

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144759.9 - phase: 0
(749 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
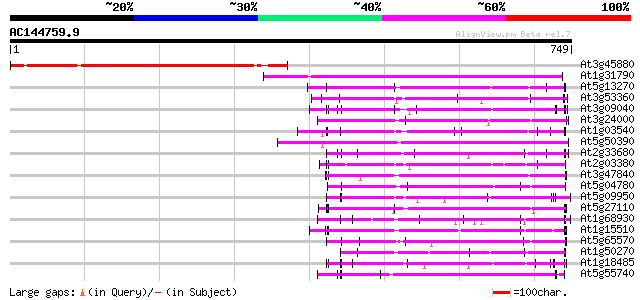
Sequences producing significant alignments: (bits) Value
At3g45880 phospholipase - like protein 452 e-127
At1g31790 unknown protein 320 1e-87
At5g13270 putative protein 157 2e-38
At3g53360 putative protein 154 1e-37
At3g09040 hypothetical protein 151 1e-36
At3g24000 hypothetical protein 150 2e-36
At1g03540 hypothetical protein 145 1e-34
At5g50390 selenium-binding protein-like 140 3e-33
At2g33680 hypothetical protein 140 3e-33
At2g03380 unknown protein 140 3e-33
At3g47840 putative protein 138 1e-32
At5g04780 selenium-binding protein-like 135 8e-32
At5g09950 selenium-binding protein-like 134 1e-31
At5g27110 putative protein 134 2e-31
At1g68930 hypothetical protein 133 4e-31
At1g15510 hypothetical protein 132 5e-31
At5g65570 putative protein 132 7e-31
At1g50270 hypothetical protein 131 1e-30
At1g18485 hypothetical protein 131 2e-30
At5g55740 selenium-binding protein-like 130 4e-30
>At3g45880 phospholipase - like protein
Length = 431
Score = 452 bits (1163), Expect = e-127
Identities = 219/371 (59%), Positives = 271/371 (73%), Gaps = 16/371 (4%)
Query: 1 MNEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSL 60
M ++IE LWREVRELSLG+K I+R +S P+P++F RN+++ +KPC+IS +I+HWP+L L
Sbjct: 1 MAKEIENLWREVRELSLGTK--IDRFDSQPSPVKFLRNYVSQSKPCVISKAITHWPALKL 58
Query: 61 WSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS 120
WS P+YLT +LS VSLHLTP G AD++T S LCFASAHV+ + FPEAL+++ S
Sbjct: 59 WSDPAYLTGALSDDVVSLHLTPNGCADAVT----GDSDLCFASAHVEKVLFPEALKVVQS 114
Query: 121 SNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFH 180
S V Y QQQNDCFR+EY ++ DCD I WATEAFG PEAVNLWIG S T FH
Sbjct: 115 SCKGLKVGYLQQQNDCFRTEYSTVALDCDGDIEWATEAFGCSPEAVNLWIGTDDSVTSFH 174
Query: 181 KDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPW 240
KDHYENLYAVV+G+KHFLL PPTDVHR YI YPAA Y Y+ +T F LE+++P R+VPW
Sbjct: 175 KDHYENLYAVVSGEKHFLLLPPTDVHRLYIEQYPAANYSYHRDTDAFKLEVEEPVRHVPW 234
Query: 241 CSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTI 300
SV+P+PSPE E KFPL+F+GP PF CTVKAGE+LYLPSMWFHHV Q+ DG TI
Sbjct: 235 SSVDPYPSPEKEASERLKFPLFFDGPKPFHCTVKAGEVLYLPSMWFHHVSQTPGDGGYTI 294
Query: 301 AVNYWYDMQFDIKYAYFNFLQSIDYRSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPS 360
AVNYWYDMQFDIKYAYFNFLQS+ Y+S S + P++ + R + D+ S+ + S
Sbjct: 295 AVNYWYDMQFDIKYAYFNFLQSLLYKS--SSLNPVL--------SWREDEDSESSDAEGS 344
Query: 361 NHQPHLLRLPL 371
P L L
Sbjct: 345 KFTPSATNLYL 355
Score = 36.2 bits (82), Expect = 0.069
Identities = 17/39 (43%), Positives = 27/39 (68%)
Query: 648 EQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVF 686
++VH ++IKLG +S+ +Q LI MY + L+ +AE VF
Sbjct: 368 KRVHVNSIKLGFESNGLIQSGLIDMYRKYRLVINAEKVF 406
Score = 31.2 bits (69), Expect = 2.2
Identities = 21/77 (27%), Positives = 33/77 (42%)
Query: 518 DVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCL 577
D G F P + L + + +VH +KLG + LI S LI Y +++ +
Sbjct: 340 DAEGSKFTPSATNLYLSSTYVSDGARFWKRVHVNSIKLGFESNGLIQSGLIDMYRKYRLV 399
Query: 578 EDANMVFNRVSRHNTLT 594
+A VF + LT
Sbjct: 400 INAEKVFRAMEGQTVLT 416
>At1g31790 unknown protein
Length = 409
Score = 320 bits (821), Expect = 1e-87
Identities = 176/402 (43%), Positives = 241/402 (59%), Gaps = 6/402 (1%)
Query: 340 PPAPPTRRRNADTTSTTSPPSNHQPHLLRLPL-RRNPKPKNLSLIHPSSQPITPPKKSKR 398
PP+PP+ + + ST N + L R PK + + QP P+
Sbjct: 9 PPSPPSLVPSFNYNSTARSVGNDVRTNFDVQLFLRKPKHQKSEPVVVIQQPQIQPQNPSS 68
Query: 399 RRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTL 458
R C +TS IL LMD+L P DIY+ L KE D A EL I+ I +T
Sbjct: 69 R--C-STSDILRLMDSLSLPGNEDIYSCLAKESARENDQRGAHELQVHIMKSSIRPTITF 125
Query: 459 LNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLD 518
+NR+L+M VSCG L+ R++FD M RDFHSWA +F+ E G+YE+A +FVSML
Sbjct: 126 INRLLLMHVSCGRLDITRQMFDRMPHRDFHSWAIVFLGCIEMGDYEDAAFLFVSMLKHSQ 185
Query: 519 VMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV--LISSSLIRFYGRFKC 576
F P WI C+LKACA + LG QVH KLG D +S SLIRFYG F+C
Sbjct: 186 KGAFKIPSWILGCVLKACAMIRDFELGKQVHALCHKLGFIDEEDSYLSGSLIRFYGEFRC 245
Query: 577 LEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
LEDAN+V +++S NT+ W AK+ + RE F E + DF +MG G+KK+ FS+VLKA
Sbjct: 246 LEDANLVLHQLSNANTVAWAAKVTNDYREGEFQEVIRDFIEMGNHGIKKNVSVFSNVLKA 305
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C + + G G+QVHA+AIKLG +SD ++C LI MYG+ G ++DAE VF+ +++E +V
Sbjct: 306 CSWVSDGGRSGQQVHANAIKLGFESDCLIRCRLIEMYGKYGKVKDAEKVFKSSKDETSVS 365
Query: 697 SLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRI 738
NAM+ Y+QNG+YIEA+K +YQMKA G++ H+ LL + +
Sbjct: 366 CWNAMVASYMQNGIYIEAIKLLYQMKATGIKAHDTLLNEAHL 407
>At5g13270 putative protein
Length = 752
Score = 157 bits (398), Expect = 2e-38
Identities = 98/319 (30%), Positives = 164/319 (50%), Gaps = 7/319 (2%)
Query: 423 IYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM 482
+YT+L+K + ++H +I G+ ++ I+ M+V CG L A+RVFD M
Sbjct: 186 MYTTLLKSLVNPRALDFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGWLVGAKRVFDQM 245
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
+V+ + L V Y + G +A+ +FV ++ + G + +++S +LKACA +
Sbjct: 246 AVKKPVACTGLMVGYTQAGRARDALKLFVDLVTE----GVEWDSFVFSVVLKACASLEEL 301
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
LG Q+H C+ KLG V + + L+ FY + E A F + N ++W+A I
Sbjct: 302 NLGKQIHACVAKLGLESEVSVGTPLVDFYIKCSSFESACRAFQEIREPNDVSWSAIISGY 361
Query: 603 CRERHFSEALGDFKKMGRVGVK-KDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDS 661
C+ F EA+ FK + +SFT++S+ +AC + + + G QVHADAIK L
Sbjct: 362 CQMSQFEEAVKTFKSLRSKNASILNSFTYTSIFQACSVLAD-CNIGGQVHADAIKRSLIG 420
Query: 662 DSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQM 721
Y + +LI MY + G L DA VFE N ++ + A + G+ G EA++ +M
Sbjct: 421 SQYGESALITMYSKCGCLDDANEVFESMDNP-DIVAWTAFISGHAYYGNASEALRLFEKM 479
Query: 722 KAAGVQPHEPLLEKLRIAC 740
+ G++P+ + AC
Sbjct: 480 VSCGMKPNSVTFIAVLTAC 498
Score = 121 bits (304), Expect = 1e-27
Identities = 94/320 (29%), Positives = 151/320 (46%), Gaps = 8/320 (2%)
Query: 398 RRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLT 457
+ RK + L MD ++ Y L + C LH ++ GIE P
Sbjct: 60 KHRKLNEAFEFLQEMDKAGVSVSSYSYQCLFEACRELRSLSHGRLLHDRM-RMGIENPSV 118
Query: 458 LL-NRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQ 516
LL N +L M+ C LE+A ++FD MS + S T+ +Y E G + A+ +F ML
Sbjct: 119 LLQNCVLQMYCECRSLEDADKLFDEMSELNAVSRTTMISAYAEQGILDKAVGLFSGMLAS 178
Query: 517 LDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKC 576
G P +++ LLK+ + G Q+H +++ G C + I + ++ Y +
Sbjct: 179 ----GDKPPSSMYTTLLKSLVNPRALDFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGW 234
Query: 577 LEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
L A VF++++ + T +V + +AL F + GV+ DSF FS VLKA
Sbjct: 235 LVGAKRVFDQMAVKKPVACTGLMVGYTQAGRARDALKLFVDLVTEGVEWDSFVFSVVLKA 294
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C ++ + G+Q+HA KLGL+S+ V L+ Y + A F+ R +V
Sbjct: 295 CASLEEL-NLGKQIHACVAKLGLESEVSVGTPLVDFYIKCSSFESACRAFQEIREPNDV- 352
Query: 697 SLNAMLMGYIQNGLYIEAVK 716
S +A++ GY Q + EAVK
Sbjct: 353 SWSAIISGYCQMSQFEEAVK 372
Score = 80.9 bits (198), Expect = 2e-15
Identities = 58/229 (25%), Positives = 112/229 (48%), Gaps = 2/229 (0%)
Query: 514 LCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGR 573
L ++D G S + + CL +AC ++ G +H + VL+ + +++ Y
Sbjct: 71 LQEMDKAGVSVSSYSYQCLFEACRELRSLSHGRLLHDRMRMGIENPSVLLQNCVLQMYCE 130
Query: 574 FKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSV 633
+ LEDA+ +F+ +S N ++ T I + + +A+G F M G K S ++++
Sbjct: 131 CRSLEDADKLFDEMSELNAVSRTTMISAYAEQGILDKAVGLFSGMLASGDKPPSSMYTTL 190
Query: 634 LKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNER 693
LK+ + G Q+HA I+ GL S++ ++ ++ MY + G L A+ VF+ ++
Sbjct: 191 LKSLVNPRAL-DFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGWLVGAKRVFDQMAVKK 249
Query: 694 NVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
V + +++GY Q G +A+K + GV+ + + AC S
Sbjct: 250 PV-ACTGLMVGYTQAGRARDALKLFVDLVTEGVEWDSFVFSVVLKACAS 297
>At3g53360 putative protein
Length = 768
Score = 154 bits (390), Expect = 1e-37
Identities = 104/324 (32%), Positives = 160/324 (49%), Gaps = 7/324 (2%)
Query: 417 FPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENAR 476
F I + Y SL+ C+ S ++H I+ + L N IL M+ CG L +AR
Sbjct: 63 FKIRLRTYISLICACSSSRSLAQGRKIHDHILNSNCKYDTILNNHILSMYGKCGSLRDAR 122
Query: 477 RVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKAC 536
VFD M R+ S+ ++ Y +NG+ AI +++ ML Q D++ F + ++KAC
Sbjct: 123 EVFDFMPERNLVSYTSVITGYSQNGQGAEAIRLYLKML-QEDLVPDQF---AFGSIIKAC 178
Query: 537 ACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWT 596
A + +V LG Q+H ++KL + H++ ++LI Y RF + DA+ VF + + ++W+
Sbjct: 179 ASSSDVGLGKQLHAQVIKLESSSHLIAQNALIAMYVRFNQMSDASRVFYGIPMKDLISWS 238
Query: 597 AKIVSSCRERHFSEALGDFKKMGRVGV-KKDSFTFSSVLKACGRMQNRGSCGEQVHADAI 655
+ I + EAL K+M GV + + F S LKAC + R G Q+H I
Sbjct: 239 SIIAGFSQLGFEFEALSHLKEMLSFGVFHPNEYIFGSSLKACSSLL-RPDYGSQIHGLCI 297
Query: 656 KLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAV 715
K L ++ CSL MY R G L A VF+ + S N ++ G NG EAV
Sbjct: 298 KSELAGNAIAGCSLCDMYARCGFLNSARRVFDQIERP-DTASWNVIIAGLANNGYADEAV 356
Query: 716 KFVYQMKAAGVQPHEPLLEKLRIA 739
QM+++G P L L A
Sbjct: 357 SVFSQMRSSGFIPDAISLRSLLCA 380
Score = 113 bits (283), Expect = 3e-25
Identities = 92/344 (26%), Positives = 153/344 (43%), Gaps = 17/344 (4%)
Query: 403 DTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRI 462
+ SH+ ++ F I+ S +K C+ P+ ++H I + +
Sbjct: 252 EALSHLKEMLSFGVFHPNEYIFGSSLKACSSLLRPDYGSQIHGLCIKSELAGNAIAGCSL 311
Query: 463 LIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGF 522
M+ CG L +ARRVFD + D SW + NG + A+ VF M
Sbjct: 312 CDMYARCGFLNSARRVFDQIERPDTASWNVIIAGLANNGYADEAVSVFSQMRSS------ 365
Query: 523 SFPPWIWSCLLKACACTMNVPL--GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDA 580
F P S CA T + L GMQ+H ++K G + + +SL+ Y D
Sbjct: 366 GFIPDAISLRSLLCAQTKPMALSQGMQIHSYIIKWGFLADLTVCNSLLTMY---TFCSDL 422
Query: 581 NMVFNRVS--RHN--TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
FN R+N +++W + + + E L FK M + D T ++L+
Sbjct: 423 YCCFNLFEDFRNNADSVSWNTILTACLQHEQPVEMLRLFKLMLVSECEPDHITMGNLLRG 482
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C + + G QVH ++K GL + +++ LI MY + G L A +F+ N R+V
Sbjct: 483 CVEISSL-KLGSQVHCYSLKTGLAPEQFIKNGLIDMYAKCGSLGQARRIFDSMDN-RDVV 540
Query: 697 SLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
S + +++GY Q+G EA+ +MK+AG++P+ + AC
Sbjct: 541 SWSTLIVGYAQSGFGEEALILFKEMKSAGIEPNHVTFVGVLTAC 584
Score = 78.2 bits (191), Expect = 2e-14
Identities = 68/260 (26%), Positives = 116/260 (44%), Gaps = 13/260 (5%)
Query: 441 IELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVF-DVMSVRDFHSWATLFVSYYE 499
+++H+ II G LT+ N +L M+ C L +F D + D SW T+ + +
Sbjct: 391 MQIHSYIIKWGFLADLTVCNSLLTMYTFCSDLYCCFNLFEDFRNNADSVSWNTILTACLQ 450
Query: 500 NGEYENAIDVFVSML---CQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLG 556
+ + + +F ML C+ D + LL+ C ++ LG QVH LK G
Sbjct: 451 HEQPVEMLRLFKLMLVSECEPDHITMGN-------LLRGCVEISSLKLGSQVHCYSLKTG 503
Query: 557 ACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFK 616
I + LI Y + L A +F+ + + ++W+ IV + EAL FK
Sbjct: 504 LAPEQFIKNGLIDMYAKCGSLGQARRIFDSMDNRDVVSWSTLIVGYAQSGFGEEALILFK 563
Query: 617 KMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCS-LIAMYGR 675
+M G++ + TF VL AC + G +++A S + CS ++ + R
Sbjct: 564 EMKSAGIEPNHVTFVGVLTACSHV-GLVEEGLKLYATMQTEHGISPTKEHCSCVVDLLAR 622
Query: 676 SGLLRDAELVFEMTRNERNV 695
+G L +AE + + E +V
Sbjct: 623 AGRLNEAERFIDEMKLEPDV 642
Score = 60.8 bits (146), Expect = 3e-09
Identities = 44/151 (29%), Positives = 72/151 (47%), Gaps = 11/151 (7%)
Query: 599 IVSSCRERHFSEALGDFKKMGRVGVKKDSF-----TFSSVLKACGRMQNRGSCGEQVHAD 653
I S C+ + EAL F K SF T+ S++ AC ++ G ++H
Sbjct: 38 INSLCKSNFYREALEAFD----FAQKNSSFKIRLRTYISLICACSSSRSLAQ-GRKIHDH 92
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIE 713
+ D+ + +++MYG+ G LRDA VF+ ERN+ S +++ GY QNG E
Sbjct: 93 ILNSNCKYDTILNNHILSMYGKCGSLRDAREVFDFMP-ERNLVSYTSVITGYSQNGQGAE 151
Query: 714 AVKFVYQMKAAGVQPHEPLLEKLRIACGSSN 744
A++ +M + P + + AC SS+
Sbjct: 152 AIRLYLKMLQEDLVPDQFAFGSIIKACASSS 182
>At3g09040 hypothetical protein
Length = 1028
Score = 151 bits (382), Expect = 1e-36
Identities = 89/303 (29%), Positives = 155/303 (50%), Gaps = 7/303 (2%)
Query: 438 ETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSY 497
+ + +H + I G+ + + + ++ M+ C +E A +VF+ + ++ W + Y
Sbjct: 344 DLGLVVHAEAIKLGLASNIYVGSSLVSMYSKCEKMEAAAKVFEALEEKNDVFWNAMIRGY 403
Query: 498 YENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGA 557
NGE +++F+ M G++ + ++ LL CA + ++ +G Q H ++K
Sbjct: 404 AHNGESHKVMELFMDMKSS----GYNIDDFTFTSLLSTCAASHDLEMGSQFHSIIIKKKL 459
Query: 558 CDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKK 617
++ + ++L+ Y + LEDA +F R+ + +TW I S ++ + SEA FK+
Sbjct: 460 AKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNVTWNTIIGSYVQDENESEAFDLFKR 519
Query: 618 MGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSG 677
M G+ D +S LKAC + G+QVH ++K GLD D + SLI MY + G
Sbjct: 520 MNLCGIVSDGACLASTLKACTHVHGLYQ-GKQVHCLSVKCGLDRDLHTGSSLIDMYSKCG 578
Query: 678 LLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLR 737
+++DA VF + E +V S+NA++ GY QN L EAV +M GV P E +
Sbjct: 579 IIKDARKVFS-SLPEWSVVSMNALIAGYSQNNLE-EAVVLFQEMLTRGVNPSEITFATIV 636
Query: 738 IAC 740
AC
Sbjct: 637 EAC 639
Score = 137 bits (344), Expect = 3e-32
Identities = 94/312 (30%), Positives = 159/312 (50%), Gaps = 15/312 (4%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
+TSL+ C S D E + H+ II + + L + N ++ M+ CG LE+AR++F+ M
Sbjct: 431 FTSLLSTCAASHDLEMGSQFHSIIIKKKLAKNLFVGNALVDMYAKCGALEDARQIFERMC 490
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSM-LCQLDVMGFSFPPWIWSCL---LKACACT 539
RD +W T+ SY ++ A D+F M LC + G +CL LKAC
Sbjct: 491 DRDNVTWNTIIGSYVQDENESEAFDLFKRMNLCGIVSDG--------ACLASTLKACTHV 542
Query: 540 MNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKI 599
+ G QVH +K G + SSLI Y + ++DA VF+ + + ++ A +
Sbjct: 543 HGLYQGKQVHCLSVKCGLDRDLHTGSSLIDMYSKCGIIKDARKVFSSLPEWSVVSMNA-L 601
Query: 600 VSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGL 659
++ + + EA+ F++M GV TF+++++AC + ++ + G Q H K G
Sbjct: 602 IAGYSQNNLEEAVVLFQEMLTRGVNPSEITFATIVEACHKPESL-TLGTQFHGQITKRGF 660
Query: 660 DSD-SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFV 718
S+ Y+ SL+ MY S + +A +F + +++ M+ G+ QNG Y EA+KF
Sbjct: 661 SSEGEYLGISLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHSQNGFYEEALKFY 720
Query: 719 YQMKAAGVQPHE 730
+M+ GV P +
Sbjct: 721 KEMRHDGVLPDQ 732
Score = 112 bits (280), Expect = 8e-25
Identities = 82/317 (25%), Positives = 150/317 (46%), Gaps = 8/317 (2%)
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVR 485
S +K CT ++H + G++ L + ++ M+ CG++++AR+VF +
Sbjct: 534 STLKACTHVHGLYQGKQVHCLSVKCGLDRDLHTGSSLIDMYSKCGIIKDARKVFSSLPEW 593
Query: 486 DFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLG 545
S L Y +N E A+ +F ML + G + ++ +++AC ++ LG
Sbjct: 594 SVVSMNALIAGYSQNN-LEEAVVLFQEMLTR----GVNPSEITFATIVEACHKPESLTLG 648
Query: 546 MQVHGCLLKLG-ACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLT-WTAKIVSSC 603
Q HG + K G + + + SL+ Y + + +A +F+ +S ++ WT +
Sbjct: 649 TQFHGQITKRGFSSEGEYLGISLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHS 708
Query: 604 RERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDS 663
+ + EAL +K+M GV D TF +VL+ C + + G +H+ L D D
Sbjct: 709 QNGFYEEALKFYKEMRHDGVLPDQATFVTVLRVCSVLSSLRE-GRAIHSLIFHLAHDLDE 767
Query: 664 YVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKA 723
+LI MY + G ++ + VF+ R NV S N+++ GY +NG +A+K M+
Sbjct: 768 LTSNTLIDMYAKCGDMKGSSQVFDEMRRRSNVVSWNSLINGYAKNGYAEDALKIFDSMRQ 827
Query: 724 AGVQPHEPLLEKLRIAC 740
+ + P E + AC
Sbjct: 828 SHIMPDEITFLGVLTAC 844
Score = 104 bits (260), Expect = 2e-22
Identities = 87/340 (25%), Positives = 143/340 (41%), Gaps = 7/340 (2%)
Query: 401 KCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLN 460
KCD S + + + P T+ +T L + PE A+ + ++ G
Sbjct: 207 KCDRISDARRVFEWIVDPNTV-CWTCLFSGYVKAGLPEEAVLVFERMRDEGHRPDHLAFV 265
Query: 461 RILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVM 520
++ ++ G L++AR +F MS D +W + + + G AI+ F +M
Sbjct: 266 TVINTYIRLGKLKDARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKS 325
Query: 521 GFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDA 580
S +L A N+ LG+ VH +KLG ++ + SSL+ Y + + +E A
Sbjct: 326 TRS----TLGSVLSAIGIVANLDLGLVVHAEAIKLGLASNIYVGSSLVSMYSKCEKMEAA 381
Query: 581 NMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRM 640
VF + N + W A I + + F M G D FTF+S+L C
Sbjct: 382 AKVFEALEEKNDVFWNAMIRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSLLSTCAAS 441
Query: 641 QNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNA 700
+ G Q H+ IK L + +V +L+ MY + G L DA +FE + NV + N
Sbjct: 442 HDL-EMGSQFHSIIIKKKLAKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNV-TWNT 499
Query: 701 MLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
++ Y+Q+ EA +M G+ L AC
Sbjct: 500 IIGSYVQDENESEAFDLFKRMNLCGIVSDGACLASTLKAC 539
Score = 97.4 bits (241), Expect = 3e-20
Identities = 84/321 (26%), Positives = 133/321 (41%), Gaps = 41/321 (12%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
++ ++ C T+ E ++H +I G+E ++ M+ C + +ARRVF+ +
Sbjct: 163 FSIVLSTCARETNVEFGRQIHCSMIKMGLERNSYCGGALVDMYAKCDRISDARRVFEWIV 222
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVP 543
+ W LF Y + G E A+ VF M +
Sbjct: 223 DPNTVCWTCLFSGYVKAGLPEEAVLVFERMRDE--------------------------- 255
Query: 544 LGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSC 603
G L ++I Y R L+DA ++F +S + + W I
Sbjct: 256 ------------GHRPDHLAFVTVINTYIRLGKLKDARLLFGEMSSPDVVAWNVMISGHG 303
Query: 604 RERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDS 663
+ + A+ F M + VK T SVL A G + N G VHA+AIKLGL S+
Sbjct: 304 KRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAIGIVANL-DLGLVVHAEAIKLGLASNI 362
Query: 664 YVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKA 723
YV SL++MY + + A VFE E+N NAM+ GY NG + ++ MK+
Sbjct: 363 YVGSSLVSMYSKCEKMEAAAKVFEALE-EKNDVFWNAMIRGYAHNGESHKVMELFMDMKS 421
Query: 724 AGVQPHEPLLEKLRIACGSSN 744
+G + L C +S+
Sbjct: 422 SGYNIDDFTFTSLLSTCAASH 442
Score = 75.5 bits (184), Expect = 1e-13
Identities = 70/301 (23%), Positives = 125/301 (41%), Gaps = 46/301 (15%)
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H++ + GI+ L N I+ ++ C + A + FD + +D +W ++ Y G+
Sbjct: 82 VHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQFDFLE-KDVTAWNSMLSMYSSIGK 140
Query: 503 YENAIDVFVSML-CQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV 561
+ FVS+ Q+ F+F S +L CA NV G Q+H ++K+G +
Sbjct: 141 PGKVLRSFVSLFENQIFPNKFTF-----SIVLSTCARETNVEFGRQIHCSMIKMGLERNS 195
Query: 562 LISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRV 621
+L+ Y + + DA VF + NT+ WT + EA+ F++M
Sbjct: 196 YCGGALVDMYAKCDRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLVFERMRDE 255
Query: 622 GVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRD 681
G + D F +V+ Y R G L+D
Sbjct: 256 GHRPDHLAFVTVINT------------------------------------YIRLGKLKD 279
Query: 682 AELVF-EMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
A L+F EM + +V + N M+ G+ + G A+++ + M+ + V+ L + A
Sbjct: 280 ARLLFGEM--SSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAI 337
Query: 741 G 741
G
Sbjct: 338 G 338
Score = 66.2 bits (160), Expect = 6e-11
Identities = 50/185 (27%), Positives = 82/185 (44%), Gaps = 3/185 (1%)
Query: 544 LGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSC 603
+G VH L LG + ++++ Y + + A F+ + + T W + +
Sbjct: 78 IGKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQFDFLEKDVT-AWNSMLSMYS 136
Query: 604 RERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDS 663
+ L F + + + FTFS VL C R N G Q+H IK+GL+ +S
Sbjct: 137 SIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNV-EFGRQIHCSMIKMGLERNS 195
Query: 664 YVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKA 723
Y +L+ MY + + DA VFE + V + GY++ GL EAV +M+
Sbjct: 196 YCGGALVDMYAKCDRISDARRVFEWIVDPNTV-CWTCLFSGYVKAGLPEEAVLVFERMRD 254
Query: 724 AGVQP 728
G +P
Sbjct: 255 EGHRP 259
Score = 43.5 bits (101), Expect = 4e-04
Identities = 24/91 (26%), Positives = 48/91 (52%), Gaps = 1/91 (1%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
+ ++++ C++ + +H+ I +L N ++ M+ CG ++ + +VFD M
Sbjct: 735 FVTVLRVCSVLSSLREGRAIHSLIFHLAHDLDELTSNTLIDMYAKCGDMKGSSQVFDEMR 794
Query: 484 VR-DFHSWATLFVSYYENGEYENAIDVFVSM 513
R + SW +L Y +NG E+A+ +F SM
Sbjct: 795 RRSNVVSWNSLINGYAKNGYAEDALKIFDSM 825
>At3g24000 hypothetical protein
Length = 633
Score = 150 bits (380), Expect = 2e-36
Identities = 101/335 (30%), Positives = 169/335 (50%), Gaps = 12/335 (3%)
Query: 412 MDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGL 471
++ + P Y +L+K+CT+ +H I+ + + N +L M+ CG
Sbjct: 51 LEGSYIPADRRFYNTLLKKCTVFKLLIQGRIVHAHILQSIFRHDIVMGNTLLNMYAKCGS 110
Query: 472 LENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSC 531
LE AR+VF+ M RDF +W TL Y ++ +A+ F ML G+S + S
Sbjct: 111 LEEARKVFEKMPQRDFVTWTTLISGYSQHDRPCDALLFFNQMLR----FGYSPNEFTLSS 166
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHN 591
++KA A G Q+HG +K G +V + S+L+ Y R+ ++DA +VF+ + N
Sbjct: 167 VIKAAAAERRGCCGHQLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRN 226
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC---GRMQNRGSCGE 648
++W A I R +AL F+ M R G + F+++S+ AC G ++ G+
Sbjct: 227 DVSWNALIAGHARRSGTEKALELFQGMLRDGFRPSHFSYASLFGACSSTGFLEQ----GK 282
Query: 649 QVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQN 708
VHA IK G ++ +L+ MY +SG + DA +F+ +R+V S N++L Y Q+
Sbjct: 283 WVHAYMIKSGEKLVAFAGNTLLDMYAKSGSIHDARKIFDRLA-KRDVVSWNSLLTAYAQH 341
Query: 709 GLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGSS 743
G EAV + +M+ G++P+E + AC S
Sbjct: 342 GFGKEAVWWFEEMRRVGIRPNEISFLSVLTACSHS 376
Score = 106 bits (264), Expect = 5e-23
Identities = 67/217 (30%), Positives = 112/217 (50%), Gaps = 2/217 (0%)
Query: 529 WSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVS 588
++ LLK C + G VH +L+ +++ ++L+ Y + LE+A VF ++
Sbjct: 63 YNTLLKKCTVFKLLIQGRIVHAHILQSIFRHDIVMGNTLLNMYAKCGSLEEARKVFEKMP 122
Query: 589 RHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGE 648
+ + +TWT I + +AL F +M R G + FT SSV+KA + RG CG
Sbjct: 123 QRDFVTWTTLISGYSQHDRPCDALLFFNQMLRFGYSPNEFTLSSVIKAAAA-ERRGCCGH 181
Query: 649 QVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQN 708
Q+H +K G DS+ +V +L+ +Y R GL+ DA+LVF+ + +V S NA++ G+ +
Sbjct: 182 QLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRNDV-SWNALIAGHARR 240
Query: 709 GLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGSSNF 745
+A++ M G +P L AC S+ F
Sbjct: 241 SGTEKALELFQGMLRDGFRPSHFSYASLFGACSSTGF 277
>At1g03540 hypothetical protein
Length = 609
Score = 145 bits (365), Expect = 1e-34
Identities = 104/345 (30%), Positives = 172/345 (49%), Gaps = 13/345 (3%)
Query: 385 PSSQPITPPKKSKRRRKCDT--TSHILPLMDALH---FPITIDIYTSLVKECTLSTDPET 439
PS P K+S+ C + + ++++ H P T +Y SL++ C
Sbjct: 20 PSISSSAPTKQSRILELCKLGQLTEAIRILNSTHSSEIPATPKLYASLLQTCNKVFSFIH 79
Query: 440 AIELHTQIITRGIELPLTLLNRILIMFVSCGL-LENARRVFDVMSVRDFHSWATLFVSYY 498
I+ H ++ G+E + N +L ++ G + RRVFD V+D SW ++ Y
Sbjct: 80 GIQFHAHVVKSGLETDRNVGNSLLSLYFKLGPGMRETRRVFDGRFVKDAISWTSMMSGYV 139
Query: 499 ENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGAC 558
E+ A++VFV M+ G + S +KAC+ V LG HG ++ G
Sbjct: 140 TGKEHVKALEVFVEMVS----FGLDANEFTLSSAVKACSELGEVRLGRCFHGVVITHGFE 195
Query: 559 DHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKM 618
+ ISS+L YG + DA VF+ + + + WTA + + + + EALG F M
Sbjct: 196 WNHFISSTLAYLYGVNREPVDARRVFDEMPEPDVICWTAVLSAFSKNDLYEEALGLFYAM 255
Query: 619 GR-VGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSG 677
R G+ D TF +VL ACG ++ R G+++H I G+ S+ V+ SL+ MYG+ G
Sbjct: 256 HRGKGLVPDGSTFGTVLTACGNLR-RLKQGKEIHGKLITNGIGSNVVVESSLLDMYGKCG 314
Query: 678 LLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
+R+A VF +++N S +A+L GY QNG + +A++ +M+
Sbjct: 315 SVREARQVFN-GMSKKNSVSWSALLGGYCQNGEHEKAIEIFREME 358
Score = 133 bits (334), Expect = 4e-31
Identities = 83/263 (31%), Positives = 139/263 (52%), Gaps = 9/263 (3%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
E+H ++IT GI + + + +L M+ CG + AR+VF+ MS ++ SW+ L Y +NG
Sbjct: 286 EIHGKLITNGIGSNVVVESSLLDMYGKCGSVREARQVFNGMSKKNSVSWSALLGGYCQNG 345
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV 561
E+E AI++F M + D+ F +LKACA V LG ++HG ++ G +V
Sbjct: 346 EHEKAIEIFREME-EKDLYCFG-------TVLKACAGLAAVRLGKEIHGQYVRRGCFGNV 397
Query: 562 LISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRV 621
++ S+LI YG+ C++ A+ V++++S N +TW A + + + EA+ F M +
Sbjct: 398 IVESALIDLYGKSGCIDSASRVYSKMSIRNMITWNAMLSALAQNGRGEEAVSFFNDMVKK 457
Query: 622 GVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRD 681
G+K D +F ++L ACG A G+ + +I + GR+GL +
Sbjct: 458 GIKPDYISFIAILTACGHTGMVDEGRNYFVLMAKSYGIKPGTEHYSCMIDLLGRAGLFEE 517
Query: 682 AELVFEMTRNERNVDSLNAMLMG 704
AE + E RN SL +L+G
Sbjct: 518 AENLLERAEC-RNDASLWGVLLG 539
Score = 130 bits (328), Expect = 2e-30
Identities = 91/319 (28%), Positives = 157/319 (48%), Gaps = 13/319 (4%)
Query: 425 TSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
+S VK C+ + H +IT G E + + + ++ +ARRVFD M
Sbjct: 167 SSAVKACSELGEVRLGRCFHGVVITHGFEWNHFISSTLAYLYGVNREPVDARRVFDEMPE 226
Query: 485 RDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVM--GFSFPPWIWSCLLKACACTMNV 542
D W + ++ +N YE A+ +F +M ++ G +F +L AC +
Sbjct: 227 PDVICWTAVLSAFSKNDLYEEALGLFYAMHRGKGLVPDGSTF-----GTVLTACGNLRRL 281
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
G ++HG L+ G +V++ SSL+ YG+ + +A VFN +S+ N+++W+A +
Sbjct: 282 KQGKEIHGKLITNGIGSNVVVESSLLDMYGKCGSVREARQVFNGMSKKNSVSWSALLGGY 341
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSD 662
C+ +A+ F++M +KD + F +VLKAC + G+++H ++ G +
Sbjct: 342 CQNGEHEKAIEIFREM----EEKDLYCFGTVLKACAGLA-AVRLGKEIHGQYVRRGCFGN 396
Query: 663 SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
V+ +LI +YG+SG + A V+ + RN+ + NAML QNG EAV F M
Sbjct: 397 VIVESALIDLYGKSGCIDSASRVYS-KMSIRNMITWNAMLSALAQNGRGEEAVSFFNDMV 455
Query: 723 AAGVQPHEPLLEKLRIACG 741
G++P + ACG
Sbjct: 456 KKGIKPDYISFIAILTACG 474
Score = 47.8 bits (112), Expect = 2e-05
Identities = 36/148 (24%), Positives = 74/148 (49%), Gaps = 3/148 (2%)
Query: 594 TWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHAD 653
T ++I+ C+ +EA+ + ++S+L+ C ++ + G Q HA
Sbjct: 28 TKQSRILELCKLGQLTEAIRILNSTHSSEIPATPKLYASLLQTCNKVFSFIH-GIQFHAH 86
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGL-LRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYI 712
+K GL++D V SL+++Y + G +R+ VF+ R ++ S +M+ GY+ ++
Sbjct: 87 VVKSGLETDRNVGNSLLSLYFKLGPGMRETRRVFD-GRFVKDAISWTSMMSGYVTGKEHV 145
Query: 713 EAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
+A++ +M + G+ +E L AC
Sbjct: 146 KALEVFVEMVSFGLDANEFTLSSAVKAC 173
Score = 46.2 bits (108), Expect = 7e-05
Identities = 41/182 (22%), Positives = 77/182 (41%), Gaps = 7/182 (3%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
+ +++K C E+H Q + RG + + + ++ ++ G +++A RV+ MS
Sbjct: 365 FGTVLKACAGLAAVRLGKEIHGQYVRRGCFGNVIVESALIDLYGKSGCIDSASRVYSKMS 424
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWI-WSCLLKACACTMNV 542
+R+ +W + + +NG E A+ F M+ + G P +I + +L AC T V
Sbjct: 425 IRNMITWNAMLSALAQNGRGEEAVSFFNDMVKK----GIK-PDYISFIAILTACGHTGMV 479
Query: 543 PLGMQVHGCLLK-LGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVS 601
G + K G S +I GR E+A + R N + ++
Sbjct: 480 DEGRNYFVLMAKSYGIKPGTEHYSCMIDLLGRAGLFEEAENLLERAECRNDASLWGVLLG 539
Query: 602 SC 603
C
Sbjct: 540 PC 541
>At5g50390 selenium-binding protein-like
Length = 701
Score = 140 bits (352), Expect = 3e-33
Identities = 102/393 (25%), Positives = 180/393 (44%), Gaps = 11/393 (2%)
Query: 358 PPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPPKKSKRRRKCDTTSHILPLMDALH- 416
P +P +R+ ++ + K + L S +T + ++ C+ L + L
Sbjct: 57 PKPKLKPEPIRIEVKES-KDQILDDTQISKSGVTICSQIEKLVLCNRFREAFELFEILEI 115
Query: 417 ---FPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLE 473
F + + Y +LV+ C ++ +++ G E ++NRIL+M V CG++
Sbjct: 116 RCSFKVGVSTYDALVEACIRLKSIRCVKRVYGFMMSNGFEPEQYMMNRILLMHVKCGMII 175
Query: 474 NARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLL 533
+ARR+FD + R+ +S+ ++ + G Y A ++F M +L ++ +L
Sbjct: 176 DARRLFDEIPERNLYSYYSIISGFVNFGNYVEAFELFKMMWEELS----DCETHTFAVML 231
Query: 534 KACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTL 593
+A A ++ +G Q+H C LKLG D+ +S LI Y + +EDA F + T+
Sbjct: 232 RASAGLGSIYVGKQLHVCALKLGVVDNTFVSCGLIDMYSKCGDIEDARCAFECMPEKTTV 291
Query: 594 TWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHAD 653
W I + EAL M GV D FT S +++ ++ + +Q HA
Sbjct: 292 AWNNVIAGYALHGYSEEALCLLYDMRDSGVSIDQFTLSIMIRISTKLA-KLELTKQAHAS 350
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIE 713
I+ G +S+ +L+ Y + G + A VF+ +N+ S NA++ GY +G +
Sbjct: 351 LIRNGFESEIVANTALVDFYSKWGRVDTARYVFDKL-PRKNIISWNALMGGYANHGRGTD 409
Query: 714 AVKFVYQMKAAGVQPHEPLLEKLRIACGSSNFS 746
AVK +M AA V P+ + AC S S
Sbjct: 410 AVKLFEKMIAANVAPNHVTFLAVLSACAYSGLS 442
>At2g33680 hypothetical protein
Length = 727
Score = 140 bits (352), Expect = 3e-33
Identities = 83/278 (29%), Positives = 146/278 (51%), Gaps = 4/278 (1%)
Query: 465 MFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSF 524
M+ GL+E+ +VF M R+ ++W+T+ Y G E AI VF L + + S
Sbjct: 162 MYCKAGLVEDGLKVFAYMPERNTYTWSTMVSGYATRGRVEEAIKVFNLFLREKEEGSDS- 220
Query: 525 PPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVF 584
++++ +L + A T+ V LG Q+H +K G V +S++L+ Y + + L +A +F
Sbjct: 221 -DYVFTAVLSSLAATIYVGLGRQIHCITIKNGLLGFVALSNALVTMYSKCESLNEACKMF 279
Query: 585 NRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRG 644
+ N++TW+A + + EA+ F +M G+K +T VL AC +
Sbjct: 280 DSSGDRNSITWSAMVTGYSQNGESLEAVKLFSRMFSAGIKPSEYTIVGVLNACSDICYLE 339
Query: 645 SCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMG 704
G+Q+H+ +KLG + + +L+ MY ++G L DA F+ + ER+V +++ G
Sbjct: 340 E-GKQLHSFLLKLGFERHLFATTALVDMYAKAGCLADARKGFDCLQ-ERDVALWTSLISG 397
Query: 705 YIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
Y+QN EA+ +MK AG+ P++P + + AC S
Sbjct: 398 YVQNSDNEEALILYRRMKTAGIIPNDPTMASVLKACSS 435
Score = 135 bits (340), Expect = 8e-32
Identities = 91/323 (28%), Positives = 151/323 (46%), Gaps = 6/323 (1%)
Query: 423 IYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM 482
++T+++ + ++H I G+ + L N ++ M+ C L A ++FD
Sbjct: 223 VFTAVLSSLAATIYVGLGRQIHCITIKNGLLGFVALSNALVTMYSKCESLNEACKMFDSS 282
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
R+ +W+ + Y +NGE A+ +F M G + +L AC+ +
Sbjct: 283 GDRNSITWSAMVTGYSQNGESLEAVKLFSRMFSA----GIKPSEYTIVGVLNACSDICYL 338
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
G Q+H LLKLG H+ +++L+ Y + CL DA F+ + + WT+ I
Sbjct: 339 EEGKQLHSFLLKLGFERHLFATTALVDMYAKAGCLADARKGFDCLQERDVALWTSLISGY 398
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSD 662
+ EAL +++M G+ + T +SVLKAC + G+QVH IK G +
Sbjct: 399 VQNSDNEEALILYRRMKTAGIIPNDPTMASVLKACSSLATL-ELGKQVHGHTIKHGFGLE 457
Query: 663 SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
+ +L MY + G L D LVF T N ++V S NAM+ G NG EA++ +M
Sbjct: 458 VPIGSALSTMYSKCGSLEDGNLVFRRTPN-KDVVSWNAMISGLSHNGQGDEALELFEEML 516
Query: 723 AAGVQPHEPLLEKLRIACGSSNF 745
A G++P + + AC F
Sbjct: 517 AEGMEPDDVTFVNIISACSHKGF 539
Score = 108 bits (270), Expect = 1e-23
Identities = 77/300 (25%), Positives = 139/300 (45%), Gaps = 5/300 (1%)
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H QII G + N ++ + CG L A +F+ + +D SW +L Y +NG
Sbjct: 36 VHGQIIRTGASTCIQHANVLVNFYAKCGKLAKAHSIFNAIICKDVVSWNSLITGYSQNGG 95
Query: 503 YENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVL 562
++ V + + ++ + + + KA + + +G Q H ++K+ + +
Sbjct: 96 ISSSYTV-MQLFREMRAQDILPNAYTLAGIFKAESSLQSSTVGRQAHALVVKMSSFGDIY 154
Query: 563 ISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVG 622
+ +SL+ Y + +ED VF + NT TW+ + EA+ F R
Sbjct: 155 VDTSLVGMYCKAGLVEDGLKVFAYMPERNTYTWSTMVSGYATRGRVEEAIKVFNLFLREK 214
Query: 623 VK--KDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLR 680
+ + F++VL + G G Q+H IK GL + +L+ MY + L
Sbjct: 215 EEGSDSDYVFTAVLSSLAATIYVG-LGRQIHCITIKNGLLGFVALSNALVTMYSKCESLN 273
Query: 681 DAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
+A +F+ + +RN + +AM+ GY QNG +EAVK +M +AG++P E + + AC
Sbjct: 274 EACKMFD-SSGDRNSITWSAMVTGYSQNGESLEAVKLFSRMFSAGIKPSEYTIVGVLNAC 332
Score = 103 bits (257), Expect = 4e-22
Identities = 73/252 (28%), Positives = 116/252 (45%), Gaps = 8/252 (3%)
Query: 438 ETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSY 497
E +LH+ ++ G E L ++ M+ G L +AR+ FD + RD W +L Y
Sbjct: 339 EEGKQLHSFLLKLGFERHLFATTALVDMYAKAGCLADARKGFDCLQERDVALWTSLISGY 398
Query: 498 YENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGA 557
+N + E A+ ++ M G + +LKAC+ + LG QVHG +K G
Sbjct: 399 VQNSDNEEALILYRRM----KTAGIIPNDPTMASVLKACSSLATLELGKQVHGHTIKHGF 454
Query: 558 CDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKK 617
V I S+L Y + LED N+VF R + ++W A I EAL F++
Sbjct: 455 GLEVPIGSALSTMYSKCGSLEDGNLVFRRTPNKDVVSWNAMISGLSHNGQGDEALELFEE 514
Query: 618 MGRVGVKKDSFTFSSVLKACGR--MQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGR 675
M G++ D TF +++ AC RG + +D I L D Y ++ + R
Sbjct: 515 MLAEGMEPDDVTFVNIISACSHKGFVERGWFYFNMMSDQIGLDPKVDHY--ACMVDLLSR 572
Query: 676 SGLLRDAELVFE 687
+G L++A+ E
Sbjct: 573 AGQLKEAKEFIE 584
Score = 79.7 bits (195), Expect = 5e-15
Identities = 50/179 (27%), Positives = 90/179 (49%), Gaps = 5/179 (2%)
Query: 541 NVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIV 600
N+ G VHG +++ GA + ++ L+ FY + L A+ +FN + + ++W + I
Sbjct: 29 NLVAGRAVHGQIIRTGASTCIQHANVLVNFYAKCGKLAKAHSIFNAIICKDVVSWNSLIT 88
Query: 601 SSCRERHFSEA---LGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKL 657
+ S + + F++M + +++T + + KA +Q+ + G Q HA +K+
Sbjct: 89 GYSQNGGISSSYTVMQLFREMRAQDILPNAYTLAGIFKAESSLQS-STVGRQAHALVVKM 147
Query: 658 GLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVK 716
D YV SL+ MY ++GL+ D VF ERN + + M+ GY G EA+K
Sbjct: 148 SSFGDIYVDTSLVGMYCKAGLVEDGLKVFAY-MPERNTYTWSTMVSGYATRGRVEEAIK 205
>At2g03380 unknown protein
Length = 689
Score = 140 bits (352), Expect = 3e-33
Identities = 94/320 (29%), Positives = 167/320 (51%), Gaps = 8/320 (2%)
Query: 423 IYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM 482
+++ +K CT D + ++H Q++ + +L +L M+ CG +++A +VF+ +
Sbjct: 144 VFSKALKACTELQDLDNGKKIHCQLV-KVPSFDNVVLTGLLDMYAKCGEIKSAHKVFNDI 202
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
++R+ W ++ Y +N E + +F M + +V+G + + L+ AC +
Sbjct: 203 TLRNVVCWTSMIAGYVKNDLCEEGLVLFNRMR-ENNVLGNEYT---YGTLIMACTKLSAL 258
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
G HGCL+K G + +SL+ Y + + +A VFN S + + WTA IV
Sbjct: 259 HQGKWFHGCLVKSGIELSSCLVTSLLDMYVKCGDISNARRVFNEHSHVDLVMWTAMIVGY 318
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSD 662
+EAL F+KM V +K + T +SVL CG ++N G VH +IK+G+ D
Sbjct: 319 THNGSVNEALSLFQKMKGVEIKPNCVTIASVLSGCGLIENL-ELGRSVHGLSIKVGI-WD 376
Query: 663 SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
+ V +L+ MY + RDA+ VFEM +E+++ + N+++ G+ QNG EA+ ++M
Sbjct: 377 TNVANALVHMYAKCYQNRDAKYVFEM-ESEKDIVAWNSIISGFSQNGSIHEALFLFHRMN 435
Query: 723 AAGVQPHEPLLEKLRIACGS 742
+ V P+ + L AC S
Sbjct: 436 SESVTPNGVTVASLFSACAS 455
Score = 129 bits (323), Expect = 8e-30
Identities = 93/325 (28%), Positives = 150/325 (45%), Gaps = 19/325 (5%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
Y +L+ CT + H ++ GIEL L+ +L M+V CG + NARRVF+ S
Sbjct: 245 YGTLIMACTKLSALHQGKWFHGCLVKSGIELSSCLVTSLLDMYVKCGDISNARRVFNEHS 304
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCL-----LKACAC 538
D W + V Y NG A+ +F M G P +C+ L C
Sbjct: 305 HVDLVMWTAMIVGYTHNGSVNEALSLFQKM------KGVEIKP---NCVTIASVLSGCGL 355
Query: 539 TMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAK 598
N+ LG VHG +K+G D ++++L+ Y + DA VF S + + W +
Sbjct: 356 IENLELGRSVHGLSIKVGIWD-TNVANALVHMYAKCYQNRDAKYVFEMESEKDIVAWNSI 414
Query: 599 IVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLG 658
I + EAL F +M V + T +S+ AC + + + G +HA ++KLG
Sbjct: 415 ISGFSQNGSIHEALFLFHRMNSESVTPNGVTVASLFSACASLGSL-AVGSSLHAYSVKLG 473
Query: 659 L--DSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVK 716
S +V +L+ Y + G + A L+F+ T E+N + +AM+ GY + G I +++
Sbjct: 474 FLASSSVHVGTALLDFYAKCGDPQSARLIFD-TIEEKNTITWSAMIGGYGKQGDTIGSLE 532
Query: 717 FVYQMKAAGVQPHEPLLEKLRIACG 741
+M +P+E + ACG
Sbjct: 533 LFEEMLKKQQKPNESTFTSILSACG 557
Score = 108 bits (271), Expect = 8e-24
Identities = 78/330 (23%), Positives = 159/330 (47%), Gaps = 15/330 (4%)
Query: 414 ALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLE 473
+LH+ + + L K T+ ++ + H + G+ +++ +++ ++ G +
Sbjct: 38 SLHYAASSPCFLLLSK----CTNIDSLRQSHGVLTGNGLMGDISIATKLVSLYGFFGYTK 93
Query: 474 NARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLL 533
+AR VFD + DF+ W + Y N E + ++ ++ GF + ++S L
Sbjct: 94 DARLVFDQIPEPDFYLWKVMLRCYCLNKESVEVVKLYDLLMKH----GFRYDDIVFSKAL 149
Query: 534 KACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTL 593
KAC ++ G ++H L+K+ + D+V++ + L+ Y + ++ A+ VFN ++ N +
Sbjct: 150 KACTELQDLDNGKKIHCQLVKVPSFDNVVL-TGLLDMYAKCGEIKSAHKVFNDITLRNVV 208
Query: 594 TWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHAD 653
WT+ I + E L F +M V + +T+ +++ AC ++ G+ H
Sbjct: 209 CWTSMIAGYVKNDLCEEGLVLFNRMRENNVLGNEYTYGTLIMACTKLSALHQ-GKWFHGC 267
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSL--NAMLMGYIQNGLY 711
+K G++ S + SL+ MY + G + +A VF +VD + AM++GY NG
Sbjct: 268 LVKSGIELSSCLVTSLLDMYVKCGDISNARRVF---NEHSHVDLVMWTAMIVGYTHNGSV 324
Query: 712 IEAVKFVYQMKAAGVQPHEPLLEKLRIACG 741
EA+ +MK ++P+ + + CG
Sbjct: 325 NEALSLFQKMKGVEIKPNCVTIASVLSGCG 354
Score = 79.7 bits (195), Expect = 5e-15
Identities = 72/286 (25%), Positives = 132/286 (45%), Gaps = 17/286 (5%)
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVR 485
S++ C L + E +H I GI + N ++ M+ C +A+ VF++ S +
Sbjct: 348 SVLSGCGLIENLELGRSVHGLSIKVGI-WDTNVANALVHMYAKCYQNRDAKYVFEMESEK 406
Query: 486 DFHSWATLFVSYYENGEYENAIDVFVSMLCQ-LDVMGFSFPPWIWSCLLKACACTMNVPL 544
D +W ++ + +NG A+ +F M + + G + + L ACA ++ +
Sbjct: 407 DIVAWNSIISGFSQNGSIHEALFLFHRMNSESVTPNGVTV-----ASLFSACASLGSLAV 461
Query: 545 GMQVHGCLLKLG--ACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
G +H +KLG A V + ++L+ FY + + A ++F+ + NT+TW+A I
Sbjct: 462 GSSLHAYSVKLGFLASSSVHVGTALLDFYAKCGDPQSARLIFDTIEEKNTITWSAMIGGY 521
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGR--MQNRGSCGEQVHADAIK--LG 658
++ +L F++M + K + TF+S+L ACG M N G++ + K
Sbjct: 522 GKQGDTIGSLELFEEMLKKQQKPNESTFTSILSACGHTGMVNE---GKKYFSSMYKDYNF 578
Query: 659 LDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMG 704
S + C ++ M R+G L A + E + +V A L G
Sbjct: 579 TPSTKHYTC-MVDMLARAGELEQALDIIEKMPIQPDVRCFGAFLHG 623
>At3g47840 putative protein
Length = 706
Score = 138 bits (348), Expect = 1e-32
Identities = 93/326 (28%), Positives = 155/326 (47%), Gaps = 12/326 (3%)
Query: 422 DIYTSLV--KECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVF 479
D YT + K C + +HT +I RG L + N + M+ CG +++ +F
Sbjct: 208 DTYTFAIALKACAGLRQVKYGKAIHTHVIVRGFVTTLCVANSLATMYTECGEMQDGLCLF 267
Query: 480 DVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPP--WIWSCLLKACA 537
+ MS RD SW +L V+Y G+ A++ F+ M PP ++ + ACA
Sbjct: 268 ENMSERDVVSWTSLIVAYKRIGQEVKAVETFIKM------RNSQVPPNEQTFASMFSACA 321
Query: 538 CTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTA 597
+ G Q+H +L LG D + +S+S+++ Y L A+++F + + ++W+
Sbjct: 322 SLSRLVWGEQLHCNVLSLGLNDSLSVSNSMMKMYSTCGNLVSASVLFQGMRCRDIISWST 381
Query: 598 KIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKL 657
I C+ E F M + G K F +S+L G M G QVHA A+
Sbjct: 382 IIGGYCQAGFGEEGFKYFSWMRQSGTKPTDFALASLLSVSGNMAVIEG-GRQVHALALCF 440
Query: 658 GLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKF 717
GL+ +S V+ SLI MY + G +++A ++F T + ++ SL AM+ GY ++G EA+
Sbjct: 441 GLEQNSTVRSSLINMYSKCGSIKEASMIFGETDRD-DIVSLTAMINGYAEHGKSKEAIDL 499
Query: 718 VYQMKAAGVQPHEPLLEKLRIACGSS 743
+ G +P + AC S
Sbjct: 500 FEKSLKVGFRPDSVTFISVLTACTHS 525
Score = 117 bits (294), Expect = 2e-26
Identities = 90/329 (27%), Positives = 155/329 (46%), Gaps = 21/329 (6%)
Query: 426 SLVKEC-TLSTDPETAIELHTQIITRGIELPLTLLNRILIMF---------VSCGLLENA 475
S V+ C T+ T+I L + + I + + N++++ F ++ G L A
Sbjct: 3 SSVRNCGTIQRFCTTSISLLQKPVEENI---VRISNQVMVKFDPNSHLRSLINAGNLRAA 59
Query: 476 RRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPP--WIWSCLL 533
R+VFD M D SW ++ Y + A+ +F +M V+ + P + S +L
Sbjct: 60 RQVFDKMPHGDIVSWTSIIKRYVTANNSDEALILFSAMR----VVDHAVSPDTSVLSVVL 115
Query: 534 KACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTL 593
KAC + N+ G +H +K V + SSL+ Y R ++ + VF+ + N +
Sbjct: 116 KACGQSSNIAYGESLHAYAVKTSLLSSVYVGSSLLDMYKRVGKIDKSCRVFSEMPFRNAV 175
Query: 594 TWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHAD 653
TWTA I + E L F +M R D++TF+ LKAC ++ + G+ +H
Sbjct: 176 TWTAIITGLVHAGRYKEGLTYFSEMSRSEELSDTYTFAIALKACAGLR-QVKYGKAIHTH 234
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIE 713
I G + V SL MY G ++D +FE +ER+V S ++++ Y + G ++
Sbjct: 235 VIVRGFVTTLCVANSLATMYTECGEMQDGLCLFE-NMSERDVVSWTSLIVAYKRIGQEVK 293
Query: 714 AVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
AV+ +M+ + V P+E + AC S
Sbjct: 294 AVETFIKMRNSQVPPNEQTFASMFSACAS 322
Score = 108 bits (271), Expect = 8e-24
Identities = 79/320 (24%), Positives = 148/320 (45%), Gaps = 6/320 (1%)
Query: 423 IYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM 482
+ + ++K C S++ LH + + + + + +L M+ G ++ + RVF M
Sbjct: 110 VLSVVLKACGQSSNIAYGESLHAYAVKTSLLSSVYVGSSLLDMYKRVGKIDKSCRVFSEM 169
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
R+ +W + G Y+ + F M ++ + ++ LKACA V
Sbjct: 170 PFRNAVTWTAIITGLVHAGRYKEGLTYFSEMSRSEELS----DTYTFAIALKACAGLRQV 225
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
G +H ++ G + +++SL Y ++D +F +S + ++WT+ IV+
Sbjct: 226 KYGKAIHTHVIVRGFVTTLCVANSLATMYTECGEMQDGLCLFENMSERDVVSWTSLIVAY 285
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSD 662
R +A+ F KM V + TF+S+ AC + +R GEQ+H + + LGL+
Sbjct: 286 KRIGQEVKAVETFIKMRNSQVPPNEQTFASMFSACASL-SRLVWGEQLHCNVLSLGLNDS 344
Query: 663 SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
V S++ MY G L A ++F+ R R++ S + ++ GY Q G E K+ M+
Sbjct: 345 LSVSNSMMKMYSTCGNLVSASVLFQGMRC-RDIISWSTIIGGYCQAGFGEEGFKYFSWMR 403
Query: 723 AAGVQPHEPLLEKLRIACGS 742
+G +P + L L G+
Sbjct: 404 QSGTKPTDFALASLLSVSGN 423
>At5g04780 selenium-binding protein-like
Length = 864
Score = 135 bits (340), Expect = 8e-32
Identities = 82/298 (27%), Positives = 155/298 (51%), Gaps = 6/298 (2%)
Query: 444 HTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEY 503
H +II +E +TLLN ++ + CG +E AR+VFD M R SW T+ Y N
Sbjct: 76 HGKIIRIDLEGDVTLLNVLINAYSKCGFVELARQVFDGMLERSLVSWNTMIGLYTRNRME 135
Query: 504 ENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLI 563
A+D+F+ M + GF F + S +L AC + ++H +K ++ +
Sbjct: 136 SEALDIFLEMRNE----GFKFSEFTISSVLSACGVNCDALECKKLHCLSVKTCIDLNLYV 191
Query: 564 SSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGV 623
++L+ Y + ++DA VF + +++TW++ + + +++ EAL +++ R+ +
Sbjct: 192 GTALLDLYAKCGMIKDAVQVFESMQDKSSVTWSSMVAGYVQNKNYEEALLLYRRAQRMSL 251
Query: 624 KKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAE 683
+++ FT SSV+ AC + G+Q+HA K G S+ +V S + MY + G LR++
Sbjct: 252 EQNQFTLSSVICACSNLAALIE-GKQMHAVICKSGFGSNVFVASSAVDMYAKCGSLRESY 310
Query: 684 LVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACG 741
++F + E+N++ N ++ G+ ++ E + +M+ G+ P+E L CG
Sbjct: 311 IIFSEVQ-EKNLELWNTIISGFAKHARPKEVMILFEKMQQDGMHPNEVTFSSLLSVCG 367
Score = 93.2 bits (230), Expect = 5e-19
Identities = 61/211 (28%), Positives = 110/211 (51%), Gaps = 2/211 (0%)
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHN 591
+L+ CA V HG ++++ V + + LI Y + +E A VF+ + +
Sbjct: 59 ILQLCARNGAVMEAKACHGKIIRIDLEGDVTLLNVLINAYSKCGFVELARQVFDGMLERS 118
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVH 651
++W I R R SEAL F +M G K FT SSVL ACG + C +++H
Sbjct: 119 LVSWNTMIGLYTRNRMESEALDIFLEMRNEGFKFSEFTISSVLSACGVNCDALEC-KKLH 177
Query: 652 ADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLY 711
++K +D + YV +L+ +Y + G+++DA VFE +++ +V + ++M+ GY+QN Y
Sbjct: 178 CLSVKTCIDLNLYVGTALLDLYAKCGMIKDAVQVFESMQDKSSV-TWSSMVAGYVQNKNY 236
Query: 712 IEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
EA+ + + ++ ++ L + AC +
Sbjct: 237 EEALLLYRRAQRMSLEQNQFTLSSVICACSN 267
Score = 89.7 bits (221), Expect = 5e-18
Identities = 60/258 (23%), Positives = 118/258 (45%), Gaps = 4/258 (1%)
Query: 425 TSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
+S++ C ++ D +LH + I+L L + +L ++ CG++++A +VF+ M
Sbjct: 158 SSVLSACGVNCDALECKKLHCLSVKTCIDLNLYVGTALLDLYAKCGMIKDAVQVFESMQD 217
Query: 485 RDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPL 544
+ +W+++ Y +N YE A+ ++ + M + S ++ AC+ +
Sbjct: 218 KSSVTWSSMVAGYVQNKNYEEALLLY----RRAQRMSLEQNQFTLSSVICACSNLAALIE 273
Query: 545 GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR 604
G Q+H + K G +V ++SS + Y + L ++ ++F+ V N W I +
Sbjct: 274 GKQMHAVICKSGFGSNVFVASSAVDMYAKCGSLRESYIIFSEVQEKNLELWNTIISGFAK 333
Query: 605 ERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSY 664
E + F+KM + G+ + TFSS+L CG GL +
Sbjct: 334 HARPKEVMILFEKMQQDGMHPNEVTFSSLLSVCGHTGLVEEGRRFFKLMRTTYGLSPNVV 393
Query: 665 VQCSLIAMYGRSGLLRDA 682
++ + GR+GLL +A
Sbjct: 394 HYSCMVDILGRAGLLSEA 411
>At5g09950 selenium-binding protein-like
Length = 995
Score = 134 bits (338), Expect = 1e-31
Identities = 89/286 (31%), Positives = 150/286 (52%), Gaps = 7/286 (2%)
Query: 442 ELHTQIITRG-IELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYEN 500
E+H +IT G ++ + + N ++ M+ CG + +ARRVF M+ +D SW ++ +N
Sbjct: 334 EVHGHVITTGLVDFMVGIGNGLVNMYAKCGSIADARRVFYFMTDKDSVSWNSMITGLDQN 393
Query: 501 GEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDH 560
G + A++ + SM + D++ SF L +CA LG Q+HG LKLG +
Sbjct: 394 GCFIEAVERYKSMR-RHDILPGSFT---LISSLSSCASLKWAKLGQQIHGESLKLGIDLN 449
Query: 561 VLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR-ERHFSEALGDFKKMG 619
V +S++L+ Y L + +F+ + H+ ++W + I + R ER EA+ F
Sbjct: 450 VSVSNALMTLYAETGYLNECRKIFSSMPEHDQVSWNSIIGALARSERSLPEAVVCFLNAQ 509
Query: 620 RVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLL 679
R G K + TFSSVL A + + G G+Q+H A+K + ++ + +LIA YG+ G +
Sbjct: 510 RAGQKLNRITFSSVLSAVSSL-SFGELGKQIHGLALKNNIADEATTENALIACYGKCGEM 568
Query: 680 RDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAG 725
E +F R+ + N+M+ GYI N L +A+ V+ M G
Sbjct: 569 DGCEKIFSRMAERRDNVTWNSMISGYIHNELLAKALDLVWFMLQTG 614
Score = 106 bits (265), Expect = 4e-23
Identities = 79/288 (27%), Positives = 136/288 (46%), Gaps = 6/288 (2%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
++H + + GI+L +++ N ++ ++ G L R++F M D SW ++ + +
Sbjct: 436 QIHGESLKLGIDLNVSVSNALMTLYAETGYLNECRKIFSSMPEHDQVSWNSIIGALARS- 494
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV 561
E ++ V G +S +L A + LG Q+HG LK D
Sbjct: 495 --ERSLPEAVVCFLNAQRAGQKLNRITFSSVLSAVSSLSFGELGKQIHGLALKNNIADEA 552
Query: 562 LISSSLIRFYGRFKCLEDANMVFNRVS-RHNTLTWTAKIVSSCRERHFSEALGDFKKMGR 620
++LI YG+ ++ +F+R++ R + +TW + I ++AL M +
Sbjct: 553 TTENALIACYGKCGEMDGCEKIFSRMAERRDNVTWNSMISGYIHNELLAKALDLVWFMLQ 612
Query: 621 VGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLR 680
G + DSF +++VL A + G +VHA +++ L+SD V +L+ MY + G L
Sbjct: 613 TGQRLDSFMYATVLSAFASVATLER-GMEVHACSVRACLESDVVVGSALVDMYSKCGRL- 670
Query: 681 DAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQP 728
D L F T RN S N+M+ GY ++G EA+K MK G P
Sbjct: 671 DYALRFFNTMPVRNSYSWNSMISGYARHGQGEEALKLFETMKLDGQTP 718
Score = 94.4 bits (233), Expect = 2e-19
Identities = 74/286 (25%), Positives = 131/286 (44%), Gaps = 13/286 (4%)
Query: 444 HTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEY 503
H+++ ++ + L N ++ ++ G +AR+VFD M +R+ SWA + Y NGE+
Sbjct: 24 HSRLYKNRLDKDVYLCNNLINAYLETGDSVSARKVFDEMPLRNCVSWACIVSGYSRNGEH 83
Query: 504 ENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKAC--ACTMNVPLGMQVHGCLLKLGACDHV 561
+ A+ M+ + G + + +L+AC ++ + G Q+HG + KL
Sbjct: 84 KEALVFLRDMVKE----GIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYAVDA 139
Query: 562 LISSSLIRFYGRFKCLED---ANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKM 618
++S+ LI Y +KC+ A F + N+++W + I + A F M
Sbjct: 140 VVSNVLISMY--WKCIGSVGYALCAFGDIEVKNSVSWNSIISVYSQAGDQRSAFRIFSSM 197
Query: 619 GRVGVKKDSFTFSS-VLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSG 677
G + +TF S V AC + EQ+ K GL +D +V L++ + +SG
Sbjct: 198 QYDGSRPTEYTFGSLVTTACSLTEPDVRLLEQIMCTIQKSGLLTDLFVGSGLVSAFAKSG 257
Query: 678 LLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKA 723
L A VF RN +LN +++G ++ EA K M +
Sbjct: 258 SLSYARKVFNQMET-RNAVTLNGLMVGLVRQKWGEEATKLFMDMNS 302
Score = 74.3 bits (181), Expect = 2e-13
Identities = 79/315 (25%), Positives = 135/315 (42%), Gaps = 14/315 (4%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSC-GLLENARRVFDVMSVRDFHSWATLFVSYYEN 500
++H + + + N ++ M+ C G + A F + V++ SW ++ Y +
Sbjct: 125 QIHGLMFKLSYAVDAVVSNVLISMYWKCIGSVGYALCAFGDIEVKNSVSWNSIISVYSQA 184
Query: 501 GEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVP---LGMQVHGCLLKLGA 557
G+ +A +F SM Q D S P L AC++ P L Q+ + K G
Sbjct: 185 GDQRSAFRIFSSM--QYDG---SRPTEYTFGSLVTTACSLTEPDVRLLEQIMCTIQKSGL 239
Query: 558 CDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKK 617
+ + S L+ + + L A VFN++ N +T +V R++ EA F
Sbjct: 240 LTDLFVGSGLVSAFAKSGSLSYARKVFNQMETRNAVTLNGLMVGLVRQKWGEEATKLFMD 299
Query: 618 MGR-VGVKKDSFTF--SSVLKACGRMQNRGSCGEQVHADAIKLGL-DSDSYVQCSLIAMY 673
M + V +S+ SS + + G +VH I GL D + L+ MY
Sbjct: 300 MNSMIDVSPESYVILLSSFPEYSLAEEVGLKKGREVHGHVITTGLVDFMVGIGNGLVNMY 359
Query: 674 GRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLL 733
+ G + DA VF ++ +V S N+M+ G QNG +IEAV+ M+ + P L
Sbjct: 360 AKCGSIADARRVFYFMTDKDSV-SWNSMITGLDQNGCFIEAVERYKSMRRHDILPGSFTL 418
Query: 734 EKLRIACGSSNFSSM 748
+C S ++ +
Sbjct: 419 ISSLSSCASLKWAKL 433
Score = 70.5 bits (171), Expect = 3e-12
Identities = 53/216 (24%), Positives = 95/216 (43%), Gaps = 6/216 (2%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
++S++ + + E ++H + I T N ++ + CG ++ ++F M+
Sbjct: 520 FSSVLSAVSSLSFGELGKQIHGLALKNNIADEATTENALIACYGKCGEMDGCEKIFSRMA 579
Query: 484 VR-DFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
R D +W ++ Y N A+D+ ML G ++++ +L A A +
Sbjct: 580 ERRDNVTWNSMISGYIHNELLAKALDLVWFML----QTGQRLDSFMYATVLSAFASVATL 635
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
GM+VH C ++ V++ S+L+ Y + L+ A FN + N+ +W + I
Sbjct: 636 ERGMEVHACSVRACLESDVVVGSALVDMYSKCGRLDYALRFFNTMPVRNSYSWNSMISGY 695
Query: 603 CRERHFSEALGDFKKMGRVG-VKKDSFTFSSVLKAC 637
R EAL F+ M G D TF VL AC
Sbjct: 696 ARHGQGEEALKLFETMKLDGQTPPDHVTFVGVLSAC 731
Score = 33.9 bits (76), Expect = 0.34
Identities = 27/90 (30%), Positives = 42/90 (46%), Gaps = 1/90 (1%)
Query: 651 HADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGL 710
H+ K LD D Y+ +LI Y +G A VF+ RN S ++ GY +NG
Sbjct: 24 HSRLYKNRLDKDVYLCNNLINAYLETGDSVSARKVFD-EMPLRNCVSWACIVSGYSRNGE 82
Query: 711 YIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
+ EA+ F+ M G+ ++ + AC
Sbjct: 83 HKEALVFLRDMVKEGIFSNQYAFVSVLRAC 112
>At5g27110 putative protein
Length = 691
Score = 134 bits (337), Expect = 2e-31
Identities = 92/328 (28%), Positives = 157/328 (47%), Gaps = 13/328 (3%)
Query: 413 DALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLL 472
D+ FP I Y +L +E +HT ++ G + + + ++ M+ L
Sbjct: 106 DSFTFPNVIKAYGALGREFL-------GRMIHTLVVKSGYVCDVVVASSLVGMYAKFNLF 158
Query: 473 ENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCL 532
EN+ +VFD M RD SW T+ +Y++GE E A+++F M + GF +
Sbjct: 159 ENSLQVFDEMPERDVASWNTVISCFYQSGEAEKALELFGRM----ESSGFEPNSVSLTVA 214
Query: 533 LKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNT 592
+ AC+ + + G ++H +K G ++S+L+ YG+ CLE A VF ++ R +
Sbjct: 215 ISACSRLLWLERGKEIHRKCVKKGFELDEYVNSALVDMYGKCDCLEVAREVFQKMPRKSL 274
Query: 593 LTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHA 652
+ W + I + + +M G + T +S+L AC R +N G+ +H
Sbjct: 275 VAWNSMIKGYVAKGDSKSCVEILNRMIIEGTRPSQTTLTSILMACSRSRNL-LHGKFIHG 333
Query: 653 DAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYI 712
I+ +++D YV CSLI +Y + G AE VF T+ + +S N M+ YI G +
Sbjct: 334 YVIRSVVNADIYVNCSLIDLYFKCGEANLAETVFSKTQKD-VAESWNVMISSYISVGNWF 392
Query: 713 EAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
+AV+ QM + GV+P + AC
Sbjct: 393 KAVEVYDQMVSVGVKPDVVTFTSVLPAC 420
Score = 130 bits (326), Expect = 4e-30
Identities = 89/322 (27%), Positives = 160/322 (49%), Gaps = 11/322 (3%)
Query: 426 SLVKECTLSTDPETAIEL-HTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
SL++ECT ST I+L H +I+T G+ + L ++ ++ +C +AR VF+ +
Sbjct: 8 SLLRECTNSTKSLRRIKLVHQRILTLGLRRDVVLCKSLINVYFTCKDHCSARHVFENFDI 67
Query: 485 R-DFHSWATLFVSYYENGEYENAIDVFVSML-CQLDVM-GFSFPPWIWSCLLKACACTMN 541
R D + W +L Y +N + + ++VF +L C + V F+FP ++KA
Sbjct: 68 RSDVYIWNSLMSGYSKNSMFHDTLEVFKRLLNCSICVPDSFTFPN-----VIKAYGALGR 122
Query: 542 VPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVS 601
LG +H ++K G V+++SSL+ Y +F E++ VF+ + + +W I
Sbjct: 123 EFLGRMIHTLVVKSGYVCDVVVASSLVGMYAKFNLFENSLQVFDEMPERDVASWNTVISC 182
Query: 602 SCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDS 661
+ +AL F +M G + +S + + + AC R+ G+++H +K G +
Sbjct: 183 FYQSGEAEKALELFGRMESSGFEPNSVSLTVAISACSRLLWLER-GKEIHRKCVKKGFEL 241
Query: 662 DSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQM 721
D YV +L+ MYG+ L A VF+ + V + N+M+ GY+ G V+ + +M
Sbjct: 242 DEYVNSALVDMYGKCDCLEVAREVFQKMPRKSLV-AWNSMIKGYVAKGDSKSCVEILNRM 300
Query: 722 KAAGVQPHEPLLEKLRIACGSS 743
G +P + L + +AC S
Sbjct: 301 IIEGTRPSQTTLTSILMACSRS 322
Score = 108 bits (271), Expect = 8e-24
Identities = 82/319 (25%), Positives = 154/319 (47%), Gaps = 10/319 (3%)
Query: 425 TSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
T + C+ E E+H + + +G EL + + ++ M+ C LE AR VF M
Sbjct: 212 TVAISACSRLLWLERGKEIHRKCVKKGFELDEYVNSALVDMYGKCDCLEVAREVFQKMPR 271
Query: 485 RDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPL 544
+ +W ++ Y G+ ++ +++ M+ + G + +L AC+ + N+
Sbjct: 272 KSLVAWNSMIKGYVAKGDSKSCVEILNRMIIE----GTRPSQTTLTSILMACSRSRNLLH 327
Query: 545 GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLED--ANMVFNRVSRHNTLTWTAKIVSS 602
G +HG +++ + ++ SLI Y FKC E A VF++ + +W I S
Sbjct: 328 GKFIHGYVIRSVVNADIYVNCSLIDLY--FKCGEANLAETVFSKTQKDVAESWNVMISSY 385
Query: 603 CRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSD 662
++ +A+ + +M VGVK D TF+SVL AC ++ G+Q+H + L++D
Sbjct: 386 ISVGNWFKAVEVYDQMVSVGVKPDVVTFTSVLPACSQLAALEK-GKQIHLSISESRLETD 444
Query: 663 SYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK 722
+ +L+ MY + G ++A +F + +++V S M+ Y +G EA+ +M+
Sbjct: 445 ELLLSALLDMYSKCGNEKEAFRIFN-SIPKKDVVSWTVMISAYGSHGQPREALYQFDEMQ 503
Query: 723 AAGVQPHEPLLEKLRIACG 741
G++P L + ACG
Sbjct: 504 KFGLKPDGVTLLAVLSACG 522
Score = 62.4 bits (150), Expect = 9e-10
Identities = 67/323 (20%), Positives = 127/323 (38%), Gaps = 52/323 (16%)
Query: 425 TSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
TS++ C+ S + +H +I + + + ++ ++ CG A VF
Sbjct: 313 TSILMACSRSRNLLHGKFIHGYVIRSVVNADIYVNCSLIDLYFKCGEANLAETVFSKTQK 372
Query: 485 RDFHSWATLFVSYYENGEYENAIDVF---VSMLCQLDVMGFSFPPWIWSCLLKACACTMN 541
SW + SY G + A++V+ VS+ + DV+ F+ +L AC+
Sbjct: 373 DVAESWNVMISSYISVGNWFKAVEVYDQMVSVGVKPDVVTFT-------SVLPACSQLAA 425
Query: 542 VPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVS 601
+ G Q+H + + L+ S+L+ Y + ++A +FN + + + ++WT I +
Sbjct: 426 LEKGKQIHLSISESRLETDELLLSALLDMYSKCGNEKEAFRIFNSIPKKDVVSWTVMISA 485
Query: 602 SCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDS 661
EAL F +M + G+K D T +VL AC
Sbjct: 486 YGSHGQPREALYQFDEMQKFGLKPDGVTLLAVLSAC------------------------ 521
Query: 662 DSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSL----NAMLMGYIQNGLYIEAVKF 717
G +GL+ + F R++ ++ + + M+ + G +EA +
Sbjct: 522 ------------GHAGLIDEGLKFFSQMRSKYGIEPIIEHYSCMIDILGRAGRLLEAYEI 569
Query: 718 VYQMKAAGVQPHEPLLEKLRIAC 740
+ Q + LL L AC
Sbjct: 570 IQQ--TPETSDNAELLSTLFSAC 590
Score = 43.9 bits (102), Expect = 3e-04
Identities = 24/90 (26%), Positives = 44/90 (48%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
+TS++ C+ E ++H I +E LL+ +L M+ CG + A R+F+ +
Sbjct: 413 FTSVLPACSQLAALEKGKQIHLSISESRLETDELLLSALLDMYSKCGNEKEAFRIFNSIP 472
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSM 513
+D SW + +Y +G+ A+ F M
Sbjct: 473 KKDVVSWTVMISAYGSHGQPREALYQFDEM 502
>At1g68930 hypothetical protein
Length = 743
Score = 133 bits (334), Expect = 4e-31
Identities = 84/283 (29%), Positives = 139/283 (48%), Gaps = 7/283 (2%)
Query: 458 LLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQL 517
+ N ++ ++CG++E+A ++F M +D SWA + +NG + AI+ F M
Sbjct: 207 MYNSLMGGLLACGMIEDALQLFRGME-KDSVSWAAMIKGLAQNGLAKEAIECFREM---- 261
Query: 518 DVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCL 577
V G + + +L AC + G Q+H C+++ DH+ + S+LI Y + KCL
Sbjct: 262 KVQGLKMDQYPFGSVLPACGGLGAINEGKQIHACIIRTNFQDHIYVGSALIDMYCKCKCL 321
Query: 578 EDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC 637
A VF+R+ + N ++WTA +V + EA+ F M R G+ D +T + AC
Sbjct: 322 HYAKTVFDRMKQKNVVSWTAMVVGYGQTGRAEEAVKIFLDMQRSGIDPDHYTLGQAISAC 381
Query: 638 GRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDS 697
+ + G Q H AI GL V SL+ +YG+ G + D+ +F N R+ S
Sbjct: 382 ANVSSLEE-GSQFHGKAITSGLIHYVTVSNSLVTLYGKCGDIDDSTRLFN-EMNVRDAVS 439
Query: 698 LNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
AM+ Y Q G +E ++ +M G++P L + AC
Sbjct: 440 WTAMVSAYAQFGRAVETIQLFDKMVQHGLKPDGVTLTGVISAC 482
Score = 117 bits (294), Expect = 2e-26
Identities = 99/368 (26%), Positives = 160/368 (42%), Gaps = 42/368 (11%)
Query: 411 LMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCG 470
++ AL +P T +Y ++V L A ++ R + L N +L+ + G
Sbjct: 32 IIRALPYPETF-LYNNIVHAYALMKSSTYA----RRVFDRIPQPNLFSWNNLLLAYSKAG 86
Query: 471 LLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWS 530
L+ F+ + RD +W L Y +G A+ + +M+ +
Sbjct: 87 LISEMESTFEKLPDRDGVTWNVLIEGYSLSGLVGAAVKAYNTMMRDFSA---NLTRVTLM 143
Query: 531 CLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRH 590
+LK + +V LG Q+HG ++KLG ++L+ S L+ Y C+ DA VF +
Sbjct: 144 TMLKLSSSNGHVSLGKQIHGQVIKLGFESYLLVGSPLLYMYANVGCISDAKKVFYGLDDR 203
Query: 591 NTL------------------------------TWTAKIVSSCRERHFSEALGDFKKMGR 620
NT+ +W A I + EA+ F++M
Sbjct: 204 NTVMYNSLMGGLLACGMIEDALQLFRGMEKDSVSWAAMIKGLAQNGLAKEAIECFREMKV 263
Query: 621 VGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLR 680
G+K D + F SVL ACG + G+Q+HA I+ YV +LI MY + L
Sbjct: 264 QGLKMDQYPFGSVLPACGGLGAINE-GKQIHACIIRTNFQDHIYVGSALIDMYCKCKCLH 322
Query: 681 DAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
A+ VF+ + ++NV S AM++GY Q G EAVK M+ +G+ P L + AC
Sbjct: 323 YAKTVFDRMK-QKNVVSWTAMVVGYGQTGRAEEAVKIFLDMQRSGIDPDHYTLGQAISAC 381
Query: 741 GSSNFSSM 748
+N SS+
Sbjct: 382 --ANVSSL 387
Score = 95.5 bits (236), Expect = 1e-19
Identities = 62/243 (25%), Positives = 117/243 (47%), Gaps = 8/243 (3%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
++H II + + + + ++ M+ C L A+ VFD M ++ SW + V Y + G
Sbjct: 291 QIHACIIRTNFQDHIYVGSALIDMYCKCKCLHYAKTVFDRMKQKNVVSWTAMVVGYGQTG 350
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV 561
E A+ +F+ M G + + ACA ++ G Q HG + G +V
Sbjct: 351 RAEEAVKIFLDM----QRSGIDPDHYTLGQAISACANVSSLEEGSQFHGKAITSGLIHYV 406
Query: 562 LISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRV 621
+S+SL+ YG+ ++D+ +FN ++ + ++WTA + + + E + F KM +
Sbjct: 407 TVSNSLVTLYGKCGDIDDSTRLFNEMNVRDAVSWTAMVSAYAQFGRAVETIQLFDKMVQH 466
Query: 622 GVKKDSFTFSSVLKACGR--MQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLL 679
G+K D T + V+ AC R + +G ++ + + S + C +I ++ RSG L
Sbjct: 467 GLKPDGVTLTGVISACSRAGLVEKGQRYFKLMTSEYGI-VPSIGHYSC-MIDLFSRSGRL 524
Query: 680 RDA 682
+A
Sbjct: 525 EEA 527
Score = 54.7 bits (130), Expect = 2e-07
Identities = 56/255 (21%), Positives = 97/255 (37%), Gaps = 62/255 (24%)
Query: 548 VHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERH 607
+HG +++ + ++++ Y K A VF+R+ + N +W +++ +
Sbjct: 28 IHGNIIRALPYPETFLYNNIVHAYALMKSSTYARRVFDRIPQPNLFSWNNLLLAYSKAGL 87
Query: 608 FSEALGDFKKM------------------GRVGVKKDSF--------------TFSSVLK 635
SE F+K+ G VG ++ T ++LK
Sbjct: 88 ISEMESTFEKLPDRDGVTWNVLIEGYSLSGLVGAAVKAYNTMMRDFSANLTRVTLMTMLK 147
Query: 636 ACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVF--------- 686
S G+Q+H IKLG +S V L+ MY G + DA+ VF
Sbjct: 148 -LSSSNGHVSLGKQIHGQVIKLGFESYLLVGSPLLYMYANVGCISDAKKVFYGLDDRNTV 206
Query: 687 -------------------EMTRN-ERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGV 726
++ R E++ S AM+ G QNGL EA++ +MK G+
Sbjct: 207 MYNSLMGGLLACGMIEDALQLFRGMEKDSVSWAAMIKGLAQNGLAKEAIECFREMKVQGL 266
Query: 727 QPHEPLLEKLRIACG 741
+ + + ACG
Sbjct: 267 KMDQYPFGSVLPACG 281
Score = 50.8 bits (120), Expect = 3e-06
Identities = 46/199 (23%), Positives = 88/199 (44%), Gaps = 9/199 (4%)
Query: 412 MDALHFPITIDIYT--SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSC 469
+D I D YT + C + E + H + IT G+ +T+ N ++ ++ C
Sbjct: 360 LDMQRSGIDPDHYTLGQAISACANVSSLEEGSQFHGKAITSGLIHYVTVSNSLVTLYGKC 419
Query: 470 GLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIW 529
G ++++ R+F+ M+VRD SW + +Y + G I +F M+ G
Sbjct: 420 GDIDDSTRLFNEMNVRDAVSWTAMVSAYAQFGRAVETIQLFDKMVQH----GLKPDGVTL 475
Query: 530 SCLLKACACTMNVPLGMQVHGCLL-KLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVS 588
+ ++ AC+ V G + + + G + S +I + R LE+A N +
Sbjct: 476 TGVISACSRAGLVEKGQRYFKLMTSEYGIVPSIGHYSCMIDLFSRSGRLEEAMRFINGMP 535
Query: 589 -RHNTLTWTAKIVSSCRER 606
+ + WT ++S+CR +
Sbjct: 536 FPPDAIGWTT-LLSACRNK 553
Score = 31.2 bits (69), Expect = 2.2
Identities = 26/120 (21%), Positives = 50/120 (41%), Gaps = 12/120 (10%)
Query: 441 IELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS-----VRDFHSWATLFV 495
I+L +++ G++ L ++ GL+E +R F +M+ V ++ +
Sbjct: 457 IQLFDKMVQHGLKPDGVTLTGVISACSRAGLVEKGQRYFKLMTSEYGIVPSIGHYSCMID 516
Query: 496 SYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKL 555
+ +G E A+ M D +G W+ LL AC N+ +G L++L
Sbjct: 517 LFSRSGRLEEAMRFINGMPFPPDAIG-------WTTLLSACRNKGNLEIGKWAAESLIEL 569
>At1g15510 hypothetical protein
Length = 866
Score = 132 bits (333), Expect = 5e-31
Identities = 83/319 (26%), Positives = 156/319 (48%), Gaps = 8/319 (2%)
Query: 425 TSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSV 484
TS++ C L D ++H +IT G + +++ N + M+++ G A ++F M
Sbjct: 301 TSVISACELLGDRRLGRDIHAYVITTGFAVDISVCNSLTQMYLNAGSWREAEKLFSRMER 360
Query: 485 RDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPL 544
+D SW T+ Y N + AID + M D + +L ACA ++
Sbjct: 361 KDIVSWTTMISGYEYNFLPDKAIDTYRMM----DQDSVKPDEITVAAVLSACATLGDLDT 416
Query: 545 GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR 604
G+++H +K +V+++++LI Y + KC++ A +F+ + R N ++WT+ I
Sbjct: 417 GVELHKLAIKARLISYVIVANNLINMYSKCKCIDKALDIFHNIPRKNVISWTSIIAGLRL 476
Query: 605 ERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSY 664
EAL ++M ++ ++ ++ T ++ L AC R+ CG+++HA ++ G+ D +
Sbjct: 477 NNRCFEALIFLRQM-KMTLQPNAITLTAALAACARI-GALMCGKEIHAHVLRTGVGLDDF 534
Query: 665 VQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAA 724
+ +L+ MY R G + A F +++V S N +L GY + G V+ +M +
Sbjct: 535 LPNALLDMYVRCGRMNTAWSQF--NSQKKDVTSWNILLTGYSERGQGSMVVELFDRMVKS 592
Query: 725 GVQPHEPLLEKLRIACGSS 743
V+P E L C S
Sbjct: 593 RVRPDEITFISLLCGCSKS 611
Score = 122 bits (306), Expect = 7e-28
Identities = 92/325 (28%), Positives = 159/325 (48%), Gaps = 17/325 (5%)
Query: 422 DIYT--SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVF 479
D+YT +++ C D E+H ++ G EL + ++N ++ M+V CG +++AR +F
Sbjct: 195 DVYTFPCVLRTCGGIPDLARGKEVHVHVVRYGYELDIDVVNALITMYVKCGDVKSARLLF 254
Query: 480 DVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIW--SCLLKACA 537
D M RD SW + Y+ENG +++F +M G S P + + ++ AC
Sbjct: 255 DRMPRRDIISWNAMISGYFENGMCHEGLELFFAM------RGLSVDPDLMTLTSVISACE 308
Query: 538 CTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTA 597
+ LG +H ++ G + + +SL + Y +A +F+R+ R + ++WT
Sbjct: 309 LLGDRRLGRDIHAYVITTGFAVDISVCNSLTQMYLNAGSWREAEKLFSRMERKDIVSWTT 368
Query: 598 KIVSSCRERHF--SEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAI 655
I S E +F +A+ ++ M + VK D T ++VL AC + + + G ++H AI
Sbjct: 369 MI--SGYEYNFLPDKAIDTYRMMDQDSVKPDEITVAAVLSACATLGDLDT-GVELHKLAI 425
Query: 656 KLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAV 715
K L S V +LI MY + + A +F +NV S +++ G N EA+
Sbjct: 426 KARLISYVIVANNLINMYSKCKCIDKALDIFH-NIPRKNVISWTSIIAGLRLNNRCFEAL 484
Query: 716 KFVYQMKAAGVQPHEPLLEKLRIAC 740
F+ QMK +QP+ L AC
Sbjct: 485 IFLRQMKMT-LQPNAITLTAALAAC 508
Score = 118 bits (295), Expect = 1e-26
Identities = 95/343 (27%), Positives = 160/343 (45%), Gaps = 7/343 (2%)
Query: 401 KCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLN 460
K + +L M L + D++ +LV+ C E ++++ ++ L + L N
Sbjct: 74 KLEEAMKLLNSMQELRVAVDEDVFVALVRLCEWKRAQEEGSKVYSIALSSMSSLGVELGN 133
Query: 461 RILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVM 520
L MFV G L +A VF MS R+ SW L Y + G ++ A+ ++ ML V
Sbjct: 134 AFLAMFVRFGNLVDAWYVFGKMSERNLFSWNVLVGGYAKQGYFDEAMCLYHRMLW---VG 190
Query: 521 GFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDA 580
G + + C+L+ C ++ G +VH +++ G + + ++LI Y + ++ A
Sbjct: 191 GVKPDVYTFPCVLRTCGGIPDLARGKEVHVHVVRYGYELDIDVVNALITMYVKCGDVKSA 250
Query: 581 NMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRM 640
++F+R+ R + ++W A I E L F M + V D T +SV+ AC +
Sbjct: 251 RLLFDRMPRRDIISWNAMISGYFENGMCHEGLELFFAMRGLSVDPDLMTLTSVISACELL 310
Query: 641 QNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNER-NVDSLN 699
+R G +HA I G D V SL MY +G R+AE +F +R ER ++ S
Sbjct: 311 GDR-RLGRDIHAYVITTGFAVDISVCNSLTQMYLNAGSWREAEKLF--SRMERKDIVSWT 367
Query: 700 AMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
M+ GY N L +A+ M V+P E + + AC +
Sbjct: 368 TMISGYEYNFLPDKAIDTYRMMDQDSVKPDEITVAAVLSACAT 410
Score = 54.7 bits (130), Expect = 2e-07
Identities = 52/257 (20%), Positives = 102/257 (39%), Gaps = 7/257 (2%)
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVR 485
+++ C D +T +ELH I + + + N ++ M+ C ++ A +F + +
Sbjct: 403 AVLSACATLGDLDTGVELHKLAIKARLISYVIVANNLINMYSKCKCIDKALDIFHNIPRK 462
Query: 486 DFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLG 545
+ SW ++ N A+ M L + + L ACA + G
Sbjct: 463 NVISWTSIIAGLRLNNRCFEALIFLRQMKMTLQPNAITL-----TAALAACARIGALMCG 517
Query: 546 MQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRE 605
++H +L+ G + ++L+ Y R + A FN + T +W +
Sbjct: 518 KEIHAHVLRTGVGLDDFLPNALLDMYVRCGRMNTAWSQFNSQKKDVT-SWNILLTGYSER 576
Query: 606 RHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYV 665
S + F +M + V+ D TF S+L C + Q G + G+ +
Sbjct: 577 GQGSMVVELFDRMVKSRVRPDEITFISLLCGCSKSQMVRQ-GLMYFSKMEDYGVTPNLKH 635
Query: 666 QCSLIAMYGRSGLLRDA 682
++ + GR+G L++A
Sbjct: 636 YACVVDLLGRAGELQEA 652
>At5g65570 putative protein
Length = 738
Score = 132 bits (332), Expect = 7e-31
Identities = 91/301 (30%), Positives = 145/301 (47%), Gaps = 7/301 (2%)
Query: 444 HTQIITRGIELPLTLLNRILI-MFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
H + G+E+ + L+ M+V G A+ V D + +D L V Y + GE
Sbjct: 188 HGLAVILGLEVSNVFVGSALVDMYVKFGKTREAKLVLDRVEEKDVVLITALIVGYSQKGE 247
Query: 503 YENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVL 562
A+ F SML V + ++ +L +C ++ G +HG ++K G +
Sbjct: 248 DTEAVKAFQSML----VEKVQPNEYTYASVLISCGNLKDIGNGKLIHGLMVKSGFESALA 303
Query: 563 ISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVG 622
+SL+ Y R ++D+ VF + N ++WT+ I + AL +F+KM R
Sbjct: 304 SQTSLLTMYLRCSLVDDSLRVFKCIEYPNQVSWTSLISGLVQNGREEMALIEFRKMMRDS 363
Query: 623 VKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDA 682
+K +SFT SS L+ C + G Q+H K G D D Y LI +YG+ G A
Sbjct: 364 IKPNSFTLSSALRGCSNLAMFEE-GRQIHGIVTKYGFDRDKYAGSGLIDLYGKCGCSDMA 422
Query: 683 ELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
LVF+ T +E +V SLN M+ Y QNG EA+ +M G+QP++ + + +AC +
Sbjct: 423 RLVFD-TLSEVDVISLNTMIYSYAQNGFGREALDLFERMINLGLQPNDVTVLSVLLACNN 481
Query: 743 S 743
S
Sbjct: 482 S 482
Score = 93.6 bits (231), Expect = 4e-19
Identities = 73/277 (26%), Positives = 137/277 (49%), Gaps = 7/277 (2%)
Query: 467 VSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPP 526
+ CG ++ AR+VFD MS R +W +L ++ + A++++ M+ +V+
Sbjct: 110 LKCGDIDYARQVFDGMSERHIVTWNSLIAYLIKHRRSKEAVEMYRLMITN-NVLP---DE 165
Query: 527 WIWSCLLKACACTMNVPLGMQVHGCLLKLG-ACDHVLISSSLIRFYGRFKCLEDANMVFN 585
+ S + KA + + HG + LG +V + S+L+ Y +F +A +V +
Sbjct: 166 YTLSSVFKAFSDLSLEKEAQRSHGLAVILGLEVSNVFVGSALVDMYVKFGKTREAKLVLD 225
Query: 586 RVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGS 645
RV + + TA IV ++ +EA+ F+ M V+ + +T++SVL +CG +++ G+
Sbjct: 226 RVEEKDVVLITALIVGYSQKGEDTEAVKAFQSMLVEKVQPNEYTYASVLISCGNLKDIGN 285
Query: 646 CGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGY 705
G+ +H +K G +S Q SL+ MY R L+ D+ VF+ V S +++ G
Sbjct: 286 -GKLIHGLMVKSGFESALASQTSLLTMYLRCSLVDDSLRVFKCIEYPNQV-SWTSLISGL 343
Query: 706 IQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
+QNG A+ +M ++P+ L C +
Sbjct: 344 VQNGREEMALIEFRKMMRDSIKPNSFTLSSALRGCSN 380
Score = 89.0 bits (219), Expect = 9e-18
Identities = 73/266 (27%), Positives = 120/266 (44%), Gaps = 13/266 (4%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
Y S++ C D +H ++ G E L +L M++ C L++++ RVF +
Sbjct: 270 YASVLISCGNLKDIGNGKLIHGLMVKSGFESALASQTSLLTMYLRCSLVDDSLRVFKCIE 329
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPP--WIWSCLLKACACTMN 541
+ SW +L +NG E A+ F M M S P + S L+ C+
Sbjct: 330 YPNQVSWTSLISGLVQNGREEMALIEFRKM------MRDSIKPNSFTLSSALRGCSNLAM 383
Query: 542 VPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVS 601
G Q+HG + K G S LI YG+ C + A +VF+ +S + ++ I S
Sbjct: 384 FEEGRQIHGIVTKYGFDRDKYAGSGLIDLYGKCGCSDMARLVFDTLSEVDVISLNTMIYS 443
Query: 602 SCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC--GRMQNRGSCGEQVHADAIKLGL 659
+ EAL F++M +G++ + T SVL AC R+ G C K+ L
Sbjct: 444 YAQNGFGREALDLFERMINLGLQPNDVTVLSVLLACNNSRLVEEG-CELFDSFRKDKIML 502
Query: 660 DSDSYVQCSLIAMYGRSGLLRDAELV 685
+D Y ++ + GR+G L +AE++
Sbjct: 503 TNDHY--ACMVDLLGRAGRLEEAEML 526
Score = 79.0 bits (193), Expect = 9e-15
Identities = 65/221 (29%), Positives = 109/221 (48%), Gaps = 16/221 (7%)
Query: 529 WSCLLKACACTMNVPLGMQVHGCLLKLGACDHV----LISSSLIRFYGRFKC--LEDANM 582
+S LL+ C ++ + +LK G + L+ +SL KC ++ A
Sbjct: 68 FSQLLRQCIDERSISGIKTIQAHMLKSGFPAEISGSKLVDASL-------KCGDIDYARQ 120
Query: 583 VFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQN 642
VF+ +S + +TW + I + R EA+ ++ M V D +T SSV KA +
Sbjct: 121 VFDGMSERHIVTWNSLIAYLIKHRRSKEAVEMYRLMITNNVLPDEYTLSSVFKAFSDLSL 180
Query: 643 RGSCGEQVHADAIKLGLD-SDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAM 701
++ H A+ LGL+ S+ +V +L+ MY + G R+A+LV + E++V + A+
Sbjct: 181 EKE-AQRSHGLAVILGLEVSNVFVGSALVDMYVKFGKTREAKLVLDRV-EEKDVVLITAL 238
Query: 702 LMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
++GY Q G EAVK M VQP+E + I+CG+
Sbjct: 239 IVGYSQKGEDTEAVKAFQSMLVEKVQPNEYTYASVLISCGN 279
>At1g50270 hypothetical protein
Length = 596
Score = 131 bits (330), Expect = 1e-30
Identities = 92/301 (30%), Positives = 145/301 (47%), Gaps = 9/301 (2%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
+ H I+ G++ + N ++ + S GL + A R+FD +D +W + + NG
Sbjct: 124 QFHAHIVKFGLDSDPFVRNSLISGYSSSGLFDFASRLFDGAEDKDVVTWTAMIDGFVRNG 183
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGA--CD 559
A+ FV M G + +LKA +V G VHG L+ G CD
Sbjct: 184 SASEAMVYFVEM----KKTGVAANEMTVVSVLKAAGKVEDVRFGRSVHGLYLETGRVKCD 239
Query: 560 HVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMG 619
V I SSL+ YG+ C +DA VF+ + N +TWTA I + R F + + F++M
Sbjct: 240 -VFIGSSLVDMYGKCSCYDDAQKVFDEMPSRNVVTWTALIAGYVQSRCFDKGMLVFEEML 298
Query: 620 RVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLL 679
+ V + T SSVL AC + G +VH IK ++ ++ +LI +Y + G L
Sbjct: 299 KSDVAPNEKTLSSVLSACAHV-GALHRGRRVHCYMIKNSIEINTTAGTTLIDLYVKCGCL 357
Query: 680 RDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIA 739
+A LVFE +E+NV + AM+ G+ +G +A Y M ++ V P+E + A
Sbjct: 358 EEAILVFERL-HEKNVYTWTAMINGFAAHGYARDAFDLFYTMLSSHVSPNEVTFMAVLSA 416
Query: 740 C 740
C
Sbjct: 417 C 417
Score = 80.1 bits (196), Expect = 4e-15
Identities = 63/225 (28%), Positives = 102/225 (45%), Gaps = 8/225 (3%)
Query: 465 MFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSF 524
M+ C ++A++VFD M R+ +W L Y ++ ++ + VF ML + DV
Sbjct: 249 MYGKCSCYDDAQKVFDEMPSRNVVTWTALIAGYVQSRCFDKGMLVFEEML-KSDVAPNEK 307
Query: 525 PPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVF 584
S +L ACA + G +VH ++K + ++LI Y + CLE+A +VF
Sbjct: 308 T---LSSVLSACAHVGALHRGRRVHCYMIKNSIEINTTAGTTLIDLYVKCGCLEEAILVF 364
Query: 585 NRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC--GRMQN 642
R+ N TWTA I + +A F M V + TF +VL AC G +
Sbjct: 365 ERLHEKNVYTWTAMINGFAAHGYARDAFDLFYTMLSSHVSPNEVTFMAVLSACAHGGLVE 424
Query: 643 RGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFE 687
G + +D Y ++ ++GR GLL +A+ + E
Sbjct: 425 EGRRLFLSMKGRFNMEPKADHYA--CMVDLFGRKGLLEEAKALIE 467
Score = 67.8 bits (164), Expect = 2e-11
Identities = 45/127 (35%), Positives = 70/127 (54%), Gaps = 3/127 (2%)
Query: 615 FKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYG 674
++ M R GV TF +LKA ++ R S Q HA +K GLDSD +V+ SLI+ Y
Sbjct: 92 YRHMRRNGVIPSRHTFPPLLKAVFKL--RDSNPFQFHAHIVKFGLDSDPFVRNSLISGYS 149
Query: 675 RSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLE 734
SGL A +F+ +++V + AM+ G+++NG EA+ + +MK GV +E +
Sbjct: 150 SSGLFDFASRLFDGAE-DKDVVTWTAMIDGFVRNGSASEAMVYFVEMKKTGVAANEMTVV 208
Query: 735 KLRIACG 741
+ A G
Sbjct: 209 SVLKAAG 215
>At1g18485 hypothetical protein
Length = 970
Score = 131 bits (329), Expect = 2e-30
Identities = 81/273 (29%), Positives = 142/273 (51%), Gaps = 11/273 (4%)
Query: 458 LLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQL 517
L RI+ M+ CG +++R VFD + ++ W + SY N Y+ ++ F+ M+
Sbjct: 122 LCTRIITMYAMCGSPDDSRFVFDALRSKNLFQWNAVISSYSRNELYDEVLETFIEMISTT 181
Query: 518 DVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCL 577
D++ F + C++KACA +V +G+ VHG ++K G + V + ++L+ FYG +
Sbjct: 182 DLLPDHFT---YPCVIKACAGMSDVGIGLAVHGLVVKTGLVEDVFVGNALVSFYGTHGFV 238
Query: 578 EDANMVFNRVSRHNTLTWTAKIVSSCRERHFSE----ALGD-FKKMGRVGVKKDSFTFSS 632
DA +F+ + N ++W + ++ + FSE LG+ ++ G D T +
Sbjct: 239 TDALQLFDIMPERNLVSWNS-MIRVFSDNGFSEESFLLLGEMMEENGDGAFMPDVATLVT 297
Query: 633 VLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNE 692
VL C R + G G+ VH A+KL LD + + +L+ MY + G + +A+++F+M N
Sbjct: 298 VLPVCAREREIG-LGKGVHGWAVKLRLDKELVLNNALMDMYSKCGCITNAQMIFKM-NNN 355
Query: 693 RNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAG 725
+NV S N M+ G+ G + QM A G
Sbjct: 356 KNVVSWNTMVGGFSAEGDTHGTFDVLRQMLAGG 388
Score = 108 bits (270), Expect = 1e-23
Identities = 76/302 (25%), Positives = 140/302 (46%), Gaps = 6/302 (1%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
ELH + + + N + + CG L A+RVF + + +SW L + ++
Sbjct: 416 ELHCYSLKQEFVYNELVANAFVASYAKCGSLSYAQRVFHGIRSKTVNSWNALIGGHAQSN 475
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV 561
+ ++D + Q+ + G + LL AC+ ++ LG +VHG +++ +
Sbjct: 476 DPRLSLDAHL----QMKISGLLPDSFTVCSLLSACSKLKSLRLGKEVHGFIIRNWLERDL 531
Query: 562 LISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRV 621
+ S++ Y L +F+ + + ++W I + ALG F++M
Sbjct: 532 FVYLSVLSLYIHCGELCTVQALFDAMEDKSLVSWNTVITGYLQNGFPDRALGVFRQMVLY 591
Query: 622 GVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRD 681
G++ + V AC + + G + HA A+K L+ D+++ CSLI MY ++G +
Sbjct: 592 GIQLCGISMMPVFGACSLLPSL-RLGREAHAYALKHLLEDDAFIACSLIDMYAKNGSITQ 650
Query: 682 AELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACG 741
+ VF + E++ S NAM+MGY +GL EA+K +M+ G P + + AC
Sbjct: 651 SSKVFNGLK-EKSTASWNAMIMGYGIHGLAKEAIKLFEEMQRTGHNPDDLTFLGVLTACN 709
Query: 742 SS 743
S
Sbjct: 710 HS 711
Score = 96.3 bits (238), Expect = 6e-20
Identities = 81/282 (28%), Positives = 127/282 (44%), Gaps = 13/282 (4%)
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVR 485
SL+ C+ E+H II +E L + +L +++ CG L + +FD M +
Sbjct: 501 SLLSACSKLKSLRLGKEVHGFIIRNWLERDLFVYLSVLSLYIHCGELCTVQALFDAMEDK 560
Query: 486 DFHSWATLFVSYYENGEYENAIDVFVSM-LCQLDVMGFSFPPWIWSCLLKACACTMNVPL 544
SW T+ Y +NG + A+ VF M L + + G S P + AC+ ++ L
Sbjct: 561 SLVSWNTVITGYLQNGFPDRALGVFRQMVLYGIQLCGISMMP-----VFGACSLLPSLRL 615
Query: 545 GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR 604
G + H LK D I+ SLI Y + + ++ VFN + +T +W A I+
Sbjct: 616 GREAHAYALKHLLEDDAFIACSLIDMYAKNGSITQSSKVFNGLKEKSTASWNAMIMGYGI 675
Query: 605 ERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQV-HADAIK--LGLDS 661
EA+ F++M R G D TF VL AC + G E + + D +K GL
Sbjct: 676 HGLAKEAIKLFEEMQRTGHNPDDLTFLGVLTAC---NHSGLIHEGLRYLDQMKSSFGLKP 732
Query: 662 DSYVQCSLIAMYGRSGLLRDA-ELVFEMTRNERNVDSLNAML 702
+ +I M GR+G L A +V E E +V ++L
Sbjct: 733 NLKHYACVIDMLGRAGQLDKALRVVAEEMSEEADVGIWKSLL 774
Score = 93.2 bits (230), Expect = 5e-19
Identities = 72/319 (22%), Positives = 144/319 (44%), Gaps = 4/319 (1%)
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
Y ++K C +D + +H ++ G+ + + N ++ + + G + +A ++FD+M
Sbjct: 190 YPCVIKACAGMSDVGIGLAVHGLVVKTGLVEDVFVGNALVSFYGTHGFVTDALQLFDIMP 249
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVP 543
R+ SW ++ + +NG E + + M+ + F +L CA +
Sbjct: 250 ERNLVSWNSMIRVFSDNGFSEESFLLLGEMMEENGDGAFMPDVATLVTVLPVCAREREIG 309
Query: 544 LGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSC 603
LG VHG +KL ++++++L+ Y + C+ +A M+F + N ++W +
Sbjct: 310 LGKGVHGWAVKLRLDKELVLNNALMDMYSKCGCITNAQMIFKMNNNKNVVSWNTMVGGFS 369
Query: 604 RERHFSEALGDFKKM--GRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDS 661
E ++M G VK D T + + C S +++H ++K
Sbjct: 370 AEGDTHGTFDVLRQMLAGGEDVKADEVTILNAVPVCFHESFLPSL-KELHCYSLKQEFVY 428
Query: 662 DSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQM 721
+ V + +A Y + G L A+ VF R+ + V+S NA++ G+ Q+ ++ QM
Sbjct: 429 NELVANAFVASYAKCGSLSYAQRVFHGIRS-KTVNSWNALIGGHAQSNDPRLSLDAHLQM 487
Query: 722 KAAGVQPHEPLLEKLRIAC 740
K +G+ P + L AC
Sbjct: 488 KISGLLPDSFTVCSLLSAC 506
Score = 89.4 bits (220), Expect = 7e-18
Identities = 67/285 (23%), Positives = 132/285 (45%), Gaps = 4/285 (1%)
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H + ++ L L N ++ M+ CG + NA+ +F + + ++ SW T+ + G+
Sbjct: 314 VHGWAVKLRLDKELVLNNALMDMYSKCGCITNAQMIFKMNNNKNVVSWNTMVGGFSAEGD 373
Query: 503 YENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVL 562
DV ML + + + + + C +P ++H LK + L
Sbjct: 374 THGTFDVLRQMLAGGEDVKADEVTILNA--VPVCFHESFLPSLKELHCYSLKQEFVYNEL 431
Query: 563 ISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVG 622
++++ + Y + L A VF+ + +W A I + +L +M G
Sbjct: 432 VANAFVASYAKCGSLSYAQRVFHGIRSKTVNSWNALIGGHAQSNDPRLSLDAHLQMKISG 491
Query: 623 VKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDA 682
+ DSFT S+L AC ++++ G++VH I+ L+ D +V S++++Y G L
Sbjct: 492 LLPDSFTVCSLLSACSKLKSL-RLGKEVHGFIIRNWLERDLFVYLSVLSLYIHCGELCTV 550
Query: 683 ELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQ 727
+ +F+ ++++ S N ++ GY+QNG A+ QM G+Q
Sbjct: 551 QALFD-AMEDKSLVSWNTVITGYLQNGFPDRALGVFRQMVLYGIQ 594
Score = 75.1 bits (183), Expect = 1e-13
Identities = 56/194 (28%), Positives = 94/194 (47%), Gaps = 8/194 (4%)
Query: 532 LLKACACTMNVPLGMQVHGCL---LKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVS 588
LL+A ++ +G ++H + +L D ++ + +I Y +D+ VF+ +
Sbjct: 90 LLQASGKRKDIEMGRKIHQLVSGSTRLRNDD--VLCTRIITMYAMCGSPDDSRFVFDALR 147
Query: 589 RHNTLTWTAKIVSSCRERHFSEALGDFKKM-GRVGVKKDSFTFSSVLKACGRMQNRGSCG 647
N W A I S R + E L F +M + D FT+ V+KAC M + G G
Sbjct: 148 SKNLFQWNAVISSYSRNELYDEVLETFIEMISTTDLLPDHFTYPCVIKACAGMSDVG-IG 206
Query: 648 EQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQ 707
VH +K GL D +V +L++ YG G + DA +F++ ERN+ S N+M+ +
Sbjct: 207 LAVHGLVVKTGLVEDVFVGNALVSFYGTHGFVTDALQLFDI-MPERNLVSWNSMIRVFSD 265
Query: 708 NGLYIEAVKFVYQM 721
NG E+ + +M
Sbjct: 266 NGFSEESFLLLGEM 279
>At5g55740 selenium-binding protein-like
Length = 830
Score = 130 bits (326), Expect = 4e-30
Identities = 98/331 (29%), Positives = 159/331 (47%), Gaps = 18/331 (5%)
Query: 412 MDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLT--LLNRILIMFVSC 469
MD + I +IY +++ C D T ++H +I+ G + +++I + C
Sbjct: 61 MDFRNLRIGPEIYGEILQGCVYERDLSTGKQIHARILKNGDFYARNEYIETKLVIFYAKC 120
Query: 470 GLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIW 529
LE A +F + VR+ SWA + G E A+ FV ML + ++ +F +
Sbjct: 121 DALEIAEVLFSKLRVRNVFSWAAIIGVKCRIGLCEGALMGFVEML-ENEIFPDNF---VV 176
Query: 530 SCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSR 589
+ KAC G VHG ++K G D V ++SSL YG+ L+DA+ VF+ +
Sbjct: 177 PNVCKACGALKWSRFGRGVHGYVVKSGLEDCVFVASSLADMYGKCGVLDDASKVFDEIPD 236
Query: 590 HNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQ 649
N + W A +V + EA+ F M + GV+ T S+ L A M G+Q
Sbjct: 237 RNAVAWNALMVGYVQNGKNEEAIRLFSDMRKQGVEPTRVTVSTCLSASANMGGVEE-GKQ 295
Query: 650 VHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNG 709
HA AI G++ D+ + SL+ Y + GL+ AE+VF+ E++V + N ++ GY+Q G
Sbjct: 296 SHAIAIVNGMELDNILGTSLLNFYCKVGLIEYAEMVFD-RMFEKDVVTWNLIISGYVQQG 354
Query: 710 LYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
L +A+ M+ LEKL+ C
Sbjct: 355 LVEDAIYMCQLMR----------LEKLKYDC 375
Score = 116 bits (290), Expect = 5e-26
Identities = 75/235 (31%), Positives = 121/235 (50%), Gaps = 8/235 (3%)
Query: 496 SYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKL 555
S +NGE + A+ S++ ++D P I+ +L+ C ++ G Q+H +LK
Sbjct: 44 SLCKNGEIKEAL----SLVTEMDFRNLRIGPEIYGEILQGCVYERDLSTGKQIHARILKN 99
Query: 556 G--ACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALG 613
G + I + L+ FY + LE A ++F+++ N +W A I CR AL
Sbjct: 100 GDFYARNEYIETKLVIFYAKCDALEIAEVLFSKLRVRNVFSWAAIIGVKCRIGLCEGALM 159
Query: 614 DFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMY 673
F +M + D+F +V KACG ++ G VH +K GL+ +V SL MY
Sbjct: 160 GFVEMLENEIFPDNFVVPNVCKACGALK-WSRFGRGVHGYVVKSGLEDCVFVASSLADMY 218
Query: 674 GRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQP 728
G+ G+L DA VF+ +RN + NA+++GY+QNG EA++ M+ GV+P
Sbjct: 219 GKCGVLDDASKVFDEI-PDRNAVAWNALMVGYVQNGKNEEAIRLFSDMRKQGVEP 272
Score = 115 bits (288), Expect = 9e-26
Identities = 81/287 (28%), Positives = 144/287 (49%), Gaps = 6/287 (2%)
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H ++ G+E + + + + M+ CG+L++A +VFD + R+ +W L V Y +NG+
Sbjct: 195 VHGYVVKSGLEDCVFVASSLADMYGKCGVLDDASKVFDEIPDRNAVAWNALMVGYVQNGK 254
Query: 503 YENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVL 562
E AI +F M Q G S L A A V G Q H + G +
Sbjct: 255 NEEAIRLFSDMRKQ----GVEPTRVTVSTCLSASANMGGVEEGKQSHAIAIVNGMELDNI 310
Query: 563 ISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVG 622
+ +SL+ FY + +E A MVF+R+ + +TW I ++ +A+ + M
Sbjct: 311 LGTSLLNFYCKVGLIEYAEMVFDRMFEKDVVTWNLIISGYVQQGLVEDAIYMCQLMRLEK 370
Query: 623 VKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDA 682
+K D T ++++ A R +N G++V I+ +SD + +++ MY + G + DA
Sbjct: 371 LKYDCVTLATLMSAAARTENL-KLGKEVQCYCIRHSFESDIVLASTVMDMYAKCGSIVDA 429
Query: 683 ELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPH 729
+ VF+ T E+++ N +L Y ++GL EA++ Y M+ GV P+
Sbjct: 430 KKVFDST-VEKDLILWNTLLAAYAESGLSGEALRLFYGMQLEGVPPN 475
Score = 73.6 bits (179), Expect = 4e-13
Identities = 67/292 (22%), Positives = 125/292 (41%), Gaps = 37/292 (12%)
Query: 438 ETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSY 497
E + H I G+EL L +L + GL+E A VFD M +D +W + Y
Sbjct: 291 EEGKQSHAIAIVNGMELDNILGTSLLNFYCKVGLIEYAEMVFDRMFEKDVVTWNLIISGY 350
Query: 498 YENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGA 557
+ G E+AI ++ L +L+ + + + L+ A A T N+ LG +V ++
Sbjct: 351 VQQGLVEDAI--YMCQLMRLEKLKYDCVTL--ATLMSAAARTENLKLGKEVQCYCIRHSF 406
Query: 558 CDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKK 617
++++S+++ Y + + DA VF+ + + W + + EAL F
Sbjct: 407 ESDIVLASTVMDMYAKCGSIVDAKKVFDSTVEKDLILWNTLLAAYAESGLSGEALRLFYG 466
Query: 618 MGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSG 677
M GV + T++ ++ + + G E D ++Q SG
Sbjct: 467 MQLEGVPPNVITWNLIILS---LLRNGQVDEA-----------KDMFLQMQ------SSG 506
Query: 678 LLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPH 729
++ N+ S M+ G +QNG EA+ F+ +M+ +G++P+
Sbjct: 507 II-------------PNLISWTTMMNGMVQNGCSEEAILFLRKMQESGLRPN 545
Score = 71.6 bits (174), Expect = 1e-12
Identities = 69/302 (22%), Positives = 127/302 (41%), Gaps = 42/302 (13%)
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
E+ I E + L + ++ M+ CG + +A++VFD +D W TL +Y E+G
Sbjct: 396 EVQCYCIRHSFESDIVLASTVMDMYAKCGSIVDAKKVFDSTVEKDLILWNTLLAAYAESG 455
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWI--WSCLLKACACTMNVPLGMQVHGCLLKLGACD 559
A+ +F M QL+ PP + W+ ++ + LL+ G D
Sbjct: 456 LSGEALRLFYGM--QLE----GVPPNVITWNLIILS----------------LLRNGQVD 493
Query: 560 HVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMG 619
F ++ + ++ N +S WT + + EA+ +KM
Sbjct: 494 EA---------KDMFLQMQSSGIIPNLIS------WTTMMNGMVQNGCSEEAILFLRKMQ 538
Query: 620 RVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIK-LGLDSDSYVQCSLIAMYGRSGL 678
G++ ++F+ + L AC + + G +H I+ L S ++ SL+ MY + G
Sbjct: 539 ESGLRPNAFSITVALSACAHLASL-HIGRTIHGYIIRNLQHSSLVSIETSLVDMYAKCGD 597
Query: 679 LRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRI 738
+ AE VF ++ + NAM+ Y G EA+ ++ G++P + +
Sbjct: 598 INKAEKVFG-SKLYSELPLSNAMISAYALYGNLKEAIALYRSLEGVGLKPDNITITNVLS 656
Query: 739 AC 740
AC
Sbjct: 657 AC 658
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,459,443
Number of Sequences: 26719
Number of extensions: 882575
Number of successful extensions: 9113
Number of sequences better than 10.0: 495
Number of HSP's better than 10.0 without gapping: 300
Number of HSP's successfully gapped in prelim test: 202
Number of HSP's that attempted gapping in prelim test: 4073
Number of HSP's gapped (non-prelim): 2458
length of query: 749
length of database: 11,318,596
effective HSP length: 107
effective length of query: 642
effective length of database: 8,459,663
effective search space: 5431103646
effective search space used: 5431103646
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144759.9


