +TELG +
Sbjct: 822 RSQSGFFFCLNGGAVSWKSTKQSTVADSTTEAEYIAASEAAKEVVWIRKFITELGVVPSI 881
Query: 1220 SA---LYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADML 1276
S LY D+ AI AK H ++KHI RYH IR +++ + + ++ N AD
Sbjct: 882 SGPIDLYCDNNGAIAQAKEPKSHQKSKHIQRRYHLIREIIDRGDVKISRVSTDANVADHF 941
Query: 1277 TKVVTIDKLKLCSTLVGL 1294
TK + K + +T +G+
Sbjct: 942 TKPLPQPKHESHTTAIGI 959
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 588 bits (1516), Expect = e-168
Identities = 366/1023 (35%), Positives = 552/1023 (53%), Gaps = 50/1023 (4%)
Query: 284 SWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKLNGSVWELKNV 343
+WV+DSGA+ H S + F + L V L + K+ G G +KLN + LKNV
Sbjct: 430 TWVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPTGPTVKISGVGT--LKLNDDIL-LKNV 486
Query: 344 RHIPNLTKNLISVGQLADE-GYTTVFHGDDWKIS---KGAMTIARGRKSGTLYKTAGACH 399
IP NLIS+ L D+ G +F + +I KG M + +GR+ LY
Sbjct: 487 LFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRM-LGQGRRVANLYLLDVGDQ 545
Query: 400 LIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICEDCILGKQKRVSFQT 459
I+V + ++WH+RLGH S + + + ++ D C C L KQ+++SF T
Sbjct: 546 SISVNAVVDISMWHRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPT 605
Query: 460 SGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSEVFEAFKR 519
S + K E +L+H DVWGP +V ++ G YF+T +DDHSR W+Y LK KSEV F
Sbjct: 606 SNKVCK-EIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEVLTVFPA 664
Query: 520 WKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGVAERMNRT 579
+ VEN+ +K+K +R+DN E KF F E GI + P TP+ N V ER ++
Sbjct: 665 FIQQVENQYKVKVKAVRSDNAPEL---KFTSFYAEKGIVSFHSCPETPEQNSVVERKHQH 721
Query: 580 LTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVKLSHLRVF 639
+ AR+L QS +P W + V T+ +LINR PS L +K P E+ +G LR F
Sbjct: 722 ILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTPYEILTGTAPVYEQLRTF 781
Query: 640 GCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIFNERVMYK 699
GC+ Y S + R+K P+S+ C+F+GY GY+L D E+ + S++V F+E V
Sbjct: 782 GCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESNTVFISRNVQFHEEVFPL 841
Query: 700 DKHNTTTNDSGLSEPVYVE----MDDVPGSPTDKSPQSGEL----AESSIRQPSDTL--- 748
K+ + + L P+ + D SP+ Q +L + +R+P L
Sbjct: 842 AKNPGSESSLKLFTPMVPVSSGIISDTTHSPSSLPSQISDLPPQISSQRVRKPPAHLNDY 901
Query: 749 -------VHPTPVPVLRRSSRPHAPNRRYIDYMLLTDGGEPEDYDEACQTTDASKWELAM 801
H P+ S+ + YI+ +T P +Y EA D +W A+
Sbjct: 902 HCNTMQSDHKYPISSTISYSKISPSHMCYINN--ITKIPIPTNYAEA---QDTKEWCEAV 956
Query: 802 KEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGID 860
E+ ++ TWE+ LP GKKA+ KWV+ +K DG+ +RYKARLV KG+ QKEG+D
Sbjct: 957 DAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLD 1016
Query: 861 YTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEG-- 918
YT+ F+PV K+ TI+ +L + AS+ +L+QLDV AFL+G+L EEI+M PEG+ E
Sbjct: 1017 YTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGI 1076
Query: 919 --KENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLL 976
N+V LK+S+YGLKQA RQW+ KF S + GF+K + DH F K Y ++I+L+
Sbjct: 1077 VLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLV 1136
Query: 977 YVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYIN 1036
YVDD+++A ++ L +L + F ++DLG K LG+++ R + + +Y
Sbjct: 1137 YVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVAR--TTAGISICQRKYAL 1194
Query: 1037 RVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRP 1096
+LQ M K VS P+ + ++ ++ E+ ++ Y +G LMY + TRP
Sbjct: 1195 ELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQ------YRRIVGKLMY-LTITRP 1247
Query: 1097 DIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYF-GKGEIKVEGYVDADFAGE 1155
DI AV + +F S P H A +L+Y++GT + L++ ++ ++G+ D+D+A
Sbjct: 1248 DITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASSDLTLKGFADSDWASC 1307
Query: 1156 VDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGF 1215
D RRSTT + VG +SW S+ Q V+ S+ E EY A+ A+ E++WL LL L
Sbjct: 1308 QDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQA 1367
Query: 1216 MQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADM 1275
LYSDS +AI++A N FH RTKHI L H +R L++ L L+ ++ AD+
Sbjct: 1368 SPPVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVADI 1427
Query: 1276 LTK 1278
LTK
Sbjct: 1428 LTK 1430
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 582 bits (1499), Expect = e-166
Identities = 368/1023 (35%), Positives = 561/1023 (53%), Gaps = 52/1023 (5%)
Query: 283 ESWVLDSGASFHASSQKEFFKNYVPGNLGKVYLGNEQSCKVVGKGEVKIKLNGSVWELKN 342
++WV+DSGA+ H S ++ F+ + V L + ++ G G V I + L+N
Sbjct: 440 DTWVIDSGATHHVSHDRKLFQTLDTSIVSFVNLPTGPNVRISGVGTVLINKDII---LQN 496
Query: 343 VRHIPNLTKNLISVGQLA-DEGYTTVFHGDDWKI---SKGAMTIARGRKSGTLYKTAGAC 398
V IP NLIS+ L D G +F +I +KG +T+ G++ G LY
Sbjct: 497 VLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKG-LTLGEGKRIGNLYVLDTQS 555
Query: 399 HLIAVATNENPNLWHKRLGHMSEKGMKVMHSKGKLPSLRSIEIDICEDCILGKQKRVSFQ 458
I+V + ++WHKRLGH S + + ++ + C C L KQK++SF
Sbjct: 556 PAISVNAVVDVSVWHKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFP 615
Query: 459 TSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDHSRKVWVYFLKHKSEVFEAFK 518
++ EL+H DVWGP +V ++ G YF+T +DDHSR W+Y LK KS+V F
Sbjct: 616 SANNICNST-FELLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFP 674
Query: 519 RWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIRMERTVPGTPQHNGVAERMNR 578
+ +VEN+ D ++K +R+DN E T+F K GI + P TP+ N V ER ++
Sbjct: 675 AFIDLVENQYDTRVKSVRSDNAKELAFTEFYK---AKGIVSFHSCPETPEQNSVVERKHQ 731
Query: 579 TLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIPEEVWSGKEVKLSHLRV 638
+ AR+L QS + +W + V T+ +LINR PS L +K P EV +GK S L+
Sbjct: 732 HILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLPDYSQLKT 791
Query: 639 FGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIFNERV-- 696
FGC+ Y S + R+K P+S+ C+F+GY GY+L D E+ + S++V F+E +
Sbjct: 792 FGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVEFHEELFP 851
Query: 697 MYKDKHNTTTNDSGLSEPVYVEMDDVPGSPTDKS----PQSGELAESSIRQPSDTLVHPT 752
+ + + TT + V+ MD + + S PQ + S R+ + H
Sbjct: 852 LASSQQSATT-----ASDVFTPMDPLSSGNSITSHLPSPQISPSTQISKRRITKFPAHLQ 906
Query: 753 PVP---VLRRSSRPHAPNRRYID----YMLLTDGGE----PEDYDEACQTTDASKWELAM 801
V + S P + + Y +ML + P+ Y EA D+ +W A+
Sbjct: 907 DYHCYFVNKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEA---KDSKEWCGAI 963
Query: 802 KEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGID 860
+E+ ++ TWE+ LP GKKA+ KWV+ VK DGS +R+KAR+V KG+ QKEG+D
Sbjct: 964 DQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLD 1023
Query: 861 YTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLE-EGK 919
YTE F+PV K+ T++ +L + AS+ YL QLD+ AFL+GDL E IYM P+G+ + +G
Sbjct: 1024 YTETFSPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGT 1083
Query: 920 E---NMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLL 976
N+VC LKKS+YGLKQA RQW++KF + + GF+K + DH F + S +I+LL+
Sbjct: 1084 SLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLV 1143
Query: 977 YVDDMLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYIN 1036
YVDD+++A + ++L L F +++LGP K LG+++ R + + LS +Y
Sbjct: 1144 YVDDIVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEG--ISLSQRKYAL 1201
Query: 1037 RVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRP 1096
+L +M D K S P+ + RLS+ E+ KE+ Y +G LMY + TRP
Sbjct: 1202 ELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLED-KEM-----YRRLVGKLMYLTI-TRP 1254
Query: 1097 DIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYFG-KGEIKVEGYVDADFAGE 1155
DI AV + +F S P AH AV +L+Y++GT + L++ + ++ ++GY DAD+
Sbjct: 1255 DITFAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTC 1314
Query: 1156 VDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGF 1215
D RRSTTG+ VG+ +SW S+ Q V+ S+ E EY A+ AS E+ WL LL L
Sbjct: 1315 PDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRV 1374
Query: 1216 MQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADM 1275
LYSDS +A+++A N FH RTKHI + H +R L++ L L+ ++ AD+
Sbjct: 1375 HSGVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADI 1434
Query: 1276 LTK 1278
LTK
Sbjct: 1435 LTK 1437
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 552 bits (1422), Expect = e-157
Identities = 409/1361 (30%), Positives = 635/1361 (46%), Gaps = 127/1361 (9%)
Query: 10 KFDGADFGFWKMQIEDYLYQKKL------------------HQPLTEKKPDSMKDDEWSL 51
K +++ WK Q E L +KL + +T ++P+ + + W
Sbjct: 20 KLTDSNYLLWKTQFESLLSSQKLIGFVNGAVNAPSQSRLVVNGEVTSEEPNPLYES-WFC 78
Query: 52 LDRQALGVVRLSLSRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTLRMAE 111
D+ + +LS V ++ T+ + +L+ + K + + L + L L E
Sbjct: 79 TDQLVRSWLFGTLSEEVLGHVHNLSTSRQIWVSLAENFNKSSVAREFSLRQNLQLLSKKE 138
Query: 112 GMSVAQHINELNIVTTQLSSVGIEFDDEVRALILLSSLPDSWSAIVTAVSSSSG------ 165
+ + E + LSS+G D+ ++ L+ L + I T + SS
Sbjct: 139 -KPFSVYCREFKTICDALSSIGKPVDESMKIFGFLNGLGRDYDPITTVIQSSLSKLPTPT 197
Query: 166 ------------SKKMKFDDVRDLV--LSEEIRRRELGESSSSSVLHTESRGRNSTRGNG 211
SK +++ + L+ I R E G + + + RGR S + G
Sbjct: 198 FNDVVSEVQGFDSKLQSYEEAASVTPHLAFNIERSESGSPQYNP--NQKGRGR-SGQNKG 254
Query: 212 RGKSKARRSKSKNHRSSHNSKSIE--CWNCGKTGHFKNQC--RLPTKNQEEKDEANVAST 267
RG R H+SS C CG+TGH +C R Q E +
Sbjct: 255 RGGYSTRGRGFSQHQSSPQVSGPRPVCQICGRTGHTALKCYNRFDNNYQAEIQAFSTLRV 314
Query: 268 SGGGDALICSLESKEESWVLDSGASFHASSQKEFFKNYVP--GNLGKVYLGNEQSCKVVG 325
S + + W DS A+ H +S ++ G+ V +G+ +
Sbjct: 315 S----------DDTGKEWHPDSAATAHVTSSTNGLQSATEYEGD-DAVLVGDGTYLPITH 363
Query: 326 KGEVKIKLNGSVWELKNVRHIPNLTKNLISVGQLADEGYTTVFHGDDWKIS----KGAMT 381
G IK + L V +PN+ K+L+SV +L D+ Y + D K+ +
Sbjct: 364 TGSTTIKSSNGKIPLNEVLVVPNIQKSLLSVSKLCDD-YPCGVYFDANKVCIIDLQTQKV 422
Query: 382 IARGRKSGTLYKTAGACHLIAVATNEN----PNLWHKRLGHMSEKGMKVMHSKGKLPSLR 437
+ G + LY +A+ +N +WH RLGH + K ++ + + + +
Sbjct: 423 VTTGPRRNGLYVLENQ-EFVALYSNRQCAATEEVWHHRLGHANSKALQHLQNSKAIQINK 481
Query: 438 SIEIDICEDCILGKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDD 497
S +CE C +GK R+ F S + L+ +H D+WGP+ V S G Y+ F+DD
Sbjct: 482 SRTSPVCEPCQMGKSSRLPFLISD-SRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDD 540
Query: 498 HSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGI 557
+SR W Y L +KSE F ++ +VEN+ + KIK ++D GGE+ K K EHGI
Sbjct: 541 YSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGI 600
Query: 558 RMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPL 617
+ P TPQ NG+AER +R L E S+ S P+KFW E+ T+ Y+INR PS L
Sbjct: 601 HHRISCPYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVL 660
Query: 618 EHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLW 677
++ P E G++ S LRVFG Y + +NK DP+S +C+F+GY GYR +
Sbjct: 661 KNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCF 720
Query: 678 DDENKKMVRSKDVIFNE-RVMYKDK-----------------HNTTTNDSGLSEPVY--- 716
K+ S++VIFNE + +K+K HN + S + PV
Sbjct: 721 YPPTGKVYISRNVIFNESELPFKEKYQSLVPQYSTPLLQAWQHNKISEISVPAAPVQLFS 780
Query: 717 --VEMDDVPGSPTDKSPQSGELAESSIRQPSDTLVHPTPVPVLRRSSR---PHA------ 765
++++ GS + Q + +S + SD V+P + + HA
Sbjct: 781 KPIDLNTYAGSQV--TEQLTDPEPTSNNEGSDEEVNPVAEEIAANQEQVINSHAMTTRSK 838
Query: 766 -----PNRRYIDYMLLTDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLP 820
PN RY + EP+ A + W A+ EE+ + TW L
Sbjct: 839 AGIQKPNTRYALITSRMNTAEPKTLASAMKHPG---WNEAVHEEINRVHMLHTWSLVPPT 895
Query: 821 IGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLS 879
L +KWV++ K DGS + KARLV KGF Q+EG+DY E F+PVV+ TIR VL
Sbjct: 896 DDMNILSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLD 955
Query: 880 IVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQ 939
+ S+ ++QLDV AFLHG+L E ++M+QP GF++ K VC L K++YGLKQAPR
Sbjct: 956 VSTSKGWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRA 1015
Query: 940 WYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDDMLVAGSNIDEIKNLKIQLS 999
W+ F +F+ GF +D F + LLLYVDD+L+ GS+ +++L L
Sbjct: 1016 WFDTFSNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALK 1075
Query: 1000 KEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTPLASHFR 1059
F MKDLGP + LG+QI D G L L Y +LQ+ M D + TPL
Sbjct: 1076 NRFSMKDLGPPRYFLGIQI-EDYANG-LFLHQTAYATDILQQAGMSDCNPMPTPLPQQL- 1132
Query: 1060 LSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAVGVVSRFMSNPGKAHWEA 1119
EL A+ Y ++ + + TRPDI AV + + M +P + +
Sbjct: 1133 --------DNLNSELFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGL 1184
Query: 1120 VKWILRYLRGTTEKCLYFGKGE-IKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMS 1178
+K ILRY++GT L + + + Y D+D AG + RRSTTG+ +G+ +SW +
Sbjct: 1185 LKRILRYIKGTIGMGLPIKRNSTLTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSA 1244
Query: 1179 RIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQE-KSALYSDSQSAIHLAKNSA 1237
+ Q V+ S+TE EY A+T A++E+ W+ LL +LG Q + +Y D+ SA++L+ N A
Sbjct: 1245 KRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPA 1304
Query: 1238 FHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTK 1278
H+R+KH YH+IR + ++ I + AD+ TK
Sbjct: 1305 LHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADVFTK 1345
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.133 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,186,700
Number of Sequences: 26719
Number of extensions: 1363191
Number of successful extensions: 5025
Number of sequences better than 10.0: 201
Number of HSP's better than 10.0 without gapping: 148
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 3941
Number of HSP's gapped (non-prelim): 428
length of query: 1302
length of database: 11,318,596
effective HSP length: 111
effective length of query: 1191
effective length of database: 8,352,787
effective search space: 9948169317
effective search space used: 9948169317
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144760.12


